

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146682.5 - phase: 0
(433 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
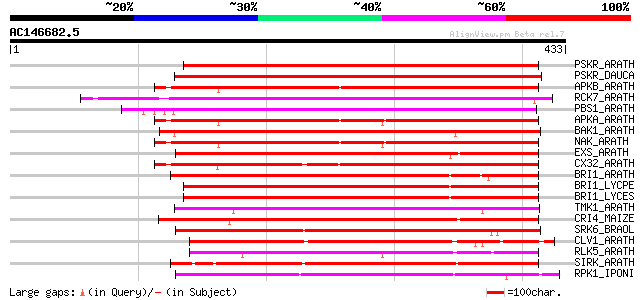
Score E
Sequences producing significant alignments: (bits) Value
PSKR_ARATH (Q9ZVR7) Putative phytosulfokine receptor precursor (... 260 4e-69
PSKR_DAUCA (Q8LPB4) Phytosulfokine receptor precursor (EC 2.7.1.... 259 9e-69
APKB_ARATH (P46573) Protein kinase APK1B, chloroplast precursor ... 256 8e-68
RCK7_ARATH (Q9LQQ8) Putative serine/threonine-protein kinase RLC... 256 1e-67
PBS1_ARATH (Q9FE20) Serine/threonine-protein kinase PBS1 (EC 2.7... 256 1e-67
APKA_ARATH (Q06548) Protein kinase APK1A, chloroplast precursor ... 255 1e-67
BAK1_ARATH (Q94F62) BRASSINOSTEROID INSENSITIVE 1-associated rec... 254 2e-67
NAK_ARATH (P43293) Probable serine/threonine-protein kinase NAK ... 252 1e-66
EXS_ARATH (Q9LYN8) Leucine-rich repeat receptor protein kinase E... 246 1e-64
CX32_ARATH (P27450) Probable serine/threonine-protein kinase Cx3... 243 5e-64
BRI1_ARATH (O22476) BRASSINOSTEROID INSENSITIVE 1 precursor (EC ... 238 2e-62
BRI1_LYCPE (Q8L899) Systemin receptor SR160 precursor (EC 2.7.1.... 234 4e-61
BRI1_LYCES (Q8GUQ5) Brassinosteroid LRR receptor kinase precurso... 233 7e-61
TMK1_ARATH (P43298) Putative receptor protein kinase TMK1 precur... 225 2e-58
CRI4_MAIZE (O24585) Putative receptor protein kinase CRINKLY4 pr... 219 1e-56
SRK6_BRAOL (Q09092) Putative serine/threonine-protein kinase rec... 215 2e-55
CLV1_ARATH (Q9SYQ8) Receptor protein kinase CLAVATA1 precursor (... 208 2e-53
RLK5_ARATH (P47735) Receptor-like protein kinase 5 precursor (EC... 200 7e-51
SIRK_ARATH (O64483) Senescence-induced receptor-like serine/thre... 199 2e-50
RPK1_IPONI (P93194) Receptor-like protein kinase precursor (EC 2... 191 2e-48
>PSKR_ARATH (Q9ZVR7) Putative phytosulfokine receptor precursor (EC
2.7.1.37) (Phytosulfokine LRR receptor kinase)
Length = 1008
Score = 260 bits (665), Expect = 4e-69
Identities = 123/277 (44%), Positives = 183/277 (65%)
Query: 136 ATRGFEEGNVIGEGGYGVVYRGVLQDGCVVAVKNLHNNKGQAEKEFKVEVEAIGKVRHKN 195
+T F++ N+IG GG+G+VY+ L DG VA+K L + GQ E+EF+ EVE + + +H N
Sbjct: 730 STNSFDQANIIGCGGFGMVYKATLPDGKKVAIKKLSGDCGQIEREFEAEVETLSRAQHPN 789
Query: 196 LVRLVGYCAEGARRMLVYEYVENGNLEQWLHGNVGPTSPLTWDIRMKIAIGTAKGLTYLH 255
LV L G+C R+L+Y Y+ENG+L+ WLH + L W R++IA G AKGL YLH
Sbjct: 790 LVLLRGFCFYKNDRLLIYSYMENGSLDYWLHERNDGPALLKWKTRLRIAQGAAKGLLYLH 849
Query: 256 EGLEPKVVHRDIKSSNILLDKNWNAKVSDFGLAKLLGSEKTHVTTRVMGTFGYVSPEYAS 315
EG +P ++HRDIKSSNILLD+N+N+ ++DFGLA+L+ +THV+T ++GT GY+ PEY
Sbjct: 850 EGCDPHILHRDIKSSNILLDENFNSHLADFGLARLMSPYETHVSTDLVGTLGYIPPEYGQ 909
Query: 316 TGMLNERSDVYSFGVLLMEIITGRSPIDYSRPPGEMNLVDWFKAMVSSRRSDELVDPLIE 375
+ + DVYSFGV+L+E++T + P+D +P G +L+ W M R+ E+ DPLI
