

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
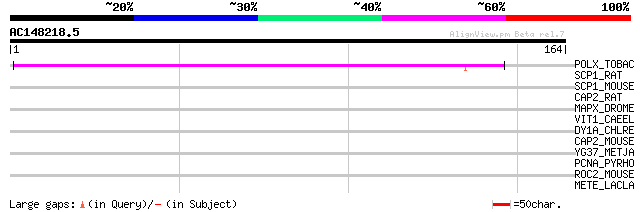
Query= AC148218.5 - phase: 0
(164 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
POLX_TOBAC (P10978) Retrovirus-related Pol polyprotein from tran... 90 2e-18
SCP1_RAT (Q03410) Synaptonemal complex protein 1 (SCP-1 protein) 35 0.099
SCP1_MOUSE (Q62209) Synaptonemal complex protein 1 (SCP-1 protein) 35 0.099
CAP2_RAT (P52481) Adenylyl cyclase-associated protein 2 (CAP 2) 31 1.1
MAPX_DROME (P23226) 205 kDa microtubule-associated protein 30 1.9
VIT1_CAEEL (P55155) Vitellogenin 1 precursor 30 3.2
DY1A_CHLRE (Q9SMH3) Dynein 1-alpha heavy chain, flagellar inner ... 30 3.2
CAP2_MOUSE (Q9CYT6) Adenylyl cyclase-associated protein 2 (CAP 2) 30 3.2
YG37_METJA (Q59031) Hypothetical protein MJ1637 precursor 29 4.2
PCNA_PYRHO (O58398) DNA polymerase sliding clamp (Proliferating ... 29 4.2
ROC2_MOUSE (P70336) Rho-associated protein kinase 2 (EC 2.7.1.37... 28 9.3
METE_LACLA (Q9CG55) 5-methyltetrahydropteroyltriglutamate--homoc... 28 9.3
>POLX_TOBAC (P10978) Retrovirus-related Pol polyprotein from
transposon TNT 1-94 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1328
Score = 90.1 bits (222), Expect = 2e-18
Identities = 51/147 (34%), Positives = 81/147 (54%), Gaps = 2/147 (1%)
Query: 2 GSKWDIEKFTGDNDFRLWKVKMEAVLIQQKCEKTLKGESSLPVTMSRAEKTEMVDKARSA 61
G K+++ KF GDN F W+ +M +LIQQ K L +S P TM + ++ ++A SA
Sbjct: 3 GVKYEVAKFNGDNGFSTWQRRMRDLLIQQGLHKVLDVDSKKPDTMKAEDWADLDERAASA 62
Query: 62 IVLCLGNKDLREATKETTATSMWSKFEYLYMTKSLAHRQVLKQQLYSFKMVKSKVVVELL 121
I L L + + E TA +W++ E LYM+K+L ++ LK+QLY+ M + + L
Sbjct: 63 IRLHLSDDVVNNIIDEDTARGIWTRLESLYMSKTLTNKLYLKKQLYALHMSEGTNFLSHL 122
Query: 122 VEFNKIIGDFEN--IKLHLEDAGALMV 146
FN +I N +K+ ED L++
Sbjct: 123 NVFNGLITQLANLGVKIEEEDKAILLL 149
>SCP1_RAT (Q03410) Synaptonemal complex protein 1 (SCP-1 protein)
Length = 997
Score = 34.7 bits (78), Expect = 0.099
Identities = 34/143 (23%), Positives = 63/143 (43%), Gaps = 9/143 (6%)
Query: 19 WKVKMEAVLIQQKCEKTLKGESSLPVTMSRAEKTEMVDKARSAIVLCLGNKDLREATKET 78
WKV +E+ L Q+ E L+ + +A + + + ++ L ++ ++ KE
Sbjct: 120 WKVSIESELKQK--ENKLQENRKIIEAQRKAIQELQFENEKVSLKLEEEIQENKDLIKEN 177
Query: 79 TATSMWSKFEYLYMTKSLAHRQVLKQQLYSFKMVKSK-VVVELLVEFNKIIGDFENIKLH 137
AT W + K R K Y ++ +++ V V+L K+I FE +++
Sbjct: 178 NATRHWCN-----LLKETCARSAEKTSKYEYEREETRQVYVDLNNNIEKMILAFEELRVQ 232
Query: 138 LEDAGALMVWCCLEDEEGDVSHL 160
E+A M + ED E + HL
Sbjct: 233 AENARLEMHFKLKEDHE-KIQHL 254
>SCP1_MOUSE (Q62209) Synaptonemal complex protein 1 (SCP-1 protein)
Length = 993
Score = 34.7 bits (78), Expect = 0.099
Identities = 34/143 (23%), Positives = 63/143 (43%), Gaps = 9/143 (6%)
Query: 19 WKVKMEAVLIQQKCEKTLKGESSLPVTMSRAEKTEMVDKARSAIVLCLGNKDLREATKET 78
WKV +E+ L Q+ E L+ + +A + + + ++ L ++ ++ KE
Sbjct: 116 WKVSIESELKQK--ENKLQENRKIIEAQRKAIQELQFENEKVSLKLEEEIQENKDLIKEN 173
Query: 79 TATSMWSKFEYLYMTKSLAHRQVLKQQLYSFKMVKSK-VVVELLVEFNKIIGDFENIKLH 137
