

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC125473.10 + phase: 0
(257 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
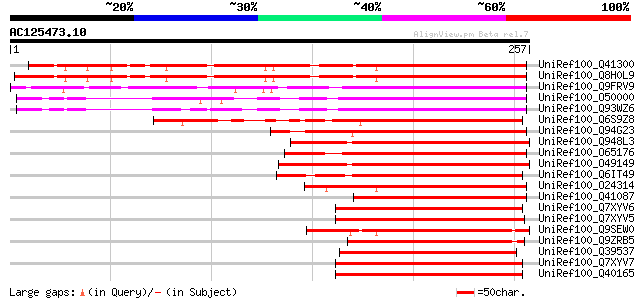
Sequences producing significant alignments: (bits) Value
UniRef100_Q41300 Abscisic stress ripening protein [Solanum chaco... 215 1e-54
UniRef100_Q8H0L9 DS2 protein [Solanum tuberosum] 213 3e-54
UniRef100_Q9FRV9 Putative ripening protein [Calystegia soldanella] 201 2e-50
UniRef100_O50000 Abscisic stress ripening protein homolog [Prunu... 198 1e-49
UniRef100_Q93WZ6 Abscisic stress ripening-like protein [Prunus p... 194 1e-48
UniRef100_Q6S9Z8 ASR protein [Ginkgo biloba] 178 1e-43
UniRef100_Q94G23 Putative transcription factor [Vitis vinifera] 158 1e-37
UniRef100_Q948L3 Drought inducible 22 kD protein [Saccharum offi... 156 6e-37
UniRef100_O65176 ABA stress ripening protein [Mesembryanthemum c... 148 2e-34
UniRef100_O49149 Abscisic acid- and stress-inducible protein [Or... 147 2e-34
UniRef100_Q6IT49 Abscisic stress ripening protein-like protein [... 144 2e-33
UniRef100_O24314 Water deficit inducible protein LP3 [Pinus taeda] 141 1e-32
UniRef100_Q41087 LP3-1 [Pinus taeda] 139 9e-32
UniRef100_Q7XYV6 ASR2 [Lycopersicon hirsutum] 138 1e-31
UniRef100_Q7XYV5 ASR2 [Lycopersicon peruvianum var. humifusum] 136 5e-31
UniRef100_Q9SEW0 Anth [Lilium longiflorum] 136 6e-31
UniRef100_Q9ZRB5 Ci21B protein [Solanum tuberosum] 136 6e-31
UniRef100_Q39537 ASR [Citrus maxima] 136 6e-31
UniRef100_Q7XYV7 ASR2 [Lycopersicon chilense] 134 2e-30
UniRef100_Q40165 ABA-and ripening-induced protein [Lycopersicon ... 134 2e-30
>UniRef100_Q41300 Abscisic stress ripening protein [Solanum chacoense]
Length = 263
Score = 215 bits (547), Expect = 1e-54
Identities = 132/269 (49%), Positives = 166/269 (61%), Gaps = 34/269 (12%)
Query: 10 GLFHHKKHDDDDNRPID-------TDSGYNKTSSY----STGDDDYNKKTS-YGNDNSGG 57
GLFHH K++++D P++ T KTS+Y S GDD Y +KT+ +G+DN G
Sbjct: 6 GLFHHHKNEEEDT-PVEKTTYEETTYGESEKTSTYGEKTSYGDDTYGEKTTTFGDDNKYG 64
Query: 58 DYETGYNKNTDNYSNNETS-GGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDS 116
+ +T Y +T Y TS GG + G KT+ G G YG+ KT+ G G YG+
