

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137602.4 - phase: 0
(130 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
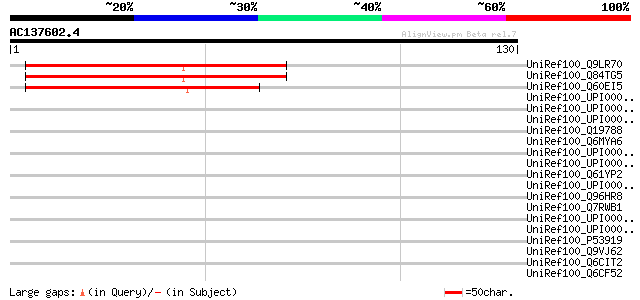
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LR70 F21B7.15 [Arabidopsis thaliana] 83 1e-15
UniRef100_Q84TG5 At1g03530 [Arabidopsis thaliana] 83 1e-15
UniRef100_Q60EI5 Hypothetical protein OSJNBa0017K09.13 [Oryza sa... 78 4e-14
UniRef100_UPI00002354D3 UPI00002354D3 UniRef100 entry 39 0.026
UniRef100_UPI00003AD62B UPI00003AD62B UniRef100 entry 38 0.058
UniRef100_UPI0000429316 UPI0000429316 UniRef100 entry 37 0.076
UniRef100_Q19788 Hypothetical protein F25H8.2 [Caenorhabditis el... 37 0.099
UniRef100_Q6MYA6 Hypothetical protein [Aspergillus fumigatus] 37 0.099
UniRef100_UPI000021F3E4 UPI000021F3E4 UniRef100 entry 37 0.13
UniRef100_UPI00001D0F80 UPI00001D0F80 UniRef100 entry 37 0.13
UniRef100_Q61YP2 Hypothetical protein CBG03454 [Caenorhabditis b... 37 0.13
UniRef100_UPI000036CF03 UPI000036CF03 UniRef100 entry 36 0.17
UniRef100_Q96HR8 Hypothetical protein LOC92345 [Homo sapiens] 36 0.17
UniRef100_Q7RWB1 Hypothetical protein [Neurospora crassa] 36 0.17
UniRef100_UPI000042D31A UPI000042D31A UniRef100 entry 35 0.38
UniRef100_UPI000042D317 UPI000042D317 UniRef100 entry 35 0.38
UniRef100_P53919 Hypothetical 54.9 kDa protein in SPC98-TOM70 in... 35 0.49
UniRef100_Q9VJ62 CG10341-PA [Drosophila melanogaster] 34 0.64
UniRef100_Q6CIT2 Similarities with sp|P53919 Saccharomyces cerev... 34 0.84
UniRef100_Q6CF52 Yarrowia lipolytica chromosome B of strain CLIB... 33 1.1
>UniRef100_Q9LR70 F21B7.15 [Arabidopsis thaliana]
Length = 801
Score = 83.2 bits (204), Expect = 1e-15
Identities = 39/71 (54%), Positives = 54/71 (75%), Gaps = 4/71 (5%)
Query: 5 PLIEGSILWITERQTSLGLIDEIFGQVKNPYYAVRYNSE----KGIREGTLISFVAEFVN 60
PL EGSILWITE++T LGL+DEIFG VK PYY VR+NSE +G+ +GT +SFVA+F
Sbjct: 405 PLTEGSILWITEKRTPLGLVDEIFGPVKCPYYIVRFNSESEVPEGVCQGTPVSFVADFAQ 464
Query: 61 YVMHPVSMMKR 71
++++ + K+
Sbjct: 465 HILNIKELQKK 475
>UniRef100_Q84TG5 At1g03530 [Arabidopsis thaliana]
Length = 801
Score = 83.2 bits (204), Expect = 1e-15
Identities = 39/71 (54%), Positives = 54/71 (75%), Gaps = 4/71 (5%)
Query: 5 PLIEGSILWITERQTSLGLIDEIFGQVKNPYYAVRYNSE----KGIREGTLISFVAEFVN 60
PL EGSILWITE++T LGL+DEIFG VK PYY VR+NSE +G+ +GT +SFVA+F
Sbjct: 405 PLTEGSILWITEKRTPLGLVDEIFGPVKCPYYIVRFNSESEVPEGVCQGTPVSFVADFAQ 464
Query: 61 YVMHPVSMMKR 71
++++ + K+
Sbjct: 465 HILNIKELQKK 475
