

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137839.10 - phase: 0
(502 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
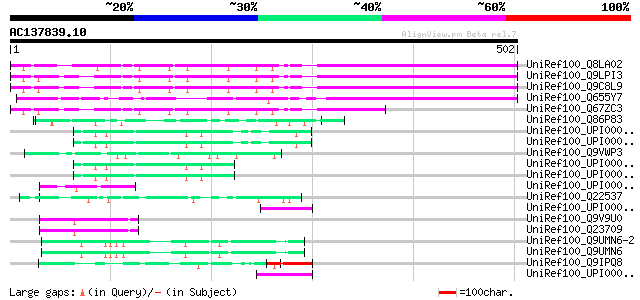
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8LA02 Contains similarity to RNA-binding protein from... 327 4e-88
UniRef100_Q9LPI3 T3F20.1 protein [Arabidopsis thaliana] 327 6e-88
UniRef100_Q9C8L9 Hypothetical protein F22G10.8 [Arabidopsis thal... 327 6e-88
UniRef100_Q655Y7 Hydroxyproline-rich glycoprotein-like [Oryza sa... 310 6e-83
UniRef100_Q67ZC3 Hypothetical protein At1g53645 [Arabidopsis tha... 144 8e-33
UniRef100_Q86P83 AT02511p [Drosophila melanogaster] 61 7e-08
UniRef100_UPI00003AE651 UPI00003AE651 UniRef100 entry 58 7e-07
UniRef100_UPI00003AE64F UPI00003AE64F UniRef100 entry 58 7e-07
UniRef100_Q9VWP3 CG7282-PA [Drosophila melanogaster] 55 5e-06
UniRef100_UPI00003AE650 UPI00003AE650 UniRef100 entry 55 6e-06
UniRef100_UPI00003AE64E UPI00003AE64E UniRef100 entry 55 6e-06
UniRef100_UPI00003222CF UPI00003222CF UniRef100 entry 54 8e-06
UniRef100_Q22537 Hypothetical protein T17H7.1 [Caenorhabditis el... 54 1e-05
UniRef100_UPI0000361AF7 UPI0000361AF7 UniRef100 entry 53 2e-05
UniRef100_Q9V9U0 CG2150-PA [Drosophila melanogaster] 52 3e-05
UniRef100_Q23709 GCR 1 protein [Drosophila melanogaster] 52 3e-05
UniRef100_Q9UMN6-2 Splice isoform 2 of Q9UMN6 [Homo sapiens] 52 3e-05
UniRef100_Q9UMN6 Myeloid/lymphoid or mixed-lineage leukemia prot... 52 3e-05
UniRef100_Q9IPQ8 EBNA-1 [Cynomolgus Epstein-Barr Virus Si-IIA] 52 4e-05
UniRef100_UPI00002D1908 UPI00002D1908 UniRef100 entry 52 5e-05
>UniRef100_Q8LA02 Contains similarity to RNA-binding protein from Arabidopsis
thaliana gi|2129727 and contains RNA recognition
PF|00076 domain [Arabidopsis thaliana]
Length = 523
Score = 327 bits (839), Expect = 4e-88
Identities = 226/560 (40%), Positives = 294/560 (52%), Gaps = 95/560 (16%)
Query: 1 MRGTIGVRLQNS---TISNATRQTLVPFST-SSGFGGGGGDGRGGGRGRGGSGTVTFNFG 56
MR IG R N TI++ +QT PF T S+ D G GRGRG
Sbjct: 1 MRSAIGRRFSNPNGFTIASLVKQT--PFLTQSTSHFSSSSDSSGRGRGRGS--------- 49
Query: 57 EKAAPGNPNPTPNVNESKPDATDSPIPPG------AGRGHGRGGTVP-DFPSFSFSSFMS 109
G P + P+ PG G GHGRG + D S +F+SF+
Sbjct: 50 -----GEDGGFPTAGRGQFGVNREPVVPGREPSSAGGYGHGRGRPIQSDSISPAFTSFVK 104
Query: 110 SIQQPGTGRGRGRGRGFD-------------PLPPQFENDSVPKKPVFIKREDNVSQTDA 156
S P GRGRG G D P PPQ + ++ +++ SQ
Sbjct: 105 S-DSPSIGRGRG-SVGSDTVSPFAAEPPRQSPPPPQQQQSQSQQQRSQPQQQQPRSQPQQ 162
Query: 157 --NDFSPPKNPVFTRSEDVR-----PVEPIDLSGDSES-DNRFVMTVPKVLPGGGRGRGK 208
ND S +PVF + ++++ P P G ++ DN F + G GRGK
Sbjct: 163 QPNDESQG-SPVFVKLQEMQDATSSPPPPESKPGQADPPDNIFNALGNEFSHPSGAGRGK 221
Query: 209 PLEEAAQ---------EAPQAPVVNRHIRVRQTPADAESDNVPR---------RQPMNRF 250
PL E+A P P + ++ +Q A D P+ R+ +
Sbjct: 222 PLVESAPIRQEDNRQIRRPPPPPQQQRVQPQQKRAPTVKDGTPKPQLSAEEAGRRARSEL 281
Query: 251 VRDDGDGS------GRGRGRGRGRDVYARGRGDRGRGGRGGRGDGRGGFKRYGDDRTSHQ 304
R + +GS GRGRGRGRG ARGRG RGRGG G R D + +
Sbjct: 282 SRGEAEGSSVGGRGGRGRGRGRG----ARGRG-RGRGGDGWRDDKK-------------E 323
Query: 305 DIARSNADGLYVGDNADGEKLAKKLGPEIMDQITEAYEEIIERVLPSPLQDEYVEAMDIN 364
+ A ++ GD+ADGEK A+K+GPE+M + E +EEI E+ LPS D ++A D N
Sbjct: 324 EEGEQEAMRIFAGDSADGEKFAEKMGPELMKTLAEGFEEICEKALPSTTHDAIIDAYDTN 383
Query: 365 CAIEFEPEYAV-EF-DNPDIDEKEPIALRDALEKMKPFLMTYEGIRSQEEWEEVIEELMQ 422
IE EPEY + +F NPDIDEK P++LR+ LEK+KPF++ YEGI+ QEEWEE I E M
Sbjct: 384 LMIECEPEYIMPDFGSNPDIDEKPPMSLRECLEKVKPFIVAYEGIKDQEEWEEAINEAMT 443
Query: 423 RVPLLKKIVDHYSGPDRVTAKKQQEELERVAKTLPTSAPSSVKEFTNRAVVSLQSNPGWG 482
+ PL+K+IVDHYSGPDRVTAKKQ EEL+R+A TLP SAP SVK F +RA ++L+SNPGWG
Sbjct: 444 QAPLMKEIVDHYSGPDRVTAKKQNEELDRIATTLPASAPDSVKRFADRAALTLKSNPGWG 503
Query: 483 FDKKCQFMDKLVFEVSQHHK 502
FDKK QFMDKLV EVSQ +K
Sbjct: 504 FDKKYQFMDKLVLEVSQSYK 523
>UniRef100_Q9LPI3 T3F20.1 protein [Arabidopsis thaliana]
Length = 829
Score = 327 bits (837), Expect = 6e-88
Identities = 229/564 (40%), Positives = 295/564 (51%), Gaps = 103/564 (18%)
Query: 1 MRGTIGVRLQNS---TISNATRQTLVPFST-----------SSGFGGGGGDGRGGGRGRG 46