Sbjct: 65 E-KTSYGDDT--YGEKPTSYGGDNTYGEKTSYGKGDDNKYGE---KTSYGEGDDNKYGEK 118
Query: 117 DTKTTTGGG----YGGG----YGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGDDRREDV 168
+ +G G YGGG YG+ + GGYGGG G+ TT + +
Sbjct: 119 TSYGDSGYGEKPSYGGGDDNKYGEKTSYGNEEGGYGGGVGE----TTNYEENESETKTSE 174
Query: 169 DYEKEEKRHSKH-EHLGELGAAAAGGFALYEKHKSEKDPENAHRHKIEEEVAAVAAVGSG 227
DY KEEK+H KH E +G LGA AAG FAL+EKHK+EKDPENAH+HKIEE +AA AA+G+G
Sbjct: 175 DY-KEEKKHHKHLEEIGGLGAVAAGAFALHEKHKAEKDPENAHKHKIEEGIAAAAAIGAG 233
Query: 228 GFAFHEHHEKKETKEEEEEAHGKKKHHFF 256
GFAFHEHHEKKE KEEEEEA GKKKHHFF
Sbjct: 234 GFAFHEHHEKKEAKEEEEEAEGKKKHHFF 262
>UniRef100_Q8H0L9 DS2 protein [Solanum tuberosum]
Length = 268
Score = 213 bits (543), Expect = 3e-54
Identities = 131/276 (47%), Positives = 169/276 (60%), Gaps = 34/276 (12%)
Query: 3 EEENRSRGLFHHKKHDDDDNRPID-------TDSGYNKTSSY----STGDDDYNKKTS-Y 50
++++ GLFHH K++++D P++ T KT +Y S GDD Y +KT+ +
Sbjct: 4 QKKHHFGGLFHHHKNEEEDT-PVEKTTYEETTYGESEKTRTYGEKTSYGDDTYGEKTTTF 62
Query: 51 GNDNSGGDYETGYNKNTDNYSNNETS-GGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGY 109
G+DN G+ +T Y +T Y TS GG + G KT+ G G YG+ KT+ G G
Sbjct: 63 GDDNKYGE-KTSYGDDT--YGEKPTSYGGDNTYGEKTSYGEGDDNKYGE---KTSYGEGD 116
Query: 110 GGGYGDSDTKTTTGGG----YGGG----YGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYG 161
YG+ + +G G YGGG YG+ + GGYGGG G+ TT
Sbjct: 117 DNQYGEKTSYGDSGYGEKPSYGGGDDNKYGEKTSYGNEEGGYGGGVGE----TTNYEENE 172
Query: 162 DDRREDVDYEKEEKRHSKH-EHLGELGAAAAGGFALYEKHKSEKDPENAHRHKIEEEVAA 220
+ + DY KEEK+H KH E +G LGA AAG FAL+EKHK+EKDPENAH+HKIEE +AA
Sbjct: 173 SETKTSEDY-KEEKKHHKHLEEIGGLGAVAAGAFALHEKHKAEKDPENAHKHKIEEGIAA 231
Query: 221 VAAVGSGGFAFHEHHEKKETKEEEEEAHGKKKHHFF 256
AA+G+GGFAFHEHHEKKE KEEEEEA GKKKHHFF
Sbjct: 232 AAAIGAGGFAFHEHHEKKEAKEEEEEAEGKKKHHFF 267
>UniRef100_Q9FRV9 Putative ripening protein [Calystegia soldanella]
Length = 248
Score = 201 bits (511), Expect = 2e-50
Identities = 125/270 (46%), Positives = 150/270 (55%), Gaps = 36/270 (13%)
Query: 1 MSEEENRSRGLFHHKKHDDDDNRPI-------DTDSGYNKTSSYSTGDDDYNKKTSYGND 53
MSEE++ LF H K +++ +P +D Y + ++Y GDD Y KK +YG D
Sbjct: 1 MSEEKHHH--LFGHHKDKEEERQPSYGENTYGSSDDSYERKNTY--GDDSYEKKNTYGGD 56