Sbjct: 910 ASVATYKGDVYSFGVVLLELLTDKRPVDMCKPKGCRDLISWVVKMKHESRASEVFDPLIY 969
Query: 376 TPPSPRALKRVLLICLRCIDLDVIKRPKMGQIVHMLE 412
+ + + + RVL I C+ + +RP Q+V L+
Sbjct: 970 SKENDKEMFRVLEIACLCLSENPKQRPTTQQLVSWLD 1006
>PSKR_DAUCA (Q8LPB4) Phytosulfokine receptor precursor (EC 2.7.1.37)
(Phytosulfokine LRR receptor kinase)
Length = 1021
Score = 259 bits (662), Expect = 9e-69
Identities = 124/287 (43%), Positives = 180/287 (62%)
Query: 129 SLKEVEMATRGFEEGNVIGEGGYGVVYRGVLQDGCVVAVKNLHNNKGQAEKEFKVEVEAI 188
SL ++ +T F + N+IG GG+G+VY+ L DG VA+K L + GQ ++EF+ EVE +
Sbjct: 732 SLDDILKSTSSFNQANIIGCGGFGLVYKATLPDGTKVAIKRLSGDTGQMDREFQAEVETL 791
Query: 189 GKVRHKNLVRLVGYCAEGARRMLVYEYVENGNLEQWLHGNVGPTSPLTWDIRMKIAIGTA 248
+ +H NLV L+GYC ++L+Y Y++NG+L+ WLH V L W R++IA G A
Sbjct: 792 SRAQHPNLVHLLGYCNYKNDKLLIYSYMDNGSLDYWLHEKVDGPPSLDWKTRLRIARGAA 851
Query: 249 KGLTYLHEGLEPKVVHRDIKSSNILLDKNWNAKVSDFGLAKLLGSEKTHVTTRVMGTFGY 308
+GL YLH+ EP ++HRDIKSSNILL + A ++DFGLA+L+ THVTT ++GT GY
Sbjct: 852 EGLAYLHQSCEPHILHRDIKSSNILLSDTFVAHLADFGLARLILPYDTHVTTDLVGTLGY 911
Query: 309 VSPEYASTGMLNERSDVYSFGVLLMEIITGRSPIDYSRPPGEMNLVDWFKAMVSSRRSDE 368
+ PEY + + DVYSFGV+L+E++TGR P+D +P G +L+ W M + +R E
Sbjct: 912 IPPEYGQASVATYKGDVYSFGVVLLELLTGRRPMDVCKPRGSRDLISWVLQMKTEKRESE 971
Query: 369 LVDPLIETPPSPRALKRVLLICLRCIDLDVIKRPKMGQIVHMLESDD 415
+ DP I + VL I RC+ + RP Q+V LE+ D
Sbjct: 972 IFDPFIYDKDHAEEMLLVLEIACRCLGENPKTRPTTQQLVSWLENID 1018
>APKB_ARATH (P46573) Protein kinase APK1B, chloroplast precursor (EC
2.7.1.-)
Length = 412
Score = 256 bits (654), Expect = 8e-68
Identities = 128/311 (41%), Positives = 203/311 (65%), Gaps = 16/311 (5%)
Query: 114 GHMMEDPNIGWGRWYSLKEVEMATRGFEEGNVIGEGGYGVVYRGVLQD----------GC 163
G +++ PN+ + ++ E++ ATR F +V+GEGG+G V++G + + G
Sbjct: 46 GEILQSPNL---KSFTFAELKAATRNFRPDSVLGEGGFGSVFKGWIDEQTLTASKPGTGV 102
Query: 164 VVAVKNLHNNKGQAEKEFKVEVEAIGKVRHKNLVRLVGYCAEGARRMLVYEYVENGNLEQ 223
V+AVK L+ + Q +E+ EV +G+ H NLV+L+GYC E R+LVYE++ G+LE
Sbjct: 103 VIAVKKLNQDGWQGHQEWLAEVNYLGQFSHPNLVKLIGYCLEDEHRLLVYEFMPRGSLEN 162
Query: 224 WLHGNVGPTSPLTWDIRMKIAIGTAKGLTYLHEGLEPKVVHRDIKSSNILLDKNWNAKVS 283
L PL+W +R+K+A+G AKGL +LH E V++RD K+SNILLD +NAK+S
Sbjct: 163 HLFRRGSYFQPLSWTLRLKVALGAAKGLAFLHNA-ETSVIYRDFKTSNILLDSEYNAKLS 221
Query: 284 DFGLAKLLGS-EKTHVTTRVMGTFGYVSPEYASTGMLNERSDVYSFGVLLMEIITGRSPI 342
DFGLAK + +K+HV+TR+MGT+GY +PEY +TG L +SDVYS+GV+L+E+++GR +
Sbjct: 222 DFGLAKDGPTGDKSHVSTRIMGTYGYAAPEYLATGHLTTKSDVYSYGVVLLEVLSGRRAV 281
Query: 343 DYSRPPGEMNLVDWFKAMVSSRRS-DELVDPLIETPPSPRALKRVLLICLRCIDLDVIKR 401
D +RPPGE LV+W + +++++R ++D ++ S +V + LRC+ ++ R
Sbjct: 282 DKNRPPGEQKLVEWARPLLANKRKLFRVIDNRLQDQYSMEEACKVATLALRCLTFEIKLR 341
Query: 402 PKMGQIVHMLE 412
P M ++V LE
Sbjct: 342 PNMNEVVSHLE 352
>RCK7_ARATH (Q9LQQ8) Putative serine/threonine-protein kinase
RLCKVII (EC 2.7.1.37)
Length = 423
