

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147875.9 + phase: 0
(122 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
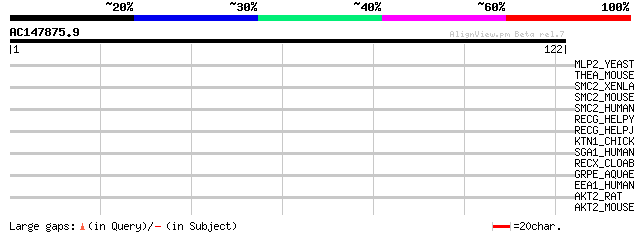
Score E
Sequences producing significant alignments: (bits) Value
MLP2_YEAST (P40457) MLP2 protein (Myosin-like protein 2) 30 0.96
THEA_MOUSE (Q8VHQ9) Brown fat inducible thioesterase (EC 3.1.2.-... 30 1.3
SMC2_XENLA (P50533) Structural maintenance of chromosome 2 (Chro... 29 1.6
SMC2_MOUSE (Q8CG48) Structural maintenance of chromosome 2-like ... 29 1.6
SMC2_HUMAN (O95347) Structural maintenance of chromosome 2-like ... 29 1.6
RECG_HELPY (O26051) ATP-dependent DNA helicase recG (EC 3.6.1.-) 29 1.6
RECG_HELPJ (Q9ZJA1) ATP-dependent DNA helicase recG (EC 3.6.1.-) 29 1.6
KTN1_CHICK (Q90631) Kinectin 29 1.6
SGA1_HUMAN (Q7Z6B7) SLIT-ROBO Rho GTPase activating protein 1 (s... 28 3.6
RECX_CLOAB (Q97GF7) Regulatory protein recX 28 4.8
GRPE_AQUAE (O66745) GrpE protein (HSP-70 cofactor) 27 8.1
EEA1_HUMAN (Q15075) Early endosome antigen 1 (Endosome-associate... 27 8.1
AKT2_RAT (P47197) RAC-beta serine/threonine-protein kinase (EC 2... 27 8.1
AKT2_MOUSE (Q60823) RAC-beta serine/threonine-protein kinase (EC... 27 8.1
>MLP2_YEAST (P40457) MLP2 protein (Myosin-like protein 2)
Length = 1679
Score = 30.0 bits (66), Expect = 0.96
Identities = 20/60 (33%), Positives = 32/60 (53%), Gaps = 1/60 (1%)
Query: 1 MKNLSSMIEESSYKAKAKMKDIETLEKKGKGQHGVMVVRRNENYEYAQVMRELEYLKKEL 60
+K + +E+ + AK+ D++T K + Q V R +E Y +M E+E LKKEL
Sbjct: 1314 LKAIKDKLEQDLHFENAKVIDLDTKLKAHELQ-SEDVSRDHEKDTYRTLMEEIESLKKEL 1372
>THEA_MOUSE (Q8VHQ9) Brown fat inducible thioesterase (EC 3.1.2.-)
(BFIT) (Adipose associated thioesterase)
Length = 594
Score = 29.6 bits (65), Expect = 1.3
Identities = 13/39 (33%), Positives = 22/39 (56%)
Query: 15 AKAKMKDIETLEKKGKGQHGVMVVRRNENYEYAQVMREL 53
+K K+K + L ++ K +HGV RR YA +++L
Sbjct: 158 SKVKLKQVIPLTEEEKTEHGVAAERRRMRLVYADTIKDL 196
>SMC2_XENLA (P50533) Structural maintenance of chromosome 2
(Chromosome-associated protein E) (Chromosome assembly
protein XCAP-E)
Length = 1203
Score = 29.3 bits (64), Expect = 1.6
Identities = 16/46 (34%), Positives = 29/46 (62%), Gaps = 1/46 (2%)
Query: 14 KAKAKMKDIET-LEKKGKGQHGVMVVRRNENYEYAQVMRELEYLKK 58
K +AK+K+I+T LE++ + R+ EY ++MRE+E+L +
Sbjct: 187 KKEAKLKEIQTILEEEITPTIHKLKEERSSYLEYQKIMREIEHLSR 232
>SMC2_MOUSE (Q8CG48) Structural maintenance of chromosome 2-like 1
protein (Chromosome-associated protein E) (XCAP-E
homolog) (FGF-inducible protein 16)