AT W + K R K Y ++ +++ V V+L K+I FE +++
Sbjct: 174 NATIHWCN-----LLKETCARSAEKTNKYEYEREETRQVYVDLNSNIEKMILAFEELRVQ 228
Query: 138 LEDAGALMVWCCLEDEEGDVSHL 160
E+A M + ED E + HL
Sbjct: 229 AENARLEMHFKLKEDHE-KIQHL 250
>CAP2_RAT (P52481) Adenylyl cyclase-associated protein 2 (CAP 2)
Length = 477
Score = 31.2 bits (69), Expect = 1.1
Identities = 25/98 (25%), Positives = 48/98 (48%), Gaps = 6/98 (6%)
Query: 2 GSKWDIEKFTGDNDFRLWKVKMEAVLIQQKCEKT---LKGESSLPVTMSRAEKTEMVDKA 58
G KW +E ND + + +++ V KC+K+ +KG+ + +T+ +K +V
Sbjct: 329 GKKWRVEYQEDRNDLVISETELKQVAYIFKCDKSTLQIKGKVN-SITVDNCKKFGLVFDH 387
Query: 59 RSAIVLCLGNKDLREATKETTATSMWSKFE--YLYMTK 94
IV + +KD++ T +K E +LY++K
Sbjct: 388 VVGIVEVINSKDIQIQVMGRVPTISINKTEGCHLYLSK 425
>MAPX_DROME (P23226) 205 kDa microtubule-associated protein
Length = 1185
Score = 30.4 bits (67), Expect = 1.9
Identities = 25/66 (37%), Positives = 32/66 (47%), Gaps = 4/66 (6%)
Query: 33 EKTLKGESSLPVTMSRAEKTEMVDKARSAIVLCLGNKDLREATKE-TTATSMWSKFEYLY 91
E L+ E + V S +EKT + A A+V G ATK +TA S SK E L
Sbjct: 767 EDILEPEKDVEVAKSLSEKTSVATVAGGAVV---GATKTHSATKAGSTAASAKSKTETLV 823
Query: 92 MTKSLA 97
M K+ A
Sbjct: 824 MKKTTA 829
>VIT1_CAEEL (P55155) Vitellogenin 1 precursor
Length = 1616
Score = 29.6 bits (65), Expect = 3.2
Identities = 27/116 (23%), Positives = 53/116 (45%), Gaps = 12/116 (10%)
Query: 33 EKTLKGESSLPVTMSRAEKTEMVDKA--------RSAIVLCLG-NKDLREATKETTATSM 83
EKTL+G+ + T+ R +K ++ K+ RS I L + + E K+T
Sbjct: 178 EKTLEGDCQVAYTVIREQKKTIITKSINFDKCTERSEIAYGLRYSSECPECEKDTVLIRP 237
Query: 84 WSKFEYLYMTKSLAHRQVLKQQLYSFKMVKSKVVVELLVEFNKIIGDFENIKLHLE 139
+ + Y+ + L +V + LY+ V + V++ ++ + +IK H+E
Sbjct: 238 QTVYTYILENEELKESEV--RSLYTVN-VNGQEVMKTETRSKLVLEENHSIKSHIE 290
>DY1A_CHLRE (Q9SMH3) Dynein 1-alpha heavy chain, flagellar inner arm
I1 complex (1-alpha DHC) (Dynein 1, subspecies f)
Length = 4625
Score = 29.6 bits (65), Expect = 3.2
Identities = 20/66 (30%), Positives = 34/66 (51%), Gaps = 12/66 (18%)
Query: 68 NKDLREATKETTATSMWSKFEYLYMTKSLAHRQVLKQQLYSFKMVKSKVVV--ELLVEFN 125
N+D++EAT+E TS + L + + L SFK +KSK + +++ +FN
Sbjct: 609 NEDIKEATRELIDTSF----------RKLRSAEGACELLQSFKSIKSKGAIQKQVMNKFN 658
Query: 126 KIIGDF 131
I+ F
Sbjct: 659 DILEQF 664
>CAP2_MOUSE (Q9CYT6) Adenylyl cyclase-associated protein 2 (CAP 2)
Length = 476
Score = 29.6 bits (65), Expect = 3.2
Identities = 24/98 (24%), Positives = 48/98 (48%), Gaps = 6/98 (6%)
Query: 2 GSKWDIEKFTGDNDFRLWKVKMEAVLIQQKCEKT---LKGESSLPVTMSRAEKTEMVDKA 58
G KW +E ND + + +++ V KC+K+ +KG+ + +T+ +K +V
Sbjct: 328 GKKWRVEYQEDRNDLVISETELKQVAYIFKCDKSTLQIKGKVN-SITVDNCKKFGLVFDH 386
Query: 59 RSAIVLCLGNKDLREATKETTATSMWSKFE--YLYMTK 94
IV + +KD++ T +K E +LY+++
Sbjct: 387 VVGIVEVINSKDIQIQVMGRVPTISINKTEGCHLYLSE 424
>YG37_METJA (Q59031) Hypothetical protein MJ1637 precursor
Length = 473
Score = 29.3 bits (64), Expect = 4.2
Identities = 21/58 (36%), Positives = 29/58 (49%), Gaps = 5/58 (8%)
