

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
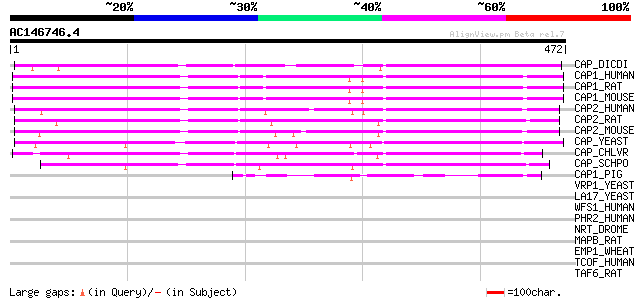
Query= AC146746.4 - phase: 0
(472 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
CAP_DICDI (P54654) Adenylyl cyclase-associated protein (CAP) 317 4e-86
CAP1_HUMAN (Q01518) Adenylyl cyclase-associated protein 1 (CAP 1) 312 1e-84
CAP1_RAT (Q08163) Adenylyl cyclase-associated protein 1 (CAP 1) 305 2e-82
CAP1_MOUSE (P40124) Adenylyl cyclase-associated protein 1 (CAP 1) 304 3e-82
CAP2_HUMAN (P40123) Adenylyl cyclase-associated protein 2 (CAP 2) 301 2e-81
CAP2_RAT (P52481) Adenylyl cyclase-associated protein 2 (CAP 2) 301 3e-81
CAP2_MOUSE (Q9CYT6) Adenylyl cyclase-associated protein 2 (CAP 2) 295 2e-79
CAP_YEAST (P17555) Adenylyl cyclase-associated protein (CAP) 249 1e-65
CAP_CHLVR (P40122) Adenylyl cyclase-associated protein (CAP) 245 2e-64
CAP_SCHPO (P36621) Adenylyl cyclase-associated protein (CAP) 233 6e-61
CAP1_PIG (P40125) Adenylyl cyclase-associated protein 1 (CAP 1) ... 87 9e-17
VRP1_YEAST (P37370) Verprolin 39 0.038
LA17_YEAST (Q12446) Proline-rich protein LAS17 37 0.084
WFS1_HUMAN (O76024) Wolframin 35 0.32
PHR2_HUMAN (O75167) Phosphatase and actin regulator 2 35 0.32
NRT_DROME (P23654) Neurotactin 35 0.32
MAPB_RAT (P15205) Microtubule-associated protein 1B (MAP 1B) (Ne... 35 0.42
EMP1_WHEAT (P25032) DNA-binding EMBP-1 protein 35 0.55
TCOF_HUMAN (Q13428) Treacle protein (Treacher Collins syndrome p... 34 0.71
TAF6_RAT (Q63801) Transcription initiation factor TFIID subunit ... 34 0.71
>CAP_DICDI (P54654) Adenylyl cyclase-associated protein (CAP)
Length = 464
Score = 317 bits (812), Expect = 4e-86
Identities = 189/479 (39%), Positives = 281/479 (58%), Gaps = 38/479 (7%)
Query: 5 LIKRLESAVTRLEA----LSTGIHHSSSASDASDAASDPS---VVAFVDLIDQYVSRLSK 57
L+KRL+ A TRLEA +++G+ SSS+S S A+ PS V F +L+DQ+++
Sbjct: 9 LLKRLDQATTRLEAVEKSIASGVASSSSSSSPSSGAAGPSSASVKEFQNLVDQHITPFVA 68
Query: 58 AADIIGGQVLDVTNRVKEAFSVQKELLIKLKQTQKPDPAGLAEFLKPLNEVIMKASSLTE 117
+ + +V + ++ +A +K L+ Q++KP L E +KPLN + + +
Sbjct: 69 LSKKLAPEVGNQVEQLVKAIDAEKALINTASQSKKPSQETLLELIKPLNNFAAEVGKIRD 128
Query: 118 GRRSD-FFNHLKAAVDSLSALAWIAFTGKDCGMSMPIAHVEESWQMAEFYSNKVLVEYRN 176
RS FFN+L A +S+ L+W+ P HV E AEFY+N++L E++
Sbjct: 129 SNRSSKFFNNLSAISESIGFLSWVVVE------PTPGPHVAEMRGSAEFYTNRILKEFKG 182
Query: 177 KDPNHVEWVKALKELYLPGLRDYVKSFHPLGPVWS-QTGKVFAPSKVNASAAPAAPSAPP 235
+ + V+WV +L L Y+K +H G W+ + G + + AS+APAAP AP
Sbjct: 183 VNQDQVDWVSNYVN-FLKDLEKYIKQYHTTGLTWNPKGGDAKSATPAPASSAPAAPVAP- 240
Query: 236 PPPASLFSSESSQASSSKPKVGMSAVFQEIGTGN-VTAGLRKVTDDMKTKNRADRSGVVG 294
+ SS SK G+ AVF E+ G+ VT+GL+KVT+DMK+KN D+S VV
Sbjct: 241 --------AVSSTPVESKKGPGLGAVFGELSKGDGVTSGLKKVTNDMKSKNFTDKSSVV- 291
Query: 295 NSVKESQAAPRAFSKVGPPK----LELQMGRKWVVENQIDQKSLVIEDCDSKQSVYVYGC 350
+AA +KV P LQ G KW +E Q++ K +VI + DS+Q+VY++ C
Sbjct: 292 ------KAADTKVAKVDAPSRPAVFALQ-GNKWSIEYQVNNKEIVIAEPDSRQTVYIFQC 344
Query: 351 KNSVLQIQGKVNNITIDNCKKTGVVFKDVVAAFEVVNSNGVEVQCQGSAPTISVDNTSGC 410
NS++QI+GKVN IT+D CKKT +VF++ +++ EVVN NGVE+Q G P+I++D TSGC
Sbjct: 345 VNSLVQIKGKVNAITLDGCKKTSIVFENAISSCEVVNCNGVEIQVTGRVPSIAIDKTSGC 404
Query: 411 QIYLSKDSLETSISTAKSSEINVLVPNVESDGDWVEHSLPQQYIHLFKEGRFETTPASH 469
QIYLSKDSLET I ++KSSE+NVL+P + D VE ++P+QY K + T SH
Sbjct: 405 QIYLSKDSLETEIVSSKSSEMNVLIPGATENDDLVELAIPEQYKTSVKGNKLHTESTSH 463
