

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146788.10 - phase: 0
(496 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
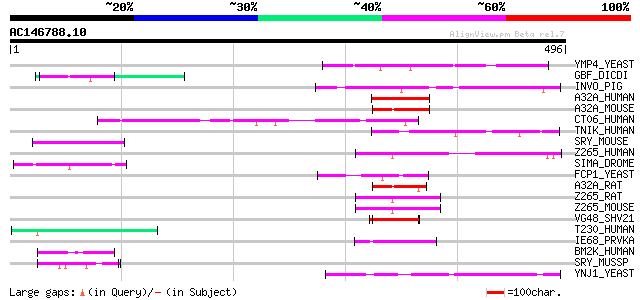
Score E
Sequences producing significant alignments: (bits) Value
YMP4_YEAST (Q04347) Hypothetical 60.1 kDa protein in SEC59-ERG5 ... 51 6e-06
GBF_DICDI (P36417) G-box binding factor (GBF) 50 1e-05
INVO_PIG (P18175) Involucrin 49 3e-05
A32A_HUMAN (P39687) Acidic leucine-rich nuclear phosphoprotein 3... 49 4e-05
A32A_MOUSE (O35381) Acidic leucine-rich nuclear phosphoprotein 3... 48 5e-05
CT06_HUMAN (Q9H501) Hypothetical protein C20orf6 48 7e-05
TNIK_HUMAN (Q9UKE5) TRAF2 and NCK interacting kinase (EC 2.7.1.37) 47 9e-05
SRY_MOUSE (Q05738) Sex-determining region Y protein (Testis-dete... 47 1e-04
Z265_HUMAN (O95218) Zinc finger protein 265 (Zinc finger, splicing) 47 1e-04
SIMA_DROME (Q24167) Similar protein 47 1e-04
FCP1_YEAST (Q03254) RNA polymerase II subunit A C-terminal domai... 47 1e-04
A32A_RAT (P49911) Acidic leucine-rich nuclear phosphoprotein 32 ... 47 1e-04
Z265_RAT (O35986) Zinc finger protein 265 (Zinc finger, splicing) 46 3e-04
Z265_MOUSE (Q9R020) Zinc finger protein 265 (Zinc finger, splici... 46 3e-04
VG48_SHV21 (Q01033) Hypothetical gene 48 protein 46 3e-04
T230_HUMAN (Q93074) Thyroid hormone receptor-associated protein ... 46 3e-04
IE68_PRVKA (P24827) Immediate-early protein RSP40 46 3e-04
BM2K_HUMAN (Q9NSY1) BMP-2 inducible protein kinase (EC 2.7.1.37)... 46 3e-04
SRY_MUSSP (Q62563) Sex-determining region Y protein (Testis-dete... 45 3e-04
YNJ1_YEAST (P53935) Hypothetical 141.5 kDa protein in YPT53-RHO2... 45 6e-04
>YMP4_YEAST (Q04347) Hypothetical 60.1 kDa protein in SEC59-ERG5
intergenic region
Length = 519
Score = 51.2 bits (121), Expect = 6e-06
Identities = 56/220 (25%), Positives = 89/220 (40%), Gaps = 28/220 (12%)
Query: 280 QQQHQHQHQQQNQQHCFHSSENGVGSLGVSRGEGLRMLKIGSGYVEEEDDE------DEE 333
Q Q + N+ E +GS G GEG L Y E D+E DE+
Sbjct: 276 QTQSNKETTSDNEDLLIKEYEGMLGSSG-DEGEGGGYLNPNINYNEVTDEEPSEASSDED 334
Query: 334 DESEDFSDEGEDESGEGCSKGHI-----------NDQDEEENDGKPSRKRARKGGFSFPR 382
D E FSD E+E K H ND +EE ++ + K +K +
Sbjct: 335 DSDERFSDSEENEPRRKKPKLHNLPELMAGYYSGNDTEEESDEDNKNVKGKKKKRDTAED 394
Query: 383 SSSSTQLVNQMNNEISGVFQDGGKSTWEKKQWIRNRIMQLEEQKIGYESQAFQLEKQRLK 442
++ Q+ N+ + + Q + WEKK + + +Q E +K ++E ++ +
Sbjct: 395 RTAREQMSNEPKRK-NRRGQRARRKIWEKKYGSQAKHVQRELEK--------EMEDRKQR 445
Query: 443 WARYSSK-KEREMERAKLENERRRLENERMVLLIRKKELE 481
Y ++ +RE + A LE R R +R KKE E
Sbjct: 446 QIEYEARVAKREAKAASLEASRSREREDRRTETNNKKEKE 485
>GBF_DICDI (P36417) G-box binding factor (GBF)
Length = 708
Score = 50.1 bits (118), Expect = 1e-05
Identities = 26/67 (38%), Positives = 32/67 (46%), Gaps = 1/67 (1%)
Query: 27 QQQQQQKQQNLPNQQQQNPHQLHHSQVVSYAPQHHDTDTNQHHHHSMKHGFPPFSSSKNK 86
QQQ Q+QQ P QQ Q HH Q + QHH Q HH M+H +++
Sbjct: 136 QQQHHQQQQQQPQHHQQMQQQQHHQQ-MQQQQQHHQQMQQQQHHQQMQHHQLQQHQHQHQ 194
Query: 87 QQQQSSQ 93
QQQQ Q
Sbjct: 195 QQQQQQQ 201
Score = 48.5 bits (114), Expect = 4e-05
Identities = 31/141 (21%), Positives = 54/141 (37%), Gaps = 13/141 (9%)
Query: 24 PLHQQQQQQKQQNLPNQQQQNPHQLHHSQVVSYAPQHHDTDTNQHHH--------HSMKH 75
P H QQ QQ+Q + QQQQ HQ Q QHH +QH H H +H
Sbjct: 147 PQHHQQMQQQQHHQQMQQQQQHHQQMQQQQHHQQMQHHQLQQHQHQHQQQQQQQQHQQQH 206
Query: 76 GFPPFSSSKNKQQQQSSQMSDEDEPNFPAEESSGGDPKRKISPWQRMKWTDTMVRLLIMA 135
+ +QQQQ Q + + P ++ ++ Q+ + ++++
Sbjct: 207 -----HQQQQQQQQQHHQQQQHHQHSQPQQQHQHNQQQQHQHNQQQHQQQQNQIQMVPQQ 261
Query: 136 VYYIGDEAGSEGTDPNKKKSS 156
+ + + + N S+
Sbjct: 262 PQSLSNSGNNNNNNNNNNNSN 282
Score = 43.1 bits (100), Expect = 0.002
Identities = 25/64 (39%), Positives = 28/64 (43%), Gaps = 3/64 (4%)
Query: 30 QQQKQQNLPNQQQQNPHQLHHSQVVSYAPQHHDTDTNQHHHHSMKHGFPPFSSSKNKQQQ 89
QQQ QQN P QQN Q HH Q PQHH Q HH M+ + +Q
Sbjct: 122 QQQAQQNQP--PQQNQQQQHHQQ-QQQQPQHHQQMQQQQHHQQMQQQQQHHQQMQQQQHH 178
Query: 90 QSSQ 93
Q Q
Sbjct: 179 QQMQ 182
Score = 33.5 bits (75), Expect = 1.3
Identities = 21/90 (23%), Positives = 43/90 (47%), Gaps = 3/90 (3%)
Query: 26 HQQQQQQKQQNLPNQQQQNPHQLHHSQVVSYAPQHHDTDTNQHHHHSMKHGFPPFSSSKN 85
H QQQQ + +QQQQN Q+ Q S + ++ + N ++++S + +++ +
Sbjct: 235 HNQQQQHQHNQQQHQQQQNQIQMVPQQPQSLSNSGNNNNNNNNNNNSNNNNNNNNNNNSH 294
Query: 86 KQQQQSSQMSDEDEPNFPAEESSGGDPKRK 115
+ + ++ N P+ + G KRK
Sbjct: 295 QLNNLTLSQNNTSGSNTPSPSTKG---KRK 321
Score = 32.7 bits (73), Expect = 2.2
Identities = 16/48 (33%), Positives = 20/48 (41%)
Query: 247 YHNSCGHGGVATSNVQHQVEGGSGTTTPPQNQPQQQHQHQHQQQNQQH 294
+H H + QHQ + +Q QQQ Q QH QQ Q H
Sbjct: 177 HHQQMQHHQLQQHQHQHQQQQQQQQHQQQHHQQQQQQQQQHHQQQQHH 224
Score = 32.3 bits (72), Expect = 2.9
Identities = 13/21 (61%), Positives = 13/21 (61%)
Query: 274 PPQNQPQQQHQHQHQQQNQQH 294
PPQ QQQH Q QQQ Q H
Sbjct: 130 PPQQNQQQQHHQQQQQQPQHH 150
Score = 32.0 bits (71), Expect = 3.8
Identities = 12/18 (66%), Positives = 15/18 (82%)
Query: 277 NQPQQQHQHQHQQQNQQH 294
+QPQQQHQH QQQ+Q +
Sbjct: 227 SQPQQQHQHNQQQQHQHN 244
Score = 30.8 bits (68), Expect = 8.4