Score = 256 bits (653), Expect = 1e-67
Identities = 150/375 (40%), Positives = 221/375 (58%), Gaps = 17/375 (4%)
Query: 56 GSTPLVSKEISMVKEIDLTRSSEKQTRIEIIED-HDEVGPKKEAEIMVEIGGVRKGDGGG 114
G T V+K K D T+ S T++ D ++E G KE ++ +++ G+ D
Sbjct: 28 GGTTSVAKSD---KRDDQTQPSSDSTKVSPYRDVNNEGGVGKEDQLSLDVKGLNLNDQVT 84
Query: 115 HMMEDPNIGWGRWYSLKEVEMATRGFEEGNVIGEGGYGVVYRGVLQD-GCVVAVKNLHNN 173
+ ++ +E+ AT F +GEGG+G V++G ++ VVA+K L N
Sbjct: 85 GKK-------AQTFTFQELAEATGNFRSDCFLGEGGFGKVFKGTIEKLDQVVAIKQLDRN 137
Query: 174 KGQAEKEFKVEVEAIGKVRHKNLVRLVGYCAEGARRMLVYEYVENGNLEQWLHGNVGPTS 233
Q +EF VEV + H NLV+L+G+CAEG +R+LVYEY+ G+LE LH
Sbjct: 138 GVQGIREFVVEVLTLSLADHPNLVKLIGFCAEGDQRLLVYEYMPQGSLEDHLHVLPSGKK 197
Query: 234 PLTWDIRMKIAIGTAKGLTYLHEGLEPKVVHRDIKSSNILLDKNWNAKVSDFGLAKLLGS 293
PL W+ RMKIA G A+GL YLH+ + P V++RD+K SNILL +++ K+SDFGLAK+ S
Sbjct: 198 PLDWNTRMKIAAGAARGLEYLHDRMTPPVIYRDLKCSNILLGEDYQPKLSDFGLAKVGPS 257
Query: 294 -EKTHVTTRVMGTFGYVSPEYASTGMLNERSDVYSFGVLLMEIITGRSPIDYSRPPGEMN 352
+KTHV+TRVMGT+GY +P+YA TG L +SD+YSFGV+L+E+ITGR ID ++ + N
Sbjct: 258 GDKTHVSTRVMGTYGYCAPDYAMTGQLTFKSDIYSFGVVLLELITGRKAIDNTKTRKDQN 317
Query: 353 LVDWFKAMVSSRRS-DELVDPLIETPPSPRALKRVLLICLRCIDLDVIKRPKMGQIV--- 408
LV W + + RR+ ++VDPL++ R L + L I C+ RP + +V
Sbjct: 318 LVGWARPLFKDRRNFPKMVDPLLQGQYPVRGLYQALAISAMCVQEQPTMRPVVSDVVLAL 377
Query: 409 HMLESDDFPFRSVSS 423
+ L S + S SS
Sbjct: 378 NFLASSKYDPNSPSS 392
>PBS1_ARATH (Q9FE20) Serine/threonine-protein kinase PBS1 (EC
2.7.1.37) (AvrPphB susceptible protein 1)
Length = 456
Score = 256 bits (653), Expect = 1e-67
Identities = 151/342 (44%), Positives = 202/342 (58%), Gaps = 18/342 (5%)
Query: 88 DHDEVGPKKEAEIMVE--IGGVRKGD-------GGGHMMED--PNIGWGR----WYSLKE 132
D G KK+++ V I G+ G GG E P G G+ ++ +E
Sbjct: 19 DESNHGQKKQSQPTVSNNISGLPSGGEKLSSKTNGGSKRELLLPRDGLGQIAAHTFAFRE 78
Query: 133 VEMATRGFEEGNVIGEGGYGVVYRGVLQD-GCVVAVKNLHNNKGQAEKEFKVEVEAIGKV 191
+ AT F +GEGG+G VY+G L G VVAVK L N Q +EF VEV + +
Sbjct: 79 LAAATMNFHPDTFLGEGGFGRVYKGRLDSTGQVVAVKQLDRNGLQGNREFLVEVLMLSLL 138
Query: 192 RHKNLVRLVGYCAEGARRMLVYEYVENGNLEQWLHGNVGPTSPLTWDIRMKIAIGTAKGL 251
H NLV L+GYCA+G +R+LVYE++ G+LE LH L W++RMKIA G AKGL
Sbjct: 139 HHPNLVNLIGYCADGDQRLLVYEFMPLGSLEDHLHDLPPDKEALDWNMRMKIAAGAAKGL 198
Query: 252 TYLHEGLEPKVVHRDIKSSNILLDKNWNAKVSDFGLAKL-LGSEKTHVTTRVMGTFGYVS 310
+LH+ P V++RD KSSNILLD+ ++ K+SDFGLAKL +K+HV+TRVMGT+GY +
Sbjct: 199 EFLHDKANPPVIYRDFKSSNILLDEGFHPKLSDFGLAKLGPTGDKSHVSTRVMGTYGYCA 258
Query: 311 PEYASTGMLNERSDVYSFGVLLMEIITGRSPIDYSRPPGEMNLVDWFKAMVSSRRS-DEL 369
PEYA TG L +SDVYSFGV+ +E+ITGR ID P GE NLV W + + + RR +L
Sbjct: 259 PEYAMTGQLTVKSDVYSFGVVFLELITGRKAIDSEMPHGEQNLVAWARPLFNDRRKFIKL 318
Query: 370 VDPLIETPPSPRALKRVLLICLRCIDLDVIKRPKMGQIVHML 411
DP ++ RAL + L + CI RP + +V L
Sbjct: 319 ADPRLKGRFPTRALYQALAVASMCIQEQAATRPLIADVVTAL 360