MR IG R N TI++ +QT PF T SSG G G G G GG
Sbjct: 307 MRSAIGRRFSNPNGFTIASLVKQT--PFLTQSTSHFSSSSDSSGRGRGRGSGEDGGFPAA 364
Query: 47 GSGTVTFNFGEKAAPGNPNPTPNVNESKPDATDSPIPPGAGRGHGRGGTVP-DFPSFSFS 105
G G FG P P P+ G GHGRG + D S +F+
Sbjct: 365 GRG----QFGVNREPVVPGREPS--------------SAGGYGHGRGRPIQSDSISPAFT 406
Query: 106 SFMSSIQQPGTGRGRGRGRGFD-------------PLPPQFENDSVPKKPVFIKREDNVS 152
SF+ S P GRGRG G D P PPQ + ++ +++ S
Sbjct: 407 SFVKS-DSPSIGRGRG-SVGSDTVSPFAAEPPRQSPPPPQQQQSQSQQQRSQPQQQQPRS 464
Query: 153 QTDA--NDFSPPKNPVFTRSEDVR-----PVEPIDLSGDSES-DNRFVMTVPKVLPGGGR 204
Q ND S +PVF + ++++ P P G ++ DN F + G
Sbjct: 465 QPQQQPNDESQG-SPVFVKLQEMQDATSSPPPPESKPGQADPPDNIFNALGNEFSHPSGA 523
Query: 205 GRGKPLEEAAQ---------EAPQAPVVNRHIRVRQTPADAESDNVPR---------RQP 246
GRGKPL E+A P P + ++ +Q A D P+ R+
Sbjct: 524 GRGKPLVESAPIRQEDNRQIRRPPPPPQQQRVQPQQKRAPTVKDGTPKPQLSAEEAGRRA 583
Query: 247 MNRFVRDDGDGS------GRGRGRGRGRDVYARGRGDRGRGGRGGRGDGRGGFKRYGDDR 300
+ R + +GS GRGRGRGRG ARGRG RGRGG G R D +
Sbjct: 584 RSELSRGEAEGSSVGGRGGRGRGRGRG----ARGRG-RGRGGDGWRDDKK---------- 628
Query: 301 TSHQDIARSNADGLYVGDNADGEKLAKKLGPEIMDQITEAYEEIIERVLPSPLQDEYVEA 360
++ A ++ GD+ADGEK A+K+GPE+M + E +EEI E+ LPS D ++A
Sbjct: 629 ---EEEGEQEAMRIFAGDSADGEKFAEKMGPELMKTLAEGFEEICEKALPSTTHDAIIDA 685
Query: 361 MDINCAIEFEPEYAV-EF-DNPDIDEKEPIALRDALEKMKPFLMTYEGIRSQEEWEEVIE 418
D N IE EPEY + +F NPDIDEK P++LR+ LEK+KPF++ YEGI+ QEEWEE I
Sbjct: 686 YDTNLMIECEPEYIMPDFGSNPDIDEKPPMSLRECLEKVKPFIVAYEGIKDQEEWEEAIN 745
Query: 419 ELMQRVPLLKKIVDHYSGPDRVTAKKQQEELERVAKTLPTSAPSSVKEFTNRAVVSLQSN 478
E M + PL+K+IVDHYSGPDRVTAKKQ EEL+R+A TLP SAP SVK F +RA ++L+SN
Sbjct: 746 EAMTQAPLMKEIVDHYSGPDRVTAKKQNEELDRIATTLPASAPDSVKRFADRAALTLKSN 805
Query: 479 PGWGFDKKCQFMDKLVFEVSQHHK 502
PGWGFDKK QFMDKLV EVSQ +K
Sbjct: 806 PGWGFDKKYQFMDKLVLEVSQSYK 829
>UniRef100_Q9C8L9 Hypothetical protein F22G10.8 [Arabidopsis thaliana]
Length = 523
Score = 327 bits (837), Expect = 6e-88
Identities = 229/564 (40%), Positives = 295/564 (51%), Gaps = 103/564 (18%)
Query: 1 MRGTIGVRLQNS---TISNATRQTLVPFST-----------SSGFGGGGGDGRGGGRGRG 46
MR IG R N TI++ +QT PF T SSG G G G G GG
Sbjct: 1 MRSAIGRRFSNPNGFTIASLVKQT--PFLTQSTSHFSSSSDSSGRGRGRGSGEDGGFPAA 58
Query: 47 GSGTVTFNFGEKAAPGNPNPTPNVNESKPDATDSPIPPGAGRGHGRGGTVP-DFPSFSFS 105
G G FG P P P+ G GHGRG + D S +F+
Sbjct: 59 GRG----QFGVNREPVVPGREPS--------------SAGGYGHGRGRPIQSDSISPAFT 100
Query: 106 SFMSSIQQPGTGRGRGRGRGFD-------------PLPPQFENDSVPKKPVFIKREDNVS 152
SF+ S P GRGRG G D P PPQ + ++ +++ S
Sbjct: 101 SFVKS-DSPSIGRGRG-SVGSDTVSPFAAEPPRQSPPPPQQQQSQSQQQRSQPQQQQPRS 158
Query: 153 QTDA--NDFSPPKNPVFTRSEDVR-----PVEPIDLSGDSES-DNRFVMTVPKVLPGGGR 204
Q ND S +PVF + ++++ P P G ++ DN F + G
Sbjct: 159 QPQQQPNDESQG-SPVFVKLQEMQDATSSPPPPESKPGQADPPDNIFNALGNEFSHPSGA 217
Query: 205 GRGKPLEEAAQ---------EAPQAPVVNRHIRVRQTPADAESDNVPR---------RQP 246
GRGKPL E+A P P + ++ +Q A D P+ R+
Sbjct: 218 GRGKPLVESAPIRQEDNRQIRRPPPPPQQQRVQPQQKRAPTVKDGTPKPQLSAEEAGRRA 277
Query: 247 MNRFVRDDGDGS------GRGRGRGRGRDVYARGRGDRGRGGRGGRGDGRGGFKRYGDDR 300
+ R + +GS GRGRGRGRG ARGRG RGRGG G R D +
Sbjct: 278 RSELSRGEAEGSSVGGRGGRGRGRGRG----ARGRG-RGRGGDGWRDDKK---------- 322
Query: 301 TSHQDIARSNADGLYVGDNADGEKLAKKLGPEIMDQITEAYEEIIERVLPSPLQDEYVEA 360
++ A ++ GD+ADGEK A+K+GPE+M + E +EEI E+ LPS D ++A
Sbjct: 323 ---EEEGEQEAMRIFAGDSADGEKFAEKMGPELMKTLAEGFEEICEKALPSTTHDAIIDA 379
Query: 361 MDINCAIEFEPEYAV-EF-DNPDIDEKEPIALRDALEKMKPFLMTYEGIRSQEEWEEVIE 418
D N IE EPEY + +F NPDIDEK P++LR+ LEK+KPF++ YEGI+ QEEWEE I
Sbjct: 380 YDTNLMIECEPEYIMPDFGSNPDIDEKPPMSLRECLEKVKPFIVAYEGIKDQEEWEEAIN 439
Query: 419 ELMQRVPLLKKIVDHYSGPDRVTAKKQQEELERVAKTLPTSAPSSVKEFTNRAVVSLQSN 478
E M + PL+K+IVDHYSGPDRVTAKKQ EEL+R+A TLP SAP SVK F +RA ++L+SN
Sbjct: 440 EAMTQAPLMKEIVDHYSGPDRVTAKKQNEELDRIATTLPASAPDSVKRFADRAALTLKSN 499
Query: 479 PGWGFDKKCQFMDKLVFEVSQHHK 502
PGWGFDKK QFMDKLV EVSQ +K
Sbjct: 500 PGWGFDKKYQFMDKLVLEVSQSYK 523
>UniRef100_Q655Y7 Hydroxyproline-rich glycoprotein-like [Oryza sativa]
Length = 436
Score = 310 bits (794), Expect = 6e-83