Identities = 14/33 (42%), Positives = 15/33 (45%)
Query: 262 QHQVEGGSGTTTPPQNQPQQQHQHQHQQQNQQH 294
QHQ + Q QQQ HQH Q QQH
Sbjct: 201 QHQQQHHQQQQQQQQQHHQQQQHHQHSQPQQQH 233
>INVO_PIG (P18175) Involucrin
Length = 347
Score = 48.9 bits (115), Expect = 3e-05
Identities = 46/223 (20%), Positives = 93/223 (41%), Gaps = 20/223 (8%)
Query: 274 PPQNQPQQQHQHQHQQQNQQHCFHSSENGVGSLGVSRGEGLRMLKIGSGYVEEEDDEDEE 333
P +PQQQ H QQQ QQ E+ V L V + + + + +V+++ + E
Sbjct: 77 PQPQEPQQQELHVEQQQQQQ------ESQVQELHVDQQQQQQESQEQELHVDQQQQQQES 130
Query: 334 DESEDFSDEGEDESG--EGCSKGHINDQDEEENDGKPSRKRARKGGFSFPRSSSSTQLVN 391
E E D+ + + + GH Q E + ++ S V+
Sbjct: 131 QEQELHVDQQQQQESQVQELHVGHHQQQQESQEQELHVDHHQQQ-----QESQEQELHVD 185
Query: 392 QMNNEISGVFQDGGKSTWEKKQWIRNRIMQLEEQKIGYESQAFQLEKQRLKWARYSSKKE 451
Q + Q+ + Q + + Q +E + + Q + + Q L ++E
Sbjct: 186 QQQQQ-----QESQEQELHVDQQQQQQESQEQELHVDHHQQQQESQVQELHVDHQQQQQE 240
Query: 452 REMERAKLENERRRLENERMVLLI--RKKELELMHIQQQQQQQ 492
+ + ++ +++ E++ L + +++EL++ +QQQQQQQ
Sbjct: 241 SQEQELHVDQHQQQQESQEQELHVDQQQQELQVQEVQQQQQQQ 283
Score = 33.9 bits (76), Expect = 0.99
Identities = 17/74 (22%), Positives = 38/74 (50%), Gaps = 10/74 (13%)
Query: 421 QLEEQKIGYESQAFQLEKQRLKWARYSSKKEREMERAKLENERRRLENERMVLLIRKKEL 480
Q++E +G+ Q + ++Q L + ++E + + ++ ++++ E++ L
Sbjct: 146 QVQELHVGHHQQQQESQEQELHVDHHQQQQESQEQELHVDQQQQQQESQEQEL------- 198
Query: 481 ELMHIQQQQQQQHS 494
H+ QQQQQQ S
Sbjct: 199 ---HVDQQQQQQES 209
Score = 33.9 bits (76), Expect = 0.99
Identities = 21/69 (30%), Positives = 30/69 (43%), Gaps = 2/69 (2%)
Query: 27 QQQQQQKQQNLPNQQQQNPHQLHHSQVVSYAPQHHDTDTNQH--HHHSMKHGFPPFSSSK 84
QQQ+ Q+Q+ +QQQQ Q+ V + Q + H HH +
Sbjct: 126 QQQESQEQELHVDQQQQQESQVQELHVGHHQQQQESQEQELHVDHHQQQQESQEQELHVD 185
Query: 85 NKQQQQSSQ 93
+QQQQ SQ
Sbjct: 186 QQQQQQESQ 194
Score = 33.5 bits (75), Expect = 1.3
Identities = 20/72 (27%), Positives = 32/72 (43%), Gaps = 4/72 (5%)
Query: 26 HQQQQQQKQQNL----PNQQQQNPHQLHHSQVVSYAPQHHDTDTNQHHHHSMKHGFPPFS 81
HQQQQ+ ++Q L QQQ++ Q H + + + + HH +
Sbjct: 171 HQQQQESQEQELHVDQQQQQQESQEQELHVDQQQQQQESQEQELHVDHHQQQQESQVQEL 230
Query: 82 SSKNKQQQQSSQ 93
++QQQQ SQ
Sbjct: 231 HVDHQQQQQESQ 242
Score = 32.3 bits (72), Expect = 2.9
Identities = 20/82 (24%), Positives = 32/82 (38%), Gaps = 4/82 (4%)
Query: 25 LHQQQQQQKQQNLPNQQQQNPHQLHHSQVVSYAPQHHDTDTNQHHHHSMKHGFPPFSSSK 84
LH QQQQ+Q++ Q+Q H HH Q Q D Q S + +
Sbjct: 198 LHVDQQQQQQES----QEQELHVDHHQQQQESQVQELHVDHQQQQQESQEQELHVDQHQQ 253
Query: 85 NKQQQQSSQMSDEDEPNFPAEE 106
++ Q+ D+ + +E
Sbjct: 254 QQESQEQELHVDQQQQELQVQE 275
Score = 31.6 bits (70), Expect = 4.9
Identities = 19/73 (26%), Positives = 29/73 (39%), Gaps = 2/73 (2%)
Query: 26 HQQQQQQKQQNLPNQQQQNPHQLHHSQVVSYAPQHHDTDTNQHHHHSMKHGFPPFSSSKN 85
HQQQQ+ + Q L QQ + ++ + QH +Q + +
Sbjct: 219 HQQQQESQVQELHVDHQQQQQESQEQEL--HVDQHQQQQESQEQELHVDQQQQELQVQEV 276
Query: 86 KQQQQSSQMSDED 98
+QQQQ Q ED
Sbjct: 277 QQQQQQQQEQQED 289
Score = 31.2 bits (69), Expect = 6.4
Identities = 19/76 (25%), Positives = 30/76 (39%), Gaps = 8/76 (10%)
Query: 22 NIPLHQQQQQQKQQNLPNQQQQNPHQLHHSQVVSYAPQHHDTDTNQHHHHSMKHGFPPFS 81
++ HQQQQ+ ++Q L QQQ Q+ Q Q D + H +
Sbjct: 247 HVDQHQQQQESQEQELHVDQQQQELQVQEVQQQQQQQQEQQEDHQKAEHLEQEEA----- 301
Query: 82 SSKNKQQQQSSQMSDE 97
++QQ Q+ E
Sbjct: 302 ---QREQQLKGQLEQE 314
Score = 30.8 bits (68), Expect = 8.4
Identities = 27/150 (18%), Positives = 54/150 (36%), Gaps = 10/150 (6%)
Query: 263 HQVEGGSGTTTPPQNQPQQQHQHQHQQQNQQHCFHSSENGVGSLGVSRGEGLRMLKIGSG 322
HQ + S +Q QQQ + Q Q+ + E+ L V + + ++
Sbjct: 171 HQQQQESQEQELHVDQQQQQQESQEQELHVDQQQQQQESQEQELHVDHHQQQQESQVQEL 230
Query: 323 YVEEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEE----------ENDGKPSRKR 372
+V+ + + E E E D+ + + + H++ Q +E + +
Sbjct: 231 HVDHQQQQQESQEQELHVDQHQQQQESQEQELHVDQQQQELQVQEVQQQQQQQQEQQEDH 290
Query: 373 ARKGGFSFPRSSSSTQLVNQMNNEISGVFQ 402
+ + QL Q+ E GV+Q
Sbjct: 291 QKAEHLEQEEAQREQQLKGQLEQEKKGVYQ 320
>A32A_HUMAN (P39687) Acidic leucine-rich nuclear phosphoprotein 32
family member A (Potent heat-stable protein phosphatase
2A inhibitor I1PP2A) (Acidic nuclear phosphoprotein
pp32) (Cerebellar leucine rich acidic nuclear protein)
(HLA-DR associated protei
Length = 249
Score = 48.5 bits (114), Expect = 4e-05
Identities = 23/53 (43%), Positives = 36/53 (67%), Gaps = 1/53 (1%)
Query: 324 VEEEDDEDEEDESEDFSDEGEDESG-EGCSKGHINDQDEEENDGKPSRKRARK 375
VE+E+DEDEE+E E+ GE+E EG + G ++D+++EE G+ R + RK
Sbjct: 186 VEDEEDEDEEEEGEEEDVSGEEEEDEEGYNDGEVDDEEDEEELGEEERGQKRK 238
>A32A_MOUSE (O35381) Acidic leucine-rich nuclear phosphoprotein 32
family member A (Potent heat-stable protein phosphatase
2A inhibitor I1PP2A) (Acidic nuclear phosphoprotein
pp32) (Cerebellar leucine rich acidic nuclear protein)
Length = 247
Score = 48.1 bits (113), Expect = 5e-05
Identities = 23/51 (45%), Positives = 34/51 (66%), Gaps = 1/51 (1%)
Query: 325 EEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEENDGKPSRKRARK 375
EEE+DE+EE E ED S E E+E EG + G ++D+++EE G+ + RK