>APKA_ARATH (Q06548) Protein kinase APK1A, chloroplast precursor (EC
2.7.1.-)
Length = 410
Score = 255 bits (652), Expect = 1e-67
Identities = 133/312 (42%), Positives = 204/312 (64%), Gaps = 18/312 (5%)
Query: 114 GHMMEDPNIGWGRWYSLKEVEMATRGFEEGNVIGEGGYGVVYRGVLQD----------GC 163
G +++ PN+ + +S E++ ATR F +V+GEGG+G V++G + + G
Sbjct: 45 GEILQSPNL---KSFSFAELKSATRNFRPDSVLGEGGFGCVFKGWIDEKSLTASRPGTGL 101
Query: 164 VVAVKNLHNNKGQAEKEFKVEVEAIGKVRHKNLVRLVGYCAEGARRMLVYEYVENGNLEQ 223
V+AVK L+ + Q +E+ EV +G+ H++LV+L+GYC E R+LVYE++ G+LE
Sbjct: 102 VIAVKKLNQDGWQGHQEWLAEVNYLGQFSHRHLVKLIGYCLEDEHRLLVYEFMPRGSLEN 161
Query: 224 WLHGNVGPTSPLTWDIRMKIAIGTAKGLTYLHEGLEPKVVHRDIKSSNILLDKNWNAKVS 283
L PL+W +R+K+A+G AKGL +LH E +V++RD K+SNILLD +NAK+S
Sbjct: 162 HLFRRGLYFQPLSWKLRLKVALGAAKGLAFLHSS-ETRVIYRDFKTSNILLDSEYNAKLS 220
Query: 284 DFGLAK--LLGSEKTHVTTRVMGTFGYVSPEYASTGMLNERSDVYSFGVLLMEIITGRSP 341
DFGLAK +G +K+HV+TRVMGT GY +PEY +TG L +SDVYSFGV+L+E+++GR
Sbjct: 221 DFGLAKDGPIG-DKSHVSTRVMGTHGYAAPEYLATGHLTTKSDVYSFGVVLLELLSGRRA 279
Query: 342 IDYSRPPGEMNLVDWFKA-MVSSRRSDELVDPLIETPPSPRALKRVLLICLRCIDLDVIK 400
+D +RP GE NLV+W K +V+ R+ ++D ++ S +V + LRC+ ++
Sbjct: 280 VDKNRPSGERNLVEWAKPYLVNKRKIFRVIDNRLQDQYSMEEACKVATLSLRCLTTEIKL 339
Query: 401 RPKMGQIVHMLE 412
RP M ++V LE
Sbjct: 340 RPNMSEVVSHLE 351
>BAK1_ARATH (Q94F62) BRASSINOSTEROID INSENSITIVE 1-associated
receptor kinase 1 precursor (EC 2.7.1.37)
(BRI1-associated receptor kinase 1) (Somatic
embryogenesis receptor-like kinase 3)
Length = 615
Score = 254 bits (650), Expect = 2e-67
Identities = 125/302 (41%), Positives = 196/302 (64%), Gaps = 5/302 (1%)
Query: 118 EDPNIGWGRW--YSLKEVEMATRGFEEGNVIGEGGYGVVYRGVLQDGCVVAVKNLHNNKG 175
EDP + G+ +SL+E+++A+ F N++G GG+G VY+G L DG +VAVK L +
Sbjct: 265 EDPEVHLGQLKRFSLRELQVASDNFSNKNILGRGGFGKVYKGRLADGTLVAVKRLKEERT 324
Query: 176 QA-EKEFKVEVEAIGKVRHKNLVRLVGYCAEGARRMLVYEYVENGNLEQWLHGNVGPTSP 234
Q E +F+ EVE I H+NL+RL G+C R+LVY Y+ NG++ L P
Sbjct: 325 QGGELQFQTEVEMISMAVHRNLLRLRGFCMTPTERLLVYPYMANGSVASCLRERPESQPP 384
Query: 235 LTWDIRMKIAIGTAKGLTYLHEGLEPKVVHRDIKSSNILLDKNWNAKVSDFGLAKLLGSE 294
L W R +IA+G+A+GL YLH+ +PK++HRD+K++NILLD+ + A V DFGLAKL+ +
Sbjct: 385 LDWPKRQRIALGSARGLAYLHDHCDPKIIHRDVKAANILLDEEFEAVVGDFGLAKLMDYK 444
Query: 295 KTHVTTRVMGTFGYVSPEYASTGMLNERSDVYSFGVLLMEIITGRSPIDYSR--PPGEMN 352
THVTT V GT G+++PEY STG +E++DV+ +GV+L+E+ITG+ D +R ++
Sbjct: 445 DTHVTTAVRGTIGHIAPEYLSTGKSSEKTDVFGYGVMLLELITGQRAFDLARLANDDDVM 504
Query: 353 LVDWFKAMVSSRRSDELVDPLIETPPSPRALKRVLLICLRCIDLDVIKRPKMGQIVHMLE 412
L+DW K ++ ++ + LVD ++ +++++ + L C ++RPKM ++V MLE
Sbjct: 505 LLDWVKGLLKEKKLEALVDVDLQGNYKDEEVEQLIQVALLCTQSSPMERPKMSEVVRMLE 564
Query: 413 SD 414
D
Sbjct: 565 GD 566
>NAK_ARATH (P43293) Probable serine/threonine-protein kinase NAK (EC
2.7.1.37)
Length = 389
Score = 252 bits (643), Expect = 1e-66
Identities = 131/312 (41%), Positives = 206/312 (65%), Gaps = 18/312 (5%)