Identities = 200/507 (39%), Positives = 265/507 (51%), Gaps = 82/507 (16%)
Query: 7 VRLQNSTISNATRQTLVPFSTSSGFGGGGGDGRGGGRGRGGSGTVTFNFGEKAAPGNPNP 66
+R + + A R +P S S+ F G G G GRGRG + + APG+P P
Sbjct: 1 MRAIGAAAAAARRHAHLPTSYSAAFSSFSGIGGGAGRGRGRG--LPPSATPPRAPGSPVP 58
Query: 67 TPNVNESKPDATDSPIPPGAGRGHGRGGTVPDFPSFSFSSFMSSIQQPGTGRGRGRGRGF 126
+ + D SP P G GRG +P +SS PG GRGRG
Sbjct: 59 DDD-DGGGADPFSSPAPIGRGRGE---AVIPS---------VSSPPLPGAGRGRGS---- 101
Query: 127 DPLPPQFENDSVPKKPVFIKREDNVSQTDANDFSPPKNPVFTRSEDVRPVEPIDLSGDSE 186
PP + PK+PV K D + ++ PP P
Sbjct: 102 ---PPPL-GEVAPKQPVPAKLFDAPAAEASSSEPPPPPP--------------------- 136
Query: 187 SDNRFVMTVPKVLPGGGRGRGKP-LEEAAQEAPQAPVVNRHIRVRQTPADAESDNVPRRQ 245
P+ LP G GRG P +++ E PQ NR IR R+ A S P
Sbjct: 137 ---------PRTLPSAGAGRGVPRMQQPPVEMPQEE--NRFIRRREEKKKAASAARPAPS 185
Query: 246 PMNRFVRDD----------GDGSGRGRGRGRGRDVYARGRGDRGRGGRGGRGDGRGGFKR 295
+ +D G G G GRGR R RG RGRG GGRG +R
Sbjct: 186 GQPKLSPEDAVKRAMELLGGGGDDDGGRGGRGRGARGRERG-RGRGRDGGRG------RR 238
Query: 296 YGDDRTSHQDIARSNADGLYVGDNADGEKLAKKLGPEIMDQITEAYEEIIERVLPSPLQD 355
D H G+Y+GDNADG++L K+LG + M EA++E + LP P QD
Sbjct: 239 SADMEEKH---------GIYLGDNADGDRLQKRLGEDKMKIFNEAFDEAADNALPDPKQD 289
Query: 356 EYVEAMDINCAIEFEPEYAVEFDNPDIDEKEPIALRDALEKMKPFLMTYEGIRSQEEWEE 415
Y+EA N IEFEPEY V F+NPDI+EK P++L D L+K+KPF++ YEGI++QEEWEE
Sbjct: 290 AYLEACHTNNMIEFEPEYHVNFNNPDIEEKPPMSLEDMLQKVKPFIVAYEGIQNQEEWEE 349
Query: 416 VIEELMQRVPLLKKIVDHYSGPDRVTAKKQQEELERVAKTLPTSAPSSVKEFTNRAVVSL 475
++++M R P +K+++D YSGPD VTAK+Q+EEL+RVA TLP + PSSVK FT++ ++SL
Sbjct: 350 AVKDVMARAPHMKELIDMYSGPDVVTAKQQEEELQRVANTLPGNIPSSVKRFTDKTLLSL 409
Query: 476 QSNPGWGFDKKCQFMDKLVFEVSQHHK 502
++NPGWGFDKKCQFMDK EVS+ +K
Sbjct: 410 KNNPGWGFDKKCQFMDKFAREVSELYK 436
>UniRef100_Q67ZC3 Hypothetical protein At1g53645 [Arabidopsis thaliana]
Length = 417
Score = 144 bits (362), Expect = 8e-33
Identities = 139/432 (32%), Positives = 186/432 (42%), Gaps = 101/432 (23%)
Query: 1 MRGTIGVRLQNS---TISNATRQTLVPFST-----------SSGFGGGGGDGRGGGRGRG 46
MR IG R N TI++ +QT PF T SSG G G G G GG
Sbjct: 1 MRSAIGRRFSNPNGFTIASLVKQT--PFLTQSTSHFSSSSDSSGRGRGRGSGEDGGFPAA 58
Query: 47 GSGTVTFNFGEKAAPGNPNPTPNVNESKPDATDSPIPPGAGRGHGRGGTVP-DFPSFSFS 105
G G FG P P P+ G GHGRG + D S +F+
Sbjct: 59 GRG----QFGVNREPVVPGREPS--------------SAGGYGHGRGRPIQSDSISPAFT 100
Query: 106 SFMSSIQQPGTGRGRGRGRGFD-------------PLPPQFENDSVPKKPVFIKREDNVS 152
SF+ S P GRGRG G D P PPQ + ++ +++ S
Sbjct: 101 SFVKS-DSPSIGRGRG-SVGSDTVSPFAAEPPRQSPPPPQQQQSQSQQQRSQPQQQQPRS 158
Query: 153 QTDA--NDFSPPKNPVFTRSEDVR-----PVEPIDLSGDSES-DNRFVMTVPKVLPGGGR 204
Q ND S +PVF + ++++ P P G ++ DN F + G
Sbjct: 159 QPQQQPNDESQG-SPVFVKLQEMQDATSSPPPPESKPGQADPPDNIFNALGNEFSHPSGA 217
Query: 205 GRGKPLEEAAQ---------EAPQAPVVNRHIRVRQTPADAESDNVPR---------RQP 246
GRGKPL E+A P P + ++ +Q A D P+ R+
Sbjct: 218 GRGKPLVESAPIRQEDNRQIRRPPPPPQQQRVQPQQKRAPTVKDGTPKPQLSAEEAGRRA 277
Query: 247 MNRFVRDDGDGS------GRGRGRGRGRDVYARGRGDRGRGGRGGRGDGRGGFKRYGDDR 300
+ R + +GS GRGRGRGRG ARGRG RGRGG G R D +
Sbjct: 278 RSELSRGEAEGSSVGGRGGRGRGRGRG----ARGRG-RGRGGDGWRDDKK---------- 322
Query: 301 TSHQDIARSNADGLYVGDNADGEKLAKKLGPEIMDQITEAYEEIIERVLPSPLQDEYVEA 360
++ A ++ GD+ADGEK A+K+GPE+M + E +EEI E+ LPS D ++A
Sbjct: 323 ---EEEGEQEAMRIFAGDSADGEKFAEKMGPELMKTLAEGFEEICEKALPSTTHDAIIDA 379
Query: 361 MDINCAIEFEPE 372
D N IE EPE
Sbjct: 380 YDTNLMIECEPE 391
>UniRef100_Q86P83 AT02511p [Drosophila melanogaster]
Length = 575
Score = 61.2 bits (147), Expect = 7e-08
Identities = 82/328 (25%), Positives = 111/328 (33%), Gaps = 78/328 (23%)
Query: 24 PFSTSSGFGGGGGDGRGG---------GRGRGGSGTVTFNFGEKAAPGNPNPTPNVNESK 74
P T GGG G+G GG G+G+GG G P + P N+
Sbjct: 229 PSDTYGAPGGGNGNGSGGRPSSSYGAPGQGQGGFG---------GRPSDSYGAPGQNQKP 279
Query: 75 PDATDSPIPPGAGRGHGRGGTVPDFPSFSFSSFMS--------SIQQPGTGRGRGRGRGF 126
D+ +P G G+G GG PS S+ + S S P +G G G G
Sbjct: 280 SDSYGAP-----GSGNGNGGR----PSSSYGAPGSGPGGRPSDSYGPPASGSGAGGAGGS 330
Query: 127 DPLPPQFENDSVPKKPVFIKREDNVS-QTDANDFSPPKNPVFTRSEDVRPVEPIDLSGDS 185
P ++ND V + + + DAND S P P + +L D