Sbjct: 187 EEEEDEEEEGEEEDVSGE-EEEDEEGYNDGEVDDEEDEEEAGEEEGSQKRK 236
>CT06_HUMAN (Q9H501) Hypothetical protein C20orf6
Length = 851
Score = 47.8 bits (112), Expect = 7e-05
Identities = 71/298 (23%), Positives = 123/298 (40%), Gaps = 49/298 (16%)
Query: 79 PFSSSKNKQQQQSSQMSDEDEPNFPAEESSGGDPKR-KISPWQRMKWTDTMVRLLIMAVY 137
P S S + ++ +SD D N E+S K+ K Q K D+ +
Sbjct: 59 PISHSTTEDLKRFYDLSDSDS-NLSGEDSKALSQKKIKKKKTQTKKEIDSKNLVEKKKET 117
Query: 138 YIGDEAGSEG-TDPNKKKSSGLLQKKGKWKSVSRAMMEKGFYVSPQQCEDKFNDLNKRYK 196
+ GSE TD + ++ K+K S +SP++ +F NK+ K
Sbjct: 118 KKANHKGSENKTDLDNSIGIKKMKTSCKFKIDSN--------ISPKKDSKEFTQKNKKEK 169
Query: 197 RVNDILGKGTACRVVENQGLLDS----MDLSPKMK-DEVRKLLNS--KHLFFREMCAYHN 249
+ +I+ T + E Q LDS + SP+++ + R+ + S + + R+ Y N
Sbjct: 170 K--NIVQHTTDSSLEEKQRTLDSGTSEIVKSPRIECSKTRREMQSVVQLIMTRDSDGYEN 227
Query: 250 SCGHGGVATSNVQHQVEGGSGTTTPPQNQPQQQHQHQHQQQNQQHCFHSSENGVGSLGVS 309
S T + + + + ++ SEN + S+G +
Sbjct: 228 S----------------------TDGEMCDKDALEEDSESVSEIGSDEESENEITSVGRA 265
Query: 310 RGEGLRMLKIGSGYVEEEDDEDEEDESEDFSDEGEDESGEGCS--KGHINDQDEEEND 365
G+ GS EEED+++EEDE ED D+ + +SG + KG+I E+E+D
Sbjct: 266 SGDD-----DGSEDDEEEDEDEEEDEDEDSEDDDKSDSGPDLARGKGNIETSSEDEDD 318
>TNIK_HUMAN (Q9UKE5) TRAF2 and NCK interacting kinase (EC 2.7.1.37)
Length = 1360
Score = 47.4 bits (111), Expect = 9e-05
Identities = 51/178 (28%), Positives = 84/178 (46%), Gaps = 23/178 (12%)
Query: 324 VEEEDDEDEEDESEDFSDEGEDE-SGEGCSKGHINDQDEEEND-GKPSRKRARKGGFSFP 381
++ +D D + DE E E SG ++++EEEND G+PS G +
Sbjct: 300 IQLKDHIDRTKKKRGEKDETEYEYSG--------SEEEEEENDSGEPSSILNLPGESTLR 351
Query: 382 RSSSSTQLVNQMNNEI---SGVFQDGGKSTWEKKQWIRNRIMQLEEQKIGYESQAFQLEK 438
R QL N+ +E + Q ++ K+Q + R ++EEQK Q +LE+
Sbjct: 352 RDFLRLQLANKERSEALRRQQLEQQQRENEEHKRQLLAERQKRIEEQK----EQRRRLEE 407
Query: 439 QRLKWARYSSKKEREM-----ERAKLENERRRLENERMVLLIRKKELELMHIQQQQQQ 491
Q+ + ++ERE E+ + E ERRR E+E+ + R+ E E ++ QQQ
Sbjct: 408 QQRREKELRKQQEREQRRHYEEQMRREEERRRAEHEQEYIR-RQLEEEQRQLEILQQQ 464
>SRY_MOUSE (Q05738) Sex-determining region Y protein
(Testis-determining factor)
Length = 395
Score = 47.0 bits (110), Expect = 1e-04
Identities = 26/83 (31%), Positives = 36/83 (43%), Gaps = 1/83 (1%)
Query: 21 DNIPLHQQQQQQKQQNLPNQQQQNPHQLHHSQVVSYAPQHHDTDTNQ-HHHHSMKHGFPP 79
D+ QQQQQQ+QQ + QQ+ HH Q + HH Q H HH + F
Sbjct: 167 DHHQQQQQQQQQQQQFHDHHQQKQQFHDHHQQQQQFHDHHHHHQEQQFHDHHQQQQQFHD 226
Query: 80 FSSSKNKQQQQSSQMSDEDEPNF 102
+ +QQQQ + + F
Sbjct: 227 HQQQQQQQQQQQFHDHHQQKQQF 249
Score = 43.9 bits (102), Expect = 0.001
Identities = 23/74 (31%), Positives = 31/74 (41%), Gaps = 1/74 (1%)
Query: 26 HQQQQQQKQQNLPNQQQQNPHQLHHSQVVSYAPQHHDTDTNQHHHHSMKHGFPPFSSSKN 85
H QQQQQ + QQQ+ HH Q + HH H HH + F +
Sbjct: 294 HPQQQQQFHDHHHQQQQKQQFHDHHQQKQQFH-DHHQQKQQFHDHHQQQQQFHDHHQQQQ 352
Query: 86 KQQQQSSQMSDEDE 99
+QQQQ Q + +
Sbjct: 353 QQQQQQQQQFHDQQ 366
Score = 42.4 bits (98), Expect = 0.003
Identities = 27/79 (34%), Positives = 32/79 (40%), Gaps = 11/79 (13%)
Query: 26 HQQQQQ---QKQQNLPNQQQQNPHQLHHSQVVSYAPQHHDTDT-NQHHHHSMKHGF---- 77
HQQQQQ QQ QQQQ HH Q + H + HHHH + F
Sbjct: 159 HQQQQQFYDHHQQQQQQQQQQQQFHDHHQQKQQFHDHHQQQQQFHDHHHHHQEQQFHDHH 218
Query: 78 ---PPFSSSKNKQQQQSSQ 93
F + +QQQQ Q
Sbjct: 219 QQQQQFHDHQQQQQQQQQQ 237
Score = 42.0 bits (97), Expect = 0.004
Identities = 26/77 (33%), Positives = 31/77 (39%), Gaps = 6/77 (7%)
Query: 26 HQQQQQQKQQNL---PNQQQQNPHQLHHSQVVSYAPQHHDTDTNQHHHHSMKHGFPPFSS 82
HQQQQQQ+QQ +QQ+Q H HH Q H H H +H F
Sbjct: 227 HQQQQQQQQQQQFHDHHQQKQQFHDHHHHQQQQQFHDHQQQQQQFHDHQQQQHQFHDHPQ 286
Query: 83 SKNK---QQQQSSQMSD 96
K + QQ Q D
Sbjct: 287 QKQQFHDHPQQQQQFHD 303
Score = 41.2 bits (95), Expect = 0.006
Identities = 30/88 (34%), Positives = 33/88 (37%), Gaps = 16/88 (18%)
Query: 26 HQQQQ-------QQKQQ--NLPNQQQQNPHQLHHSQVVSYAPQHHDTDTNQHHHHSMKHG 76
HQQQQ QQKQQ + P QQQQ H Q HH H HH K
Sbjct: 274 HQQQQHQFHDHPQQKQQFHDHPQQQQQFHDHHHQQQQKQQFHDHHQQKQQFHDHHQQKQQ 333
Query: 77 F-------PPFSSSKNKQQQQSSQMSDE 97
F F +QQQQ Q +
Sbjct: 334 FHDHHQQQQQFHDHHQQQQQQQQQQQQQ 361
Score = 40.8 bits (94), Expect = 0.008
Identities = 24/76 (31%), Positives = 29/76 (37%), Gaps = 8/76 (10%)
Query: 26 HQQQQQQKQQNLPNQQQQNPHQL--HHSQVVSYAPQHHDTDTNQHHHHSMKHGFPPFSSS 83
H QQQQQ + QQQQ Q HH Q + HH Q H H +
Sbjct: 217 HHQQQQQFHDHQQQQQQQQQQQFHDHHQQKQQFHDHHHHQQQQQFHDHQQQ------QQQ 270
Query: 84 KNKQQQQSSQMSDEDE 99
+ QQQ Q D +
Sbjct: 271 FHDHQQQQHQFHDHPQ 286
Score = 38.1 bits (87), Expect = 0.053
Identities = 26/83 (31%), Positives = 33/83 (39%), Gaps = 10/83 (12%)
Query: 27 QQQQQQKQQNLPNQQQQNPHQLHHSQVVSYAPQ-----HHDTDTNQHHHHSMKHGFPPFS 81
QQQQQQ+Q + +QQQQ + H Q Q HH H HH + F
Sbjct: 147 QQQQQQQQFHNHHQQQQQFYDHHQQQQQQQQQQQQFHDHHQQKQQFHDHHQQQQQFHDHH 206
Query: 82 SSKNKQQ-----QQSSQMSDEDE 99
+QQ QQ Q D +
Sbjct: 207 HHHQEQQFHDHHQQQQQFHDHQQ 229
Score = 36.2 bits (82), Expect = 0.20
Identities = 24/82 (29%), Positives = 31/82 (37%), Gaps = 8/82 (9%)
Query: 26 HQQQQQQKQQNLPNQQQQNPHQLHHSQVVSYAPQHHDTDTNQ---HHHHSMKHGFPPFSS 82
H QQQQQ + + Q+Q H H Q + Q Q H HH K F