Query: 88 EYLYMTK-----SLAHRQVLKQQLYSFKMVKSKVVVELLVEFNKIIGDFENIKLHLED 140
EYLY+TK SL +L L +FK V +KV ++L E K ++ LED
Sbjct: 199 EYLYLTKNIKIESLFEEVILNNNLDNFKKVYAKVYSQILEEDVKKYLRIGSLPFALED 256
>PCNA_PYRHO (O58398) DNA polymerase sliding clamp (Proliferating
cell nuclear antigen homolog) (PCNA)
Length = 249
Score = 29.3 bits (64), Expect = 4.2
Identities = 29/125 (23%), Positives = 52/125 (41%), Gaps = 20/125 (16%)
Query: 38 GESSLPVTMSRAEKTEMVDKARSAIVLCLGNKDLREATKETTATSMW------------S 85
GE ++ V M +K KA+ ++L G ++ E + + TAT +
Sbjct: 65 GEETIGVNMDHLKKVLKRGKAKDTLILRKGEENFLEISLQGTATRTFRLPLIDVEEIEVE 124
Query: 86 KFEYLYMTKSLAHRQVLKQQLYSFKMVKSKVVVELLVEFNKIIGDFE------NIKLHLE 139
+ Y K + +VLK+ + +V ++ + + N+ I E +KL LE
Sbjct: 125 LPDLPYTAKVVVLGEVLKEAVKDASLVSDS--IKFMAKENEFIMRAEGETQEVEVKLTLE 182
Query: 140 DAGAL 144
D G L
Sbjct: 183 DEGLL 187
>ROC2_MOUSE (P70336) Rho-associated protein kinase 2 (EC 2.7.1.37)
(Rho-associated, coiled-coil containing protein kinase 2)
(p164 ROCK-2)
Length = 1388
Score = 28.1 bits (61), Expect = 9.3
Identities = 33/135 (24%), Positives = 58/135 (42%), Gaps = 16/135 (11%)
Query: 13 DNDFRLWKVK---MEAVLIQQKCEKTLKGESSLPVTMSRAEKTEMVDKA-----RSAIVL 64
D+ +L K+K M A I+ + EK L E +L KT+ V+K R V
Sbjct: 998 DSQEQLSKLKDEEMSAAAIKAQFEKQLLNERTL--------KTQAVNKLAEIMNRKEPVK 1049
Query: 65 CLGNKDLREATKETTATSMWSKFEYLYMTKSLAHRQVLKQQLYSFKMVKSKVVVELLVEF 124
+ D+R KE M K E +T+ + Q ++ + +S++ +EL +
Sbjct: 1050 RGSDTDVRRKEKENRKLHMELKSEREKLTQQMIKYQKELNEMQAQIAEESQIRIELQMTL 1109
Query: 125 NKIIGDFENIKLHLE 139
+ D E ++ L+
Sbjct: 1110 DSKDSDIEQLRSQLQ 1124
>METE_LACLA (Q9CG55)
5-methyltetrahydropteroyltriglutamate--homocysteine
methyltransferase (EC 2.1.1.14) (Methionine synthase,
vitamin-B12 independent isozyme) (Cobalamin-independent
methionine synthase)
Length = 759
Score = 28.1 bits (61), Expect = 9.3
Identities = 28/111 (25%), Positives = 52/111 (46%), Gaps = 5/111 (4%)
Query: 29 QQKCEKTLKGESSLPVT-MSRAEKTEMVDKARSAIVLCLGNK----DLREATKETTATSM 83
Q + K +KG + PVT ++ + E + S + L L + DL +A +
Sbjct: 540 QSQTSKPVKGMLTGPVTILNWSFPREDISLKASTLQLALAVQEEVLDLEKAGIKIIQIDE 599
Query: 84 WSKFEYLYMTKSLAHRQVLKQQLYSFKMVKSKVVVELLVEFNKIIGDFENI 134
+ E L + KS +++ L + +F++V SKV E + + +FE+I
Sbjct: 600 AALREKLPLRKSDWYKEYLDWAIPAFRLVHSKVKPETQIHTHMCYSEFEDI 650
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.317 0.131 0.381
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,012,818
Number of Sequences: 164201
Number of extensions: 624829
Number of successful extensions: 1652
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 11
Number of HSP's that attempted gapping in prelim test: 1651
Number of HSP's gapped (non-prelim): 12
length of query: 164
length of database: 59,974,054
effective HSP length: 102
effective length of query: 62
effective length of database: 43,225,552
effective search space: 2679984224
effective search space used: 2679984224
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC148218.5