Length = 1191
Score = 29.3 bits (64), Expect = 1.6
Identities = 17/46 (36%), Positives = 29/46 (62%), Gaps = 1/46 (2%)
Query: 14 KAKAKMKDIET-LEKKGKGQHGVMVVRRNENYEYAQVMRELEYLKK 58
K +AK+K+I+T LE++ + R+ EY +VMRE+E+L +
Sbjct: 187 KKEAKLKEIKTILEEEITPTIQKLKEERSSYLEYQKVMREIEHLSR 232
>SMC2_HUMAN (O95347) Structural maintenance of chromosome 2-like 1
protein (Chromosome-associated protein E) (hCAP-E)
(XCAP-E homolog) (PRO0324)
Length = 1197
Score = 29.3 bits (64), Expect = 1.6
Identities = 17/46 (36%), Positives = 29/46 (62%), Gaps = 1/46 (2%)
Query: 14 KAKAKMKDIET-LEKKGKGQHGVMVVRRNENYEYAQVMRELEYLKK 58
K +AK+K+I+T LE++ + R+ EY +VMRE+E+L +
Sbjct: 187 KKEAKLKEIKTILEEEITPTIQKLKEERSSYLEYQKVMREIEHLSR 232
>RECG_HELPY (O26051) ATP-dependent DNA helicase recG (EC 3.6.1.-)
Length = 623
Score = 29.3 bits (64), Expect = 1.6
Identities = 17/56 (30%), Positives = 27/56 (47%)
Query: 3 NLSSMIEESSYKAKAKMKDIETLEKKGKGQHGVMVVRRNENYEYAQVMRELEYLKK 58
N+ S++E KD+ LE+ G GV+ V E YA+V++ Y K+
Sbjct: 12 NVKSLLEALLVYTPKGYKDLNLLERFETGLSGVLEVGILEKRNYAKVLKIFAYSKR 67
>RECG_HELPJ (Q9ZJA1) ATP-dependent DNA helicase recG (EC 3.6.1.-)
Length = 623
Score = 29.3 bits (64), Expect = 1.6
Identities = 17/56 (30%), Positives = 27/56 (47%)
Query: 3 NLSSMIEESSYKAKAKMKDIETLEKKGKGQHGVMVVRRNENYEYAQVMRELEYLKK 58
N+ S++E KD+ LE+ G GV+ V E YA+V++ Y K+
Sbjct: 12 NVKSLLEALLVYTPKGYKDLSLLERFETGLSGVLEVGILEKKNYAKVLKIFAYSKR 67
>KTN1_CHICK (Q90631) Kinectin
Length = 1364
Score = 29.3 bits (64), Expect = 1.6
Identities = 17/62 (27%), Positives = 32/62 (51%)
Query: 1 MKNLSSMIEESSYKAKAKMKDIETLEKKGKGQHGVMVVRRNENYEYAQVMRELEYLKKEL 60
M++ + +E + K ++K++ETL+K+ + + E Y +REL+ L EL
Sbjct: 1184 MRSSVASLEHEVERLKEEIKEVETLKKEREHLESELEKAEIERSTYVSEVRELKDLLTEL 1243
Query: 61 FK 62
K
Sbjct: 1244 QK 1245
>SGA1_HUMAN (Q7Z6B7) SLIT-ROBO Rho GTPase activating protein 1
(srGAP1) (Rho-GTPase-activating protein 13)
Length = 1085
Score = 28.1 bits (61), Expect = 3.6
Identities = 15/55 (27%), Positives = 31/55 (56%), Gaps = 2/55 (3%)
Query: 7 MIEESSYKAKAKMKDIETLEKKGKGQHG--VMVVRRNENYEYAQVMRELEYLKKE 59
M S A++K+K+ E E+K G+ G V +R E ++ ++++E +K++
Sbjct: 170 MYHAESISAESKLKEAEKQEEKQIGRSGDPVFHIRLEERHQRRSSVKKIEKMKEK 224
>RECX_CLOAB (Q97GF7) Regulatory protein recX
Length = 214
Score = 27.7 bits (60), Expect = 4.8
Identities = 16/55 (29%), Positives = 29/55 (52%), Gaps = 1/55 (1%)
Query: 4 LSSMIEESSYKAKAKMKDIETLEKKGKGQHGVMVVRRNENYEYAQVMRELEYLKK 58
L +IEE +Y AK + + +EK K + + V R + + + R +E+L+K
