

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
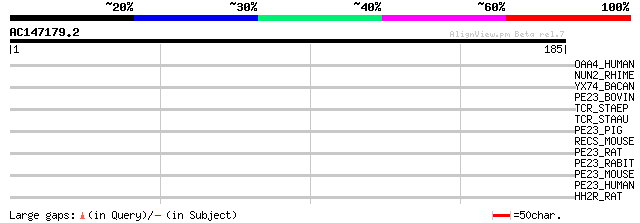
Query= AC147179.2 + phase: 0
(185 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
OAA4_HUMAN (Q9H209) Olfactory receptor 10A4 (HP2) (Olfactory rec... 35 0.13
NUN2_RHIME (P56911) NADH-quinone oxidoreductase chain N 2 (EC 1.... 35 0.13
YX74_BACAN (Q81N43) Hypothetical UPF0078 protein BA3374/GBAA3374... 32 1.1
PE23_BOVIN (P34979) Prostaglandin E2 receptor, EP3 subtype (Pros... 31 1.9
TCR_STAEP (P62967) Tetracycline resistance protein 30 2.4
TCR_STAAU (P02983) Tetracycline resistance protein (Tetracycline... 30 2.4
PE23_PIG (P50131) Prostaglandin E2 receptor, EP3 subtype (Prosta... 29 7.1
RECS_MOUSE (Q8BJZ3) RECS1 protein (Responsive to centrifugal for... 28 9.2
PE23_RAT (P34980) Prostaglandin E2 receptor, EP3 subtype (Prosta... 28 9.2
PE23_RABIT (P46069) Prostaglandin E2 receptor, EP3 subtype (Pros... 28 9.2
PE23_MOUSE (P30557) Prostaglandin E2 receptor, EP3 subtype (Pros... 28 9.2
PE23_HUMAN (P43115) Prostaglandin E2 receptor, EP3 subtype (Pros... 28 9.2
HH2R_RAT (P25102) Histamine H2 receptor (H2R) (Gastric receptor I) 28 9.2
>OAA4_HUMAN (Q9H209) Olfactory receptor 10A4 (HP2) (Olfactory
receptor-like protein JCG5)
Length = 315
Score = 34.7 bits (78), Expect = 0.13
Identities = 21/62 (33%), Positives = 30/62 (47%), Gaps = 5/62 (8%)
Query: 28 YVLLGAASSCIFLTLSLRLVPSVCGFFLILLHIFTIAGAVSGCAAVGANRWYSAHMVATV 87
+ GAA C+ T++ ++C LH I G +S CA + A W+S VATV
Sbjct: 105 FFFFGAAECCLLATMAYDRYVAICD----PLHYPVIMGHIS-CAQLAAASWFSGFSVATV 159
Query: 88 LT 89
T
Sbjct: 160 QT 161
>NUN2_RHIME (P56911) NADH-quinone oxidoreductase chain N 2 (EC
1.6.99.5) (NADH dehydrogenase I, chain N 2) (NDH-1,
chain N 2)
Length = 479
Score = 34.7 bits (78), Expect = 0.13
Identities = 31/106 (29%), Positives = 45/106 (42%), Gaps = 12/106 (11%)
Query: 66 AVSGCAAVGANRWYSAHMVATVLTAIFQGSVSVLVFTRTSDFLGELQSYVREEDGSVILK 125
A+ G A G + S + + T + G ++ R DF GE+ ED + K
Sbjct: 311 AIFGVVAGGQDGIASVMLYLLIYTFMNLGIFGAVIMMRNGDFSGEV-----IEDYAGFAK 365
Query: 126 LSGGLAIL----IFCLEWVVLTLAFFLKYY---ACVEGGNTCRTVV 164
GLA+L +F L + T FF K+Y A VE G V+
Sbjct: 366 FHPGLALLMLLYLFSLAGIPPTAGFFAKFYVLVALVERGFVALAVI 411
>YX74_BACAN (Q81N43) Hypothetical UPF0078 protein
BA3374/GBAA3374/BAS3128
Length = 190
Score = 31.6 bits (70), Expect = 1.1
Identities = 17/45 (37%), Positives = 24/45 (52%)
Query: 29 VLLGAASSCIFLTLSLRLVPSVCGFFLILLHIFTIAGAVSGCAAV 73
+L G + + L LS + P VCG IL HIF++ + G AV
Sbjct: 60 ILKGTFACLLPLILSSTINPIVCGLLAILGHIFSVFASFKGGKAV 104