>UniRef100_Q60EI5 Hypothetical protein OSJNBa0017K09.13 [Oryza sativa]
Length = 681
Score = 78.2 bits (191), Expect = 4e-14
Identities = 37/64 (57%), Positives = 48/64 (74%), Gaps = 4/64 (6%)
Query: 5 PLIEGSILWITERQTSLGLIDEIFGQVKNPYYAVRYNSEK----GIREGTLISFVAEFVN 60
PL EGSILWITE + LG++DE+FG VKNPYY VRYNS + I GT +SFVAEF +
Sbjct: 291 PLNEGSILWITESRIPLGIVDELFGPVKNPYYLVRYNSAEEVPADISAGTAVSFVAEFAD 350
Query: 61 YVMH 64
++++
Sbjct: 351 HILN 354
>UniRef100_UPI00002354D3 UPI00002354D3 UniRef100 entry
Length = 680
Score = 38.9 bits (89), Expect = 0.026
Identities = 19/59 (32%), Positives = 37/59 (62%), Gaps = 6/59 (10%)
Query: 9 GSILWITERQTSLGLIDEIFGQVKNPYYAVRYNS-----EKGIREGTLISFVAEFVNYV 62
GS+L + +R+ + G++ E G+V+NP YAVR+ + + G+ GT++ +V + +V
Sbjct: 318 GSLLCLEDRRVA-GVVSETLGRVENPLYAVRFATTADVEKHGLSRGTVVYYVVDHSTFV 375
>UniRef100_UPI00003AD62B UPI00003AD62B UniRef100 entry
Length = 482
Score = 37.7 bits (86), Expect = 0.058
Identities = 17/43 (39%), Positives = 31/43 (71%), Gaps = 1/43 (2%)
Query: 5 PLIEGSILWITERQTSLGLIDEIFGQVKNPYYAVRYNSEKGIR 47
P+ E SI++ +RQ + G + E+FG V++P+Y +R+NS + I+
Sbjct: 220 PINEESIIFKEDRQAA-GKVFEVFGPVQHPFYVLRFNSAEHIK 261
>UniRef100_UPI0000429316 UPI0000429316 UniRef100 entry
Length = 535
Score = 37.4 bits (85), Expect = 0.076
Identities = 22/66 (33%), Positives = 39/66 (58%), Gaps = 9/66 (13%)
Query: 5 PLIEGSILWITERQTSLGLIDEIFGQVKNPYYAVRYNS-----EKGIREGTLISF---VA 56
P+ E ++++ ++RQ + G I EIFG V +P+Y +R+NS KGI+ + F +
Sbjct: 268 PVNEDTVIFKSDRQAA-GKIFEIFGPVAHPFYVLRFNSSDHIESKGIKINDTMYFAPSMK 326
Query: 57 EFVNYV 62
+F Y+
Sbjct: 327 DFTQYI 332
>UniRef100_Q19788 Hypothetical protein F25H8.2 [Caenorhabditis elegans]
Length = 390
Score = 37.0 bits (84), Expect = 0.099
Identities = 15/27 (55%), Positives = 21/27 (77%)
Query: 16 ERQTSLGLIDEIFGQVKNPYYAVRYNS 42
++ +LG I +IFGQVKNP Y +R+NS
Sbjct: 198 QKGNALGQIYDIFGQVKNPQYVIRFNS 224
>UniRef100_Q6MYA6 Hypothetical protein [Aspergillus fumigatus]
Length = 671
Score = 37.0 bits (84), Expect = 0.099
Identities = 21/62 (33%), Positives = 36/62 (57%), Gaps = 6/62 (9%)
Query: 6 LIEGSILWITERQTSLGLIDEIFGQVKNPYYAVRYNS-----EKGIREGTLISFVAEFVN 60
L GS+L + +R T G++ E G+V+ P YAVR+ + E+G+ +G + +V E
Sbjct: 316 LESGSLLCLEDR-TVAGVVSETLGRVEKPLYAVRFPNAAAIEERGLSKGKNVYYVEEHST 374
Query: 61 YV 62
+V
Sbjct: 375 FV 376
>UniRef100_UPI000021F3E4 UPI000021F3E4 UniRef100 entry
Length = 243
Score = 36.6 bits (83), Expect = 0.13
Identities = 17/42 (40%), Positives = 30/42 (70%), Gaps = 1/42 (2%)