>CAP1_HUMAN (Q01518) Adenylyl cyclase-associated protein 1 (CAP 1)
Length = 474
Score = 312 bits (800), Expect = 1e-84
Identities = 195/482 (40%), Positives = 274/482 (56%), Gaps = 24/482 (4%)
Query: 3 ETLIKRLESAVTRLEALS-TGIHHSSSASDASDAASDPSVVAFVDLIDQYVSRLSKAADI 61
+ L++RLE AV RLEA+S T H A S A + P V AF L+ V+ K +
Sbjct: 4 QNLVERLERAVGRLEAVSHTSDMHRGYADSPSKAGAAPYVQAFDSLLAGPVAEYLKISKE 63
Query: 62 IGGQVLDVTNRVKEAFSVQKELLIKLKQTQKPDPAGLAEFLKPLNEVIMKASSLTEGRR- 120
IGG V V +++ LL+ Q Q+P L++ L P++E I + + E R
Sbjct: 64 IGGDVQKHAEMVHTGLKLERALLVTASQCQQPAENKLSDLLAPISEQIKEVITFREKNRG 123
Query: 121 SDFFNHLKAAVDSLSALAWIAFTGKDCGMSMPIAHVEESWQMAEFYSNKVLVEYRNKDPN 180
S FNHL A +S+ AL W+A K P +V+E A FY+N+VL EY++ D
Sbjct: 124 SKLFNHLSAVSESIQALGWVAMAPK------PGPYVKEMNDAAMFYTNRVLKEYKDVDKK 177
Query: 181 HVEWVKALKELYLPGLRDYVKSFHPLGPVWSQTGKVFAPSKVNASAAPAAPSAPPPPPAS 240
HV+WVKA ++ L+ Y+K FH G WS+TG V A + P+A S PPPPP
Sbjct: 178 HVDWVKAYLSIWTE-LQAYIKEFHTTGLAWSKTGPV-AKELSGLPSGPSAGSGPPPPPPG 235
Query: 241 LFSSESSQASSSKPKVGMSAVFQEIGTG-NVTAGLRKVTDDMKT-KNRA--DRSGVVGNS 296
S +S S SA+F +I G ++T L+ V+DDMKT KN A +SG V +
Sbjct: 236 PPPPPVSTSSGSDESASRSALFAQINQGESITHALKHVSDDMKTHKNPALKAQSGPVRSG 295
Query: 297 VK-------ESQAAPRAFSKVGPPKLELQMGRKWVVENQIDQKSLVIEDCDSKQSVYVYG 349
K ++ +P+ +K P LEL+ G+KW VENQ + +LVIED + KQ Y+Y
Sbjct: 296 PKPFSAPKPQTSPSPKRATKKEPAVLELE-GKKWRVENQENVSNLVIEDTELKQVAYIYK 354
Query: 350 CKNSVLQIQGKVNNITIDNCKKTGVVFKDVVAAFEVVNSNGVEVQCQGSAPTISVDNTSG 409
C N+ LQI+GK+N+IT+DNCKK G+VF DVV E++NS V+VQ G PTIS++ T G
Sbjct: 355 CVNTTLQIKGKINSITVDNCKKLGLVFDDVVGIVEIINSKDVKVQVMGKVPTISINKTDG 414
Query: 410 CQIYLSKDSLETSISTAKSSEINVLVPNVESDGDWVEHSLPQQYIHLFKEGRFETTPASH 469
C YLSK+SL+ I +AKSSE+NVL+P GD+ E +P+Q+ L+ + TT
Sbjct: 415 CHAYLSKNSLDCEIVSAKSSEMNVLIPT--EGGDFNEFPVPEQFKTLWNGQKLVTTVTEI 472
Query: 470 SG 471
+G
Sbjct: 473 AG 474
>CAP1_RAT (Q08163) Adenylyl cyclase-associated protein 1 (CAP 1)
Length = 473
Score = 305 bits (781), Expect = 2e-82
Identities = 189/481 (39%), Positives = 269/481 (55%), Gaps = 23/481 (4%)
Query: 3 ETLIKRLESAVTRLEALSTGIHHSSSASDASDAASDPSVVAFVDLIDQYVSRLSKAADII 62
+ L++RLE AV RLEA+S D+ + P V AF L+ V+ K + I
Sbjct: 4 QNLVERLERAVGRLEAVSHTSDMHCGYGDSPSKGAVPYVQAFDSLLANPVAEYLKMSKEI 63
Query: 63 GGQVLDVTNRVKEAFSVQKELLIKLKQTQKPDPAGLAEFLKPLNEVIMKASSLTEGRR-S 121
GG V V +++ LL+ Q Q+P L++ L P++E I + + E R S
Sbjct: 64 GGDVQKHAEMVHTGLKLERALLVTASQCQQPAGNKLSDLLAPISEQIQEVITFREKNRGS 123
Query: 122 DFFNHLKAAVDSLSALAWIAFTGKDCGMSMPIAHVEESWQMAEFYSNKVLVEYRNKDPNH 181
FFNHL A +S+ AL W+A K P V+E A FY+N+VL EYR+ D H
Sbjct: 124 KFFNHLSAVSESIQALGWVALAAK------PGPFVKEMNDAAMFYTNRVLKEYRDVDKKH 177
Query: 182 VEWVKALKELYLPGLRDYVKSFHPLGPVWSQTGKVFAPSKVNASAAPAAPSAPPPPPASL 241
V+WV+A ++ L+ Y+K FH G WS+TG V A + P+ S PPPPP
Sbjct: 178 VDWVRAYLSIWTE-LQAYIKEFHTTGLAWSKTGPV-AKELSGLPSGPSVGSGPPPPPPGP 235
Query: 242 FSSESSQASSSKPKVGMSAVFQEIGTG-NVTAGLRKVTDDMKT-KNRA--DRSGVVGNSV 297
+S S SA+F +I G ++T L+ V+DDMKT KN A +SG V +
Sbjct: 236 PPPPVPTSSGSDDSASRSALFAQINQGESITHALKHVSDDMKTHKNPALKAQSGPVRSGP 295
Query: 298 K-------ESQAAPRAFSKVGPPKLELQMGRKWVVENQIDQKSLVIEDCDSKQSVYVYGC 350
K ++ +P+ +K P LEL+ G+KW VENQ + +LVI+D + KQ Y+Y C
Sbjct: 296 KPFSAPKPQTSPSPKPATKKEPALLELE-GKKWRVENQENVSNLVIDDTELKQVAYIYKC 354
Query: 351 KNSVLQIQGKVNNITIDNCKKTGVVFKDVVAAFEVVNSNGVEVQCQGSAPTISVDNTSGC 410
N+ LQI+GK+N+IT+DNCKK G+VF DVV E++NS V+VQ G PTIS++ T GC