>PE23_BOVIN (P34979) Prostaglandin E2 receptor, EP3 subtype
(Prostanoid EP3 receptor) (PGE receptor, EP3 subtype)
Length = 417
Score = 30.8 bits (68), Expect = 1.9
Identities = 19/60 (31%), Positives = 30/60 (49%), Gaps = 8/60 (13%)
Query: 50 VCGFFLILLHIFTIAG-------AVSGCAAVGANRWYSAHMVATVLTAIFQGS-VSVLVF 101
+C FF + + +F ++ AV A A WYS+HM +V A+ G ++VL F
Sbjct: 128 LCTFFGLTMTVFGLSSLFIASAMAVERALATRAPHWYSSHMKTSVTRAVLLGVWLAVLAF 187
>TCR_STAEP (P62967) Tetracycline resistance protein
Length = 459
Score = 30.4 bits (67), Expect = 2.4
Identities = 27/104 (25%), Positives = 49/104 (46%), Gaps = 13/104 (12%)
Query: 47 VPSVCGFFLILLHIFTIAGAVSGCAAVGANRWYSAHMVATVLTAIFQGSVSVLVFTRTSD 106
+P + G F L +AG +S + Y ++ + IF G++SV+VF
Sbjct: 256 IPFMLGLFSGGLIFSIVAGFISMVPYM-MKTIYHVNVATIGNSVIFPGTMSVIVF----- 309
Query: 107 FLGELQSYVREEDGSVILKLSGGLAILIFCLEWVVLTLAFFLKY 150
G ++ + GS+ + + G L+I I LT+AFF+++
Sbjct: 310 --GYFGGFLVDRKGSLFVFILGSLSISI-----SFLTIAFFVEF 346
>TCR_STAAU (P02983) Tetracycline resistance protein (Tetracycline
efflux protein)
Length = 459
Score = 30.4 bits (67), Expect = 2.4
Identities = 27/104 (25%), Positives = 49/104 (46%), Gaps = 13/104 (12%)
Query: 47 VPSVCGFFLILLHIFTIAGAVSGCAAVGANRWYSAHMVATVLTAIFQGSVSVLVFTRTSD 106
+P + G F L +AG +S + Y ++ + IF G++SV+VF
Sbjct: 256 IPFMLGLFSGGLIFSIVAGFISMVPYM-MKTIYHVNVATIGNSVIFPGTMSVIVF----- 309
Query: 107 FLGELQSYVREEDGSVILKLSGGLAILIFCLEWVVLTLAFFLKY 150
G ++ + GS+ + + G L+I I LT+AFF+++
Sbjct: 310 --GYFGGFLVDRKGSLFVFILGSLSISI-----SFLTIAFFVEF 346
>PE23_PIG (P50131) Prostaglandin E2 receptor, EP3 subtype
(Prostanoid EP3 receptor) (PGE receptor, EP3 subtype)
Length = 373
Score = 28.9 bits (63), Expect = 7.1
Identities = 18/60 (30%), Positives = 29/60 (48%), Gaps = 8/60 (13%)
Query: 50 VCGFFLILLHIFTIAG-------AVSGCAAVGANRWYSAHMVATVLTAIFQGS-VSVLVF 101
+C FF + + F ++ AV A+ A WYS+HM + A+ G ++VL F
Sbjct: 106 LCTFFGLTMTAFGLSSLFIASAMAVERALAIRAPHWYSSHMKTSATRAVLLGVWLAVLAF 165
>RECS_MOUSE (Q8BJZ3) RECS1 protein (Responsive to centrifugal force
and shear stress gene 1 protein)
Length = 309
Score = 28.5 bits (62), Expect = 9.2
Identities = 19/54 (35%), Positives = 31/54 (57%), Gaps = 3/54 (5%)
Query: 55 LILLHIFTIA-GAVSGCAAVGANRWYSAHMVATVLTAIFQGSVSVLVFTRTSDF 107
+ILL IFT+A G V+G + + A ++A ++TA+ SV++ F DF
Sbjct: 163 IILLTIFTLALGFVTG--TISSMYENKAVIIAMIITAVVSISVTIFCFQTKVDF 214
>PE23_RAT (P34980) Prostaglandin E2 receptor, EP3 subtype
(Prostanoid EP3 receptor) (PGE receptor, EP3 subtype)
Length = 365
Score = 28.5 bits (62), Expect = 9.2
Identities = 16/59 (27%), Positives = 27/59 (45%), Gaps = 7/59 (11%)