Query: 5 PLIEGSILWITERQTSLGLIDEIFGQVKNPYYAVRYNSEKGI 46
P+ E ++++ ++RQ + G I EIFG V +P+Y +R+NS + I
Sbjct: 42 PVNEDTVIFKSDRQAA-GKIFEIFGPVAHPFYVLRFNSSEHI 82
>UniRef100_UPI00001D0F80 UPI00001D0F80 UniRef100 entry
Length = 509
Score = 36.6 bits (83), Expect = 0.13
Identities = 17/42 (40%), Positives = 30/42 (70%), Gaps = 1/42 (2%)
Query: 5 PLIEGSILWITERQTSLGLIDEIFGQVKNPYYAVRYNSEKGI 46
P+ E ++++ ++RQ + G I EIFG V +P+Y +R+NS + I
Sbjct: 262 PVNEDTVIFKSDRQAA-GKIFEIFGPVAHPFYVLRFNSSEHI 302
>UniRef100_Q61YP2 Hypothetical protein CBG03454 [Caenorhabditis briggsae]
Length = 405
Score = 36.6 bits (83), Expect = 0.13
Identities = 13/29 (44%), Positives = 23/29 (78%)
Query: 16 ERQTSLGLIDEIFGQVKNPYYAVRYNSEK 44
E++ ++G + +IFGQVK+P Y +R+NS +
Sbjct: 198 EKRNAIGQVYDIFGQVKSPQYVIRFNSSE 226
>UniRef100_UPI000036CF03 UPI000036CF03 UniRef100 entry
Length = 495
Score = 36.2 bits (82), Expect = 0.17
Identities = 22/66 (33%), Positives = 39/66 (58%), Gaps = 9/66 (13%)
Query: 5 PLIEGSILWITERQTSLGLIDEIFGQVKNPYYAVRYNS-----EKGIREGTLISF---VA 56
P+ E ++++ ++RQ + G I EIFG V +P+Y +R+NS KGI+ + F +
Sbjct: 222 PVNEETVIFKSDRQAA-GKIFEIFGPVAHPFYVLRFNSSDHIESKGIKIKETMYFAPSMK 280
Query: 57 EFVNYV 62
+F Y+
Sbjct: 281 DFTQYI 286
>UniRef100_Q96HR8 Hypothetical protein LOC92345 [Homo sapiens]
Length = 494
Score = 36.2 bits (82), Expect = 0.17
Identities = 22/66 (33%), Positives = 39/66 (58%), Gaps = 9/66 (13%)
Query: 5 PLIEGSILWITERQTSLGLIDEIFGQVKNPYYAVRYNS-----EKGIREGTLISF---VA 56
P+ E ++++ ++RQ + G I EIFG V +P+Y +R+NS KGI+ + F +
Sbjct: 222 PVNEETVIFKSDRQAA-GKIFEIFGPVAHPFYVLRFNSSDHIESKGIKIKETMYFAPSMK 280
Query: 57 EFVNYV 62
+F Y+
Sbjct: 281 DFTQYI 286
>UniRef100_Q7RWB1 Hypothetical protein [Neurospora crassa]
Length = 701
Score = 36.2 bits (82), Expect = 0.17
Identities = 22/59 (37%), Positives = 32/59 (53%), Gaps = 6/59 (10%)
Query: 9 GSILWITERQTSLGLIDEIFGQVKNPYYAVRYNSEKGIRE-----GTLISFVAEFVNYV 62
GS+L E +T +G + EI G VK+P Y + ++ I+E GT I + E NYV
Sbjct: 271 GSVL-CKEDRTVIGALAEIIGNVKSPLYTCAFANQDEIKELGLEVGTQIFYSVEHANYV 328
>UniRef100_UPI000042D31A UPI000042D31A UniRef100 entry
Length = 612
Score = 35.0 bits (79), Expect = 0.38
Identities = 14/39 (35%), Positives = 28/39 (70%)
Query: 6 LIEGSILWITERQTSLGLIDEIFGQVKNPYYAVRYNSEK 44
+++ ++ E +T +G + EIFG+V+ P Y+V++NSE+
Sbjct: 320 ILQEKSIFCFEDRTVIGPLFEIFGRVQQPVYSVKFNSEE 358
>UniRef100_UPI000042D317 UPI000042D317 UniRef100 entry
Length = 612
Score = 35.0 bits (79), Expect = 0.38
Identities = 14/39 (35%), Positives = 28/39 (70%)