Query: 114 GHMMEDPNIGWGRWYSLKEVEMATRGFEEGNVIGEGGYGVVYRGVLQD----------GC 163
G ++++ N+ + +SL E++ ATR F +V+GEGG+G V++G + + G
Sbjct: 45 GEILQNANL---KNFSLSELKSATRNFRPDSVVGEGGFGCVFKGWIDESSLAPSKPGTGI 101
Query: 164 VVAVKNLHNNKGQAEKEFKVEVEAIGKVRHKNLVRLVGYCAEGARRMLVYEYVENGNLEQ 223
V+AVK L+ Q +E+ E+ +G++ H NLV+L+GYC E R+LVYE++ G+LE
Sbjct: 102 VIAVKRLNQEGFQGHREWLAEINYLGQLDHPNLVKLIGYCLEEEHRLLVYEFMTRGSLEN 161
Query: 224 WLHGNVGPTSPLTWDIRMKIAIGTAKGLTYLHEGLEPKVVHRDIKSSNILLDKNWNAKVS 283
L PL+W+ R+++A+G A+GL +LH +P+V++RD K+SNILLD N+NAK+S
Sbjct: 162 HLFRRGTFYQPLSWNTRVRMALGAARGLAFLHNA-QPQVIYRDFKASNILLDSNYNAKLS 220
Query: 284 DFGLAK--LLGSEKTHVTTRVMGTFGYVSPEYASTGMLNERSDVYSFGVLLMEIITGRSP 341
DFGLA+ +G + +HV+TRVMGT GY +PEY +TG L+ +SDVYSFGV+L+E+++GR
Sbjct: 221 DFGLARDGPMG-DNSHVSTRVMGTQGYAAPEYLATGHLSVKSDVYSFGVVLLELLSGRRA 279
Query: 342 IDYSRPPGEMNLVDWFKA-MVSSRRSDELVDPLIETPPSPRALKRVLLICLRCIDLDVIK 400
ID ++P GE NLVDW + + + RR ++DP ++ S ++ ++ L CI +D
Sbjct: 280 IDKNQPVGEHNLVDWARPYLTNKRRLLRVMDPRLQGQYSLTRALKIAVLALDCISIDAKS 339
Query: 401 RPKMGQIVHMLE 412
RP M +IV +E
Sbjct: 340 RPTMNEIVKTME 351
>EXS_ARATH (Q9LYN8) Leucine-rich repeat receptor protein kinase EXS
precursor (EC 2.7.1.37) (Extra sporogenous cells protein)
(EXCESS MICROSPOROCYTES1 protein)
Length = 1192
Score = 246 bits (627), Expect = 1e-64
Identities = 120/285 (42%), Positives = 180/285 (63%), Gaps = 3/285 (1%)
Query: 130 LKEVEMATRGFEEGNVIGEGGYGVVYRGVLQDGCVVAVKNLHNNKGQAEKEFKVEVEAIG 189
L ++ AT F + N+IG+GG+G VY+ L VAVK L K Q +EF E+E +G
Sbjct: 907 LGDIVEATDHFSKKNIIGDGGFGTVYKACLPGEKTVAVKKLSEAKTQGNREFMAEMETLG 966
Query: 190 KVRHKNLVRLVGYCAEGARRMLVYEYVENGNLEQWLHGNVGPTSPLTWDIRMKIAIGTAK 249
KV+H NLV L+GYC+ ++LVYEY+ NG+L+ WL G L W R+KIA+G A+
Sbjct: 967 KVKHPNLVSLLGYCSFSEEKLLVYEYMVNGSLDHWLRNQTGMLEVLDWSKRLKIAVGAAR 1026
Query: 250 GLTYLHEGLEPKVVHRDIKSSNILLDKNWNAKVSDFGLAKLLGSEKTHVTTRVMGTFGYV 309
GL +LH G P ++HRDIK+SNILLD ++ KV+DFGLA+L+ + ++HV+T + GTFGY+
Sbjct: 1027 GLAFLHHGFIPHIIHRDIKASNILLDGDFEPKVADFGLARLISACESHVSTVIAGTFGYI 1086
Query: 310 SPEYASTGMLNERSDVYSFGVLLMEIITGRSPI--DYSRPPGEMNLVDWFKAMVSSRRSD 367
PEY + + DVYSFGV+L+E++TG+ P D+ G NLV W ++ ++
Sbjct: 1087 PPEYGQSARATTKGDVYSFGVILLELVTGKEPTGPDFKESEGG-NLVGWAIQKINQGKAV 1145
Query: 368 ELVDPLIETPPSPRALKRVLLICLRCIDLDVIKRPKMGQIVHMLE 412
+++DPL+ + + R+L I + C+ KRP M ++ L+
Sbjct: 1146 DVIDPLLVSVALKNSQLRLLQIAMLCLAETPAKRPNMLDVLKALK 1190
>CX32_ARATH (P27450) Probable serine/threonine-protein kinase Cx32,
chloroplast precursor (EC 2.7.1.37)
Length = 419
Score = 243 bits (621), Expect = 5e-64
Identities = 129/311 (41%), Positives = 202/311 (64%), Gaps = 19/311 (6%)
Query: 114 GHMMEDPNIGWGRWYSLKEVEMATRGFEEGNVIGEGGYGVVYRGVLQ----------DGC 163
G ++E PN+ + Y+ +++ AT+ F+ +++G+GG+G VYRG + G
Sbjct: 63 GKLLESPNL---KVYNFLDLKTATKNFKPDSMLGQGGFGKVYRGWVDATTLAPSRVGSGM 119