Sbjct: 355 VNTTLQIKGKINSITVDNCKKLGLVFDDVVGIGEIINSRDVKVQVMGKVPTISINKTDGC 414
Query: 411 QIYLSKDSLETSISTAKSSEINVLVPNVESDGDWVEHSLPQQYIHLFKEGRFETTPASHS 470
YLSK+SL+ I +AKSSE+NVL+P GD+ E +P+Q+ L+ + TT +
Sbjct: 415 HAYLSKNSLDCEIVSAKSSEMNVLIPT--EGGDFNEFPVPEQFKTLWNGQKLVTTVTEIA 472
Query: 471 G 471
G
Sbjct: 473 G 473
>CAP1_MOUSE (P40124) Adenylyl cyclase-associated protein 1 (CAP 1)
Length = 473
Score = 304 bits (779), Expect = 3e-82
Identities = 189/481 (39%), Positives = 268/481 (55%), Gaps = 23/481 (4%)
Query: 3 ETLIKRLESAVTRLEALSTGIHHSSSASDASDAASDPSVVAFVDLIDQYVSRLSKAADII 62
+ L++RLE AV RLEA+S D+ + P V AF L+ V+ K + I
Sbjct: 4 QNLVERLERAVGRLEAVSHTSDMHCGYGDSPSKGAVPYVQAFDSLLANPVAEYLKMSKEI 63
Query: 63 GGQVLDVTNRVKEAFSVQKELLIKLKQTQKPDPAGLAEFLKPLNEVIMKASSLTEGRR-S 121
GG V V +++ LL Q Q+P L++ L P++E I + + E R S
Sbjct: 64 GGDVQKHAEMVHTGLKLERALLATASQCQQPAGNKLSDLLAPISEQIQEVITFREKNRGS 123
Query: 122 DFFNHLKAAVDSLSALAWIAFTGKDCGMSMPIAHVEESWQMAEFYSNKVLVEYRNKDPNH 181
FFNHL A +S+ AL W+A K P V+E A FY+N+VL EYR+ D H
Sbjct: 124 KFFNHLSAVSESIQALGWVALAAK------PGPFVKEMNDAAMFYTNRVLKEYRDVDKKH 177
Query: 182 VEWVKALKELYLPGLRDYVKSFHPLGPVWSQTGKVFAPSKVNASAAPAAPSAPPPPPASL 241
V+WV+A ++ L+ Y+K FH G WS+TG V A + P+ S PPPPP
Sbjct: 178 VDWVRAYLSIWTE-LQAYIKEFHTTGLAWSKTGPV-AKELSGLPSGPSVGSGPPPPPPGP 235
Query: 242 FSSESSQASSSKPKVGMSAVFQEIGTG-NVTAGLRKVTDDMKT-KNRA--DRSGVVGNSV 297
+S S SA+F +I G ++T L+ V+DDMKT KN A +SG V +
Sbjct: 236 PPPPIPTSSGSDDSASRSALFAQINQGESITHALKHVSDDMKTHKNPALKAQSGPVRSGP 295
Query: 298 K-------ESQAAPRAFSKVGPPKLELQMGRKWVVENQIDQKSLVIEDCDSKQSVYVYGC 350
K ++ +P+ +K P LEL+ G+KW VENQ + +LVI+D + KQ Y+Y C
Sbjct: 296 KPFSAPKPQTSPSPKPATKKEPALLELE-GKKWRVENQENVSNLVIDDTELKQVAYIYKC 354
Query: 351 KNSVLQIQGKVNNITIDNCKKTGVVFKDVVAAFEVVNSNGVEVQCQGSAPTISVDNTSGC 410
N+ LQI+GK+N+IT+DNCKK G+VF DVV E++NS V+VQ G PTIS++ T GC
Sbjct: 355 VNTTLQIKGKINSITVDNCKKLGLVFDDVVGIVEIINSRDVKVQVMGKVPTISINKTDGC 414
Query: 411 QIYLSKDSLETSISTAKSSEINVLVPNVESDGDWVEHSLPQQYIHLFKEGRFETTPASHS 470
YLSK+SL+ I +AKSSE+NVL+P GD+ E +P+Q+ L+ + TT +
Sbjct: 415 HAYLSKNSLDCEIVSAKSSEMNVLIPT--EGGDFNEFPVPEQFKTLWNGQKLVTTVTEIA 472
Query: 471 G 471
G
Sbjct: 473 G 473
>CAP2_HUMAN (P40123) Adenylyl cyclase-associated protein 2 (CAP 2)
Length = 477
Score = 301 bits (772), Expect = 2e-81
Identities = 188/481 (39%), Positives = 272/481 (56%), Gaps = 32/481 (6%)
Query: 5 LIKRLESAVTRLEALSTGIHH---SSSASDASDAASDPSVVAFVDLIDQYVSRLSKAADI 61
L++RLE AV+RLE+LS H + + A PSV AF L+D V+ K + I
Sbjct: 7 LVERLERAVSRLESLSAESHRPPGNCGEVNGVIAGVAPSVEAFDKLMDSMVAEFLKNSRI 66
Query: 62 IGGQVLDVTNRVKEAFSVQKELLIKLKQTQKPDPAGLAEFLKPLNEVIMKASSLTEGRR- 120
+ G V V AF Q+ L+ Q Q+P +A LKP++E I + + E R
Sbjct: 67 LAGDVETHAEMVHSAFQAQRAFLLMASQYQQPHENDVAALLKPISEKIQEIQTFRERNRG 126
Query: 121 SDFFNHLKAAVDSLSALAWIAFTGKDCGMSMPIAHVEESWQMAEFYSNKVLVEYRNKDPN 180
S+ FNHL A +S+ AL WIA + K P +V+E A FY+N+VL +Y++ D
Sbjct: 127 SNMFNHLSAVSESIPALGWIAVSPK------PGPYVKEMNDAATFYTNRVLKDYKHSDLR 180
Query: 181 HVEWVKALKELYLPGLRDYVKSFHPLGPVWSQTGKV------FAPSKVNASAAPAAPSAP 234
HV+WVK+ ++ L+ Y+K H G WS+TG V F+ P P P
Sbjct: 181 HVDWVKSYLNIWSE-LQAYIKEHHTTGLTWSKTGPVASTVSAFSVLSSGPGLPPPPPPLP 239
Query: 235 PPPPASLFSSESSQASSSKPKVGMSAVFQEIGTGN-VTAGLRKVTDDMKT-KNRADRS-- 290
PP P LF +E + SS + SA+F ++ G +T GLR VTDD KT KN + R+
Sbjct: 240 PPGPPPLFENEGKKEESSPSR---SALFAQLNQGEAITKGLRHVTDDQKTYKNPSLRAQG 296
