

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136472.7 + phase: 0
(123 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
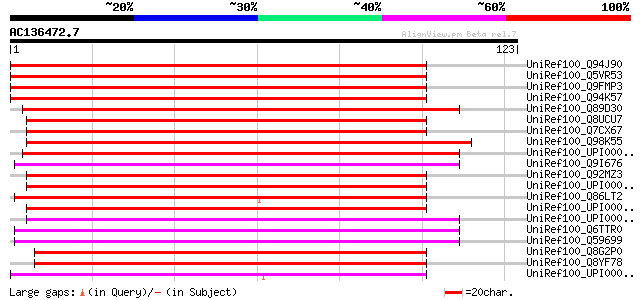
UniRef100_Q94J90 Putative dihydropyrimidine amidohydrolase [Oryz... 176 8e-44
UniRef100_Q5VR53 Putative dihydropyrimidine amidohydrolase [Oryz... 176 8e-44
UniRef100_Q9FMP3 Dihydropyrimidinase [Arabidopsis thaliana] 174 4e-43
UniRef100_Q94K57 Putative dihydropyrimidinase [Arabidopsis thali... 174 4e-43
UniRef100_Q89D30 Dihydropyrimidinase [Bradyrhizobium japonicum] 105 2e-22
UniRef100_Q8UCU7 Dihydropyrimidinase [Agrobacterium tumefaciens] 104 5e-22
UniRef100_Q7CX67 AGR_C_4328p [Agrobacterium tumefaciens] 104 5e-22
UniRef100_Q98K55 Dihydropyrimidinase [Rhizobium loti] 102 2e-21
UniRef100_UPI000032F29A UPI000032F29A UniRef100 entry 101 3e-21
UniRef100_Q9I676 Dihydropyrimidinase [Pseudomonas aeruginosa] 101 4e-21
UniRef100_Q92MZ3 PUTATIVE D-HYDANTOINASE (DIHYDROPYRIMIDINASE) P... 100 8e-21
UniRef100_UPI0000301EAB UPI0000301EAB UniRef100 entry 100 1e-20
UniRef100_Q86LT2 Dihydropyrimidine amidohydrolase [Dictyostelium... 99 2e-20
UniRef100_UPI0000320D5E UPI0000320D5E UniRef100 entry 98 4e-20
UniRef100_UPI00002EBB34 UPI00002EBB34 UniRef100 entry 97 6e-20
UniRef100_Q6TTR0 D-hydantoinase [Pseudomonas putida] 97 1e-19
UniRef100_Q59699 D-hydantoinase [Pseudomonas putida] 96 2e-19
UniRef100_Q8G2P0 D-hydantoinase [Brucella suis] 95 3e-19
UniRef100_Q8YF78 D-HYDANTOINASE [Brucella melitensis] 94 7e-19
UniRef100_UPI0000363DC9 UPI0000363DC9 UniRef100 entry 94 9e-19
>UniRef100_Q94J90 Putative dihydropyrimidine amidohydrolase [Oryza sativa]
Length = 539
Score = 176 bits (447), Expect = 8e-44
Identities = 82/101 (81%), Positives = 92/101 (90%)
Query: 1 MSIDAMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKA 60
MSIDAM+E+AKA++ GQRVIGEPV+SGL LDDSWLW PDF A+KYVMSPPIR+ GH+KA
Sbjct: 293 MSIDAMDEIAKAKREGQRVIGEPVVSGLVLDDSWLWDPDFMIASKYVMSPPIREAGHNKA 352
Query: 61 LQAALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNG 101
LQ ALS+G+LQLVGTDHC FNSTQKAFG DDFRKIPNGVNG
Sbjct: 353 LQVALSSGILQLVGTDHCTFNSTQKAFGSDDFRKIPNGVNG 393
>UniRef100_Q5VR53 Putative dihydropyrimidine amidohydrolase [Oryza sativa]
Length = 511
Score = 176 bits (447), Expect = 8e-44
Identities = 82/101 (81%), Positives = 92/101 (90%)
Query: 1 MSIDAMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKA 60
MSIDAM+E+AKA++ GQRVIGEPV+SGL LDDSWLW PDF A+KYVMSPPIR+ GH+KA
Sbjct: 265 MSIDAMDEIAKAKREGQRVIGEPVVSGLVLDDSWLWDPDFMIASKYVMSPPIREAGHNKA 324
Query: 61 LQAALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNG 101
LQ ALS+G+LQLVGTDHC FNSTQKAFG DDFRKIPNGVNG
Sbjct: 325 LQVALSSGILQLVGTDHCTFNSTQKAFGSDDFRKIPNGVNG 365