Query: 6 LIEGSILWITERQTSLGLIDEIFGQVKNPYYAVRYNSEK 44
+++ ++ E +T +G + EIFG+V+ P Y+V++NSE+
Sbjct: 320 ILQEKSIFCFEDRTVIGPLFEIFGRVQQPVYSVKFNSEE 358
>UniRef100_P53919 Hypothetical 54.9 kDa protein in SPC98-TOM70 intergenic region
[Saccharomyces cerevisiae]
Length = 492
Score = 34.7 bits (78), Expect = 0.49
Identities = 15/39 (38%), Positives = 26/39 (66%), Gaps = 1/39 (2%)
Query: 6 LIEGSILWITERQTSLGLIDEIFGQVKNPYYAVRYNSEK 44
L EGSI + +R T +G++ E+FG ++NP+Y ++ K
Sbjct: 164 LKEGSIFCLEDR-TLIGMLTEVFGPLQNPFYRIKLPDSK 201
>UniRef100_Q9VJ62 CG10341-PA [Drosophila melanogaster]
Length = 564
Score = 34.3 bits (77), Expect = 0.64
Identities = 14/37 (37%), Positives = 25/37 (66%)
Query: 10 SILWITERQTSLGLIDEIFGQVKNPYYAVRYNSEKGI 46
++L++ + + LG + ++ GQV +P Y VR+NS K I
Sbjct: 315 TVLFLEKGRKVLGEVFDVLGQVSDPLYCVRFNSNKQI 351
>UniRef100_Q6CIT2 Similarities with sp|P53919 Saccharomyces cerevisiae YNL124w
singleton [Kluyveromyces lactis]
Length = 485
Score = 33.9 bits (76), Expect = 0.84
Identities = 17/41 (41%), Positives = 28/41 (67%), Gaps = 1/41 (2%)
Query: 6 LIEGSILWITERQTSLGLIDEIFGQVKNPYYAVRYNSEKGI 46
L EGSIL + E+ T +G + E+FG ++ P+Y V ++ +K I
Sbjct: 164 LKEGSILCLQEK-TLVGPVCEVFGPLQAPFYRVSFDKDKEI 203
>UniRef100_Q6CF52 Yarrowia lipolytica chromosome B of strain CLIB99 of Yarrowia
lipolytica [Yarrowia lipolytica]
Length = 723
Score = 33.5 bits (75), Expect = 1.1
Identities = 19/52 (36%), Positives = 34/52 (64%), Gaps = 5/52 (9%)
Query: 16 ERQTSLGLIDEIFGQVKNPYYAVRYNSEK---GIRE--GTLISFVAEFVNYV 62
E +T +G++ E FG VK P+Y+VR +S++ ++E G+ I +VA N++
Sbjct: 434 EDKTIVGVLFETFGLVKMPFYSVRVSSDEEAAALKEKVGSPIYYVAGHANFL 485
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.322 0.138 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 213,893,012
Number of Sequences: 2790947
Number of extensions: 7602223
Number of successful extensions: 17754
Number of sequences better than 10.0: 72
Number of HSP's better than 10.0 without gapping: 22
Number of HSP's successfully gapped in prelim test: 50
Number of HSP's that attempted gapping in prelim test: 17727
Number of HSP's gapped (non-prelim): 73
length of query: 130
length of database: 848,049,833
effective HSP length: 106
effective length of query: 24
effective length of database: 552,209,451
effective search space: 13253026824
effective search space used: 13253026824
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 67 (30.4 bits)
Medicago: description of AC137602.4