Query: 164 VVAVKNLHNNKGQAEKEFKVEVEAIGKVRHKNLVRLVGYCAEGARRMLVYEYVENGNLEQ 223
+VA+K L++ Q E++ EV +G + H+NLV+L+GYC E +LVYE++ G+LE
Sbjct: 120 IVAIKRLNSESVQGFAEWRSEVNFLGMLSHRNLVKLLGYCREDKELLLVYEFMPKGSLES 179
Query: 224 WLHGNVGPTSPLTWDIRMKIAIGTAKGLTYLHEGLEPKVVHRDIKSSNILLDKNWNAKVS 283
L P WD+R+KI IG A+GL +LH L+ +V++RD K+SNILLD N++AK+S
Sbjct: 180 HLFRR---NDPFPWDLRIKIVIGAARGLAFLHS-LQREVIYRDFKASNILLDSNYDAKLS 235
Query: 284 DFGLAKL-LGSEKTHVTTRVMGTFGYVSPEYASTGMLNERSDVYSFGVLLMEIITGRSPI 342
DFGLAKL EK+HVTTR+MGT+GY +PEY +TG L +SDV++FGV+L+EI+TG +
Sbjct: 236 DFGLAKLGPADEKSHVTTRIMGTYGYAAPEYMATGHLYVKSDVFAFGVVLLEIMTGLTAH 295
Query: 343 DYSRPPGEMNLVDWFKAMVSSR-RSDELVDPLIETPPSPRALKRVLLICLRCIDLDVIKR 401
+ RP G+ +LVDW + +S++ R +++D I+ + + + I L CI+ D R
Sbjct: 296 NTKRPRGQESLVDWLRPELSNKHRVKQIMDKGIKGQYTTKVATEMARITLSCIEPDPKNR 355
Query: 402 PKMGQIVHMLE 412
P M ++V +LE
Sbjct: 356 PHMKEVVEVLE 366
>BRI1_ARATH (O22476) BRASSINOSTEROID INSENSITIVE 1 precursor (EC
2.7.1.37) (AtBRI1) (Brassinosteroid LRR receptor kinase)
Length = 1196
Score = 238 bits (608), Expect = 2e-62
Identities = 125/290 (43%), Positives = 184/290 (63%), Gaps = 5/290 (1%)
Query: 126 RWYSLKEVEMATRGFEEGNVIGEGGYGVVYRGVLQDGCVVAVKNLHNNKGQAEKEFKVEV 185
R + ++ AT GF ++IG GG+G VY+ +L+DG VA+K L + GQ ++EF E+
Sbjct: 869 RKLTFADLLQATNGFHNDSLIGSGGFGDVYKAILKDGSAVAIKKLIHVSGQGDREFMAEM 928
Query: 186 EAIGKVRHKNLVRLVGYCAEGARRMLVYEYVENGNLEQWLHGNVGPTSPLTWDIRMKIAI 245
E IGK++H+NLV L+GYC G R+LVYE+++ G+LE LH L W R KIAI
Sbjct: 929 ETIGKIKHRNLVPLLGYCKVGDERLLVYEFMKYGSLEDVLHDPKKAGVKLNWSTRRKIAI 988
Query: 246 GTAKGLTYLHEGLEPKVVHRDIKSSNILLDKNWNAKVSDFGLAKLLGSEKTHVTTRVM-G 304
G+A+GL +LH P ++HRD+KSSN+LLD+N A+VSDFG+A+L+ + TH++ + G
Sbjct: 989 GSARGLAFLHHNCSPHIIHRDMKSSNVLLDENLEARVSDFGMARLMSAMDTHLSVSTLAG 1048
Query: 305 TFGYVSPEYASTGMLNERSDVYSFGVLLMEIITGRSPIDYSRPPGEMNLVDWFKAMVSSR 364
T GYV PEY + + + DVYS+GV+L+E++TG+ P D S G+ NLV W K R
Sbjct: 1049 TPGYVPPEYYQSFRCSTKGDVYSYGVVLLELLTGKRPTD-SPDFGDNNLVGWVKQHAKLR 1107
Query: 365 RSDELVDP--LIETPPSPRALKRVLLICLRCIDLDVIKRPKMGQIVHMLE 412
SD + DP + E P L + L + + C+D +RP M Q++ M +
Sbjct: 1108 ISD-VFDPELMKEDPALEIELLQHLKVAVACLDDRAWRRPTMVQVMAMFK 1156
>BRI1_LYCPE (Q8L899) Systemin receptor SR160 precursor (EC 2.7.1.37)
(Brassinosteroid LRR receptor kinase)
Length = 1207
Score = 234 bits (596), Expect = 4e-61
Identities = 122/279 (43%), Positives = 176/279 (62%), Gaps = 3/279 (1%)
Query: 136 ATRGFEEGNVIGEGGYGVVYRGVLQDGCVVAVKNLHNNKGQAEKEFKVEVEAIGKVRHKN 195
AT GF +++G GG+G VY+ L+DG VVA+K L + GQ ++EF E+E IGK++H+N
Sbjct: 884 ATNGFHNDSLVGSGGFGDVYKAQLKDGSVVAIKKLIHVSGQGDREFTAEMETIGKIKHRN 943
Query: 196 LVRLVGYCAEGARRMLVYEYVENGNLEQWLHGNVGPTSPLTWDIRMKIAIGTAKGLTYLH 255
LV L+GYC G R+LVYEY++ G+LE LH L W R KIAIG A+GL +LH
Sbjct: 944 LVPLLGYCKVGEERLLVYEYMKYGSLEDVLHDRKKTGIKLNWPARRKIAIGAARGLAFLH 1003