Query: 54 NSGGDYETGYNKNTDNYSNNETSGGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGYG--- 110
+S YE D+Y T GG S K T GG + K T GG
Sbjct: 57 DS---YERKNTYGEDSYEKKNTYGGDESYERKNTYGGDES-----YEKKNTYGGDESYER 108
Query: 111 -GGYGDSDTKTTTGG--GYGG-GYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGDDRRE 166
YGD++ + GYG YG+ + GGG YG + GG R
Sbjct: 109 KNNYGDNNESSYENKPTGYGKTSYGED----SYGGGQTDKYGSATGIEEEGG------RT 158
Query: 167 DVDYEKEEKRHSKHEHLGELGAAAAGGFALYEKHKSEKDPENAHRHKIEEEVAAVAAVGS 226
DYEKE+K H E LG LG AAG +ALYEKH+++KDPENAHRHKIEEEVAA AVGS
Sbjct: 159 HEDYEKEKKHHKHLEELGGLGTVAAGAYALYEKHEAKKDPENAHRHKIEEEVAAAVAVGS 218
Query: 227 GGFAFHEHHEKKETKEEEEEAHGKKKHHFF 256
GGFAFHEHHEKKETKEEEEEA GKKKHHFF
Sbjct: 219 GGFAFHEHHEKKETKEEEEEAEGKKKHHFF 248
>UniRef100_O50000 Abscisic stress ripening protein homolog [Prunus armeniaca]
Length = 200
Score = 198 bits (504), Expect = 1e-49
Identities = 120/259 (46%), Positives = 139/259 (53%), Gaps = 67/259 (25%)
Query: 4 EENRSRGLFHHKKHDDDDNRPIDTDSGYNKTSSYSTGDDDYNKKTSYGNDNSGGDYETGY 63
EE GLFHH K D++RPI+T S Y ++ YS
Sbjct: 3 EEKHHHGLFHHHK---DEDRPIET-SDYPQSGGYS------------------------- 33
Query: 64 NKNTDNYSNNETSGGYGSGGAKTTTGGGYG---GGYGDSDTKT---TTGGGYGGGYGDSD 117
T GYG GG GGGYG GGYGD+ + G GYGG
Sbjct: 34 -------DEGRTGSGYGGGGGGYGGGGGYGDGGGGYGDNTAYSGEGRPGSGYGG------ 80
Query: 118 TKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGDDRREDVDYEKEEKRH 177
GGGYG SD GG Y ++ TTG ++DY+KEEK H
Sbjct: 81 -----GGGYGESADYSD---------GGRYKETAAYGTTG-----TNESEIDYKKEEKHH 121
Query: 178 SKHEHLGELGAAAAGGFALYEKHKSEKDPENAHRHKIEEEVAAVAAVGSGGFAFHEHHEK 237
EHL E GAAAAG FAL+EKH+S+KDPE+AH+HKIEEE+AA AAVGSGGFAFHEHHEK
Sbjct: 122 KHLEHLSEPGAAAAGVFALHEKHESKKDPEHAHKHKIEEEIAAAAAVGSGGFAFHEHHEK 181
Query: 238 KETKEEEEEAHGKKKHHFF 256
KE KEEEEE+HGKK HH F
Sbjct: 182 KEAKEEEEESHGKKHHHLF 200
>UniRef100_Q93WZ6 Abscisic stress ripening-like protein [Prunus persica]
Length = 193
Score = 194 bits (494), Expect = 1e-48
Identities = 119/253 (47%), Positives = 137/253 (54%), Gaps = 62/253 (24%)
Query: 4 EENRSRGLFHHKKHDDDDNRPIDTDSGYNKTSSYSTGDDDYNKKTSYGNDNSGGDYETGY 63
EE GLFHH K D++RPI+T S Y ++ YS
Sbjct: 3 EEKHHHGLFHHHK---DEDRPIET-SDYPQSGGYS------------------------- 33
Query: 64 NKNTDNYSNNETSGGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTG 123
T GYG GG G GGGYGD +T + G G GYG G
Sbjct: 34 -------DEGRTGSGYGGGGGY----GDGGGGYGD-NTAYSGEGRPGSGYGG-------G 74
Query: 124 GGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGDDRREDVDYEKEEKRHSKHEHL 183
GGYG SD GG Y ++ TTG ++DY+KEEK H EHL
Sbjct: 75 GGYGESADYSD---------GGRYKETAAYGTTG-----THESEIDYKKEEKHHKHLEHL 120
Query: 184 GELGAAAAGGFALYEKHKSEKDPENAHRHKIEEEVAAVAAVGSGGFAFHEHHEKKETKEE 243
E GAAAAG FAL+EKH+S+KDPE+AHRHKIEEE+AA AAVGSGGFAFHEHHEKKE KEE
Sbjct: 121 SEAGAAAAGVFALHEKHESKKDPEHAHRHKIEEEIAAAAAVGSGGFAFHEHHEKKEAKEE 180
Query: 244 EEEAHGKKKHHFF 256
EEE+HGKK HH F
Sbjct: 181 EEESHGKKHHHLF 193
>UniRef100_Q6S9Z8 ASR protein [Ginkgo biloba]
Length = 181
Score = 178 bits (452), Expect = 1e-43
Identities = 99/188 (52%), Positives = 117/188 (61%), Gaps = 32/188 (17%)
Query: 72 NNETSGGYGSGGA--KTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGG 129
N ++ GGY +GG + GYGG YG SD T G GY D+ G G
Sbjct: 18 NVDSEGGYNTGGDAYNKPSDDGYGGAYGSSDYNT------GSGYNDT----------GSG 61
Query: 130 YGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGDDRREDVDYEK---EEKRHSKHEHLGEL 186
Y D TG GYG YG+ T+TT Y + DYEK EEK H + EH+GEL
Sbjct: 62 YND------TGSGYG--YGEKVTETTV---YATEDSSQDDYEKAMKEEKHHKRMEHVGEL 110
Query: 187 GAAAAGGFALYEKHKSEKDPENAHRHKIEEEVAAVAAVGSGGFAFHEHHEKKETKEEEEE 246
G AAG +A+YEKH+++KDPE+AHRHKIEEEVAA AAVG+GG+AFHEHHEKKE KEE EE
Sbjct: 111 GTMAAGAYAMYEKHEAKKDPEHAHRHKIEEEVAAAAAVGAGGYAFHEHHEKKEDKEEAEE 170
Query: 247 AHGKKKHH 254
A GKK HH
Sbjct: 171 ASGKKHHH 178
>UniRef100_Q94G23 Putative transcription factor [Vitis vinifera]
Length = 149
Score = 158 bits (399), Expect = 1e-37
Identities = 80/133 (60%), Positives = 91/133 (68%), Gaps = 13/133 (9%)
Query: 130 YGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGDDRREDV------DYEKEEKRHSKHEHL 183
Y D+ TT Y D+ T GY + E + DY KEEK H EHL
Sbjct: 24 YSDNAYSDTT-------YSDTSYATDGVSGYAAETTEVLADDPAPDYRKEEKHHKHLEHL 76
Query: 184 GELGAAAAGGFALYEKHKSEKDPENAHRHKIEEEVAAVAAVGSGGFAFHEHHEKKETKEE 243
GELG AAAG +AL+EKHKSEKDPE+AH+HKIEEE+AA AAVG+GGFAFHEHHEKKE KEE
Sbjct: 77 GELGVAAAGAYALHEKHKSEKDPEHAHKHKIEEEIAAAAAVGAGGFAFHEHHEKKEAKEE 136
Query: 244 EEEAHGKKKHHFF 256