>UniRef100_Q9FMP3 Dihydropyrimidinase [Arabidopsis thaliana]
Length = 531
Score = 174 bits (441), Expect = 4e-43
Identities = 81/101 (80%), Positives = 90/101 (88%)
Query: 1 MSIDAMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKA 60
MS+DAM+E+AKARKSGQ+VIGEPV+SGL LDD WLW PDF A+KYVMSPPIR GH KA
Sbjct: 284 MSVDAMDEIAKARKSGQKVIGEPVVSGLILDDHWLWDPDFTIASKYVMSPPIRPVGHGKA 343
Query: 61 LQAALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNG 101
LQ ALSTG+LQLVGTDHC FNSTQKA G+DDFR+IPNGVNG
Sbjct: 344 LQDALSTGILQLVGTDHCTFNSTQKALGLDDFRRIPNGVNG 384
>UniRef100_Q94K57 Putative dihydropyrimidinase [Arabidopsis thaliana]
Length = 248
Score = 174 bits (441), Expect = 4e-43
Identities = 81/101 (80%), Positives = 90/101 (88%)
Query: 1 MSIDAMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKA 60
MS+DAM+E+AKARKSGQ+VIGEPV+SGL LDD WLW PDF A+KYVMSPPIR GH KA
Sbjct: 1 MSVDAMDEIAKARKSGQKVIGEPVVSGLILDDHWLWDPDFTIASKYVMSPPIRPVGHGKA 60
Query: 61 LQAALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNG 101
LQ ALSTG+LQLVGTDHC FNSTQKA G+DDFR+IPNGVNG
Sbjct: 61 LQDALSTGILQLVGTDHCTFNSTQKALGLDDFRRIPNGVNG 101
>UniRef100_Q89D30 Dihydropyrimidinase [Bradyrhizobium japonicum]
Length = 484
Score = 105 bits (262), Expect = 2e-22
Identities = 52/106 (49%), Positives = 68/106 (64%)
Query: 4 DAMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKALQA 63
+A E +A+AR +G+RV GEP++ L LD H D+D +A+ VMSPP R + H +L A
Sbjct: 244 EAHEAIARARAAGKRVYGEPLIQHLLLDAREYQHKDWDHSAQRVMSPPFRDKQHQDSLWA 303
Query: 64 ALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNGDHSNLRYL 109
L +G LQ+V TDHC F + QK FG+ DFRKIPNG G L L
Sbjct: 304 GLQSGSLQVVATDHCAFTTEQKRFGLGDFRKIPNGTGGLEDRLALL 349
>UniRef100_Q8UCU7 Dihydropyrimidinase [Agrobacterium tumefaciens]
Length = 485
Score = 104 bits (259), Expect = 5e-22
Identities = 50/97 (51%), Positives = 66/97 (67%)
Query: 5 AMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKALQAA 64
A E + +AR++G RV GEP++ L LD+S + D+D AA+ VMSPP R + H +L A
Sbjct: 243 AHEAIRRARQNGMRVYGEPLIQHLILDESEYANADWDHAARRVMSPPFRNRQHQDSLWAG 302
Query: 65 LSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNG 101
L++G LQ V TDHC F + QK FG+ DFRKIPNG G
Sbjct: 303 LASGSLQCVATDHCAFTTEQKRFGLGDFRKIPNGTGG 339
>UniRef100_Q7CX67 AGR_C_4328p [Agrobacterium tumefaciens]
Length = 506
Score = 104 bits (259), Expect = 5e-22
Identities = 50/97 (51%), Positives = 66/97 (67%)
Query: 5 AMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKALQAA 64
A E + +AR++G RV GEP++ L LD+S + D+D AA+ VMSPP R + H +L A
Sbjct: 264 AHEAIRRARQNGMRVYGEPLIQHLILDESEYANADWDHAARRVMSPPFRNRQHQDSLWAG 323
Query: 65 LSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNG 101