Sbjct: 195 HHQQQQQFHDHHHHHQEQQFHDHHQQQQQFHDHQQQQQQQQQQQFHDHHQQKQQFHDHHH 254
Query: 83 SKNKQ-----QQQSSQMSDEDE 99
+ +Q QQQ Q D +
Sbjct: 255 HQQQQQFHDHQQQQQQFHDHQQ 276
Score = 34.7 bits (78), Expect = 0.58
Identities = 30/95 (31%), Positives = 36/95 (37%), Gaps = 21/95 (22%)
Query: 26 HQQQQQ------QKQQNLPNQQQQ-----NPHQL-----HHSQVVSYAPQHHDTDTNQ-- 67
HQQQQQ Q+QQ +QQQQ +P Q H Q + HH Q
Sbjct: 255 HQQQQQFHDHQQQQQQFHDHQQQQHQFHDHPQQKQQFHDHPQQQQQFHDHHHQQQQKQQF 314
Query: 68 HHHHSMKHGFPPFSSSKNK---QQQQSSQMSDEDE 99
H HH K F K + QQ Q D +
Sbjct: 315 HDHHQQKQQFHDHHQQKQQFHDHHQQQQQFHDHHQ 349
Score = 31.6 bits (70), Expect = 4.9
Identities = 24/81 (29%), Positives = 31/81 (37%), Gaps = 11/81 (13%)
Query: 26 HQQQQQ-------QKQQNLPNQQQQNPHQLHHSQVVSYAPQHHDTDTNQHHHHSMKHGFP 78
HQQ+QQ Q+QQ + QQQ Q H Q + H Q H H +
Sbjct: 243 HQQKQQFHDHHHHQQQQQFHDHQQQQ-QQFHDHQQQQHQFHDHPQQKQQFHDHPQQQ--- 298
Query: 79 PFSSSKNKQQQQSSQMSDEDE 99
+ QQQQ Q D +
Sbjct: 299 QQFHDHHHQQQQKQQFHDHHQ 319
Score = 30.8 bits (68), Expect = 8.4
Identities = 23/72 (31%), Positives = 29/72 (39%), Gaps = 10/72 (13%)
Query: 22 NIPLHQQQQQQKQQNLPNQQQQNPHQLHHSQVVSYAPQHHDTDTNQ---HHHHSMKHGFP 78
+IP QQQQ+Q QQQ + H Q + Q Q H HH K F
Sbjct: 137 DIPTGHLQQQQQQ---QQQQQFHNHHQQQQQFYDHHQQQQQQQQQQQQFHDHHQQKQQF- 192
Query: 79 PFSSSKNKQQQQ 90
++QQQQ
Sbjct: 193 ---HDHHQQQQQ 201
>Z265_HUMAN (O95218) Zinc finger protein 265 (Zinc finger, splicing)
Length = 337
Score = 46.6 bits (109), Expect = 1e-04
Identities = 46/193 (23%), Positives = 84/193 (42%), Gaps = 31/193 (16%)
Query: 310 RGEGLRMLKIGSGYVEEEDDEDEEDESEDFS----DEGEDESGEGCSKGHINDQDEEEND 365
RG+ + I ++E + +EEDE ED S DE EDE SK +++ +EE+++
Sbjct: 135 RGKAVGPASILKEVEDKESEGEEEDEDEDLSKYKLDEDEDEDDADLSKYNLDASEEEDSN 194
Query: 366 GKPSRKRAR-KGGFSFPRSSSSTQLVNQMNNEISGVFQDGGKSTWEKKQWIRNRIMQLEE 424
K S +R+R K S RSSS + + R+R
Sbjct: 195 KKKSNRRSRSKSRSSHSRSSSRSSSPSSS----------------------RSRSRSRSR 232
Query: 425 QKIGYESQAFQLEKQRLKWARYSSKKEREMERAKLENERRRLENERMVLLIRKKE--LEL 482
+S++ ++R + S+ A L +E+ ++ ++L ++E L+L
Sbjct: 233 SSSSSQSRSRSSSRERSRSRGSKSRSSSRSTGALLPHEKDLIQVHHLLLRGTEREVVLDL 292
Query: 483 MH--IQQQQQQQH 493
+H I ++ +Q H
Sbjct: 293 LHLVIAKKDEQDH 305
>SIMA_DROME (Q24167) Similar protein
Length = 1507
Score = 46.6 bits (109), Expect = 1e-04
Identities = 34/110 (30%), Positives = 48/110 (42%), Gaps = 12/110 (10%)
Query: 4 ANAMFSTITPSGLLGVVDNIPLHQQQQQQKQQNLPNQQQQNPHQLHHSQ---------VV 54
AN+ S +TP+ + P HQQQQQ Q QQQQ H HH +
Sbjct: 738 ANSPLSPLTPNSTATASN--PSHQQQQQHHNQQQQQQQQQQHHPQHHDNSNSSSNIDPLF 795
Query: 55 SYAPQHHDTDTNQHHHHSMKHGFPPFSSSKNKQQQQSSQMSDEDEPNFPA 104
+Y + +DT +QH H P SS +S ++ ED+ +F A
Sbjct: 796 NYREESNDTSCSQHLHSPSITSKSPEDSSLPSLCSPNS-LTQEDDFSFEA 844
Score = 35.8 bits (81), Expect = 0.26
Identities = 17/50 (34%), Positives = 24/50 (48%)
Query: 248 HNSCGHGGVATSNVQHQVEGGSGTTTPPQNQPQQQHQHQHQQQNQQHCFH 297
+ S + A S + + T + P +Q QQQH +Q QQQ QQ H
Sbjct: 729 NTSSSNNSYANSPLSPLTPNSTATASNPSHQQQQQHHNQQQQQQQQQQHH 778
Score = 32.0 bits (71), Expect = 3.8
Identities = 48/271 (17%), Positives = 98/271 (35%), Gaps = 52/271 (19%)
Query: 263 HQVEGGSGTTTPPQNQPQQQHQH----------------------QHQQQNQQHCFHSSE 300
HQ G+ Q QPQ Q QH QH QQ Q F S
Sbjct: 897 HQQYAGNTGYQQQQQQPQLQQQHFSNSLCSSPASTVSSLSPSPVQQHHQQQQAAVFTSDS 956
Query: 301 NGVGSLGVSRGEGLRMLKIGSG--YVEEEDDEDEEDESEDFSDEGEDESGEGCSKGHIND 358
+ + +L G G + GSG EE ++ ++ + + G + + +
Sbjct: 957 SELAALLCGSGNGTLSILAGSGVTVAEECNERLQQHQQQQQQTSGNEFRTFQQLQQELQL 1016
Query: 359 QDEEENDGKPSRKRARKG--------GFSFPRSSSSTQLVNQMNNEISGVFQ-DGGKSTW 409
Q+E++ + +++ ++ + Q+ + + + F+ D K +
Sbjct: 1017 QEEQQQRQQQQQQQQQQQQQQQLLSLNIECKKEKYDVQMGGSLCHPMEDAFENDYSKDSA 1076
Query: 410 EKKQWIRNRIMQLEEQKIGYES--------QAFQLEKQRLKWARYSSKKEREMERAKLEN 461
W ++ ++ + + + A QL +Q ++++++ + +
Sbjct: 1077 NLDCWDLIQMQVVDTEPVSPNAASPTPCKVSAIQLLQQ-----------QQQLQQQQQQQ 1125
Query: 462 ERRRLENERMVLLIRKKELELMHIQQQQQQQ 492
+ L ++ + KEL QQQQQQQ
Sbjct: 1126 QNIILNAVPLITIQNNKELMQQQQQQQQQQQ 1156
>FCP1_YEAST (Q03254) RNA polymerase II subunit A C-terminal domain
phosphatase (EC 3.1.3.16) (CTD phosphatase FCP1)
Length = 732
Score = 46.6 bits (109), Expect = 1e-04
Identities = 29/105 (27%), Positives = 50/105 (47%), Gaps = 21/105 (20%)
Query: 276 QNQPQQQHQHQHQQQNQQHCFHSSENGVGSLGVSRGEGLRMLKIGSGYVEEEDDED---- 331
Q Q Q++ ++ + Q QQH S EN L + G+ ++ +DDED
Sbjct: 608 QTQLQKRQEYLEETQEQQHMLTSQEN------------LNLFAAGTSWLNNDDDEDIPDT 655
Query: 332 --EEDESEDFSDEGEDESGEGCSKGHINDQDEEENDGKPSRKRAR 374
++DE +D DE +DE+ S+G + E+N S+K+ +
Sbjct: 656 ASDDDEDDDHDDESDDENN---SEGIDRKRSIEDNHDDTSQKKTK 697
>A32A_RAT (P49911) Acidic leucine-rich nuclear phosphoprotein 32
family member A (Leucine-rich acidic nuclear protein)
Length = 247
Score = 46.6 bits (109), Expect = 1e-04
Identities = 24/52 (46%), Positives = 35/52 (67%), Gaps = 5/52 (9%)
Query: 325 EEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEEN----DGKPSRKR 372
EEE+DE+EE E ED S E E+E EG + G ++D+++EE+ +G RKR
Sbjct: 187 EEEEDEEEEGEEEDVSGE-EEEDEEGYNDGEVDDEEDEEDAAEEEGSQKRKR 237