Query: 50 VCGFFLILLHIFTIAG-------AVSGCAAVGANRWYSAHMVATVLTAIFQGSVSVLVF 101
+C FF + + +F ++ AV A+ A WY++HM + +SVL F
Sbjct: 106 LCTFFGLTMTVFGLSSLLVASAMAVERALAIRAPHWYASHMKTRATPVLLGVWLSVLAF 164
>PE23_RABIT (P46069) Prostaglandin E2 receptor, EP3 subtype
(Prostanoid EP3 receptor) (PGE receptor, EP3 subtype)
Length = 411
Score = 28.5 bits (62), Expect = 9.2
Identities = 17/60 (28%), Positives = 29/60 (48%), Gaps = 8/60 (13%)
Query: 50 VCGFFLILLHIFTIAG-------AVSGCAAVGANRWYSAHMVATVLTAIFQGS-VSVLVF 101
+C FF + + +F ++ AV A+ A WY++HM A+ G ++VL F
Sbjct: 125 LCTFFGLTMTVFGLSSLFIASAMAVERALAIRAPHWYASHMKTRATRAVLLGVWLAVLAF 184
>PE23_MOUSE (P30557) Prostaglandin E2 receptor, EP3 subtype
(Prostanoid EP3 receptor) (PGE receptor, EP3 subtype)
Length = 365
Score = 28.5 bits (62), Expect = 9.2
Identities = 16/59 (27%), Positives = 27/59 (45%), Gaps = 7/59 (11%)
Query: 50 VCGFFLILLHIFTIAG-------AVSGCAAVGANRWYSAHMVATVLTAIFQGSVSVLVF 101
+C FF + + +F ++ AV A+ A WY++HM + +SVL F
Sbjct: 106 LCTFFGLTMTVFGLSSLLVASAMAVERALAIRAPHWYASHMKTRATPVLLGVWLSVLAF 164
>PE23_HUMAN (P43115) Prostaglandin E2 receptor, EP3 subtype
(Prostanoid EP3 receptor) (PGE receptor, EP3 subtype)
Length = 390
Score = 28.5 bits (62), Expect = 9.2
Identities = 17/60 (28%), Positives = 29/60 (48%), Gaps = 8/60 (13%)
Query: 50 VCGFFLILLHIFTIAG-------AVSGCAAVGANRWYSAHMVATVLTAIFQGS-VSVLVF 101
+C FF + + +F ++ AV A+ A WY++HM A+ G ++VL F
Sbjct: 129 LCTFFGLTMTVFGLSSLFIASAMAVERALAIRAPHWYASHMKTRATRAVLLGVWLAVLAF 188
>HH2R_RAT (P25102) Histamine H2 receptor (H2R) (Gastric receptor
I)
Length = 358
Score = 28.5 bits (62), Expect = 9.2
Identities = 19/46 (41%), Positives = 25/46 (54%), Gaps = 3/46 (6%)
Query: 32 GAASSCIFLTLSLRLVPSVCGFFLILLHIFTIAGAVSGCAAVGANR 77
G SC +++L++ SV LIL+ TIAG V C AV NR
Sbjct: 5 GTVHSCCLDSMALKVTISVVLTTLILI---TIAGNVVVCLAVSLNR 47
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.328 0.142 0.449
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,981,111
Number of Sequences: 164201
Number of extensions: 767728
Number of successful extensions: 2352
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 12
Number of HSP's that attempted gapping in prelim test: 2349
Number of HSP's gapped (non-prelim): 14
length of query: 185
length of database: 59,974,054
effective HSP length: 104
effective length of query: 81
effective length of database: 42,897,150
effective search space: 3474669150
effective search space used: 3474669150
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 62 (28.5 bits)
Medicago: description of AC147179.2