L++G LQ V TDHC F + QK FG+ DFRKIPNG G
Sbjct: 324 LASGSLQCVATDHCAFTTEQKRFGLGDFRKIPNGTGG 360
>UniRef100_Q98K55 Dihydropyrimidinase [Rhizobium loti]
Length = 483
Score = 102 bits (255), Expect = 2e-21
Identities = 51/108 (47%), Positives = 67/108 (61%)
Query: 5 AMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKALQAA 64
A E + +AR+ G RV GEP++ L LD++ ++ D+D AA+ VMSPP R + H +L A
Sbjct: 241 AHEAIRRARQKGMRVFGEPLVQHLTLDETEYFNKDWDHAARRVMSPPFRNKLHQDSLWAG 300
Query: 65 LSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNGDHSNLRYLRPK 112
L G LQ+V TDHC F + QK FG+ DF KIPNG G L L K
Sbjct: 301 LQAGSLQVVATDHCAFTTKQKRFGVGDFTKIPNGTGGLEDRLPVLWTK 348
>UniRef100_UPI000032F29A UPI000032F29A UniRef100 entry
Length = 484
Score = 101 bits (252), Expect = 3e-21
Identities = 49/106 (46%), Positives = 68/106 (63%)
Query: 4 DAMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKALQA 63
++ E + +AR++G+RV GEP++ L LD+S ++ D+D AA+ VMSPP R + H +L A
Sbjct: 241 ESHEAIRRARQAGKRVWGEPLIQHLTLDESEYFNKDWDHAARRVMSPPFRNKMHQDSLWA 300
Query: 64 ALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNGDHSNLRYL 109
L G L +V TDHC F + QK FG+ DF KIPNG G L L
Sbjct: 301 GLQAGSLSVVATDHCAFTTEQKRFGVGDFTKIPNGTGGLEDRLPML 346
>UniRef100_Q9I676 Dihydropyrimidinase [Pseudomonas aeruginosa]
Length = 479
Score = 101 bits (251), Expect = 4e-21
Identities = 54/108 (50%), Positives = 64/108 (59%)
Query: 2 SIDAMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKAL 61
S +A++E+A AR GQ V GE + L LDDS HPD+ TAA YVMSPP R H +AL
Sbjct: 242 SREALDEIAYARAKGQPVYGEVLAGHLLLDDSVYRHPDWATAAGYVMSPPFRPVEHQEAL 301
Query: 62 QAALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNGDHSNLRYL 109
L +G L TDHC F + QKA G DDF KIPNG G + L
Sbjct: 302 WRGLQSGNLHTTATDHCCFCAEQKAMGRDDFSKIPNGTAGIEDRMALL 349
>UniRef100_Q92MZ3 PUTATIVE D-HYDANTOINASE (DIHYDROPYRIMIDINASE) PROTEIN [Rhizobium
meliloti]
Length = 484
Score = 100 bits (249), Expect = 8e-21
Identities = 47/97 (48%), Positives = 64/97 (65%)
Query: 5 AMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKALQAA 64
A E + +AR G RV GEP++ L LD++ + D+D AA+ VMSPP R + H +L A
Sbjct: 242 AHEAIRRARAKGMRVFGEPLIQHLTLDETEYFDKDWDHAARRVMSPPFRNKLHQDSLWAG 301
Query: 65 LSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNG 101
L++G LQ+V TDHC F + QK FG+ DF +IPNG G
Sbjct: 302 LASGSLQVVATDHCAFTTEQKRFGVGDFTRIPNGTGG 338
>UniRef100_UPI0000301EAB UPI0000301EAB UniRef100 entry
Length = 365
Score = 99.8 bits (247), Expect = 1e-20
Identities = 47/97 (48%), Positives = 65/97 (66%)
Query: 5 AMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKALQAA 64
A E + +AR+ G RV GEP++ L LD+S ++ D++ AA+ VMSPP R + H L A
Sbjct: 242 AHEAIRRARQKGMRVFGEPLIQFLTLDESEYFNKDWEHAARRVMSPPFRDKSHQDGLWAG 301