>Z265_RAT (O35986) Zinc finger protein 265 (Zinc finger, splicing)
Length = 332
Score = 45.8 bits (107), Expect = 3e-04
Identities = 30/81 (37%), Positives = 45/81 (55%), Gaps = 5/81 (6%)
Query: 310 RGEGLRMLKIGSGYVEEEDDEDEEDESEDFS----DEGEDESGEGCSKGHINDQDEEEND 365
RG+ + I ++E + +EEDE ED S DE EDE SK +++ +EE+++
Sbjct: 135 RGKAVGPASILKEVEDKESEGEEEDEDEDLSKYKLDEDEDEDDADLSKYNLDASEEEDSN 194
Query: 366 GKPSRKRAR-KGGFSFPRSSS 385
K S +R+R K S RSSS
Sbjct: 195 KKKSNRRSRSKSRSSHSRSSS 215
>Z265_MOUSE (Q9R020) Zinc finger protein 265 (Zinc finger, splicing)
(Fragment)
Length = 326
Score = 45.8 bits (107), Expect = 3e-04
Identities = 30/81 (37%), Positives = 45/81 (55%), Gaps = 5/81 (6%)
Query: 310 RGEGLRMLKIGSGYVEEEDDEDEEDESEDFS----DEGEDESGEGCSKGHINDQDEEEND 365
RG+ + I ++E + +EEDE ED S DE EDE SK +++ +EE+++
Sbjct: 135 RGKAVGPASILKEVEDKESEGEEEDEDEDLSKYKLDEDEDEDDADLSKYNLDASEEEDSN 194
Query: 366 GKPSRKRAR-KGGFSFPRSSS 385
K S +R+R K S RSSS
Sbjct: 195 KKKSNRRSRSKSRSSHSRSSS 215
>VG48_SHV21 (Q01033) Hypothetical gene 48 protein
Length = 797
Score = 45.8 bits (107), Expect = 3e-04
Identities = 22/44 (50%), Positives = 27/44 (61%)
Query: 322 GYVEEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEEND 365
G E ++ EDE DE ED DEGEDE EG +G D+ E+E D
Sbjct: 629 GEDEGDEGEDEGDEGEDEGDEGEDEGDEGEDEGDEGDEGEDEGD 672
Score = 45.1 bits (105), Expect = 4e-04
Identities = 21/42 (50%), Positives = 27/42 (64%)
Query: 325 EEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEENDG 366
E ED+ DE DE ED DEGEDE EG +G D+ +E ++G
Sbjct: 656 EGEDEGDEGDEGEDEGDEGEDEGDEGEDEGDEGDEGDEGDEG 697
Score = 43.1 bits (100), Expect = 0.002
Identities = 21/55 (38%), Positives = 30/55 (54%)
Query: 322 GYVEEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEENDGKPSRKRARKG 376
G E ED++DEEDE ED DEG++ EG D+ +E ++GK +G
Sbjct: 462 GEDEGEDEDDEEDEGEDEGDEGDEGEDEGDEGDEGEDEGDEGDEGKDEGDEGDEG 516
Score = 43.1 bits (100), Expect = 0.002
Identities = 23/44 (52%), Positives = 27/44 (61%), Gaps = 3/44 (6%)
Query: 322 GYVEEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEEND 365
G E ED+ DE DE ED DEGEDE EG +G D+ E+E D
Sbjct: 601 GEDEGEDEGDEGDEGEDEGDEGEDEGDEGEDEG---DEGEDEGD 641
Score = 42.0 bits (97), Expect = 0.004
Identities = 22/44 (50%), Positives = 27/44 (61%), Gaps = 3/44 (6%)
Query: 322 GYVEEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEEND 365
G E ++ EDE DE ED DEGEDE EG +G D+ E+E D
Sbjct: 615 GEDEGDEGEDEGDEGEDEGDEGEDEGDEGEDEG---DEGEDEGD 655
Score = 42.0 bits (97), Expect = 0.004
Identities = 20/43 (46%), Positives = 27/43 (62%)
Query: 325 EEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEENDGK 367
E ++ EDE DE ED DEGEDE EG ++ DE E++G+
Sbjct: 663 EGDEGEDEGDEGEDEGDEGEDEGDEGDEGDEGDEGDEGEDEGE 705
Score = 42.0 bits (97), Expect = 0.004
Identities = 22/44 (50%), Positives = 27/44 (61%), Gaps = 3/44 (6%)
Query: 322 GYVEEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEEND 365
G E ++ EDE DE ED DEGEDE EG +G D+ E+E D
Sbjct: 622 GEDEGDEGEDEGDEGEDEGDEGEDEGDEGEDEG---DEGEDEGD 662
Score = 42.0 bits (97), Expect = 0.004
Identities = 21/42 (50%), Positives = 29/42 (69%), Gaps = 1/42 (2%)
Query: 326 EEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEENDGK 367
E++ EDE DE ED DEGEDE EG +G ++ DE E++G+
Sbjct: 533 EDEWEDEGDEGEDEGDEGEDEGDEGEDEGE-DEGDEGEDEGE 573
Score = 42.0 bits (97), Expect = 0.004
Identities = 21/44 (47%), Positives = 25/44 (56%)
Query: 322 GYVEEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEEND 365
G E ++ EDE DE ED DEGEDE EG D+ E+E D
Sbjct: 636 GEDEGDEGEDEGDEGEDEGDEGEDEGDEGDEGEDEGDEGEDEGD 679
Score = 41.6 bits (96), Expect = 0.005
Identities = 21/41 (51%), Positives = 26/41 (63%), Gaps = 3/41 (7%)
Query: 325 EEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEEND 365
E ++ EDE DE ED DEGEDE EG +G D+ E+E D
Sbjct: 611 EGDEGEDEGDEGEDEGDEGEDEGDEGEDEG---DEGEDEGD 648
Score = 40.4 bits (93), Expect = 0.011
Identities = 18/41 (43%), Positives = 25/41 (60%)
Query: 325 EEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEEND 365
E ED+EDE+DE E+ DEG+D EG +G + +E D
Sbjct: 420 EGEDEEDEKDEKEEGEDEGDDGEDEGEDEGEDEGDEGDEGD 460
Score = 40.0 bits (92), Expect = 0.014
Identities = 28/93 (30%), Positives = 40/93 (42%), Gaps = 4/93 (4%)
Query: 322 GYVEEEDDEDEEDESEDFSDEGED---ESGEGCSKGHINDQDEEEND-GKPSRKRARKGG 377
G E ED+ DE DE ++ DEGED E EG +G D+ E+E D G +G
Sbjct: 445 GEDEGEDEGDEGDEGDEGEDEGEDEDDEEDEGEDEGDEGDEGEDEGDEGDEGEDEGDEGD 504
Query: 378 FSFPRSSSSTQLVNQMNNEISGVFQDGGKSTWE 410
+ ++ + G D G+ WE
Sbjct: 505 EGKDEGDEGDEGKDEGDEGDEGDEGDEGEDEWE 537
Score = 40.0 bits (92), Expect = 0.014
Identities = 25/56 (44%), Positives = 33/56 (58%), Gaps = 4/56 (7%)
Query: 322 GYVEEEDDEDE-EDESEDFSDEGEDESGEGCSKGHINDQDEEENDGKPSRKRARKG 376
G E +D EDE EDE ED DEG DE EG +G D+D+EE++G+ +G
Sbjct: 434 GEDEGDDGEDEGEDEGEDEGDEG-DEGDEGEDEG--EDEDDEEDEGEDEGDEGDEG 486
Score = 39.3 bits (90), Expect = 0.024
Identities = 32/112 (28%), Positives = 46/112 (40%), Gaps = 12/112 (10%)