Query: 291 GVVGNSVKE---SQAAPRAF-SKVGPPKLELQMGRKWVVENQIDQKSLVIEDCDSKQSVY 346
G + K S +P+++ S+ P LEL+ G+KW VE Q D+ LVI + + KQ Y
Sbjct: 297 GQTQSPTKSHTPSPTSPKSYPSQKHAPVLELE-GKKWRVEYQEDRNDLVISETELKQVAY 355
Query: 347 VYGCKNSVLQIQGKVNNITIDNCKKTGVVFKDVVAAFEVVNSNGVEVQCQGSAPTISVDN 406
++ C+ S +QI+GKVN+I IDNCKK G+VF +VV EV+NS +++Q G PTIS++
Sbjct: 356 IFKCEKSTIQIKGKVNSIIIDNCKKLGLVFDNVVGIVEVINSQDIQIQVMGRVPTISINK 415
Query: 407 TSGCQIYLSKDSLETSISTAKSSEINVLVPNVESDGDWVEHSLPQQYIHLFKEGRFETTP 466
T GC IYLS+D+L+ I +AKSSE+N+L+P DGD+ E +P+Q+ + + T P
Sbjct: 416 TEGCHIYLSEDALDCEIVSAKSSEMNILIP---QDGDYREFPIPEQFKTAWDGSKLITEP 472
Query: 467 A 467
A
Sbjct: 473 A 473
>CAP2_RAT (P52481) Adenylyl cyclase-associated protein 2 (CAP 2)
Length = 477
Score = 301 bits (770), Expect = 3e-81
Identities = 183/478 (38%), Positives = 271/478 (56%), Gaps = 26/478 (5%)
Query: 5 LIKRLESAVTRLEALSTGIHHSSSASDASDAASD----PSVVAFVDLIDQYVSRLSKAAD 60
L++RLE AV+RLE LS+G+H ++ S PSV AF LI+ V+ K +
Sbjct: 7 LMQRLEFAVSRLERLSSGLHEPPGGCGEVNSLSRGIVAPSVEAFDKLINSMVAEFLKNSR 66
Query: 61 IIGGQVLDVTNRVKEAFSVQKELLIKLKQTQKPDPAGLAEFLKPLNEVIMKASSLTEGRR 120
++ G V V AF Q+ L+ + Q Q+P +A LKP++E I + + E R
Sbjct: 67 VLAGDVETHAEMVHGAFQAQRAFLLMVSQHQQPQENEVAVLLKPISEKIQEIQTFRERNR 126
Query: 121 -SDFFNHLKAAVDSLSALAWIAFTGKDCGMSMPIAHVEESWQMAEFYSNKVLVEYRNKDP 179
SD FNHL A +S++AL WIA + K P +V+E A FY+N+VL +Y++ D
Sbjct: 127 GSDMFNHLSAVSESIAALGWIAVSPK------PGPYVKEMNDAATFYTNRVLRDYKHSDL 180
Query: 180 NHVEWVKALKELYLPGLRDYVKSFHPLGPVWSQTGKVFAPSK---VNASAAPAAPSAPPP 236
HV+WV++ +++ L+ Y+K H G WS+TG V + + + +S P PPP
Sbjct: 181 RHVDWVRSYLKIWSE-LQAYIKEHHTTGLTWSKTGPVASTASAFSILSSGPGLPPPPPPP 239
Query: 237 PPASLFSSESSQASSSKPKVGMSAVFQEIGTGN-VTAGLRKVTDDMKT-KNRADRS-GVV 293
PP ++ +P SA+F ++ G +T GLR VTDD K KN + R+ G +
Sbjct: 240 PPPGPPPPFENEGGKEEPSPSRSALFAQLNQGEAITKGLRHVTDDKKIYKNPSLRAQGQI 299
Query: 294 GNSVKESQAAPRAFSKVGP----PKLELQMGRKWVVENQIDQKSLVIEDCDSKQSVYVYG 349
+ K +P + P P LEL+ G+KW VE Q D+ LVI + + KQ Y++
Sbjct: 300 RSPTKTRTPSPTSSKSNSPQKHAPVLELE-GKKWRVEYQEDRNDLVISETELKQVAYIFK 358
Query: 350 CKNSVLQIQGKVNNITIDNCKKTGVVFKDVVAAFEVVNSNGVEVQCQGSAPTISVDNTSG 409
C S LQI+GKVN+IT+DNCKK G+VF VV EV+NS +++Q G PTIS++ T G
Sbjct: 359 CDKSTLQIKGKVNSITVDNCKKFGLVFDHVVGIVEVINSKDIQIQVMGRVPTISINKTEG 418
Query: 410 CQIYLSKDSLETSISTAKSSEINVLVPNVESDGDWVEHSLPQQYIHLFKEGRFETTPA 467
C +YLSKD+L+ I +AKSSE+NVLVP + D+ E +P+Q+ ++ + T PA
Sbjct: 419 CHLYLSKDALDCEIVSAKSSEMNVLVPQGD---DYREFPIPEQFKTIWDGSKLITEPA 473
>CAP2_MOUSE (Q9CYT6) Adenylyl cyclase-associated protein 2 (CAP 2)
Length = 476
Score = 295 bits (754), Expect = 2e-79
Identities = 183/480 (38%), Positives = 270/480 (56%), Gaps = 31/480 (6%)
Query: 5 LIKRLESAVTRLEALSTGIH---HSSSASDASDAASDPSVVAFVDLIDQYVSRLSKAADI 61
L++RLE AV RLE LS G+ + + PSV AF LI+ V+ K + +
Sbjct: 7 LMERLERAVIRLEQLSAGLDGPPRGCGEVNGVNGGVAPSVEAFDKLINSMVAEFLKNSRV 66
Query: 62 IGGQVLDVTNRVKEAFSVQKELLIKLKQTQKPDPAGLAEFLKPLNEVIMKASSLTEGRR- 120
+ G V V AF Q+ L+ + Q Q+P +A LKP++E I + + E R
Sbjct: 67 LAGDVETHAEMVHGAFQAQRAFLLMVSQYQQPQENEVAVLLKPISEKIQEIQTFRERNRG 126
Query: 121 SDFFNHLKAAVDSLSALAWIAFTGKDCGMSMPIAHVEESWQMAEFYSNKVLVEYRNKDPN 180
S+ FNHL A +S++AL WIA + K P +V+E A FY+N+VL +Y++ D
Sbjct: 127 SNMFNHLSAVSESIAALGWIAVSPK------PGPYVKEMNDAATFYTNRVLKDYKHSDLR 180
Query: 181 HVEWVKALKELYLPGLRDYVKSFHPLGPVWSQTGKVFAPSKVNA--SAAPAAPSAPPPPP 238
HV+WV++ ++ L+ Y++ H G WS+TG V + + + S+ P P PPPPP