Sbjct: 331 GPGGADYDNDIVEYEADQQGYRPQIRYEGDANDGSGPSGPGGPGGQ--------NLGADG 382
Query: 186 ESDNRFVMTVPKVLPGGGRGRGKPLEEAAQEAPQAPVVNRHIRVRQTPADAESDNVPRRQ 245
S R PG G G G + Q + + R D + +
Sbjct: 383 YSSGR---------PGNGNGNGNGGYSGGRPGGQDLGPSGYSGGRPGGQDLGAGGYSNGK 433
Query: 246 PMNRFVRDDGDGSGRGRGRGRGRDVYARGR-------------GDRGRGGRGGRGDGR-- 290
P + + G GR G+ GRD Y+ GR G G G GG GR
Sbjct: 434 PGGQDLGPGGYSGGRPGGQDLGRDGYSGGRPGGQDLGASGYSNGRPGGNGNGGSDGGRVI 493
Query: 291 ----------GGFKRYGDDRTSHQDIAR 308
GG + Y R QD+ R
Sbjct: 494 IGGRVIGGQDGGDQGYSGGRPGGQDLGR 521
Score = 50.4 bits (119), Expect = 1e-04
Identities = 82/322 (25%), Positives = 108/322 (33%), Gaps = 65/322 (20%)
Query: 26 STSSGFGGGGGDGRG----GGRGRGGSGTVTFNFGEKAAPGNPNPTPNVNESKPDATDSP 81
S+S G GGG GR G G G G + +G GN P+ + P +
Sbjct: 169 SSSYGAPGGGNGGRPSDTYGAPGGGNGGRPSDTYGAPGG-GNNGGRPSSSYGAPGGGNGG 227
Query: 82 IP------PGAGRGHGRGGTVPDFPSFSFSSFMSSIQQPGTGRGRGRGRGFDPLPPQFEN 135
P PG G G+G GG PS S+ + PG G+G GR D +N
Sbjct: 228 RPSDTYGAPGGGNGNGSGGR----PSSSYGA-------PGQGQGGFGGRPSDSYGAPGQN 276
Query: 136 DSVPKKPVFIKREDNVSQTDANDFSPPKNPVFTRSEDVRPVEPIDLSGDSESDNRFVMTV 195
Q ++ + P + RP G
Sbjct: 277 -----------------QKPSDSYGAPGSG---NGNGGRPSSSYGAPGSGPGGRPSDSYG 316
Query: 196 PKVLPGGGRGRGKPLEEAAQEAPQAPVVNRHIRVRQTPADAESDNVPRRQPMNRFVRDDG 255
P P G G G A P + I + E+D R P R+ D
Sbjct: 317 P---PASGSGAGG----AGGSGPGGADYDNDI------VEYEADQQGYR-PQIRYEGDAN 362
Query: 256 DGSGRGR-----GRGRGRDVYARGRGDRGRG-GRGGRGDGRGGFKRYGDDRTSHQDIARS 309
DGSG G+ G D Y+ GR G G G GG GR G + G S R
Sbjct: 363 DGSGPSGPGGPGGQNLGADGYSSGRPGNGNGNGNGGYSGGRPGGQDLGPSGYSG---GRP 419
Query: 310 NADGLYVGDNADGEKLAKKLGP 331
L G ++G+ + LGP
Sbjct: 420 GGQDLGAGGYSNGKPGGQDLGP 441
>UniRef100_UPI00003AE651 UPI00003AE651 UniRef100 entry
Length = 457
Score = 57.8 bits (138), Expect = 7e-07
Identities = 76/286 (26%), Positives = 96/286 (32%), Gaps = 68/286 (23%)
Query: 64 PNPTPNVNESKPDATDSPIP----PGAGRGHGRGG--------------TVPDFPSFSFS 105
P P P+V SKP+AT P+P P A HGRG T P FP +
Sbjct: 135 PPPRPDVG-SKPEATPPPVPSTPRPIASSLHGRGSAPAPVLNRQPSLGPTPPPFPGSRAA 193
Query: 106 SFMSSIQQPGTGRGRGRGRGFDPLPPQFENDSVPKKPVFIKREDNVSQTDANDFSPPKNP 165
S++QPG G G PLPP V +KP T + D + P P
Sbjct: 194 GSAGSLRQPGPGPATPYS-GRPPLPPTPGRSPVDEKPPPPPPPSGHRPTASRDMALPPPP 252
Query: 166 VFTRSEDV------------------RPVEPIDLSGDSESD------NRFVMTVPKVLPG 201
V RP P G + SD R + VP P
Sbjct: 253 PQNSKPPVPASPRPPLGVPAPPPPPSRPGPPPVPPGPASSDEMPRLPQRNLSLVPPAAPS 312
Query: 202 GGRGRGKPLEEAAQEAPQAPVVNRHIRVRQTPADAESDNVPRRQPMNRFVRDDGDGSGRG 261
G GR PL E P P +R P+ A +P P++R
Sbjct: 313 SGSGRSGPLPPPPSERPPPP-------IRDPPSRA--GPLPPPPPISR-------NGSTS 356
Query: 262 RGRGRGRDVYARGRGDRGRGG--------RGGRGDGRGGFKRYGDD 299
R + +R + RGG R G R GF+ GDD
Sbjct: 357 RALPAAPQLPSRAGLENQRGGPRPPLPPDRPGTAALRNGFQESGDD 402
>UniRef100_UPI00003AE64F UPI00003AE64F UniRef100 entry
Length = 433
Score = 57.8 bits (138), Expect = 7e-07
Identities = 76/286 (26%), Positives = 96/286 (32%), Gaps = 68/286 (23%)
Query: 64 PNPTPNVNESKPDATDSPIP----PGAGRGHGRGG--------------TVPDFPSFSFS 105
P P P+V SKP+AT P+P P A HGRG T P FP +
Sbjct: 111 PPPRPDVG-SKPEATPPPVPSTPRPIASSLHGRGSAPAPVLNRQPSLGPTPPPFPGSRAA 169
Query: 106 SFMSSIQQPGTGRGRGRGRGFDPLPPQFENDSVPKKPVFIKREDNVSQTDANDFSPPKNP 165
S++QPG G G PLPP V +KP T + D + P P
Sbjct: 170 GSAGSLRQPGPGPATPYS-GRPPLPPTPGRSPVDEKPPPPPPPSGHRPTASRDMALPPPP 228
Query: 166 VFTRSEDV------------------RPVEPIDLSGDSESD------NRFVMTVPKVLPG 201
V RP P G + SD R + VP P
Sbjct: 229 PQNSKPPVPASPRPPLGVPAPPPPPSRPGPPPVPPGPASSDEMPRLPQRNLSLVPPAAPS 288
Query: 202 GGRGRGKPLEEAAQEAPQAPVVNRHIRVRQTPADAESDNVPRRQPMNRFVRDDGDGSGRG 261
G GR PL E P P +R P+ A +P P++R
Sbjct: 289 SGSGRSGPLPPPPSERPPPP-------IRDPPSRA--GPLPPPPPISR-------NGSTS 332
Query: 262 RGRGRGRDVYARGRGDRGRGG--------RGGRGDGRGGFKRYGDD 299
R + +R + RGG R G R GF+ GDD
Sbjct: 333 RALPAAPQLPSRAGLENQRGGPRPPLPPDRPGTAALRNGFQESGDD 378
>UniRef100_Q9VWP3 CG7282-PA [Drosophila melanogaster]
Length = 1868
Score = 55.1 bits (131), Expect = 5e-06
Identities = 75/283 (26%), Positives = 111/283 (38%), Gaps = 43/283 (15%)