Query: 266 EGGSGTTTPPQNQPQQQHQHQHQQQNQQHCFHSSENGVGSLGVSRG----EGLRMLKIGS 321
EG G + + + + + + + + E G G G EG G
Sbjct: 455 EGDEGDEGEDEGEDEDDEEDEGEDEGDEGDEGEDEGDEGDEGEDEGDEGDEGKDEGDEGD 514
Query: 322 GYVEEEDDEDE-------EDESEDFSDEGEDESGEGCSKGHINDQDEEENDG 366
+E D+ DE EDE ED DEGEDE EG +G +DE E++G
Sbjct: 515 EGKDEGDEGDEGDEGDEGEDEWEDEGDEGEDEGDEGEDEGD-EGEDEGEDEG 565
Score = 39.3 bits (90), Expect = 0.024
Identities = 19/45 (42%), Positives = 28/45 (62%), Gaps = 4/45 (8%)
Query: 325 EEEDDEDE----EDESEDFSDEGEDESGEGCSKGHINDQDEEEND 365
+EED++DE EDE +D DEGEDE + +G D+ E+E +
Sbjct: 423 DEEDEKDEKEEGEDEGDDGEDEGEDEGEDEGDEGDEGDEGEDEGE 467
Score = 38.9 bits (89), Expect = 0.031
Identities = 22/55 (40%), Positives = 32/55 (58%), Gaps = 4/55 (7%)
Query: 325 EEEDDEDE---EDESEDFSDEGEDESGEGCSKGHINDQDEEENDGKPSRKRARKG 376
E+E DE E EDE ++ DEGEDE EG +G ++ DE E++G+ +G
Sbjct: 562 EDEGDEGEDEGEDEGDEGEDEGEDEGDEGEDEGE-DEGDEGEDEGEDEGDEGDEG 615
Score = 38.9 bits (89), Expect = 0.031
Identities = 21/44 (47%), Positives = 30/44 (67%), Gaps = 2/44 (4%)
Query: 325 EEEDDEDE-EDESEDFSDEGEDESGEGCSKGHINDQDEEENDGK 367
E ED+ DE EDE ++ DEGEDE EG +G ++ DE E++G+
Sbjct: 542 EGEDEGDEGEDEGDEGEDEGEDEGDEGEDEGE-DEGDEGEDEGE 584
Score = 38.9 bits (89), Expect = 0.031
Identities = 20/42 (47%), Positives = 26/42 (61%), Gaps = 1/42 (2%)
Query: 325 EEEDDEDEEDESEDFS-DEGEDESGEGCSKGHINDQDEEEND 365
E+E+ EDE D+ ED DEGEDE EG D+ E+E+D
Sbjct: 430 EKEEGEDEGDDGEDEGEDEGEDEGDEGDEGDEGEDEGEDEDD 471
Score = 38.9 bits (89), Expect = 0.031
Identities = 17/41 (41%), Positives = 28/41 (67%), Gaps = 2/41 (4%)
Query: 325 EEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEEND 365
EEED+ ++E++ +D +EGEDE +G +G D+ E+E D
Sbjct: 416 EEEDEGEDEEDEKDEKEEGEDEGDDGEDEG--EDEGEDEGD 454
Score = 38.5 bits (88), Expect = 0.040
Identities = 21/46 (45%), Positives = 30/46 (64%), Gaps = 4/46 (8%)
Query: 325 EEEDDEDE---EDESEDFSDEGEDESGEGCSKGHINDQDEEENDGK 367
E+E DE E EDE ++ DEGEDE EG +G ++ DE E++G+
Sbjct: 551 EDEGDEGEDEGEDEGDEGEDEGEDEGDEGEDEGE-DEGDEGEDEGE 595
Score = 37.7 bits (86), Expect = 0.069
Identities = 24/53 (45%), Positives = 28/53 (52%), Gaps = 9/53 (16%)
Query: 322 GYVEEEDDEDE-EDESEDFSDEGEDE--------SGEGCSKGHINDQDEEEND 365
G E ED+ DE EDE ED DEGEDE EG +G D+ E+E D
Sbjct: 568 GEDEGEDEGDEGEDEGEDEGDEGEDEGEDEGDEGEDEGEDEGDEGDEGEDEGD 620
Score = 37.7 bits (86), Expect = 0.069
Identities = 19/41 (46%), Positives = 25/41 (60%), Gaps = 2/41 (4%)
Query: 325 EEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEEND 365
E ++ EDE DE ED DEGEDE + +G D+ E+E D
Sbjct: 539 EGDEGEDEGDEGEDEGDEGEDEGEDEGDEG--EDEGEDEGD 577
Score = 37.7 bits (86), Expect = 0.069
Identities = 20/44 (45%), Positives = 25/44 (56%), Gaps = 3/44 (6%)
Query: 325 EEEDDEDE---EDESEDFSDEGEDESGEGCSKGHINDQDEEEND 365
E+E DE E EDE ++ DEGEDE EG D+ E+E D
Sbjct: 584 EDEGDEGEDEGEDEGDEGEDEGEDEGDEGDEGEDEGDEGEDEGD 627
Score = 37.4 bits (85), Expect = 0.090
Identities = 22/57 (38%), Positives = 31/57 (53%), Gaps = 5/57 (8%)
Query: 325 EEEDDEDE---EDESEDFSDEGEDESGEGCSKGHI--NDQDEEENDGKPSRKRARKG 376
E+E DE E EDE ++ DEGEDE EG +G ++ DE E++G +G
Sbjct: 573 EDEGDEGEDEGEDEGDEGEDEGEDEGDEGEDEGEDEGDEGDEGEDEGDEGEDEGDEG 629
Score = 37.4 bits (85), Expect = 0.090
Identities = 19/41 (46%), Positives = 25/41 (60%), Gaps = 3/41 (7%)
Query: 325 EEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEEND 365
E ED+ DE ++ D DEGEDE EG +G D+ E+E D
Sbjct: 649 EGEDEGDEGEDEGDEGDEGEDEGDEGEDEG---DEGEDEGD 686
Score = 37.4 bits (85), Expect = 0.090
Identities = 22/45 (48%), Positives = 27/45 (59%), Gaps = 3/45 (6%)
Query: 322 GYVEEEDDEDE-EDESEDFSDEGEDESGEGCSKGHINDQDEEEND 365
G E ED+ DE EDE ED DEGEDE + +G D+ E+E D
Sbjct: 557 GEDEGEDEGDEGEDEGEDEGDEGEDEGEDEGDEG--EDEGEDEGD 599
Score = 37.0 bits (84), Expect = 0.12
Identities = 21/42 (50%), Positives = 26/42 (61%), Gaps = 3/42 (7%)
Query: 325 EEEDDEDE-EDESEDFSDEGEDESGEGCSKGHINDQDEEEND 365
E ED+ DE EDE ED DEGEDE + +G D+ E+E D
Sbjct: 549 EGEDEGDEGEDEGEDEGDEGEDEGEDEGDEG--EDEGEDEGD 588
Score = 36.6 bits (83), Expect = 0.15
Identities = 19/44 (43%), Positives = 26/44 (58%), Gaps = 3/44 (6%)
Query: 322 GYVEEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEEND 365
G E ++ EDE ++ D DEGEDE EG +G D+ E+E D
Sbjct: 594 GEDEGDEGEDEGEDEGDEGDEGEDEGDEGEDEG---DEGEDEGD 634
Score = 33.5 bits (75), Expect = 1.3
Identities = 19/53 (35%), Positives = 26/53 (48%), Gaps = 2/53 (3%)
Query: 325 EEEDDEDEEDESEDFSDEGE--DESGEGCSKGHINDQDEEENDGKPSRKRARK 375
E ED+ DE ++ D DEG+ DE EG +G +DE + K A K
Sbjct: 673 EGEDEGDEGEDEGDEGDEGDEGDEGDEGEDEGEDEGEDEGDEGTKDKEGNANK 725
Score = 33.5 bits (75), Expect = 1.3
Identities = 18/43 (41%), Positives = 25/43 (57%), Gaps = 3/43 (6%)
Query: 325 EEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEENDGK 367
+E D+ DE DE ++ DEGED EG +G +D+E N K
Sbjct: 686 DEGDEGDEGDEGDEGEDEGED---EGEDEGDEGTKDKEGNANK 725
Score = 32.7 bits (73), Expect = 2.2
Identities = 19/46 (41%), Positives = 28/46 (60%), Gaps = 2/46 (4%)
Query: 322 GYVEEEDDEDE-EDESEDFSDEGEDESGEGCSKGHINDQDEEENDG 366