Sbjct: 181 HVDWVRSYLNIWSE-LQAYIREHHTTGLTWSKTGPVASTASAFSILSSGPGLPPPPPPPP 239
Query: 239 AS----LFSSESSQASSSKPKVGMSAVFQEIGTGN-VTAGLRKVTDDMKT-KNRADRS-G 291
F +E + +P SA+F ++ G +T GLR VTDD KT KN + R+ G
Sbjct: 240 PPGPPPPFENEDKK---EEPSPSRSALFAQLNQGEAITKGLRHVTDDKKTYKNPSLRAQG 296
Query: 292 VVGNSVKESQAAPRAFSKVGP----PKLELQMGRKWVVENQIDQKSLVIEDCDSKQSVYV 347
+ + K +P + P P LEL+ G+KW VE Q D+ LVI + + KQ Y+
Sbjct: 297 QIRSPTKTHTPSPTSPKSNSPQKHTPVLELE-GKKWRVEYQEDRNDLVISETELKQVAYI 355
Query: 348 YGCKNSVLQIQGKVNNITIDNCKKTGVVFKDVVAAFEVVNSNGVEVQCQGSAPTISVDNT 407
+ C S LQI+GKVN+IT+DNCKK G+VF VV EV+NS +++Q G PTIS++ T
Sbjct: 356 FKCDKSTLQIKGKVNSITVDNCKKFGLVFDHVVGIVEVINSKDIQIQVMGRVPTISINKT 415
Query: 408 SGCQIYLSKDSLETSISTAKSSEINVLVPNVESDGDWVEHSLPQQYIHLFKEGRFETTPA 467
GC +YLS+D+L+ I +AKSSE+NVLVP D D+ E +P+Q+ ++ + T PA
Sbjct: 416 EGCHLYLSEDALDCEIVSAKSSEMNVLVP---QDDDYREFPIPEQFKTIWDGSKLVTEPA 472
>CAP_YEAST (P17555) Adenylyl cyclase-associated protein (CAP)
Length = 526
Score = 249 bits (635), Expect = 1e-65
Identities = 166/521 (31%), Positives = 260/521 (49%), Gaps = 64/521 (12%)
Query: 5 LIKRLESAVTRLEALS-------------------------TGIHHSSSASDASDAASDP 39
L+KRLE A RLE ++ + SA +A + DP
Sbjct: 16 LLKRLEEATARLEDVTIYQEGYIQNKLEASKNNKPSDSGADANTTNEPSAENAPEVEQDP 75
Query: 40 S-VVAFVDLIDQYVSRLSKAADIIGGQVLDVTNRVKEAFSVQKELLIKLKQTQKPDPAG- 97
+ AF I + + L + + I VLD +K F Q L +++KPD +
Sbjct: 76 KCITAFQSYIGENIDPLVELSGKIDTVVLDALQLLKGGFQSQLTFLRAAVRSRKPDYSSQ 135
Query: 98 -LAEFLKPLNEVIMKASSLTEG-RRSDFFNHLKAAVDSLSALAWIAFTGKDCGMSMPIAH 155
A+ L+P+NE I+K L E R+S +F +L A + +W+A + P++
Sbjct: 136 TFADSLRPINENIIKLGQLKESNRQSKYFAYLSALSEGAPLFSWVA-------VDTPVSM 188
Query: 156 VEESWQMAEFYSNKVLVEYRNKDPNHVEWVKALKELYLPGLRDYVKSFHPLGPVWSQTGK 215
V + A+F++N++L EYR DPN VEWVK + L+ Y+K +H G W + G
Sbjct: 189 VTDFKDAAQFWTNRILKEYRESDPNAVEWVKKFLASF-DNLKAYIKEYHTTGVSWKKDGM 247
Query: 216 VFA---------------PSKVNASAAPAAPSAPPPPPASLF--SSESSQASSSKPKVGM 258
FA PS +A+AAPA P PP PPAS+F S+++ SS K G+
Sbjct: 248 DFADAMAQSTKNTGATSSPSPASATAAPAPPPPPPAPPASVFEISNDTPATSSDANKGGI 307
Query: 259 SAVFQEIGTG-NVTAGLRKVTDDMKTKNRAD--RSGVVGNSVKESQAAPR-----AFSKV 310
AVF E+ G N+T GL+KV +T + +S V ++ +S PR
Sbjct: 308 GAVFAELNQGENITKGLKKVDKSQQTHKNPELRQSSTVSSTGSKSGPPPRPKKPSTLKTK 367
Query: 311 GPPKLELQMGRKWVVENQIDQKSLVIEDCDSKQSVYVYGCKNSVLQIQGKVNNITIDNCK 370
PP+ EL +G KW +EN ++ ++ D + +S+++ C ++QI+GKVN I++ +
Sbjct: 368 RPPRKEL-VGNKWFIENYENETESLVIDANKDESIFIGKCSQVLVQIKGKVNAISLSETE 426
Query: 371 KTGVVFKDVVAAFEVVNSNGVEVQCQGSAPTISVDNTSGCQIYLSKDSLETSISTAKSSE 430
VV ++ +V+ SN +Q S P IS+D + G IYLSK+SL T I T+ S+
Sbjct: 427 SCSVVLDSSISGMDVIKSNKFGIQVNHSLPQISIDKSDGGNIYLSKESLNTEIYTSCSTA 486
Query: 431 INVLVPNVESDGDWVEHSLPQQYIHLFKEGRFETTPASHSG 471
INV +P + D D+VE +P+Q H F +G+F++ H+G
Sbjct: 487 INVNLP-IGEDDDYVEFPIPEQMKHSFADGKFKSAVFEHAG 526
>CAP_CHLVR (P40122) Adenylyl cyclase-associated protein (CAP)
Length = 481
Score = 245 bits (626), Expect = 2e-64
Identities = 160/477 (33%), Positives = 237/477 (49%), Gaps = 43/477 (9%)
Query: 3 ETLIKRLESAVTRLEALSTGIHHSSSASDASDAASDPSVVAFVDLI---DQYVSRLSKAA 59
E L+ RLE+ RLEA++ S AA D F D +++S
Sbjct: 2 EQLVSRLEAVTNRLEAVA-----SRGGGSTIKAAVDDDPAWFDDFKVFNAKFLSGFVNDT 56
Query: 60 DIIGGQVLDVTNRVKEAFSVQKELLIKLKQTQKPDPAGLAEFLKPLNEVIMKASSLTEGR 119
+GG++ + V EAF L+ + +P L LKPL+E I K E
Sbjct: 57 IALGGELEQMGKFVNEAFHCHLVLMEVAARHNRPSQTDLEGLLKPLSEAISKVQDFREKN 116