Query: 65 LSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNG 101
L +G LQ+V TDH F++ QK G++DF KIPNG NG
Sbjct: 302 LQSGSLQVVATDHAAFSTEQKRMGVNDFTKIPNGSNG 338
>UniRef100_Q86LT2 Dihydropyrimidine amidohydrolase [Dictyostelium discoideum]
Length = 503
Score = 99.0 bits (245), Expect = 2e-20
Identities = 52/101 (51%), Positives = 64/101 (62%), Gaps = 1/101 (0%)
Query: 2 SIDAMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDK-A 60
SI A + + K RK G RV GEP+ +GL +D S +W+ D+ AA +VM PPIR K
Sbjct: 250 SIGAADVICKHRKEGVRVYGEPIAAGLGVDGSHMWNHDWRHAAAFVMGPPIRPDPRTKGV 309
Query: 61 LQAALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNG 101
L L+ G L VGTD+C F + QKA G DDF KIPNGVNG
Sbjct: 310 LMDYLARGDLDCVGTDNCTFCADQKAMGKDDFTKIPNGVNG 350
>UniRef100_UPI0000320D5E UPI0000320D5E UniRef100 entry
Length = 485
Score = 98.2 bits (243), Expect = 4e-20
Identities = 46/97 (47%), Positives = 64/97 (65%)
Query: 5 AMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKALQAA 64
A E + +AR++G+RV GEP++ L LD+ + D+D AA+ VMSPP R + H +L +
Sbjct: 242 AHEAIRRARQNGKRVFGEPLIQHLVLDEGEYQNKDWDHAARRVMSPPFRNKLHQDSLWSG 301
Query: 65 LSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNG 101
L+ G LQ+V TDHC F + QK G+ DF KIPNG G
Sbjct: 302 LAAGSLQVVATDHCAFTTEQKRMGLGDFTKIPNGTGG 338
>UniRef100_UPI00002EBB34 UPI00002EBB34 UniRef100 entry
Length = 297
Score = 97.4 bits (241), Expect = 6e-20
Identities = 50/105 (47%), Positives = 63/105 (59%)
Query: 5 AMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKALQAA 64
A E + +AR+ G RV GEP++ L LD+S ++ D+ AA+ VMSPP R + H L A
Sbjct: 107 AHEAIRRARQKGIRVYGEPLIQFLTLDESEYFNKDWAHAARRVMSPPFRAKEHQDGLWAG 166
Query: 65 LSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNGDHSNLRYL 109
L G LQ+V TDH F S QK G+DDF KIPNG G L L
Sbjct: 167 LQAGSLQVVATDHAAFTSDQKKMGVDDFTKIPNGTGGLEERLAVL 211
>UniRef100_Q6TTR0 D-hydantoinase [Pseudomonas putida]
Length = 479
Score = 96.7 bits (239), Expect = 1e-19
Identities = 52/108 (48%), Positives = 64/108 (59%)
Query: 2 SIDAMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKAL 61
S +A++E+A AR GQ V GE + L LDDS PD+ TAA YVMSPP R + H +AL
Sbjct: 242 SREALDEIAYARGKGQPVYGEVLPGHLLLDDSVYRDPDWATAAGYVMSPPFRPREHQEAL 301
Query: 62 QAALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNGDHSNLRYL 109
L +G L TDHC F + QKA G DDF +IPNG G + L
Sbjct: 302 WRGLQSGNLHTTATDHCCFCAEQKAMGRDDFSRIPNGTAGIEDRMAVL 349
>UniRef100_Q59699 D-hydantoinase [Pseudomonas putida]
Length = 495
Score = 95.5 bits (236), Expect = 2e-19
Identities = 51/108 (47%), Positives = 63/108 (58%)
Query: 2 SIDAMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKAL 61
S +A++E+ AR GQ V GE + L LDDS PD+ TAA YVMSPP R + H +AL