G E ED+ DE EDE ++ DEG++ EG +G +DE E++G
Sbjct: 532 GEDEWEDEGDEGEDEGDEGEDEGDEGEDEGEDEGD-EGEDEGEDEG 576
Score = 32.3 bits (72), Expect = 2.9
Identities = 21/50 (42%), Positives = 28/50 (56%), Gaps = 4/50 (8%)
Query: 325 EEEDDE-DEEDESEDFSDEGE--DESGEGCSKGHINDQDEEENDGKPSRK 371
+E D+ DE DE ED DEG+ DE EG +G +DE E++G K
Sbjct: 669 DEGDEGEDEGDEGEDEGDEGDEGDEGDEG-DEGEDEGEDEGEDEGDEGTK 717
>T230_HUMAN (Q93074) Thyroid hormone receptor-associated protein
complex 230 kDa component (Trap230) (Activator-recruited
cofactor 240 KDa component) (ARC240) (CAG repeat protein
45) (OPA-containing protein) (Trinucleotide repeat
containing 11)
Length = 2212
Score = 45.8 bits (107), Expect = 3e-04
Identities = 34/138 (24%), Positives = 51/138 (36%), Gaps = 7/138 (5%)
Query: 2 MEANAMFSTITPSGLLGVVDNI-------PLHQQQQQQKQQNLPNQQQQNPHQLHHSQVV 54
++ M ST+TP GV + QQQQQQ+QQ QQQQ Q +
Sbjct: 2056 LQQTPMISTMTPMSAQGVQAGVRSTAILPEQQQQQQQQQQQQQQQQQQQQQQQQQQYHIR 2115
Query: 55 SYAPQHHDTDTNQHHHHSMKHGFPPFSSSKNKQQQQSSQMSDEDEPNFPAEESSGGDPKR 114
Q Q + + +QQQ Q + P P +S ++
Sbjct: 2116 QQQQQQILRQQQQQQQQQQQQQQQQQQQQQQQQQQHQQQQQQQAAPPQPQPQSQPQFQRQ 2175
Query: 115 KISPWQRMKWTDTMVRLL 132
+ Q+ + T +VR L
Sbjct: 2176 GLQQTQQQQQTAALVRQL 2193
Score = 32.3 bits (72), Expect = 2.9
Identities = 14/73 (19%), Positives = 40/73 (54%)
Query: 420 MQLEEQKIGYESQAFQLEKQRLKWARYSSKKEREMERAKLENERRRLENERMVLLIRKKE 479
M + + G S A E+Q+ + + +++++ ++ + + ++ + ++ ++R+++
Sbjct: 2068 MSAQGVQAGVRSTAILPEQQQQQQQQQQQQQQQQQQQQQQQQQQYHIRQQQQQQILRQQQ 2127
Query: 480 LELMHIQQQQQQQ 492
+ QQQQQQQ
Sbjct: 2128 QQQQQQQQQQQQQ 2140
Score = 32.0 bits (71), Expect = 3.8
Identities = 22/62 (35%), Positives = 25/62 (39%), Gaps = 20/62 (32%)
Query: 252 GHGGVATSNVQHQ------------------VEGGSGTTT--PPQNQPQQQHQHQHQQQN 291
GHG +T HQ V+ G +T P Q Q QQQ Q Q QQQ
Sbjct: 2042 GHGLTSTQRFSHQTLQQTPMISTMTPMSAQGVQAGVRSTAILPEQQQQQQQQQQQQQQQQ 2101
Query: 292 QQ 293
QQ
Sbjct: 2102 QQ 2103
>IE68_PRVKA (P24827) Immediate-early protein RSP40
Length = 364
Score = 45.8 bits (107), Expect = 3e-04
Identities = 29/75 (38%), Positives = 43/75 (56%), Gaps = 4/75 (5%)
Query: 309 SRGEGLRMLKIGSGYVEEEDDEDEEDESEDFSDEGEDESGE-GCSKGHINDQDEEEN-DG 366
S GEG G G E+ED++++EDE ED ++ EDE GE G G ++DE+E+ +G
Sbjct: 285 SDGEGSGSDDGGDG--EDEDEDEDEDEDEDDGEDEEDEEGEDGGEDGEDGEEDEDEDGEG 342
Query: 367 KPSRKRARKGGFSFP 381
+ K A + G P
Sbjct: 343 EEGGKDAARRGTRAP 357
Score = 39.7 bits (91), Expect = 0.018
Identities = 19/40 (47%), Positives = 25/40 (62%), Gaps = 7/40 (17%)
Query: 327 EDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEENDG 366
EDD ++EDE ED +E EDE GE ++DEEE +G
Sbjct: 208 EDDGEDEDEEEDGEEEDEDEEGE-------EEEDEEEEEG 240
Score = 36.2 bits (82), Expect = 0.20
Identities = 23/77 (29%), Positives = 34/77 (43%), Gaps = 22/77 (28%)
Query: 323 YVEEEDDEDEEDES-------------------EDFSD---EGEDESGEGCSKGHINDQD 360
Y E+++ EDEEDE ED SD G D+ G+G + D+D
Sbjct: 249 YEEDDEAEDEEDEEDGDDFDGASVGDDDVFEPPEDGSDGEGSGSDDGGDGEDEDEDEDED 308
Query: 361 EEENDGKPSRKRARKGG 377
E+E+DG+ + G
Sbjct: 309 EDEDDGEDEEDEEGEDG 325
Score = 35.4 bits (80), Expect = 0.34
Identities = 16/43 (37%), Positives = 27/43 (62%), Gaps = 2/43 (4%)
Query: 325 EEEDDEDEEDESEDFSDEGEDESGEGCSKGH--INDQDEEEND 365
+EE+D +EEDE E+ +E ++E EG G + ++D+E D
Sbjct: 215 DEEEDGEEEDEDEEGEEEEDEEEEEGDEDGETDVYEEDDEAED 257
Score = 34.3 bits (77), Expect = 0.76
Identities = 19/60 (31%), Positives = 31/60 (51%), Gaps = 1/60 (1%)
Query: 326 EEDDEDEEDESEDFSDEGEDESGEGCSKG-HINDQDEEENDGKPSRKRARKGGFSFPRSS 384
E++DE+E+ E ED +EGE+E E +G + D E D + + + G F +S
Sbjct: 212 EDEDEEEDGEEEDEDEEGEEEEDEEEEEGDEDGETDVYEEDDEAEDEEDEEDGDDFDGAS 271
Score = 32.3 bits (72), Expect = 2.9
Identities = 15/45 (33%), Positives = 25/45 (55%)
Query: 322 GYVEEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEENDG 366
G E+ED+E EE+E E+ + ED + + + +E+E DG
Sbjct: 220 GEEEDEDEEGEEEEDEEEEEGDEDGETDVYEEDDEAEDEEDEEDG 264
Score = 31.6 bits (70), Expect = 4.9
Identities = 22/63 (34%), Positives = 34/63 (53%), Gaps = 2/63 (3%)
Query: 304 GSLGVSRGEGLRMLKIGSGYVEEEDDEDEEDESEDFSDE-GEDESGEGCSKGHINDQDEE 362
GS+ GE + G E+E+ E+EEDE E+ DE GE + E + +++DEE
Sbjct: 204 GSVCEDDGEDEDEEEDGEEEDEDEEGEEEEDEEEEEGDEDGETDVYEEDDEAE-DEEDEE 262
Query: 363 END 365
+ D
Sbjct: 263 DGD 265
>BM2K_HUMAN (Q9NSY1) BMP-2 inducible protein kinase (EC 2.7.1.37)
(BIKe) (HRIHFB2017)
Length = 1161
Score = 45.8 bits (107), Expect = 3e-04
Identities = 26/68 (38%), Positives = 32/68 (46%), Gaps = 10/68 (14%)
Query: 26 HQQQQQQKQQNLPNQQQQNPHQLHHSQVVSYAPQHHDTDTNQHHHHSMKHGFPPFSSSKN 85
HQQQQQQ+QQ QQQQ Q Q QHH + HHHH ++ +
Sbjct: 459 HQQQQQQQQQQQQQQQQQQQQQQQQQQ------QHH----HHHHHHLLQDAYMQQYQHAT 508
Query: 86 KQQQQSSQ 93
+QQQ Q
Sbjct: 509 QQQQMLQQ 516
Score = 32.7 bits (73), Expect = 2.2
Identities = 72/363 (19%), Positives = 121/363 (32%), Gaps = 39/363 (10%)
Query: 29 QQQQKQQNLPNQQQQNPHQLHHSQVVSYAPQHHDTDTNQHHHHSMKHGFPPFSSSKNKQQ 88
QQ Q P+QQQQ Q Q Q QHHHH H + +Q
Sbjct: 449 QQLHLQHRHPHQQQQQQQQ-QQQQQQQQQQQQQQQQQQQHHHHHHHH---LLQDAYMQQY 504