Query: 256 EGLEPKVVHRDIKSSNILLDKNWNAKVSDFGLAKLLGSEKTHVTTRVM-GTFGYVSPEYA 314
P ++HRD+KSSN+LLD+N A+VSDFG+A+L+ + TH++ + GT GYV PEY
Sbjct: 1004 HNCIPHIIHRDMKSSNVLLDENLEARVSDFGMARLMSAMDTHLSVSTLAGTPGYVPPEYY 1063
Query: 315 STGMLNERSDVYSFGVLLMEIITGRSPIDYSRPPGEMNLVDWFKAMVSSRRSDELVDPLI 374
+ + + DVYS+GV+L+E++TG+ P D S G+ NLV W K + +D L+
Sbjct: 1064 QSFRCSTKGDVYSYGVVLLELLTGKQPTD-SADFGDNNLVGWVKLHAKGKITDVFDRELL 1122
Query: 375 ETPPSPR-ALKRVLLICLRCIDLDVIKRPKMGQIVHMLE 412
+ S L + L + C+D KRP M Q++ M +
Sbjct: 1123 KEDASIEIELLQHLKVACACLDDRHWKRPTMIQVMAMFK 1161
>BRI1_LYCES (Q8GUQ5) Brassinosteroid LRR receptor kinase precursor (EC
2.7.1.37) (tBRI1) (Altered brassinolide sensitivity 1)
(Systemin receptor SR160)
Length = 1207
Score = 233 bits (594), Expect = 7e-61
Identities = 122/279 (43%), Positives = 176/279 (62%), Gaps = 3/279 (1%)
Query: 136 ATRGFEEGNVIGEGGYGVVYRGVLQDGCVVAVKNLHNNKGQAEKEFKVEVEAIGKVRHKN 195
AT GF +++G GG+G VY+ L+DG VVA+K L + GQ ++EF E+E IGK++H+N
Sbjct: 884 ATNGFHNDSLVGSGGFGDVYKAQLKDGSVVAIKKLIHVSGQGDREFTAEMETIGKIKHRN 943
Query: 196 LVRLVGYCAEGARRMLVYEYVENGNLEQWLHGNVGPTSPLTWDIRMKIAIGTAKGLTYLH 255
LV L+GYC G R+LVYEY++ G+LE LH L W R KIAIG A+GL +LH
Sbjct: 944 LVPLLGYCKVGEERLLVYEYMKYGSLEDVLHDRKKIGIKLNWPARRKIAIGAARGLAFLH 1003
Query: 256 EGLEPKVVHRDIKSSNILLDKNWNAKVSDFGLAKLLGSEKTHVTTRVM-GTFGYVSPEYA 314
P ++HRD+KSSN+LLD+N A+VSDFG+A+L+ + TH++ + GT GYV PEY
Sbjct: 1004 HNCIPHIIHRDMKSSNVLLDENLEARVSDFGMARLMSAMDTHLSVSTLAGTPGYVPPEYY 1063
Query: 315 STGMLNERSDVYSFGVLLMEIITGRSPIDYSRPPGEMNLVDWFKAMVSSRRSDELVDPLI 374
+ + + DVYS+GV+L+E++TG+ P D S G+ NLV W K + +D L+
Sbjct: 1064 QSFRCSTKGDVYSYGVVLLELLTGKQPTD-SADFGDNNLVGWVKLHAKGKITDVFDRELL 1122
Query: 375 ETPPSPR-ALKRVLLICLRCIDLDVIKRPKMGQIVHMLE 412
+ S L + L + C+D KRP M Q++ M +
Sbjct: 1123 KEDASIEIELLQHLKVACACLDDRHWKRPTMIQVMAMFK 1161
>TMK1_ARATH (P43298) Putative receptor protein kinase TMK1 precursor
(EC 2.7.1.-)
Length = 942
Score = 225 bits (573), Expect = 2e-58
Identities = 119/291 (40%), Positives = 174/291 (58%), Gaps = 6/291 (2%)
Query: 129 SLKEVEMATRGFEEGNVIGEGGYGVVYRGVLQDGCVVAVKNLHNN--KGQAEKEFKVEVE 186
S++ + T F N++G GG+GVVY+G L DG +AVK + N G+ EFK E+
Sbjct: 577 SIQVLRSVTNNFSSDNILGSGGFGVVYKGELHDGTKIAVKRMENGVIAGKGFAEFKSEIA 636
Query: 187 AIGKVRHKNLVRLVGYCAEGARRMLVYEYVENGNLEQWL-HGNVGPTSPLTWDIRMKIAI 245
+ KVRH++LV L+GYC +G ++LVYEY+ G L + L + PL W R+ +A+
Sbjct: 637 VLTKVRHRHLVTLLGYCLDGNEKLLVYEYMPQGTLSRHLFEWSEEGLKPLLWKQRLTLAL 696
Query: 246 GTAKGLTYLHEGLEPKVVHRDIKSSNILLDKNWNAKVSDFGLAKLLGSEKTHVTTRVMGT 305
A+G+ YLH +HRD+K SNILL + AKV+DFGL +L K + TR+ GT
Sbjct: 697 DVARGVEYLHGLAHQSFIHRDLKPSNILLGDDMRAKVADFGLVRLAPEGKGSIETRIAGT 756
Query: 306 FGYVSPEYASTGMLNERSDVYSFGVLLMEIITGRSPIDYSRPPGEMNLVDWFKAMVSSRR 365
FGY++PEYA TG + + DVYSFGV+LME+ITGR +D S+P ++LV WFK M ++
Sbjct: 757 FGYLAPEYAVTGRVTTKVDVYSFGVILMELITGRKSLDESQPEESIHLVSWFKRMYINKE 816