Query: 15 SNATRQTLVPFSTSSGFGGGGGDGRGGGRGRGGSGTVTFNFGEKAAPGNPNPTPNVNESK 74
S +T+ S +SG GGGGG G GGG G GG G G G + P V++ +
Sbjct: 141 SKQKTRTISTSSANSGSGGGGGGGGGGGGGGGGGG------GSLLVQG--SQPPGVSDKQ 192
Query: 75 PDATD-SPIPPGAGRGHGRGGTVPDFPSFSFS--------SFMSSIQQ--PGTGRGRGRG 123
P S + P +G G P PS S S S +S++ + P G RGRG
Sbjct: 193 PGKDGCSKMSPSSGNSTG-----PGAPSLSGSLGGASSTPSLLSTVVKTPPTGGAKRGRG 247
Query: 124 RGFDPLPPQFENDSVPKKPVFIKREDNVSQTDANDFSPPKNP--VFTRSEDVRPVEPIDL 181
R D +PP+ S + SQ K P V + R V +
Sbjct: 248 RS-DSMPPRSTTPSSVVAHSGRTKSPAASQPQLQQ-QMKKRPTRVVPGTTTPRRVSDASM 305
Query: 182 SGDSESDNRFVMTVP-----KVLPGGGR----GRGKPLEEAAQEAPQAPVV--NRHIRVR 230
+ +S+SD+ + P K P G+ G+G+ A+ AP A +
Sbjct: 306 ASESDSDSDEPVRRPKRQSAKDKPQAGKAQPPGKGRLASSASSTAPAAHPSDDSEEDEEE 365
Query: 231 QTPADAESDNVPRRQPMNRFVRDDGDGSGR----GRGRGRGRD 269
+ P+ A + + ++Q +R G R G +GRD
Sbjct: 366 EEPSAARAASSKQQQQQASSLRGSRAGGNRAMSSGAASAKGRD 408
>UniRef100_UPI00003AE650 UPI00003AE650 UniRef100 entry
Length = 411
Score = 54.7 bits (130), Expect = 6e-06
Identities = 57/201 (28%), Positives = 69/201 (33%), Gaps = 44/201 (21%)
Query: 64 PNPTPNVNESKPDATDSPIP----PGAGRGHGRGG--------------TVPDFPSFSFS 105
P P P+V SKP+AT P+P P A HGRG T P FP +
Sbjct: 111 PPPRPDVG-SKPEATPPPVPSTPRPIASSLHGRGSAPAPVLNRQPSLGPTPPPFPGSRAA 169
Query: 106 SFMSSIQQPGTGRGRGRGRGFDPLPPQFENDSVPKKPVFIKREDNVSQTDANDFSPPKNP 165
S++QPG G G PLPP V +KP T + D + P P
Sbjct: 170 GSAGSLRQPGPGPATPYS-GRPPLPPTPGRSPVDEKPPPPPPPSGHRPTASRDMALPPPP 228
Query: 166 VFTRSEDV------------------RPVEPIDLSGDSESD------NRFVMTVPKVLPG 201
V RP P G + SD R + VP P
Sbjct: 229 PQNSKPPVPASPRPPLGVPAPPPPPSRPGPPPVPPGPASSDEMPRLPQRNLSLVPPAAPS 288
Query: 202 GGRGRGKPLEEAAQEAPQAPV 222
G GR PL E P P+
Sbjct: 289 SGSGRSGPLPPPPSERPPPPI 309
>UniRef100_UPI00003AE64E UPI00003AE64E UniRef100 entry
Length = 496
Score = 54.7 bits (130), Expect = 6e-06
Identities = 57/201 (28%), Positives = 69/201 (33%), Gaps = 44/201 (21%)
Query: 64 PNPTPNVNESKPDATDSPIP----PGAGRGHGRGG--------------TVPDFPSFSFS 105
P P P+V SKP+AT P+P P A HGRG T P FP +
Sbjct: 163 PPPRPDVG-SKPEATPPPVPSTPRPIASSLHGRGSAPAPVLNRQPSLGPTPPPFPGSRAA 221
Query: 106 SFMSSIQQPGTGRGRGRGRGFDPLPPQFENDSVPKKPVFIKREDNVSQTDANDFSPPKNP 165
S++QPG G G PLPP V +KP T + D + P P
Sbjct: 222 GSAGSLRQPGPGPATPYS-GRPPLPPTPGRSPVDEKPPPPPPPSGHRPTASRDMALPPPP 280
Query: 166 VFTRSEDV------------------RPVEPIDLSGDSESD------NRFVMTVPKVLPG 201
V RP P G + SD R + VP P
Sbjct: 281 PQNSKPPVPASPRPPLGVPAPPPPPSRPGPPPVPPGPASSDEMPRLPQRNLSLVPPAAPS 340
Query: 202 GGRGRGKPLEEAAQEAPQAPV 222
G GR PL E P P+
Sbjct: 341 SGSGRSGPLPPPPSERPPPPI 361
>UniRef100_UPI00003222CF UPI00003222CF UniRef100 entry
Length = 310
Score = 54.3 bits (129), Expect = 8e-06
Identities = 40/99 (40%), Positives = 46/99 (46%), Gaps = 14/99 (14%)
Query: 30 GFGGGGGDGRGGGRGRGGSGTVTFNFGEKAAPGNP---NPTPNVNESKPDATDSPIPPGA 86
G GGGGG RGGG G GG F E +P P +P AT PI GA
Sbjct: 203 GGGGGGGSDRGGGGGAGG-------FREDKSPVTPYTASPLDGAGSITVTATGFPITVGA 255
Query: 87 GRGHGRGGTVPDFPSFSFS-SFMSSIQQPGTGRGRGRGR 124
G G GGT P + S S S S+I G G+G G G+
Sbjct: 256 G---GAGGTTPSGAASSGSNSIFSTITAAGGGKGAGAGQ 291
>UniRef100_Q22537 Hypothetical protein T17H7.1 [Caenorhabditis elegans]
Length = 682
Score = 53.5 bits (127), Expect = 1e-05
Identities = 84/289 (29%), Positives = 101/289 (34%), Gaps = 56/289 (19%)
Query: 10 QNSTISNATRQTLVPFSTSSGFGGGGGDGRGGGRGRGGSGTVTFNFGEKAAPGNPNPTPN 69
+NS S++ Q F G GG GG G+G G GG G P
Sbjct: 286 ENSQHSDSNSQ----FDFPRGPGGRGGRGQGPDFGPGGQG-----------GRGQGPDFG 330
Query: 70 VNESKPDATDSPIPPGAGRGHGRGGTVPDF-PSFSFSSFMSSIQQPGTGRGRGRGRGFDP 128
+ P S P G G G G+G PDF P F S PG GRG+G F P
Sbjct: 331 PQDDFPGRRGSGGPGGRG-GRGQG---PDFEPQDDFPGRRGS-GGPGRRGGRGQGPDFGP 385
Query: 129 LPPQFENDSVPKKPVFIKREDNVSQTDANDFSPPKNPVFTRSEDVRPVEPIDLSGDSESD 188
D P + + DF P + + D P + D SG S
Sbjct: 386 ------QDDFPGRRGSGGPGGRGGRGQGPDFGPGRQGGRGQGPDFGPQD--DFSGRRGSG 437
Query: 189 NRFVMTVPKVLPGGGRGRGKPLEEAAQEAPQAPVVNRHIRVRQTPADAESDNVPRRQ--- 245
PGG GRG QE P Q P D+ P R+
Sbjct: 438 G----------PGGRGGRG-------QEPDFGP--GGQGGRGQGPDFGPQDDFPGRRGSG 478