Query: 89 QQSSQMSDEDEPNFPAEESSGGDPKRKISPWQRMKWTDTMVRLLIMAVYYIG-DEAGSEG 147
Q ++Q + F P P ++ + ++A + +A E
Sbjct: 505 QHATQQQQMLQQQFLMHSVYQPQPSASQYPTMMPQYQQAFFQQQMLAQHQPSQQQASPEY 564
Query: 148 TDPNKKKSSGLLQKKGKWKSVSRAMMEKGFYVSPQQCEDKFNDLN----KRYKRVNDI-- 201
++ S L+ + +M+ Y + + DK N K D+
Sbjct: 565 LTSPQEFSPALVSYTSSLPAQVGTIMDSS-YSANRSVADKEAIANFTNQKNISNPPDMSG 623
Query: 202 ---LGKGTACRVVENQGLLDSMDLSPKMKDEVRKLLNSKHLFFREMCAYHNS-------- 250
G+ ++ E + L DL + E R S + A HN
Sbjct: 624 WNPFGEDNFSKLTEEELLDREFDLLRSNRLEERA---SSDKNVDSLSAPHNHPPEDPFGS 680
Query: 251 ---CGHGGVATSNVQH-QVEGGSGTTTPPQNQPQQQHQHQHQQQNQQHCFHSSENGVGSL 306
H G +H + +GT P +N + ++Q+ +S G
Sbjct: 681 VPFISHSGSPEKKAEHSSINQENGTANPIKN-----GKTSPASKDQRTGKKTSVQGQVQK 735
Query: 307 GVSRGEGLRMLKIGSGYVEEEDDEDEED----ESEDFSDEGEDESGEGCSKGHINDQDEE 362
G E S EE+++D+E+ E DF+D+ + G ++ +DEE
Sbjct: 736 GNDESESDFESDPPSPKSSEEEEQDDEEVLQGEQGDFNDDDTEPENLGHRPLLMDSEDEE 795
Query: 363 END 365
E +
Sbjct: 796 EEE 798
>SRY_MUSSP (Q62563) Sex-determining region Y protein
(Testis-determining factor)
Length = 355
Score = 45.4 bits (106), Expect = 3e-04
Identities = 29/82 (35%), Positives = 36/82 (43%), Gaps = 15/82 (18%)
Query: 26 HQQQQQQKQQNLPNQQQQ---NPHQL-------HHSQVVSYAPQHHDTDTNQHHHHSMKH 75
HQQQQQQ+ + P QQQQ +P Q HH Q HH H HH +
Sbjct: 249 HQQQQQQQFHDHPQQQQQFHDHPQQKQQFHDHHHHQQQKQQFHDHHQQKQQFHDHHQQQQ 308
Query: 76 GFPPFSSSKNKQQQQSSQMSDE 97
F ++QQQQ Q D+
Sbjct: 309 QF-----HDHQQQQQQQQFHDQ 325
Score = 44.3 bits (103), Expect = 7e-04
Identities = 27/77 (35%), Positives = 32/77 (41%), Gaps = 8/77 (10%)
Query: 26 HQQQQQQKQQNLPNQQQQNPHQLHHSQVVSYAPQHHDTDTNQ---HHHHSMKHGFPPFSS 82
H QQQ++Q + QQQQ H HH Q HH Q H HH K F
Sbjct: 191 HDHQQQKQQFHDHQQQQQQFHDHHHQQQQQQFHDHHHHQQQQQQFHDHHQQKQQF----- 245
Query: 83 SKNKQQQQSSQMSDEDE 99
+ QQQQ Q D +
Sbjct: 246 HDHHQQQQQQQFHDHPQ 262
Score = 42.0 bits (97), Expect = 0.004
Identities = 23/64 (35%), Positives = 26/64 (39%)
Query: 26 HQQQQQQKQQNLPNQQQQNPHQLHHSQVVSYAPQHHDTDTNQHHHHSMKHGFPPFSSSKN 85
HQQQQQQ + QQQQ H HH Q H Q H H + F
Sbjct: 203 HQQQQQQFHDHHHQQQQQQFHDHHHHQQQQQQFHDHHQQKQQFHDHHQQQQQQQFHDHPQ 262
Query: 86 KQQQ 89
+QQQ
Sbjct: 263 QQQQ 266
Score = 37.7 bits (86), Expect = 0.069
Identities = 25/78 (32%), Positives = 33/78 (42%), Gaps = 5/78 (6%)
Query: 22 NIPLHQQQQQQKQQNLPNQQQQNPHQLHHSQVVSYAPQHHDTDTNQHHHHSMKHGFPPFS 81
+IP QQQQ+Q + +QQ+Q H HH Q + HH H H + F
Sbjct: 137 DIPTGHPQQQQQQFHNHHQQKQQFHD-HHQQKQQFH-DHHQQQQQFHDHQQQQQQFHDHQ 194
Query: 82 SSKNK---QQQQSSQMSD 96
K + QQQ Q D
Sbjct: 195 QQKQQFHDHQQQQQQFHD 212
Score = 36.2 bits (82), Expect = 0.20
Identities = 20/64 (31%), Positives = 27/64 (41%), Gaps = 2/64 (3%)
Query: 26 HQQQQQQKQQNLPNQQQQNPHQLHHSQVVSYAPQHHDTDTNQHHHHSMKHGFPPFSSSKN 85
H QQQQQ + QQQ + HQ Q + Q + HHH + F +
Sbjct: 173 HHQQQQQFHDHQQQQQQFHDHQQQKQQFHDH--QQQQQQFHDHHHQQQQQQFHDHHHHQQ 230
Query: 86 KQQQ 89
+QQQ
Sbjct: 231 QQQQ 234
Score = 35.4 bits (80), Expect = 0.34
Identities = 22/74 (29%), Positives = 28/74 (37%), Gaps = 4/74 (5%)
Query: 26 HQQQQQQKQQNLPNQQQQNPHQLHHSQVVSYAPQHHDTDTNQHHHHSMKHGFPPFSSSKN 85
H QQQQ+Q + +QQ+Q H H Q H H H K F +
Sbjct: 226 HHHQQQQQQFHDHHQQKQQFHDHHQQQQQQQFHDHPQQQQQFHDHPQQKQQF----HDHH 281
Query: 86 KQQQQSSQMSDEDE 99
QQQ Q D +
Sbjct: 282 HHQQQKQQFHDHHQ 295
>YNJ1_YEAST (P53935) Hypothetical 141.5 kDa protein in YPT53-RHO2
intergenic region
Length = 1240
Score = 44.7 bits (104), Expect = 6e-04
Identities = 45/210 (21%), Positives = 86/210 (40%), Gaps = 25/210 (11%)
Query: 283 HQHQHQQQNQQHCFHSSENGVGSLGVSRGEGLRMLKIGSGYVEEEDDEDEEDESEDFSDE 342
+ H H++QN H + S + +S E + +++ D +D D
Sbjct: 509 YDHDHKRQNHPHHHYHSTSTHSEDELSEEEYISDIEL------PHDPHKHFHRDDDILDG 562
Query: 343 GEDESGEGCSKGHINDQDEEENDGKPSRKRARKGGFSFPRSSSSTQLVNQMNNEISGVFQ 402
EDE ++E+EN+G G R +L+ I+ + Q
Sbjct: 563 DEDEP-----------EEEDENEGDDEEDTYDSGLDETDRLEEGRKLIQIA---ITKLLQ 608
Query: 403 DGGKSTWEKKQWIRNRIMQLEEQKIGYESQAFQLEKQRLKWARYSSKKEREMERAKLENE 462
+++ +KQ NR+ L+E + + + EK++ K + KK + + E
Sbjct: 609 SRIMASYHEKQADNNRLKLLQELEEEKRKKREKEEKKQKKREKEKEKKRLQQLAKEEEKR 668
Query: 463 RRRLENERMVLLIRKKELELMHIQQQQQQQ 492
+R E ER+ KKELE +++++ Q+
Sbjct: 669 KREEEKERL-----KKELEEREMRRREAQR 693
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.310 0.127 0.367
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 64,019,269
Number of Sequences: 164201
Number of extensions: 3105033
Number of successful extensions: 44696
Number of sequences better than 10.0: 1025
Number of HSP's better than 10.0 without gapping: 542
Number of HSP's successfully gapped in prelim test: 507
Number of HSP's that attempted gapping in prelim test: 21474
Number of HSP's gapped (non-prelim): 9611
length of query: 496
length of database: 59,974,054
effective HSP length: 114
effective length of query: 382
effective length of database: 41,255,140
effective search space: 15759463480
effective search space used: 15759463480
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 68 (30.8 bits)
Medicago: description of AC146788.10