Query: 120 RSDF-FNHLKAAVDSLSALAWIAFTGKDCGMSMPIAHVEESWQMAEFYSNKVLVEYRNKD 178
RS FNHL A + L L W+ K P+ ++++ + A+FY+NK+L E+R D
Sbjct: 117 RSSKQFNHLSAISEGLPFLGWVGVAPK------PVLYIQQMEESAQFYTNKLLKEFRESD 170
Query: 179 PNHVEWVKALKELYLPGLRDYVKSFHPLGPVWSQTGKVFAPSKVNASA----APAAPSA- 233
P W + +L L G YVK H G +W++ P+ + SA P PS
Sbjct: 171 PKQANWATSFIQL-LKGFAAYVKDHHQAGLMWNKEKSAATPAALAVSAHKPPVPPPPSGF 229
Query: 234 --PPPPPASLFSSESSQASSSKPKVGMSAVFQEIGTGN-VTAGLRKVTDDMKTKNRADRS 290
PPPPP + + S + S +F ++ G+ VTAGL+KVTDDMKT +
Sbjct: 230 APPPPPPIQAPTVTHAVTGSHSSEDSRSQLFAQLSKGSEVTAGLKKVTDDMKTHKNPELR 289
Query: 291 GVVGNSVKESQAAPRAFSKVG--------------PPKLELQMGRKWVVENQIDQKSLVI 336
+K S PR ++ P +LQ +KWV+ENQ +L I
Sbjct: 290 NQP--PLKSSALDPRPYTPPNLKKFSAPVSKPAKKPALFQLQ-NKKWVIENQDGNTNLEI 346
Query: 337 EDCDSKQSVYVYGCKNSVLQIQGKVNNITIDNCKKTGVVFKDVVAAFEVVNSNGVEVQCQ 396
+C+ KQ+VY+Y C S + I GKVN+I +D+C+K + F DV++ FE N +VQ
Sbjct: 347 SECNDKQTVYMYKCHASKVHINGKVNSIILDSCEKCVIEFTDVISTFEFTNCKACKVQID 406
Query: 397 GSAPTISVDNTSGCQIYLSKDSLETSISTAKSSEINVLVPNVESDGDWVEHSLPQQY 453
G APTIS++ T G Q++++ LE+ I TAKSSE+N+ V ++ DGD E+ LP+Q+
Sbjct: 407 GFAPTISIEKTDGAQVFINPKCLESQIVTAKSSEMNICV--MKPDGDLTEYPLPEQF 461
>CAP_SCHPO (P36621) Adenylyl cyclase-associated protein (CAP)
Length = 551
Score = 233 bits (595), Expect = 6e-61
Identities = 145/458 (31%), Positives = 238/458 (51%), Gaps = 35/458 (7%)
Query: 27 SSASDASDAASDPSVVAFVDLIDQYVSRLSKAADIIGGQVLDVTNRVKEAFSVQKELLIK 86
S S ++ A S+ A+ + +Y+S+ + + IGG + + + V++AF++ +++L
Sbjct: 92 SLTSTSAVEAVPASISAYDEFCSKYLSKYMELSKKIGGLIAEQSEHVEKAFNLLRQVLSV 151
Query: 87 LKQTQKPDPAG--LAEFLKPLNEVIMKASSLT-EGRRSDFFNHLKAAVDSLSALAWIAFT 143
+ QKPD L EFLKP+ ++ +++ E R + FN L + +S L W+
Sbjct: 152 ALKAQKPDMDSPELLEFLKPIQSELLTITNIRDEHRTAPEFNQLSTVMSGISILGWVTVE 211
Query: 144 GKDCGMSMPIAHVEESWQMAEFYSNKVLVEYRNKDPNHVEWVKALKELYLPGLRDYVKSF 203
P++ + E ++FY+N+V+ E++ KD +EWV++ L L L YVK+
Sbjct: 212 ------PTPLSFMSEMKDSSQFYANRVMKEFKGKDDLQIEWVRSYLTL-LTELITYVKTH 264
Query: 204 HPLGPVWS---------------QTGKVFAPSKVNASAAPAAPSAPPPPPASLFSSESSQ 248
G WS K APS +++ P PPPPP++ F +S++
Sbjct: 265 FKTGLTWSTKQDAVPLKTALANLSASKTQAPSSGDSANGGLPPPPPPPPPSNDFWKDSNE 324
Query: 249 ASSSKPKVGMSAVFQEIGTGN-VTAGLRKVT-DDMKTKNRADRS-----GVVGNSVKESQ 301
+ + K M AVF EI G +T+GLRKV +M KN R G +
Sbjct: 325 PAPADNKGDMGAVFAEINKGEGITSGLRKVDKSEMTHKNPNLRKTGPTPGPKPKIKSSAP 384
Query: 302 AAPRAFSKVGPPKLELQMGRKWVVENQIDQKSLVIEDCDSKQSVYVYGCKNSVLQIQGKV 361
+ P + V PP++EL+ KW VENQ+D S+V++ + SV ++GC N + I+GK+
Sbjct: 385 SKPAETAPVKPPRIELE-NTKWFVENQVDNHSIVLDSVELNHSVQIFGCSNCTIIIKGKL 443
Query: 362 NNITIDNCKKTGVVFKDVVAAFEVVNSNGVEVQCQGSAPTISVDNTSGCQIYLSKDSLET 421
N +++ NCK+T VV +VAAF++ + Q P I +D G IYLSK SL +
Sbjct: 444 NTVSMSNCKRTSVVVDTLVAAFDIAKCSNFGCQVMNHVPMIVIDQCDGGSIYLSKSSLSS 503
Query: 422 SISTAKSSEINVLVPNVESDGDWVEHSLPQQYIHLFKE 459
+ T+KS+ +N+ VPN E GD+ E ++P+Q H E
Sbjct: 504 EVVTSKSTSLNINVPNEE--GDYAERAVPEQIKHKVNE 539
>CAP1_PIG (P40125) Adenylyl cyclase-associated protein 1 (CAP 1)
(ASP-56 protein) (Fragments)
Length = 233
Score = 87.0 bits (214), Expect = 9e-17
Identities = 84/267 (31%), Positives = 117/267 (43%), Gaps = 77/267 (28%)
Query: 190 ELYLPGLRDYVKSFHPLGPVWSQTGKVFAPSKVNASAAPAAPSAPPPPPASLFSSESSQA 249
E+ GL++Y K GPV A + P+A S PPP
Sbjct: 31 EMVHTGLKEY-KDVDKXGPV--------AKELSGLPSGPSAGSGPPP------------- 68