Query: 246 -PMNRFVRDDGDGSGRGRGRGRGRDVYARGR----GDRGRGGRGGRGDG 289
P R R G G G GRG+D + + G RG GG GGRG G
Sbjct: 479 GPEGRDGRGQGPDFGPGSQGGRGQDSDSGSQDAFPGRRGSGGPGGRGQG 527
Score = 50.4 bits (119), Expect = 1e-04
Identities = 77/281 (27%), Positives = 95/281 (33%), Gaps = 43/281 (15%)
Query: 30 GFGGGGGDGRGGGRGRGGSGTVTFNFGEKAAPGNPNPTPNVNESKPD---ATDSPIPPGA 86
G G GG G GGRG+G +F + G P + PD D P G+
Sbjct: 337 GRRGSGGPGGRGGRGQGPDFEPQDDFPGRRGSGGPGRRGGRGQG-PDFGPQDDFPGRRGS 395
Query: 87 GRGHGRGGTV--PDFPSFSFSSFMSSIQQPGTGRGRGRGRGFDPLPPQFENDSVPKKPVF 144
G GRGG PDF PG GRG+G F P D +
Sbjct: 396 GGPGGRGGRGQGPDFG-------------PGRQGGRGQGPDFGP------QDDFSGRRGS 436
Query: 145 IKREDNVSQTDANDFSPPKNPVFTRSEDVRPVEPID----LSGDSESDNRFVMTVPKVLP 200
+ DF P + D P + G D R P P
Sbjct: 437 GGPGGRGGRGQEPDFGPGGQGGRGQGPDFGPQDDFPGRRGSGGPEGRDGRG--QGPDFGP 494
Query: 201 GGGRGRGKPLEEAAQEA-PQAPVVNRHIRVRQTPADAESDNVPRRQ----PMNRFVRDDG 255
G GRG+ + +Q+A P Q P D+ P R+ P R R G
Sbjct: 495 GSQGGRGQDSDSGSQDAFPGRRGSGGPGGRGQGPDFGPQDDFPGRRGSGGPEGRDGRGQG 554
Query: 256 DGSGRGRGRGRGRD-------VYARGRGDRGRGGRGGRGDG 289
G G GRG+D + RG G GG GGRG G
Sbjct: 555 PDFGPGSQGGRGQDSDSGSQDAFPGRRGPGGPGGLGGRGQG 595
Score = 39.7 bits (91), Expect = 0.20
Identities = 65/255 (25%), Positives = 91/255 (35%), Gaps = 54/255 (21%)
Query: 108 MSSIQQPGTGR-------GRGRGRGFDPLPPQFENDSVPKKPVFIKREDNVSQTDANDFS 160
M+ QQ G+ + GRG G GF P ++ + + T+ N FS
Sbjct: 210 MNRFQQTGSQQNFQSRRGGRGDGPGFVPGTQDNNQRGSGERGQRQNFGPSDNLTNGNQFS 269
Query: 161 PPKNPVFTRSEDVRPVEPIDLSGDSESDNRFVMTVPKVLPG-GGRGRGKPLEEAAQEAPQ 219
+ F R + + S S+S+++F P+ G GGRG+G Q
Sbjct: 270 KKQ---FARGPSSMNSDLSENSQHSDSNSQF--DFPRGPGGRGGRGQGPDFGPGGQGGRG 324
Query: 220 APVVNRHIRVRQTPADAESDNVPRRQ----PMNRFVRDDGD---------------GSGR 260
Q P D+ P R+ P R R G G GR
Sbjct: 325 -----------QGPDFGPQDDFPGRRGSGGPGGRGGRGQGPDFEPQDDFPGRRGSGGPGR 373
Query: 261 GRGRGRG-----RDVYARGRGDRGRGGRGGRGD------GRGGFKRYGDDRTSHQDIARS 309
GRG+G +D + RG G GGRGGRG GR G + G D D +
Sbjct: 374 RGGRGQGPDFGPQDDFPGRRGSGGPGGRGGRGQGPDFGPGRQGGRGQGPDFGPQDDFSGR 433
Query: 310 NADGLYVGDNADGEK 324
G G G++
Sbjct: 434 RGSGGPGGRGGRGQE 448
>UniRef100_UPI0000361AF7 UPI0000361AF7 UniRef100 entry
Length = 381
Score = 53.1 bits (126), Expect = 2e-05
Identities = 30/53 (56%), Positives = 31/53 (57%), Gaps = 1/53 (1%)
Query: 249 RFVRDDGDGSGRGRGRGRGRDVYARGRGDRGRGGRGGRGDG-RGGFKRYGDDR 300
RF RD GD S G RGRGR Y G DRGR G G G G RGG+K YG R
Sbjct: 93 RFGRDQGDRSEGGGYRGRGRGGYDSGGYDRGRRGPPGMGGGDRGGYKNYGGSR 145
Score = 38.1 bits (87), Expect = 0.59
Identities = 39/121 (32%), Positives = 49/121 (40%), Gaps = 20/121 (16%)
Query: 14 ISNATRQTLVPFSTSSGFGGGGGD--GRGGGRGR---GGSGTVTFNFGEKAAPGNPNPTP 68
+S ATR+ G GGGGG GRGG RGR GG G +F+ P +
Sbjct: 244 VSFATRRAEFTQRGGGGGGGGGGGRGGRGGFRGRGGGGGGGGPSFDMKGGDWPCPNSSCG 303
Query: 69 NVNESKP---DATDSPIPPGAGRGHGRGGTVPDFPSFSFSSFMSSIQQPGTGRGRGRGRG 125
N+N ++ + +P P AG G G G F F RGRG G
Sbjct: 304 NINFARRHECNKCGAPKPGDAGFGSGDRGGRGGFGGDRGGGF------------RGRGGG 351
Query: 126 F 126
F
Sbjct: 352 F 352
Score = 36.6 bits (83), Expect = 1.7
Identities = 21/45 (46%), Positives = 26/45 (57%), Gaps = 4/45 (8%)
Query: 259 GRGRGRG---RGRDVYARGRGDRGRGGRGGRGDGRGGFKRYGDDR 300
G G+G+G G + R +GDR GG G RG GRGG+ G DR
Sbjct: 79 GGGQGQGGSAEGSGRFGRDQGDRSEGG-GYRGRGRGGYDSGGYDR 122
Score = 35.0 bits (79), Expect = 5.0
Identities = 25/42 (59%), Positives = 25/42 (59%), Gaps = 5/42 (11%)
Query: 253 DDGDGSGRGRGRGR-GRDVYARGRGDRGRGGRGGRGDGRGGF 293
D G GSG GRG G D RG G RGRGG G RG RGGF
Sbjct: 323 DAGFGSGDRGGRGGFGGD---RGGGFRGRGG-GFRGGDRGGF 360
>UniRef100_Q9V9U0 CG2150-PA [Drosophila melanogaster]
Length = 188
Score = 52.4 bits (124), Expect = 3e-05
Identities = 37/101 (36%), Positives = 45/101 (43%), Gaps = 4/101 (3%)
Query: 30 GFGGGGGDGRGGGRGRGGSGTVTFNFGEKAAPG---NPNPTPNVNESKPDATDSPIPPGA 86
GFGGG G G GGG G GG G + G + + G +P V + + G
Sbjct: 40 GFGGGFGGGLGGGGGGGGGGYQAVSGGFQTSEGQNVDPQLLEQVRQILLNEESKQGGGGG 99
Query: 87 GRGHGRGGTVPDFPSFSFSSFMSSIQQPGTGRGRGRGRGFD 127
G G G GG P PS S+ +S PG G G GR G D
Sbjct: 100 GGGGGGGGGYPSGPSSSYGPPSTSYGAPGIG-GGGRVVGID 139
