

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC123572.11 + phase: 0
(475 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
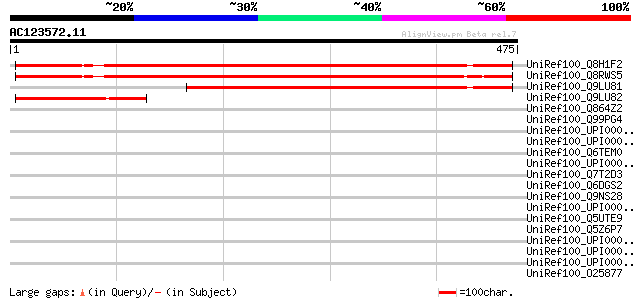
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8H1F2 Hypothetical protein At3g26090 [Arabidopsis tha... 585 e-166
UniRef100_Q8RWS5 Hypothetical protein At3g26090 [Arabidopsis tha... 582 e-165
UniRef100_Q9LU81 Arabidopsis thaliana genomic DNA, chromosome 3,... 402 e-110
UniRef100_Q9LU82 Arabidopsis thaliana genomic DNA, chromosome 3,... 145 2e-33
UniRef100_Q864Z2 Regulator of G-protein signalling 5 [Sus scrofa] 42 0.050
UniRef100_Q99PG4 Regulator of G-protein signaling 18 [Mus musculus] 41 0.086
UniRef100_UPI00002F972C UPI00002F972C UniRef100 entry 40 0.19
UniRef100_UPI0000362778 UPI0000362778 UniRef100 entry 39 0.25
UniRef100_Q6TEM0 Regulator of G-protein signalling 4 [Brachydani... 39 0.25
UniRef100_UPI000018124E UPI000018124E UniRef100 entry 39 0.33
UniRef100_Q7T2D3 Regulator of G-protein signaling 5 [Brachydanio... 39 0.33
UniRef100_Q6DGS2 Zgc:92793 [Brachydanio rerio] 39 0.43
UniRef100_Q9NS28 Regulator of G-protein signaling 18 [Homo sapiens] 38 0.73
UniRef100_UPI0000368562 UPI0000368562 UniRef100 entry 37 0.95
UniRef100_Q5UTE9 Regulator of G-protein signalling 18 [Oncorhync... 37 0.95
UniRef100_Q5Z6P7 Putative nitrite transporter [Oryza sativa] 37 0.95
UniRef100_UPI000024D1ED UPI000024D1ED UniRef100 entry 37 1.2
UniRef100_UPI000035F36C UPI000035F36C UniRef100 entry 36 2.1
UniRef100_UPI000036848E UPI000036848E UniRef100 entry 36 2.1
UniRef100_O25877 Nicotinamide mononucleotide transporter [Helico... 36 2.1
>UniRef100_Q8H1F2 Hypothetical protein At3g26090 [Arabidopsis thaliana]
Length = 459
Score = 585 bits (1508), Expect = e-166
Identities = 286/468 (61%), Positives = 365/468 (77%), Gaps = 17/468 (3%)
Query: 6 CAVKGGCPTDYVAVTVSILSFILLLIWSIFPFIVHKVPRTKGSGFWIPVIQVVASFNLLL 65
CA+ GGCP+DYVAV +S++ F +LL S+ P ++HK PRT S FWIPVIQV++SFNLL
Sbjct: 5 CALHGGCPSDYVAVAISVICFFVLLSRSVLPCLIHKAPRTNSSSFWIPVIQVISSFNLLF 64
Query: 66 SIMTISSNLKRAIGGSHVTCGQVSFHNHTLC-LFPVWGEGPLGFGLLLSSRITQAFQLYF 124
SIM +S NL R + H C L+ VW EGPLGFGLL+S RITQAFQLYF