+EEAHGKK HH F
Sbjct: 137 DEEAHGKKHHHLF 149
>UniRef100_Q948L3 Drought inducible 22 kD protein [Saccharum officinarum]
Length = 142
Score = 156 bits (394), Expect = 6e-37
Identities = 76/118 (64%), Positives = 89/118 (75%), Gaps = 2/118 (1%)
Query: 140 GGGYGGGYGDSDTKTTTGGGYGDDRREDVDYEKEEKRHSKHEHLGELGAAAAGGFALYEK 199
GGGYG ++T T G+D + Y+KEEK H +HLGE GA AAG FALYEK
Sbjct: 27 GGGYGEAAEYTETTVTEVVSTGEDEYDK--YKKEEKEHKHKQHLGEAGAIAAGAFALYEK 84
Query: 200 HKSEKDPENAHRHKIEEEVAAVAAVGSGGFAFHEHHEKKETKEEEEEAHGKKKHHFFG 257
H+++KDPE+AHRHKIEEEVAA AAVGSGGFAFHEHHEKK+ +E EEA G+KKHHFFG
Sbjct: 85 HEAKKDPEHAHRHKIEEEVAAAAAVGSGGFAFHEHHEKKKDHKEAEEAGGEKKHHFFG 142
Score = 33.9 bits (76), Expect = 4.2
Identities = 17/52 (32%), Positives = 25/52 (47%), Gaps = 9/52 (17%)
Query: 4 EENRSRGLFHHKKHDDDDNRPIDTDSGYNKTSSY---------STGDDDYNK 46
EE LFHHKK +++ GY + + Y STG+D+Y+K
Sbjct: 3 EEKHHHHLFHHKKDEEEQVEQPAGGGGYGEAAEYTETTVTEVVSTGEDEYDK 54
>UniRef100_O65176 ABA stress ripening protein [Mesembryanthemum crystallinum]
Length = 143
Score = 148 bits (373), Expect = 2e-34
Identities = 70/120 (58%), Positives = 82/120 (68%), Gaps = 8/120 (6%)
Query: 137 TTTGGGYGGGYGDSDTKTTTGGGYGDDRREDVDYEKEEKRHSKHEHLGELGAAAAGGFAL 196
T +G GYG GY ++ T + DY KEE+ H EH+GELGA AAG FAL
Sbjct: 32 TYSGDGYGTGYSETTTAVIA--------EPEKDYAKEERHHKHKEHMGELGAVAAGAFAL 83
Query: 197 YEKHKSEKDPENAHRHKIEEEVAAVAAVGSGGFAFHEHHEKKETKEEEEEAHGKKKHHFF 256
+EKHK EKDPE+AHRHKIEEE+AA A VG+GG+ FHEHHEKKE KEE +E GK HH F
Sbjct: 84 HEKHKIEKDPEHAHRHKIEEEIAAAARVGAGGYVFHEHHEKKEAKEERKEHEGKHHHHLF 143
>UniRef100_O49149 Abscisic acid- and stress-inducible protein [Oryza sativa]
Length = 138
Score = 147 bits (372), Expect = 2e-34
Identities = 73/124 (58%), Positives = 87/124 (69%), Gaps = 2/124 (1%)
Query: 134 DTKTTTGGGYGGGYGDSDTKTTTGGGYGDDRREDVDYEKEEKRHSKHEHLGELGAAAAGG 193
D T YG G S+T TT G D E Y+KEEK+H +HLGE GA AAG
Sbjct: 17 DEPATGVDSYGEGVYTSETVTTEVVAGGQDEYER--YKKEEKQHKHKQHLGEAGALAAGA 74
Query: 194 FALYEKHKSEKDPENAHRHKIEEEVAAVAAVGSGGFAFHEHHEKKETKEEEEEAHGKKKH 253
FALYEKH+++KDPENAHRHKI EE+AA AAVG+GG+AFHEHHEKK+ + EE+ G+KKH
Sbjct: 75 FALYEKHEAKKDPENAHRHKITEEIAATAAVGAGGYAFHEHHEKKKDHKSAEESTGEKKH 134
Query: 254 HFFG 257
H FG