Query: 250 SSSKPKVGMSAVFQEIGTG-NVTAGLRKVTDDMKT-KNRADR--SGVVGNSVKESQAAPR 305
SA+F +I G ++T L+ V DD KT KN AD+ SG V K P
Sbjct: 69 ---------SALFAQIHQGESITHALKHVADDEKTHKNPADKAQSGPVRXGPK-----PF 114
Query: 306 AFSKVGPPKLELQMGRKWVVENQIDQKSLVIEDCDSKQSVYVYGCKNSVLQIQGKVNNIT 365
+ SK G + +KW EN + +LVI+D + K NS LQI+ +N+I+
Sbjct: 115 SASKPGISPSPKPVTKKWRXENXENVSNLVIDDTELKSV-------NSTLQIKXXINSIS 167
Query: 366 IDNCKKTGVVFKDVVAAFEVVNSNGVEVQCQGSAPTISVDNTSGCQIYLSKDSLETSIST 425
+DN K P IS++ G IYLSK+SL+ I +
Sbjct: 168 VDNYK----------------------------VPXISINKXDGRHIYLSKNSLDCEIVS 199
Query: 426 AKSSEINVLVPNVESDGDWVEHSLPQQ 452
AKSSE+NVL+P GD+ E +P+Q
Sbjct: 200 AKSSEMNVLIPT--EGGDFNEFPVPEQ 224
>VRP1_YEAST (P37370) Verprolin
Length = 817
Score = 38.5 bits (88), Expect = 0.038
Identities = 32/113 (28%), Positives = 49/113 (43%), Gaps = 14/113 (12%)
Query: 225 SAAPAAPSAPPPPPASLFSSESSQASSSKPKVGMSAVFQEIGTGN---VTAGLRKVTDDM 281
S A + PSAPPPPP ++ ASS + K+ S+ + G A ++K DD
Sbjct: 432 SVATSVPSAPPPPPT--LTTNKPSASSKQSKISSSSSSSAVTPGGPLPFLAEIQKKRDDR 489
Query: 282 KT-------KNRADRSGVVGNSVKESQAAPRAFS-KVGPPKLELQMGRKWVVE 326
+ + V+G+S K+ P S + PPK Q G ++ E
Sbjct: 490 FVVGGDTGYTTQDKQEDVIGSS-KDDNVRPSPISPSINPPKQSSQNGMSFLDE 541
Score = 33.5 bits (75), Expect = 1.2
Identities = 14/32 (43%), Positives = 21/32 (64%)
Query: 223 NASAAPAAPSAPPPPPASLFSSESSQASSSKP 254
N +++ AP PPPPP FS+ S+ ++SS P
Sbjct: 390 NKASSMPAPPPPPPPPPGAFSTSSALSASSIP 421
>LA17_YEAST (Q12446) Proline-rich protein LAS17
Length = 633
Score = 37.4 bits (85), Expect = 0.084
Identities = 24/59 (40%), Positives = 32/59 (53%), Gaps = 4/59 (6%)
Query: 200 VKSFHPLGP-VWSQTGKVFAPSKVNASAAPAA---PSAPPPPPASLFSSESSQASSSKP 254
V +F+ L P + + TG+ P N A A P APPPPPASL S+ Q++ S P
Sbjct: 432 VTTFNTLTPQMTAATGQPAVPLPQNTQAPSQATNVPVAPPPPPASLGQSQIPQSAPSAP 490
>WFS1_HUMAN (O76024) Wolframin
Length = 890
Score = 35.4 bits (80), Expect = 0.32
Identities = 24/111 (21%), Positives = 45/111 (39%), Gaps = 8/111 (7%)
Query: 222 VNASAAPAAPSAPPPPPASLFSSES--------SQASSSKPKVGMSAVFQEIGTGNVTAG 273
++++ AP PS P PPPA + S Q S +P+ G + A
Sbjct: 1 MDSNTAPLGPSCPQPPPAPQPQARSRLNATASLEQERSERPRAPGPQAGPGPGVRDAAAP 60
Query: 274 LRKVTDDMKTKNRADRSGVVGNSVKESQAAPRAFSKVGPPKLELQMGRKWV 324
+++ RAD +G ++ +K G PK + ++G+ ++
Sbjct: 61 AEPQAQHTRSRERADGTGPTKGDMEIPFEEVLERAKAGDPKAQTEVGKHYL 111
>PHR2_HUMAN (O75167) Phosphatase and actin regulator 2
Length = 634
Score = 35.4 bits (80), Expect = 0.32
Identities = 29/113 (25%), Positives = 40/113 (34%), Gaps = 20/113 (17%)
Query: 211 SQTGKVFAPSKVNASAAPAAPSAPPPPPAS--------------------LFSSESSQAS 250
S G P K N A PPP PAS +S S+ ++
Sbjct: 184 SHKGDEVPPIKKNTKAPGKQAPVPPPKPASRNTTREAAGSSHSKKTTGSKASASPSTSST 243
Query: 251 SSKPKVGMSAVFQEIGTGNVTAGLRKVTDDMKTKNRADRSGVVGNSVKESQAA 303
SS+PK V + GT T G RK T + + + G S + + A
Sbjct: 244 SSRPKASKETVSSKAGTVGTTKGKRKTDKQPITSHLSSDTTTSGTSDLKGEPA 296
>NRT_DROME (P23654) Neurotactin
Length = 846
Score = 35.4 bits (80), Expect = 0.32
Identities = 30/113 (26%), Positives = 49/113 (42%), Gaps = 8/113 (7%)
Query: 194 PGLRDYVKSFHPLGPVWSQ------TGKVFAPSKVNASAAPAAPSAPPPPPASLFSSESS 247
PGL + ++SF P + G A S+ A A PA ++ PP L ++
Sbjct: 171 PGLGERLRSFFARKPSAEKEKKQLVNGDADAKSEATAEATPAEDASDAPPKRGLLNAIKL 230
Query: 248 QASSSKPKVGMSAVFQEIGTGNV-TAGLRKVTDDMKTKNRADRSGVVGNSVKE 299
++ PK S E+G G A + + D +K ++ DR+ V N +E