Sbjct: 65 SIM-MSVNLLRFR----------TKHWWRYCYLWAVWIEGPLGFGLLMSCRITQAFQLYF 113
Query: 125 IFVKRRLPLIRSFLLIPLILLPWIIGAAVIHIKKPLSNRCHMSVQWTIPVVCLHALYVAT 184
IFVK+RLP ++S++ +PL+LLPWI GAA+IH KPL+++CHM +QWT PV LHALYV
Sbjct: 114 IFVKKRLPPVKSYIFLPLVLLPWIFGAAIIHATKPLNDKCHMGLQWTFPVAGLHALYVLA 173
Query: 185 LVGVTAAVHHIEFRFDELRDLWRGILVSSVSVAVWVTAYILNEIHDNISWLQVVSRFLLL 244
L+ T AV H+EFRFDELRDLW+GILVS+ S+ +WVTA++LNEIH+ ISWLQV SRF+LL
Sbjct: 174 LIAFTRAVRHVEFRFDELRDLWKGILVSATSIVIWVTAFVLNEIHEEISWLQVASRFVLL 233
Query: 245 VLASILVLAFFSISSSQPLLSQISLRRRESREFRTMGQALGIPDSGVLTQSEPISRVDPN 304
V ILV+ FFSISS+QPLLSQISL++R++ EF+ MGQALGIPDSG+L + E VDPN
Sbjct: 234 VTGGILVVVFFSISSNQPLLSQISLKKRQNFEFQRMGQALGIPDSGLLFRKEEFRPVDPN 293
Query: 305 EPLDKLLLNKKFRQSFMGFADSCLAGESVHFFDEVHELSKISEHDCVRRIYMARHIIEKY 364
EPLDKLLLNK+FR SFM FADSC AGE++HFF+EV+E KI E D +RRIYMARHI+EK+
Sbjct: 294 EPLDKLLLNKRFRHSFMEFADSCYAGETLHFFEEVYEHGKIPEDDSIRRIYMARHIMEKF 353
Query: 365 MVAGAAMEINISHRSKQEILSTSDLARADLFHNALNEIVHLMKTNLAKDYWSSMFFLKFQ 424
+VAGA ME+N+SH+++QEIL+T DL DLF NALNE++ L+K NL +DYWSS++F+KF+
Sbjct: 354 IVAGAEMELNLSHKTRQEILTTQDLTHTDLFKNALNEVMQLIKMNLVRDYWSSIYFIKFK 413
Query: 425 EECDMRCNGYELEQMTGWNY-SPRLSSVHGTDDPFHHDHLLKNSECGN 471
EE +E G+++ SPRLSSV G+DDPF+ +H+ K+S C +
Sbjct: 414 EEESC----HEAMHKEGYSFSSPRLSSVQGSDDPFYQEHMSKSSRCSS 457
>UniRef100_Q8RWS5 Hypothetical protein At3g26090 [Arabidopsis thaliana]
Length = 459
Score = 582 bits (1501), Expect = e-165
Identities = 285/467 (61%), Positives = 365/467 (78%), Gaps = 15/467 (3%)
Query: 6 CAVKGGCPTDYVAVTVSILSFILLLIWSIFPFIVHKVPRTKGSGFWIPVIQVVASFNLLL 65
CA+ GGCP+DYVAV +S++ F +LL S+ P ++HK PRT S FWIPVIQV++SFNLL
Sbjct: 5 CALHGGCPSDYVAVAISVICFFVLLSRSVLPCLIHKAPRTNSSSFWIPVIQVISSFNLLF 64
Query: 66 SIMTISSNLKRAIGGSHVTCGQVSFHNHTLC-LFPVWGEGPLGFGLLLSSRITQAFQLYF 124
SIM +S NL R + H C L+ VW EGPLGFGLL+S RITQAFQLYF
Sbjct: 65 SIM-MSVNLLRFR----------TKHWWRYCYLWAVWIEGPLGFGLLMSCRITQAFQLYF 113
Query: 125 IFVKRRLPLIRSFLLIPLILLPWIIGAAVIHIKKPLSNRCHMSVQWTIPVVCLHALYVAT 184
IFVK+RLP ++S++ +PL+LLPWI GAA+IH KPL+++CHM +QWT PV LHALYV
Sbjct: 114 IFVKKRLPPVKSYIFLPLVLLPWIFGAAIIHATKPLNDKCHMGLQWTFPVAGLHALYVLA 173
Query: 185 LVGVTAAVHHIEFRFDELRDLWRGILVSSVSVAVWVTAYILNEIHDNISWLQVVSRFLLL 244