Sbjct: 135 HLFG 138
>UniRef100_Q6IT49 Abscisic stress ripening protein-like protein [Musa acuminata]
Length = 143
Score = 144 bits (363), Expect = 2e-33
Identities = 69/122 (56%), Positives = 86/122 (69%), Gaps = 7/122 (5%)
Query: 133 SDTKTTTGGGYGGGYGDSDTKTTTGGGYGDDRREDVDYEKEEKRHSKHEHLGELGAAAAG 192
S+T + G Y GY T+T D+ + Y+KEEK H EHLGE+GA AAG
Sbjct: 26 SETAYSGGDDYASGY----TETVVAESASDEYEK---YKKEEKHHKHKEHLGEMGAVAAG 78
Query: 193 GFALYEKHKSEKDPENAHRHKIEEEVAAVAAVGSGGFAFHEHHEKKETKEEEEEAHGKKK 252
FALYEKH+++KDP++AH+HKIEEE+AA AVGSGG+AFHEHHEK++ K E EEA GKK
Sbjct: 79 AFALYEKHEAKKDPDHAHKHKIEEEIAAAVAVGSGGYAFHEHHEKRDAKNEAEEASGKKH 138
Query: 253 HH 254
HH
Sbjct: 139 HH 140
>UniRef100_O24314 Water deficit inducible protein LP3 [Pinus taeda]
Length = 153
Score = 141 bits (356), Expect = 1e-32
Identities = 68/114 (59%), Positives = 84/114 (73%), Gaps = 4/114 (3%)
Query: 147 YGDSDTKTT---TGGGYGDDRREDVDYEKEEKRHSKH-EHLGELGAAAAGGFALYEKHKS 202
YGD ++ G DD+ + + ++E++H KH E LG LG AAG FAL+EKH S
Sbjct: 35 YGDEVIQSADVYAAGEVNDDKFAEYEKARKEEKHHKHLEELGGLGTVAAGAFALHEKHAS 94
Query: 203 EKDPENAHRHKIEEEVAAVAAVGSGGFAFHEHHEKKETKEEEEEAHGKKKHHFF 256
+KDPENAHRHKIEEE+AA AAVG+GG+ FHEHHEKKE+KEEE+EA GKK HH F
Sbjct: 95 KKDPENAHRHKIEEEIAAAAAVGAGGYVFHEHHEKKESKEEEKEAEGKKHHHLF 148
Score = 33.9 bits (76), Expect = 4.2
Identities = 27/88 (30%), Positives = 41/88 (45%), Gaps = 9/88 (10%)
Query: 1 MSEEENRSRGLFHHKKHDDDDNRPIDTDSGYNKTSSYSTGDDDYNKKTSYG----NDNSG 56
MSEE++ L HHKK D+ +N P + T++Y GD+ Y ND+
Sbjct: 1 MSEEKHHHH-LLHHKKEDESENVPSEVVCA-ETTTAY--GDEVIQSADVYAAGEVNDDKF 56
Query: 57 GDYETGYNKNTDNYSNNETSGGYGSGGA 84
+YE K ++ + E GG G+ A
Sbjct: 57 AEYEKA-RKEEKHHKHLEELGGLGTVAA 83
>UniRef100_Q41087 LP3-1 [Pinus taeda]
Length = 126
Score = 139 bits (349), Expect = 9e-32
Identities = 62/86 (72%), Positives = 72/86 (83%)
Query: 171 EKEEKRHSKHEHLGELGAAAAGGFALYEKHKSEKDPENAHRHKIEEEVAAVAAVGSGGFA 230
+K+EK H EHLGE+G AAG FAL+EKH +KDPE+AHRHKIEEEVAA AAVG+GG+
Sbjct: 41 KKDEKHHKHMEHLGEMGTVAAGAFALHEKHADKKDPEHAHRHKIEEEVAAAAAVGAGGYV 100
Query: 231 FHEHHEKKETKEEEEEAHGKKKHHFF 256
FHEHHEKKE+KEEE+EA GKK HH F
Sbjct: 101 FHEHHEKKESKEEEKEAEGKKHHHLF 126
>UniRef100_Q7XYV6 ASR2 [Lycopersicon hirsutum]