Sbjct: 52 LQDIIEEDNY-LYAKSRALSFIEKTFKTEKEIRVKLREKEFNEKIIERVIEFLRK 105
>GRPE_AQUAE (O66745) GrpE protein (HSP-70 cofactor)
Length = 182
Score = 26.9 bits (58), Expect = 8.1
Identities = 15/57 (26%), Positives = 27/57 (47%), Gaps = 10/57 (17%)
Query: 2 KNLSSMIEESSYKAKAKMKDIETLEKKGKGQHGVMVVRRNENYEYAQVMRELEYLKK 58
K + +EE + K + I+ LE + +N N A++ RE++YLK+
Sbjct: 6 KEIEKELEELRKREKELEEKIQKLE----------TIAKNSNLRVAELQREIDYLKE 52
>EEA1_HUMAN (Q15075) Early endosome antigen 1 (Endosome-associated
protein p162) (Zinc finger FYVE domain containing
protein 2)
Length = 1411
Score = 26.9 bits (58), Expect = 8.1
Identities = 19/63 (30%), Positives = 31/63 (49%), Gaps = 2/63 (3%)
Query: 1 MKNLSSMIEESSYKAKAKMKDIETLEKKGKGQHGVMVVRRNENYEYAQVMRELEYLKKEL 60
+KN S E+ + K KA + D+E K+ K H + V N E ++ + LE K+
Sbjct: 869 LKNSKSEFEKENQKGKAAILDLEKTCKELK--HQLQVQMENTLKEQKELKKSLEKEKEAS 926
Query: 61 FKL 63
+L
Sbjct: 927 HQL 929
>AKT2_RAT (P47197) RAC-beta serine/threonine-protein kinase (EC
2.7.1.37) (RAC-PK-beta) (Protein kinase Akt-2) (Protein
kinase B, beta) (PKB beta)
Length = 481
Score = 26.9 bits (58), Expect = 8.1
Identities = 18/60 (30%), Positives = 33/60 (55%), Gaps = 5/60 (8%)
Query: 5 SSMIEESSYKAKAK--MKDIETLEKKGKGQHGVMVVRRNE---NYEYAQVMRELEYLKKE 59
S M+E + KA+AK M D + L+ GKG G +++ R + Y +++R+ + K+
Sbjct: 133 SEMMEVAVSKARAKVTMNDFDYLKLLGKGTFGKVILVREKATGRYYAMKILRKEVIIAKD 192
>AKT2_MOUSE (Q60823) RAC-beta serine/threonine-protein kinase (EC
2.7.1.37) (RAC-PK-beta) (Protein kinase Akt-2) (Protein
kinase B, beta) (PKB beta)
Length = 481
Score = 26.9 bits (58), Expect = 8.1
Identities = 18/60 (30%), Positives = 33/60 (55%), Gaps = 5/60 (8%)
Query: 5 SSMIEESSYKAKAK--MKDIETLEKKGKGQHGVMVVRRNE---NYEYAQVMRELEYLKKE 59
S M+E + KA+AK M D + L+ GKG G +++ R + Y +++R+ + K+
Sbjct: 133 SEMMEVAVNKARAKVTMNDFDYLKLLGKGTFGKVILVREKATGRYYAMKILRKEVIIAKD 192
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.313 0.129 0.347
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,467,983
Number of Sequences: 164201
Number of extensions: 232068
Number of successful extensions: 746
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 736
Number of HSP's gapped (non-prelim): 21
length of query: 122
length of database: 59,974,054
effective HSP length: 98
effective length of query: 24
effective length of database: 43,882,356
effective search space: 1053176544
effective search space used: 1053176544
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 58 (26.9 bits)
Medicago: description of AC147875.9