L+ T AV H+EFRFDELRDLW+GILVS+ S+ +WVTA++LNEIH+ ISWLQV SRF+LL
Sbjct: 174 LIAFTRAVRHVEFRFDELRDLWKGILVSATSIVIWVTAFVLNEIHEEISWLQVASRFVLL 233
Query: 245 VLASILVLAFFSISSSQPLLSQISLRRRESREFRTMGQALGIPDSGVLTQSEPISRVDPN 304
V ILV+ FFSISS+QPLLSQISL++R++ EF+ MGQALGIPDSG+L + E VDPN
Sbjct: 234 VTGGILVVVFFSISSNQPLLSQISLKKRQNFEFQRMGQALGIPDSGLLFRKEEFRPVDPN 293
Query: 305 EPLDKLLLNKKFRQSFMGFADSCLAGESVHFFDEVHELSKISEHDCVRRIYMARHIIEKY 364
EPLDKLLLNK+FR SFM FADSC AGE++HFF+EV+E KI E D +RRIYMARHI+EK+
Sbjct: 294 EPLDKLLLNKRFRHSFMEFADSCYAGETLHFFEEVYEHGKIPEDDSIRRIYMARHIMEKF 353
Query: 365 MVAGAAMEINISHRSKQEILSTSDLARADLFHNALNEIVHLMKTNLAKDYWSSMFFLKFQ 424
+VAGA ME+N+SH+++QEIL+T DL DLF NALNE++ L+K NL +DYWSS++F+KF+
Sbjct: 354 IVAGAEMELNLSHKTRQEILTTQDLTHTDLFKNALNEVMQLIKMNLVRDYWSSIYFIKFK 413
Query: 425 EECDMRCNGYELEQMTGWNYSPRLSSVHGTDDPFHHDHLLKNSECGN 471
EE C+ ++ ++ SPRLSSV G+DDPF+ +H+ K+S C +
Sbjct: 414 EE--ESCHEAMHKERYSFS-SPRLSSVQGSDDPFYQEHMSKSSRCSS 457
>UniRef100_Q9LU81 Arabidopsis thaliana genomic DNA, chromosome 3, P1 clone: MPE11
[Arabidopsis thaliana]
Length = 305
Score = 402 bits (1033), Expect = e-110
Identities = 193/307 (62%), Positives = 246/307 (79%), Gaps = 5/307 (1%)
Query: 166 MSVQWTIPVVCLHALYVATLVGVTAAVHHIEFRFDELRDLWRGILVSSVSVAVWVTAYIL 225
M +QWT PV LHALYV L+ T AV H+EFRFDELRDLW+GILVS+ S+ +WVTA++L
Sbjct: 1 MGLQWTFPVAGLHALYVLALIAFTRAVRHVEFRFDELRDLWKGILVSATSIVIWVTAFVL 60
Query: 226 NEIHDNISWLQVVSRFLLLVLASILVLAFFSISSSQPLLSQISLRRRESREFRTMGQALG 285
NEIH+ ISWLQV SRF+LLV ILV+ FFSISS+QPLLSQISL++R++ EF+ MGQALG
Sbjct: 61 NEIHEEISWLQVASRFVLLVTGGILVVVFFSISSNQPLLSQISLKKRQNFEFQRMGQALG 120
Query: 286 IPDSGVLTQSEPISRVDPNEPLDKLLLNKKFRQSFMGFADSCLAGESVHFFDEVHELSKI 345
IPDSG+L + E VDPNEPLDKLLLNK+FR SFM FADSC AGE++HFF+EV+E KI
Sbjct: 121 IPDSGLLFRKEEFRPVDPNEPLDKLLLNKRFRHSFMEFADSCYAGETLHFFEEVYEHGKI 180
Query: 346 SEHDCVRRIYMARHIIEKYMVAGAAMEINISHRSKQEILSTSDLARADLFHNALNEIVHL 405
E D +RRIYMARHI+EK++VAGA ME+N+SH+++QEIL+T DL DLF NALNE++ L
Sbjct: 181 PEDDSIRRIYMARHIMEKFIVAGAEMELNLSHKTRQEILTTQDLTHTDLFKNALNEVMQL 240
Query: 406 MKTNLAKDYWSSMFFLKFQEECDMRCNGYELEQMTGWNY-SPRLSSVHGTDDPFHHDHLL 464
+K NL +DYWSS++F+KF+EE +E G+++ SPRLSSV G+DDPF+ +H+