>UniRef100_Q23709 GCR 1 protein [Drosophila melanogaster]
Length = 188
Score = 52.4 bits (124), Expect = 3e-05
Identities = 37/101 (36%), Positives = 45/101 (43%), Gaps = 4/101 (3%)
Query: 30 GFGGGGGDGRGGGRGRGGSGTVTFNFGEKAAPG---NPNPTPNVNESKPDATDSPIPPGA 86
GFGGG G G GGG G GG G + G + + G +P V + + G
Sbjct: 40 GFGGGFGGGLGGGGGGGGGGYQAVSGGFQTSEGQNVDPQLLEQVRQILLNEESKQGGGGG 99
Query: 87 GRGHGRGGTVPDFPSFSFSSFMSSIQQPGTGRGRGRGRGFD 127
G G G GG P PS S+ +S PG G G GR G D
Sbjct: 100 GGGGGGGGGYPSGPSSSYGPPSTSYGAPGIG-GGGRVVGID 139
>UniRef100_Q9UMN6-2 Splice isoform 2 of Q9UMN6 [Homo sapiens]
Length = 582
Score = 52.4 bits (124), Expect = 3e-05
Identities = 82/306 (26%), Positives = 103/306 (32%), Gaps = 77/306 (25%)
Query: 32 GGGGGDGRGG-GRGRGGSGTVTFNFGEKAAPGNPNPTPN------------VNESKPDAT 78
G GGG GRGG G G G PG P + + +
Sbjct: 26 GAGGGGGRGGRGNGAERVRVALRRGGGATGPGGAEPGEDTALLRLLGLRRGLRRLRRLWA 85
Query: 79 DSPIPPGAGRGHGRG-----GTVPD------------FPSFSF------SSFMSSI--QQ 113
+ G GRG GRG G VP+ F F SS S++ Q+
Sbjct: 86 GPRVQRGRGRGRGRGWGPSRGCVPEEESSDGESDEEEFQGFHSDEDVAPSSLRSALRSQR 145
Query: 114 PGTGRGRGRGRGFDPLPPQFENDSVPKKPVFIKREDNVSQTDANDFSPPKNPVFTRSED- 172
RGRGR PLPP D P +PPK P R E+
Sbjct: 146 GRAPRGRGRKHKTTPLPPPRLADVAP--------------------TPPKTPARKRGEEG 185
Query: 173 ----VRPVEPIDLSGDSESDNRFVMTVPKVLPGGGRGR--GKPLEEAAQEAPQAPVVNRH 226
V+ + + + R P P RGR G+P ++ Q VV
Sbjct: 186 TERMVQALTELLRRAQAPQAPRSRACEPST-PRRSRGRPPGRPAGPCRRK--QQAVVVAE 242
Query: 227 IRVRQTPADAESDNVPRRQPMNRFVRDDGDGSGRGRGRGRGRDVYARGRGDRGRGGRGGR 286
V + VP + + +G G G G R RG G RGGRGGR
Sbjct: 243 AAVTIPKPEPPPPVVPVKHQTGSWKCKEGPGPGPGTPR--------RG-GQSSRGGRGGR 293
Query: 287 GDGRGG 292
G GRGG
Sbjct: 294 GRGRGG 299
Score = 35.4 bits (80), Expect = 3.8
Identities = 20/35 (57%), Positives = 21/35 (59%), Gaps = 1/35 (2%)
Query: 255 GDGSGRGRGRGRGRDVYARGRGDRGRGGRGGRGDG 289
G GS G G RGR R RG G GGRGGRG+G
Sbjct: 6 GGGSCPGPGSARGR-FPGRPRGAGGGGGRGGRGNG 39
>UniRef100_Q9UMN6 Myeloid/lymphoid or mixed-lineage leukemia protein 4 [Homo sapiens]
Length = 2715
Score = 52.4 bits (124), Expect = 3e-05
Identities = 82/306 (26%), Positives = 103/306 (32%), Gaps = 77/306 (25%)
Query: 32 GGGGGDGRGG-GRGRGGSGTVTFNFGEKAAPGNPNPTPN------------VNESKPDAT 78
G GGG GRGG G G G PG P + + +
Sbjct: 26 GAGGGGGRGGRGNGAERVRVALRRGGGATGPGGAEPGEDTALLRLLGLRRGLRRLRRLWA 85
Query: 79 DSPIPPGAGRGHGRG-----GTVPD------------FPSFSF------SSFMSSI--QQ 113
+ G GRG GRG G VP+ F F SS S++ Q+
Sbjct: 86 GPRVQRGRGRGRGRGWGPSRGCVPEEESSDGESDEEEFQGFHSDEDVAPSSLRSALRSQR 145
Query: 114 PGTGRGRGRGRGFDPLPPQFENDSVPKKPVFIKREDNVSQTDANDFSPPKNPVFTRSED- 172
RGRGR PLPP D P +PPK P R E+
Sbjct: 146 GRAPRGRGRKHKTTPLPPPRLADVAP--------------------TPPKTPARKRGEEG 185
Query: 173 ----VRPVEPIDLSGDSESDNRFVMTVPKVLPGGGRGR--GKPLEEAAQEAPQAPVVNRH 226
V+ + + + R P P RGR G+P ++ Q VV
Sbjct: 186 TERMVQALTELLRRAQAPQAPRSRACEPST-PRRSRGRPPGRPAGPCRRK--QQAVVVAE 242
Query: 227 IRVRQTPADAESDNVPRRQPMNRFVRDDGDGSGRGRGRGRGRDVYARGRGDRGRGGRGGR 286
V + VP + + +G G G G R RG G RGGRGGR
Sbjct: 243 AAVTIPKPEPPPPVVPVKHQTGSWKCKEGPGPGPGTPR--------RG-GQSSRGGRGGR 293
Query: 287 GDGRGG 292
G GRGG
Sbjct: 294 GRGRGG 299
Score = 35.4 bits (80), Expect = 3.8
Identities = 20/35 (57%), Positives = 21/35 (59%), Gaps = 1/35 (2%)
Query: 255 GDGSGRGRGRGRGRDVYARGRGDRGRGGRGGRGDG 289
G GS G G RGR R RG G GGRGGRG+G
Sbjct: 6 GGGSCPGPGSARGR-FPGRPRGAGGGGGRGGRGNG 39
>UniRef100_Q9IPQ8 EBNA-1 [Cynomolgus Epstein-Barr Virus Si-IIA]
Length = 588
Score = 52.0 bits (123), Expect = 4e-05
Identities = 69/249 (27%), Positives = 87/249 (34%), Gaps = 73/249 (29%)
Query: 29 SGFGGGGGDGRGGGRGRGGSGTVTFNFGEKAAPGNPNPTPNVNESKPDATDSPIPPGAGR 88
SG GG GG G GGGRGRGGS + G + G GR
Sbjct: 228 SGAGGAGGSGGGGGRGRGGSRGRGGSRGRGGSRGRGG-------------------SRGR 268
Query: 89 GHGRGGTVPDFPSFSFSSFMSSIQQPGTGRGRGRGRGFDPLPPQFENDSVPKKPVFIKRE 148
G GRGG+ + G GRGRGRGRG P + + P+ P
Sbjct: 269 GRGRGGS----------------RGRGRGRGRGRGRGQGPR----QGEKRPRSPSGSSSS 308
Query: 149 DNVSQTDANDFSPPKNPVFTRSEDVRPVEPIDLSGDS----ESDNRFVMTVPKVLPGGGR 204
+ S++ ++ R+ LS S D +TVP GR