Length = 114
Score = 138 bits (348), Expect = 1e-31
Identities = 63/93 (67%), Positives = 76/93 (80%)
Query: 162 DDRREDVDYEKEEKRHSKHEHLGELGAAAAGGFALYEKHKSEKDPENAHRHKIEEEVAAV 221
+D VDYEKE K HS E +GELGA AAG FAL+EKHK++KDPENAH+HKIEEE+AAV
Sbjct: 19 EDEGGPVDYEKEVKHHSHLEKIGELGAVAAGAFALHEKHKAKKDPENAHKHKIEEEIAAV 78
Query: 222 AAVGSGGFAFHEHHEKKETKEEEEEAHGKKKHH 254
AAVG+GGFAFHEHH+KK+ K+E++E G HH
Sbjct: 79 AAVGAGGFAFHEHHQKKDAKKEKKEVEGGHHHH 111
>UniRef100_Q7XYV5 ASR2 [Lycopersicon peruvianum var. humifusum]
Length = 112
Score = 136 bits (343), Expect = 5e-31
Identities = 62/94 (65%), Positives = 76/94 (79%)
Query: 162 DDRREDVDYEKEEKRHSKHEHLGELGAAAAGGFALYEKHKSEKDPENAHRHKIEEEVAAV 221
+D VDYEKE K HS E +GELGA AAG F L+EKHK++KDPE+AH+HKIEEE+AAV
Sbjct: 19 EDEGGPVDYEKEVKHHSHLEKIGELGAVAAGAFGLHEKHKAKKDPEHAHKHKIEEEIAAV 78
Query: 222 AAVGSGGFAFHEHHEKKETKEEEEEAHGKKKHHF 255
AAVG+GGFAFHEHH+KK+ K+E +EA G HH+
Sbjct: 79 AAVGAGGFAFHEHHQKKDAKKEGKEAEGGHHHHY 112
>UniRef100_Q9SEW0 Anth [Lilium longiflorum]
Length = 142
Score = 136 bits (342), Expect = 6e-31
Identities = 68/113 (60%), Positives = 85/113 (75%), Gaps = 5/113 (4%)
Query: 148 GDSDTKTTTGGGYGDDRRED--VDYEKEEKRHSKH-EHLGELGAAAAGGFALYEKHKSEK 204
G + T +T+ YG ++ + DYEKE K+H KH EHLGE+ A AG FALYEKH+++K
Sbjct: 32 GLNSTYSTSSDVYGGNQSQANYEDYEKE-KKHIKHKEHLGEMATAGAGAFALYEKHEAKK 90
Query: 205 DPENAHRHKIEEEVAAVAAVGSGGFAFHEHHEKKETKEEEEEAHGKKKHHFFG 257
DPE+AHRHK+EEE+AA AA G+GG+ FHEHHEKK K+E EE G KKHHFFG
Sbjct: 91 DPEHAHRHKLEEEIAAAAAAGAGGYTFHEHHEKKTLKKENEEVEG-KKHHFFG 142
>UniRef100_Q9ZRB5 Ci21B protein [Solanum tuberosum]
Length = 108
Score = 136 bits (342), Expect = 6e-31
Identities = 63/88 (71%), Positives = 77/88 (86%), Gaps = 2/88 (2%)
Query: 168 VDYEKEEKRHSKHEHLGELGAAAAGGFALYEKHKSEKDPENAHRHKIEEEVAAVAAVGSG 227
VDYEKE K HS E +GELGA AAG FAL+EKHK++KDPENAH+HKIEEE+AAVAAVG+G
Sbjct: 23 VDYEKEVKHHSHLEKIGELGAVAAGAFALHEKHKAKKDPENAHKHKIEEEIAAVAAVGAG 82
Query: 228 GFAFHEHHEKKETKEEEEEAHGKKKHHF 255
GFAFHEHH+KK++K+E++EA G HH+
Sbjct: 83 GFAFHEHHQKKDSKKEKKEAEG--GHHY 108
>UniRef100_Q39537 ASR [Citrus maxima]
Length = 98
Score = 136 bits (342), Expect = 6e-31
Identities = 61/88 (69%), Positives = 71/88 (80%)