Sbjct: 241 IKMNLVRDYWSSIYFIKFKEEESC----HEAMHKEGYSFSSPRLSSVQGSDDPFYQEHMS 296
Query: 465 KNSECGN 471
K+S C +
Sbjct: 297 KSSRCSS 303
>UniRef100_Q9LU82 Arabidopsis thaliana genomic DNA, chromosome 3, P1 clone: MPE11
[Arabidopsis thaliana]
Length = 125
Score = 145 bits (366), Expect = 2e-33
Identities = 73/123 (59%), Positives = 88/123 (71%), Gaps = 2/123 (1%)
Query: 6 CAVKGGCPTDYVAVTVSILSFILLLIWSIFPFIVHKVPRTKGSGFWIPVIQVVASFNLLL 65
CA+ GGCP+DYVAV +S++ F +LL S+ P ++HK PRT S FWIPVIQV++SFNLL
Sbjct: 5 CALHGGCPSDYVAVAISVICFFVLLSRSVLPCLIHKAPRTNSSSFWIPVIQVISSFNLLF 64
Query: 66 SIMTISSNLKRAIGGSHVTCGQVSFHNHTLCLFPVWGEGPLGFGLLLSSRITQAFQLYFI 125
SIM I + R IGG GQV T+ +W EGPLGFGLL+S RITQAFQLYFI
Sbjct: 65 SIMLICFDSGRNIGGDIAIFGQVCL--ETMIPHLLWIEGPLGFGLLMSCRITQAFQLYFI 122
Query: 126 FVK 128
FVK
Sbjct: 123 FVK 125
>UniRef100_Q864Z2 Regulator of G-protein signalling 5 [Sus scrofa]
Length = 181
Score = 41.6 bits (96), Expect = 0.050
Identities = 36/138 (26%), Positives = 59/138 (42%), Gaps = 3/138 (2%)
Query: 287 PDSGVLTQSEPISR-VDPNEPLDKLLLNKKFRQSFMGFADSCLAGESVHFFDEVHELSKI 345
PD V Q + + + LDKLL N SF F S + E++ F+ + KI
Sbjct: 44 PDKPVKIQKPSLDEALQWRDSLDKLLQNNYGLASFKSFLKSEFSEENLEFWMACEDYKKI 103
Query: 346 SEHDCVRRIYMARHIIEKYMVAGAAMEINISHRSKQEILSTSDLARADLFHNALNEIVHL 405
V+ A+ I E+++ + A E+NI H +K+ + F A I L
Sbjct: 104 KSP--VKMAEKAKKIYEEFIQSEAPKEVNIDHFTKEITMKNLVEPSPSSFDVAQKRIYAL 161
Query: 406 MKTNLAKDYWSSMFFLKF 423
M+ + + S F+ +F
Sbjct: 162 MEKDSLPRFVRSEFYQEF 179
>UniRef100_Q99PG4 Regulator of G-protein signaling 18 [Mus musculus]
Length = 235
Score = 40.8 bits (94), Expect = 0.086
Identities = 33/133 (24%), Positives = 57/133 (42%), Gaps = 2/133 (1%)
Query: 293 TQSEPISRVDPNEPLDKLLLNKKFRQSFMGFADSCLAGESVHFFDEVHELSKISEHDCVR 352
T+ P V E DKLL ++ +F F + + E++ F+ + K E +
Sbjct: 73 TRVSPEEAVKWAESFDKLLSHRDGVDAFTRFLKTEFSEENIEFWVACEDFKKCKEPQQI- 131
Query: 353 RIYMARHIIEKYMVAGAAMEINISHRSKQEILSTSDLARADLFHNALNEIVHLMKTNLAK 412
I A+ I EK++ A E+NI +K+ I + F A + + LM+ + K
Sbjct: 132 -ILKAKAIYEKFIQNDAPKEVNIDFHTKEVIAKSIAQPTLHSFDTAQSRVYQLMEHDSYK 190
Query: 413 DYWSSMFFLKFQE 425
+ S +L E
Sbjct: 191 RFLKSETYLHLIE 203
>UniRef100_UPI00002F972C UPI00002F972C UniRef100 entry