Sbjct: 309 GSSSRSSSSG----------RASSGGSSSGGTLSNGSFYGFPGDRPLTVTVPG--SALGR 356
Query: 205 GRGKPLEEAAQEAPQAPVVNRHIRVRQTPADAESDNVPRRQPMNRFVRDDGDGSGR---- 260
RG + E P A + Q P E D P P + G G G+
Sbjct: 357 YRGTDGTDGGDEPPGA--------MEQGP---EED--PGEGPSRQHTTSGGRGGGKKGGW 403
Query: 261 -GRGRGRGR 268
GR RG GR
Sbjct: 404 FGRHRGEGR 412
Score = 48.9 bits (115), Expect = 3e-04
Identities = 28/47 (59%), Positives = 30/47 (63%), Gaps = 2/47 (4%)
Query: 255 GDGSGRGRGRGRGRDVYARGR-GDRGRGGRGGRGDGRGGFKRYGDDR 300
G G GRGRG RGR +RGR G RGRGG GRG GRGG + G R
Sbjct: 237 GGGGGRGRGGSRGRG-GSRGRGGSRGRGGSRGRGRGRGGSRGRGRGR 282
Score = 43.1 bits (100), Expect = 0.018
Identities = 26/47 (55%), Positives = 27/47 (57%), Gaps = 4/47 (8%)
Query: 255 GDGSGRGRGRGRGRDVYARGRG-DRGRGGRGGRGDGRGGFKRYGDDR 300
G G RGRG RGR RGRG RGRG GRG GRG R G+ R
Sbjct: 255 GRGGSRGRGGSRGR---GRGRGGSRGRGRGRGRGRGRGQGPRQGEKR 298
Score = 42.4 bits (98), Expect = 0.031
Identities = 24/37 (64%), Positives = 25/37 (66%), Gaps = 2/37 (5%)
Query: 255 GDGSGRGRGRGRGRDVYARGRGDRGRGGRGGRGDGRG 291
G G RGRG RGR +RGRG RGRGG GRG GRG
Sbjct: 249 GRGGSRGRGGSRGRGG-SRGRG-RGRGGSRGRGRGRG 283
Score = 42.0 bits (97), Expect = 0.041
Identities = 27/58 (46%), Positives = 30/58 (51%), Gaps = 4/58 (6%)
Query: 257 GSGRGRGRGRGRDVYARGRG-DRGRGGRGGRGDGRGGFKRYGDDRTSHQDIARSNADG 313
GSG G GRGRG +RGRG RGRGG GRG RG + G R + R G
Sbjct: 235 GSGGGGGRGRGG---SRGRGGSRGRGGSRGRGGSRGRGRGRGGSRGRGRGRGRGRGRG 289
Score = 39.3 bits (90), Expect = 0.27
Identities = 24/49 (48%), Positives = 24/49 (48%), Gaps = 7/49 (14%)
Query: 252 RDDGDGSGRGRGRGRGRDVYARGRGDRGRGGRGGRGDGRGGFKRYGDDR 300
R G G GRGRGRGR RG G G GG G G R GDDR
Sbjct: 41 RGRGGSRGHGRGRGRGR---GRGGGQGGTVASGGSGSG----PRLGDDR 82
Score = 38.9 bits (89), Expect = 0.35
Identities = 31/99 (31%), Positives = 43/99 (43%), Gaps = 5/99 (5%)
Query: 26 STSSGFGGGGGDGRGGGRGRGGSGTVTFNFGEKAAPGNPNPTPNVNESKPDATDSPIPPG 85
S G G GG GRG GRGRG GEK P +P+ + + S ++ G
Sbjct: 265 SRGRGRGRGGSRGRGRGRGRGRGRGQGPRQGEK-RPRSPSGSSSSGSSSRSSSSGRASSG 323
Query: 86 AGRGHGRGGTVPDFPSFSFSSFMS-SIQQPGTGRGRGRG 123
G GGT+ + + F ++ PG+ GR RG
Sbjct: 324 ---GSSSGGTLSNGSFYGFPGDRPLTVTVPGSALGRYRG 359
Score = 37.7 bits (86), Expect = 0.78
Identities = 22/48 (45%), Positives = 25/48 (51%), Gaps = 2/48 (4%)
Query: 255 GDGSGRGRGRGRGRDVYARGRGDRGRGGRGGRGD--GRGGFKRYGDDR 300
G G+G G G G + G G RGRGG GRG GRGG + G R
Sbjct: 219 GSGAGGAGGSGAGGAGGSGGGGGRGRGGSRGRGGSRGRGGSRGRGGSR 266
Score = 35.0 bits (79), Expect = 5.0
Identities = 26/78 (33%), Positives = 31/78 (39%), Gaps = 10/78 (12%)
Query: 27 TSSGFGGGGGDGRGG----------GRGRGGSGTVTFNFGEKAAPGNPNPTPNVNESKPD 76
++SG GGGG GRGG GRGRGG T G + + +P
Sbjct: 31 STSGSGGGGTRGRGGSRGHGRGRGRGRGRGGGQGGTVASGGSGSGPRLGDDRRPDGQRPS 90
Query: 77 ATDSPIPPGAGRGHGRGG 94
S I G G G GG
Sbjct: 91 KRRSCIGCRGGAGGGSGG 108
>UniRef100_UPI00002D1908 UPI00002D1908 UniRef100 entry
Length = 512
Score = 51.6 bits (122), Expect = 5e-05
Identities = 29/57 (50%), Positives = 33/57 (57%), Gaps = 1/57 (1%)
Query: 245 QPMNRFVRDDGDGSGRGRGRGRGRDVYARGRGDRGRGGRGGRGDG-RGGFKRYGDDR 300
+P RF RD G+ S G RGRGR Y G +RGR G G G G RGG+K YG R
Sbjct: 108 RPGPRFGRDQGERSDGGGYRGRGRGGYDSGGYERGRRGPPGMGGGDRGGYKNYGGSR 164
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.313 0.137 0.403
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 995,191,345
Number of Sequences: 2790947
Number of extensions: 55030235
Number of successful extensions: 435834
Number of sequences better than 10.0: 3572
Number of HSP's better than 10.0 without gapping: 1182
Number of HSP's successfully gapped in prelim test: 2625
Number of HSP's that attempted gapping in prelim test: 296184
Number of HSP's gapped (non-prelim): 59154
length of query: 502
length of database: 848,049,833
effective HSP length: 132
effective length of query: 370
effective length of database: 479,644,829
effective search space: 177468586730
effective search space used: 177468586730
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 77 (34.3 bits)
Medicago: description of AC137839.10