Query: 164 RREDVDYEKEEKRHSKHEHLGELGAAAAGGFALYEKHKSEKDPENAHRHKIEEEVAAVAA 223
+ E+ DY+KEEK H EHLG LG A AG F E HK+EKDP++AHRHKIEEE+AA A
Sbjct: 4 KEEEFDYKKEEKHHKHLEHLGGLGTAGAGAFPRLESHKAEKDPDHAHRHKIEEEIAAAAG 63
Query: 224 VGSGGFAFHEHHEKKETKEEEEEAHGKK 251
+GSGGFAFHEHHEKKE KEE++EAHGKK
Sbjct: 64 LGSGGFAFHEHHEKKEAKEEDQEAHGKK 91
>UniRef100_Q7XYV7 ASR2 [Lycopersicon chilense]
Length = 115
Score = 134 bits (338), Expect = 2e-30
Identities = 62/93 (66%), Positives = 75/93 (79%)
Query: 162 DDRREDVDYEKEEKRHSKHEHLGELGAAAAGGFALYEKHKSEKDPENAHRHKIEEEVAAV 221
+D VDYEKE K HS E +GELGA AAG FAL+EKHK++KD ENAH+HKIEEE+AAV
Sbjct: 19 EDEGGPVDYEKEVKHHSHLEKIGELGAVAAGAFALHEKHKAKKDLENAHKHKIEEEIAAV 78
Query: 222 AAVGSGGFAFHEHHEKKETKEEEEEAHGKKKHH 254
AAVG+GGFAFHEHH+KK+ K+E++E G HH
Sbjct: 79 AAVGAGGFAFHEHHQKKDAKKEKKEVEGGHHHH 111
>UniRef100_Q40165 ABA-and ripening-induced protein [Lycopersicon esculentum]
Length = 114
Score = 134 bits (337), Expect = 2e-30
Identities = 61/93 (65%), Positives = 75/93 (80%)
Query: 162 DDRREDVDYEKEEKRHSKHEHLGELGAAAAGGFALYEKHKSEKDPENAHRHKIEEEVAAV 221
+D VDYEKE K HS E +GELGA AAG AL+EKHK++KDPE+AH+HKIEEE+AAV
Sbjct: 19 EDEGGPVDYEKEVKHHSHLEKIGELGAVAAGALALHEKHKAKKDPEHAHKHKIEEEIAAV 78
Query: 222 AAVGSGGFAFHEHHEKKETKEEEEEAHGKKKHH 254
AAVG+GGFAFHEHH+KK+ K+E++E G HH
Sbjct: 79 AAVGAGGFAFHEHHQKKDAKKEKKEVEGGHHHH 111
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.303 0.131 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 608,704,631
Number of Sequences: 2790947
Number of extensions: 37611484
Number of successful extensions: 452875
Number of sequences better than 10.0: 8577
Number of HSP's better than 10.0 without gapping: 3282
Number of HSP's successfully gapped in prelim test: 5668
Number of HSP's that attempted gapping in prelim test: 171080
Number of HSP's gapped (non-prelim): 74547
length of query: 257
length of database: 848,049,833
effective HSP length: 125
effective length of query: 132
effective length of database: 499,181,458
effective search space: 65891952456
effective search space used: 65891952456
T: 11
A: 40
X1: 17 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 43 (21.7 bits)
S2: 73 (32.7 bits)
Medicago: description of AC125473.10