Length = 303
Score = 39.7 bits (91), Expect = 0.19
Identities = 29/101 (28%), Positives = 47/101 (45%), Gaps = 2/101 (1%)
Query: 307 LDKLLLNKKFRQSFMGFADSCLAGESVHFFDEVHELSKISEHDCVRRIYMARHIIEKYMV 366
L++LL +K + F F S + E++ F+ + KI R AR I ++Y+
Sbjct: 183 LERLLSSKNGIKVFQAFLKSEFSDENIEFWLVCEDYKKIGSS--FRMSSRARKIFKRYIQ 240
Query: 367 AGAAMEINISHRSKQEILSTSDLARADLFHNALNEIVHLMK 407
A AA EINI H+++ I F +A + LM+
Sbjct: 241 ADAAREINIDHKTRDLIRRNVAAPTRVCFDDAQRIVYSLME 281
>UniRef100_UPI0000362778 UPI0000362778 UniRef100 entry
Length = 126
Score = 39.3 bits (90), Expect = 0.25
Identities = 29/103 (28%), Positives = 48/103 (46%), Gaps = 2/103 (1%)
Query: 305 EPLDKLLLNKKFRQSFMGFADSCLAGESVHFFDEVHELSKISEHDCVRRIYMARHIIEKY 364
E +DK+L K Q F F S + E++ F+ + KI R A+ I + Y
Sbjct: 9 ESVDKVLGCKDGIQIFQAFLKSEFSDENIEFWMVCEDYKKI--RSSFRMSSRAKKIFKCY 66
Query: 365 MVAGAAMEINISHRSKQEILSTSDLARADLFHNALNEIVHLMK 407
+ A A EINI H+++ I + F++A + +LM+
Sbjct: 67 IQAEATREINIDHKTRDLIRRNVAAPTRECFNDAQRIVYNLME 109
>UniRef100_Q6TEM0 Regulator of G-protein signalling 4 [Brachydanio rerio]
Length = 215
Score = 39.3 bits (90), Expect = 0.25
Identities = 27/134 (20%), Positives = 60/134 (44%), Gaps = 9/134 (6%)
Query: 298 ISRVDPNEP------LDKLLLNKKFRQSFMGFADSCLAGESVHFFDEVHELSKISEHDCV 351
++R+ P E L+ N R++F F S + E++ F+ + +
Sbjct: 56 VNRITPAETEKWKTSFTNLIKNDDGRKAFASFLQSEYSQENIEFWVACEDFKQTPAD--- 112
Query: 352 RRIYMARHIIEKYMVAGAAMEINISHRSKQEILSTSDLARADLFHNALNEIVHLMKTNLA 411
+ AR+I E+Y+ A + E+N+ ++++ ++ F A ++I LM+ +
Sbjct: 113 KMNLKARNIFERYIEADSPREVNLDSVTREQTRKNLEMCDVSCFDEAQSKIFTLMEKDSY 172
Query: 412 KDYWSSMFFLKFQE 425
+ + S FL+ +
Sbjct: 173 RRFLRSRLFLELSQ 186
>UniRef100_UPI000018124E UPI000018124E UniRef100 entry
Length = 235
Score = 38.9 bits (89), Expect = 0.33
Identities = 29/121 (23%), Positives = 54/121 (43%), Gaps = 2/121 (1%)
Query: 305 EPLDKLLLNKKFRQSFMGFADSCLAGESVHFFDEVHELSKISEHDCVRRIYMARHIIEKY 364
E DKLL ++ +F F + + E++ F+ + K +E + I A+ I EK+
Sbjct: 85 ESFDKLLSHRDGVDAFTRFLKTEFSEENIEFWVACEDFKKCTEPQ--QLILKAKSIYEKF 142
Query: 365 MVAGAAMEINISHRSKQEILSTSDLARADLFHNALNEIVHLMKTNLAKDYWSSMFFLKFQ 424
+ A E+N+ +K+ I + F A + + LM+ + K + S +L