Sbjct: 242 SREALDEITYARAKGQPVYGEVLPGHLLLDDSVYRDPDWATAAGYVMSPPFRPREHQEAL 301
Query: 62 QAALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNGDHSNLRYL 109
L +G L TDHC F + QKA G DDF +IPNG G + L
Sbjct: 302 WRGLQSGNLHTTATDHCCFCAEQKAMGRDDFSRIPNGTAGIEDRMAVL 349
>UniRef100_Q8G2P0 D-hydantoinase [Brucella suis]
Length = 489
Score = 95.1 bits (235), Expect = 3e-19
Identities = 46/95 (48%), Positives = 62/95 (64%)
Query: 7 EEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKALQAALS 66
E + +AR+ G RV GEP++ L LD+S + D+D AA+ VMSPP R + + +L A L+
Sbjct: 248 EAIRRARQKGMRVFGEPLIQHLTLDESEYHNRDWDYAARRVMSPPFRDKLNQDSLWAGLA 307
Query: 67 TGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNG 101
G LQ V TDHC F + QK +GI +F KIPNG G
Sbjct: 308 AGSLQCVATDHCAFTTEQKRYGIGNFTKIPNGTGG 342
>UniRef100_Q8YF78 D-HYDANTOINASE [Brucella melitensis]
Length = 489
Score = 94.0 bits (232), Expect = 7e-19
Identities = 46/95 (48%), Positives = 62/95 (64%)
Query: 7 EEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKALQAALS 66
E + +AR+ G RV GEP++ L LD+S + D+D AA+ VMSPP R + + +L A L+
Sbjct: 248 EAIRRARQKGIRVFGEPLIQHLTLDESEYHNRDWDYAARRVMSPPFRDKLNQDSLWAGLA 307
Query: 67 TGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNG 101
G LQ V TDHC F + QK +GI +F KIPNG G
Sbjct: 308 AGSLQCVATDHCAFTTEQKRYGIGNFTKIPNGTGG 342
>UniRef100_UPI0000363DC9 UPI0000363DC9 UniRef100 entry
Length = 479
Score = 93.6 bits (231), Expect = 9e-19
Identities = 50/102 (49%), Positives = 60/102 (58%), Gaps = 1/102 (0%)
Query: 1 MSIDAMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKA 60
MS A V+KARK G V GEP+ + L D + WH D+ AAKYVM PP+R
Sbjct: 239 MSKSAANVVSKARKDGNVVFGEPIAASLGTDGTNYWHKDWAHAAKYVMGPPLRPDPSTPG 298
Query: 61 -LQAALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNG 101
L L+ L + GTD+C F+ QKA G DDF KIPNGVNG
Sbjct: 299 YLMDLLANDDLTVTGTDNCTFSVCQKALGKDDFTKIPNGVNG 340
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.320 0.137 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 207,444,988
Number of Sequences: 2790947
Number of extensions: 7641495
Number of successful extensions: 17634
Number of sequences better than 10.0: 415
Number of HSP's better than 10.0 without gapping: 275
Number of HSP's successfully gapped in prelim test: 140
Number of HSP's that attempted gapping in prelim test: 17225
Number of HSP's gapped (non-prelim): 415
length of query: 123
length of database: 848,049,833
effective HSP length: 99
effective length of query: 24
effective length of database: 571,746,080
effective search space: 13721905920
effective search space used: 13721905920
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 67 (30.4 bits)
Medicago: description of AC136472.7