Sbjct: 231 PIANMIPK-KKSNDDVELGLGKAGLASMETLDDSLKDQDTVDRAPVKTNGTEE 282
>MAPB_RAT (P15205) Microtubule-associated protein 1B (MAP 1B)
(Neuraxin) [Contains: MAP1 light chain LC1]
Length = 2459
Score = 35.0 bits (79), Expect = 0.42
Identities = 39/138 (28%), Positives = 59/138 (42%), Gaps = 10/138 (7%)
Query: 215 KVFAPSKVNASAAPAAPSAPPPPPASL--FSSESSQASSSKPKVGMSAVFQEIGTGNVTA 272
K + S V + PSA P P +L S + S+ +S K K + + T V A
Sbjct: 2242 KTKSSSPVKKGDGKSKPSAASPKPGALKESSDKVSRVASPKKKESVEKAMKTTTTPEVKA 2301
Query: 273 GLRKVTDDMKTKNRADRSGVVGNSVKESQAAP----RAFSKVGPPKLELQMGRKWVVENQ 328
R D +TKN A+ S SVK + A P A S PP L + + + + N
Sbjct: 2302 -TRGEEKDKETKNAANAS--ASKSVKTATAGPGTTKTAKSSTVPPGLPVYLDLCY-IPNH 2357
Query: 329 IDQKSLVIEDCDSKQSVY 346
+ K++ +E +S Y
Sbjct: 2358 SNSKNVDVEFFKRVRSSY 2375
>EMP1_WHEAT (P25032) DNA-binding EMBP-1 protein
Length = 354
Score = 34.7 bits (78), Expect = 0.55
Identities = 38/168 (22%), Positives = 65/168 (38%), Gaps = 22/168 (13%)
Query: 181 HVEWVKALKELYLPGLR--DYVKSFHPLGPVWSQTGKVFAPSKVNASAAPAAPSAPPPPP 238
H EW ++ Y + + PL P Q G V A + A A A P PPPP
Sbjct: 26 HAEWAASMHAYYAAAASAAGHPYAAWPLPPQAQQHGLVAAGAGA-AYGAGAVPHVPPPPA 84
Query: 239 AS------------LFSSESSQASSSKPKVGMSAVFQEIGTGNVTAGLRKVTDDMKTKNR 286
+ + ES+ A+ +VG + Q + +G + + + D ++
Sbjct: 85 GTRHAHASMAAGVPYMAGESASAAGKGKRVGKT---QRVPSGEINSS--SGSGDAGSQGS 139
Query: 287 ADRSGVVGNSVKESQAAPRAFSKVGPPKLELQMGRKWVVENQIDQKSL 334
+++ N S +A R K G K E + + V+N + + L
Sbjct: 140 SEKGDAGANQKGSSSSAKR--RKSGAAKTEGEPSQAATVQNAVTEPPL 185
>TCOF_HUMAN (Q13428) Treacle protein (Treacher Collins syndrome
protein)
Length = 1411
Score = 34.3 bits (77), Expect = 0.71
Identities = 28/109 (25%), Positives = 42/109 (37%), Gaps = 8/109 (7%)
Query: 221 KVNASAAPA----APSAPPPPPASLFSSESSQASSSKPKVGMSAVFQEIGTGNVTAGLRK 276
+V A++APA A P PP + +SQ + KP+ + +E K
Sbjct: 222 QVRAASAPAKGTPGKGATPAPPGKA-GAVASQTKAGKPEEDSESSSEESSDSEEETPAAK 280
Query: 277 VTDDMKTKNRADRSGVVGNSVKESQ---AAPRAFSKVGPPKLELQMGRK 322
K + + G KES AAP K GP + Q G++
Sbjct: 281 ALLQAKASGKTSQVGAASAPAKESPRKGAAPAPPGKTGPAVAKAQAGKR 329
>TAF6_RAT (Q63801) Transcription initiation factor TFIID subunit 6
(Transcription initiation factor TFIID 70 kDa subunit)
(TAF(II)70) (TAFII-70) (TAFII-80) (TAFII80) (p80)
Length = 678
Score = 34.3 bits (77), Expect = 0.71
Identities = 20/58 (34%), Positives = 28/58 (47%), Gaps = 1/58 (1%)
Query: 214 GKVFAPSKVNASAAPAAPSAPPPPPASLFSSESSQASSSKPKVGMSAVFQEIGTGNVT 271
G + P + SA AAP P PPP SS ++SS +V +S G+G+ T
Sbjct: 511 GSIALPVQTLVSARAAAPPQPSPPPTKFIVMSSSSSASSTQQV-LSLSTSAPGSGSTT 567
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.312 0.128 0.363
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 54,093,987
Number of Sequences: 164201
Number of extensions: 2249299
Number of successful extensions: 11921
Number of sequences better than 10.0: 101
Number of HSP's better than 10.0 without gapping: 32
Number of HSP's successfully gapped in prelim test: 72
Number of HSP's that attempted gapping in prelim test: 11508
Number of HSP's gapped (non-prelim): 277
length of query: 472
length of database: 59,974,054
effective HSP length: 114
effective length of query: 358
effective length of database: 41,255,140
effective search space: 14769340120
effective search space used: 14769340120
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 68 (30.8 bits)
Medicago: description of AC146746.4