Sbjct: 143 IKNDAPKEVNLDFHTKEVITKSIAQPTLHSFDAAQSRVCQLMEHDSYKRFLKSEIYLHLI 202
Query: 425 E 425
E
Sbjct: 203 E 203
>UniRef100_Q7T2D3 Regulator of G-protein signaling 5 [Brachydanio rerio]
Length = 182
Score = 38.9 bits (89), Expect = 0.33
Identities = 29/116 (25%), Positives = 49/116 (42%), Gaps = 2/116 (1%)
Query: 292 LTQSEPISRVDPNEPLDKLLLNKKFRQSFMGFADSCLAGESVHFFDEVHELSKISEHDCV 351
L ++ P E LDK+L N +F F S + E++ F++ + K +
Sbjct: 51 LQKATPEEAAQWRESLDKVLSNSYGLATFKSFLRSEFSEENIEFWEACEDFKKTKNP--L 108
Query: 352 RRIYMARHIIEKYMVAGAAMEINISHRSKQEILSTSDLARADLFHNALNEIVHLMK 407
+ A+ I E ++ G E+NI H +K L + F A + I LM+
Sbjct: 109 KMATKAKKIYEDFIQTGGPKEVNIDHFTKDVTLRNLVDLSSSTFELAQSRIYTLME 164
>UniRef100_Q6DGS2 Zgc:92793 [Brachydanio rerio]
Length = 459
Score = 38.5 bits (88), Expect = 0.43
Identities = 26/103 (25%), Positives = 50/103 (48%), Gaps = 3/103 (2%)
Query: 307 LDKLLLNKKFRQSFMGFADSCLAGESVHFFDEVHELSKISEHDCVRRIYMARHIIEKYMV 366
+D++L + R+ F+ F +S + E++ F+ V EL + + R+ + I E+++
Sbjct: 334 IDEVLKDPVGREQFLKFLESEFSSENLRFWLAVQELKRRPIREVPTRV---QEIWEEFLA 390
Query: 367 AGAAMEINISHRSKQEILSTSDLARADLFHNALNEIVHLMKTN 409
AGA IN+ +S + F +A I LMK++
Sbjct: 391 AGAPSAINVDSKSYDKTTQNVKDPGRYAFEDAQEHIYKLMKSD 433
>UniRef100_Q9NS28 Regulator of G-protein signaling 18 [Homo sapiens]
Length = 235
Score = 37.7 bits (86), Expect = 0.73
Identities = 29/133 (21%), Positives = 56/133 (41%), Gaps = 2/133 (1%)
Query: 293 TQSEPISRVDPNEPLDKLLLNKKFRQSFMGFADSCLAGESVHFFDEVHELSKISEHDCVR 352
T+ P V E DKLL ++ ++F F + + E++ F+ + K +
Sbjct: 73 TRVSPEEAVKWGESFDKLLSHRDGLEAFTRFLKTEFSEENIEFWIACEDFKKSKGPQQIH 132
Query: 353 RIYMARHIIEKYMVAGAAMEINISHRSKQEILSTSDLARADLFHNALNEIVHLMKTNLAK 412
A+ I EK++ A E+N+ +K+ I ++ F A + + LM+ +
Sbjct: 133 --LKAKAIYEKFIQTDAPKEVNLDFHTKEVITNSITQPTLHSFDAAQSRVYQLMEQDSYT 190
Query: 413 DYWSSMFFLKFQE 425
+ S +L E
Sbjct: 191 RFLKSDIYLDLME 203
>UniRef100_UPI0000368562 UPI0000368562 UniRef100 entry
Length = 235
Score = 37.4 bits (85), Expect = 0.95
Identities = 29/133 (21%), Positives = 56/133 (41%), Gaps = 2/133 (1%)
Query: 293 TQSEPISRVDPNEPLDKLLLNKKFRQSFMGFADSCLAGESVHFFDEVHELSKISEHDCVR 352
T+ P V E DKLL ++ ++F F + + E++ F+ + K +
Sbjct: 73 TRVSPEEAVKWGESFDKLLSHRDGLEAFTRFLKTEFSEENIEFWIACEDFKKSKGPQQIH 132
Query: 353 RIYMARHIIEKYMVAGAAMEINISHRSKQEILSTSDLARADLFHNALNEIVHLMKTNLAK 412
A+ I EK++ A E+N+ +K+ I ++ F A + + LM+ +
Sbjct: 133 --LKAKAIYEKFIQTDAPKEVNLDFHTKEVITNSITQPTLHSFDVAQSRVYQLMEQDSYT 190
Query: 413 DYWSSMFFLKFQE 425
+ S +L E
Sbjct: 191 RFLKSDIYLDLME 203
>UniRef100_Q5UTE9 Regulator of G-protein signalling 18 [Oncorhynchus mykiss]
Length = 236
Score = 37.4 bits (85), Expect = 0.95
Identities = 28/118 (23%), Positives = 53/118 (44%), Gaps = 2/118 (1%)
Query: 304 NEPLDKLLLNKKFRQSFMGFADSCLAGESVHFFDEVHELSKISEHDCVRRIYMARHIIEK 363
++ + L+ + +SF F + + E++ F+ E I +R I A+H+
Sbjct: 84 SDSFEDLIRHSDGVESFSQFLRTEFSEENIEFWLACEEYKTIDSE--IRLISKAKHMYSV 141
Query: 364 YMVAGAAMEINISHRSKQEILSTSDLARADLFHNALNEIVHLMKTNLAKDYWSSMFFL 421
++ A A E+NI + SK I T F A +++ LMK + + +S +L
Sbjct: 142 FIEAEAPKEVNIDYASKVAIQKTMASPTNTCFDAAQSKVYSLMKKDCYPRFLTSDIYL 199
>UniRef100_Q5Z6P7 Putative nitrite transporter [Oryza sativa]
Length = 597
Score = 37.4 bits (85), Expect = 0.95
Identities = 27/122 (22%), Positives = 57/122 (46%), Gaps = 22/122 (18%)
Query: 166 MSVQWTIPVVCLHALYVATLVGVTAAVHHIEFRFDE----LRDLWRGILVSSVSVAVWVT 221
+S W +P LH + A +V H+EF +D+ +R + + S+S+ +V+
Sbjct: 460 LSAYWLVPQYALHGMAEAF-----NSVGHLEFMYDQSPESMRSMATALFWLSISLGSYVS 514
Query: 222 AYILNEIH-------------DNISWLQVVSRFLLLVLASILVLAFFSISSSQPLLSQIS 268
+++ +H DNI+ ++ + ++ L +L LA+++I + L +
Sbjct: 515 TMLISAVHRWSAGADGSNWLPDNINRGRLDYFYWIVALLQVLNLAYYAICARCYLFKPLQ 574
Query: 269 LR 270
LR
Sbjct: 575 LR 576
>UniRef100_UPI000024D1ED UPI000024D1ED UniRef100 entry
Length = 130
Score = 37.0 bits (84), Expect = 1.2
Identities = 30/113 (26%), Positives = 50/113 (43%), Gaps = 3/113 (2%)
Query: 296 EPISRVDP-NEPLDKLLLNKKFRQSFMGFADSCLAGESVHFFDEVHELSKISEHDCVRRI 354
EP+ VD E ++KLL K + +F F S + E++ F+ E K++ I
Sbjct: 2 EPVDDVDSWGESIEKLLSCKAGQAAFQDFLKSEYSEENILFWLACEEYKKVTNEP--EMI 59
Query: 355 YMARHIIEKYMVAGAAMEINISHRSKQEILSTSDLARADLFHNALNEIVHLMK 407
MA I +++ A +INI H +++ I F A + LM+
Sbjct: 60 SMANQIYTEFVQTEAPRQINIDHVTRELIKQNVHAPTKVCFDEAQRIVFGLME 112
>UniRef100_UPI000035F36C UPI000035F36C UniRef100 entry
Length = 181
Score = 36.2 bits (82), Expect = 2.1
Identities = 24/80 (30%), Positives = 37/80 (46%), Gaps = 2/80 (2%)
Query: 305 EPLDKLLLNKKFRQSFMGFADSCLAGESVHFFDEVHELSKISEHDCVRRIYMARHIIEKY 364
E LD++L N +F F S + E+V F+ + K D V+ A+ I E +
Sbjct: 64 ESLDRVLNNSYGLNAFRSFLQSEFSDENVEFWMSCEDFKKTK--DPVKMAAKAKKIYEDF 121
Query: 365 MVAGAAMEINISHRSKQEIL 384
+ A E+NI H +K L
Sbjct: 122 IQAEGPREVNIDHFTKDMTL 141
>UniRef100_UPI000036848E UPI000036848E UniRef100 entry
Length = 181
Score = 36.2 bits (82), Expect = 2.1
Identities = 30/116 (25%), Positives = 48/116 (40%), Gaps = 2/116 (1%)
Query: 305 EPLDKLLLNKKFRQSFMGFADSCLAGESVHFFDEVHELSKISEHDCVRRIYMARHIIEKY 364
+ LDKLL N SF F S + E++ F+ + KI + A+ I E++
Sbjct: 63 DSLDKLLQNNYGLASFKSFLKSEFSEENLEFWIACEDYKKIKSP--AKMAEKAKQIYEEF 120
Query: 365 MVAGAAMEINISHRSKQEILSTSDLARADLFHNALNEIVHLMKTNLAKDYWSSMFF 420
+ A E+NI H +K + F A I LM+ + + S F+
Sbjct: 121 IQTEAPKEVNIDHFTKDITMKNLVEPSLSSFDMAQKRIHALMEKDSLPRFVRSEFY 176
>UniRef100_O25877 Nicotinamide mononucleotide transporter [Helicobacter pylori]
Length = 220
Score = 36.2 bits (82), Expect = 2.1
Identities = 25/93 (26%), Positives = 43/93 (45%), Gaps = 8/93 (8%)
Query: 165 HMSVQWTIP---VVCLHALYVATLVGVTA----AVHHIEFRFDELRDLWRGILVSSVSVA 217
+++ QW + ++CL T+ G+ A H + +L WR IL+ V V
Sbjct: 69 YVAYQWKLNADVILCLFLYMPVTIYGLFAWKKTEQHEGVIKAQKLSKNWRFILILGVGVL 128
Query: 218 VWVTAYILNEIHDNISWLQVVSRFLLLVLASIL 250
V+A EI N W + + F++ ++A IL
Sbjct: 129 TCVSALFFKEIKTNFLWAESFN-FVIFIIAFIL 160
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.327 0.140 0.432
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 764,321,480
Number of Sequences: 2790947
Number of extensions: 30151575
Number of successful extensions: 110260
Number of sequences better than 10.0: 59
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 55
Number of HSP's that attempted gapping in prelim test: 110248
Number of HSP's gapped (non-prelim): 59
length of query: 475
length of database: 848,049,833
effective HSP length: 131
effective length of query: 344
effective length of database: 482,435,776
effective search space: 165957906944
effective search space used: 165957906944
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 77 (34.3 bits)
Medicago: description of AC123572.11