Query: 366 SD--ELVDPLIETPPSPRA-LKRVLLICLRCIDLDVIKRPKMGQIVHMLES 413
+ + +D I+ A + V + C + +RP MG V++L S
Sbjct: 817 ASFKKAIDTTIDLDEETLASVHTVAELAGHCCAREPYQRPDMGHAVNILSS 867
>CRI4_MAIZE (O24585) Putative receptor protein kinase CRINKLY4
precursor (EC 2.7.1.-)
Length = 901
Score = 219 bits (557), Expect = 1e-56
Identities = 117/300 (39%), Positives = 187/300 (62%), Gaps = 6/300 (2%)
Query: 117 MEDPNIGWGRWYSLKEVEMATRGFEEGNVIGEGGYGVVYRGVLQDGCVVAVKNL--HNNK 174
MED I + +S +E+E AT GF E + +G+G + V++G+L+DG VVAVK ++
Sbjct: 482 MEDLKIRRAQEFSYEELEQATGGFSEDSQVGKGSFSCVFKGILRDGTVVAVKRAIKASDV 541
Query: 175 GQAEKEFKVEVEAIGKVRHKNLVRLVGYCAEGARRMLVYEYVENGNLEQWLHG-NVGPTS 233
++ KEF E++ + ++ H +L+ L+GYC +G+ R+LVYE++ +G+L Q LHG +
Sbjct: 542 KKSSKEFHNELDLLSRLNHAHLLNLLGYCEDGSERLLVYEFMAHGSLYQHLHGKDPNLKK 601
Query: 234 PLTWDIRMKIAIGTAKGLTYLHEGLEPKVVHRDIKSSNILLDKNWNAKVSDFGLAKLLGS 293
L W R+ IA+ A+G+ YLH P V+HRDIKSSNIL+D++ NA+V+DFGL+ L +
Sbjct: 602 RLNWARRVTIAVQAARGIEYLHGYACPPVIHRDIKSSNILIDEDHNARVADFGLSILGPA 661
Query: 294 EK-THVTTRVMGTFGYVSPEYASTGMLNERSDVYSFGVLLMEIITGRSPIDYSRPPGEMN 352
+ T ++ GT GY+ PEY L +SDVYSFGV+L+EI++GR ID G N
Sbjct: 662 DSGTPLSELPAGTLGYLDPEYYRLHYLTTKSDVYSFGVVLLEILSGRKAIDMQFEEG--N 719
Query: 353 LVDWFKAMVSSRRSDELVDPLIETPPSPRALKRVLLICLRCIDLDVIKRPKMGQIVHMLE 412
+V+W ++ + ++DP++ P ALK++ + +C+ + RP M ++ LE
Sbjct: 720 IVEWAVPLIKAGDIFAILDPVLSPPSDLEALKKIASVACKCVRMRGKDRPSMDKVTTALE 779
>SRK6_BRAOL (Q09092) Putative serine/threonine-protein kinase
receptor precursor (EC 2.7.1.37) (S-receptor kinase)
(SRK)
Length = 849
Score = 215 bits (547), Expect = 2e-55
Identities = 117/293 (39%), Positives = 180/293 (60%), Gaps = 9/293 (3%)
Query: 130 LKEVEMATRGFEEGNVIGEGGYGVVYRGVLQDGCVVAVKNLHNNKGQAEKEFKVEVEAIG 189
++ V AT F N +G+GG+G+VY+G L DG +AVK L Q EF EV I
Sbjct: 518 METVVKATENFSSCNKLGQGGFGIVYKGRLLDGKEIAVKRLSKTSVQGTDEFMNEVTLIA 577
Query: 190 KVRHKNLVRLVGYCAEGARRMLVYEYVENGNLEQWLHGNVGPTSPLTWDIRMKIAIGTAK 249
+++H NLV+++G C EG +ML+YEY+EN +L+ +L G S L W+ R I G A+
Sbjct: 578 RLQHINLVQVLGCCIEGDEKMLIYEYLENLSLDSYLFGKT-RRSKLNWNERFDITNGVAR 636
Query: 250 GLTYLHEGLEPKVVHRDIKSSNILLDKNWNAKVSDFGLAKLLGSEKTHVTT-RVMGTFGY 308
GL YLH+ +++HRD+K SNILLDKN K+SDFG+A++ ++T T +V+GT+GY
Sbjct: 637 GLLYLHQDSRFRIIHRDLKVSNILLDKNMIPKISDFGMARIFERDETEANTMKVVGTYGY 696
Query: 309 VSPEYASTGMLNERSDVYSFGVLLMEIITGRSPIDYSRPPGEMNLVDWFKAMVSSRRSDE 368
+SPEYA G+ +E+SDV+SFGV+++EI++G+ + E +L+ + + R+ E
Sbjct: 697 MSPEYAMYGIFSEKSDVFSFGVIVLEIVSGKKNRGFYNLDYENDLLSYVWSRWKEGRALE 756
Query: 369 LVDPLI----ETPPS---PRALKRVLLICLRCIDLDVIKRPKMGQIVHMLESD 414
+VDP+I + PS P+ + + + I L C+ RP M +V M S+
Sbjct: 757 IVDPVIVDSLSSQPSIFQPQEVLKCIQIGLLCVQELAEHRPAMSSVVWMFGSE 809
>CLV1_ARATH (Q9SYQ8) Receptor protein kinase CLAVATA1 precursor (EC
2.7.1.-)
Length = 980
Score = 208 bits (529), Expect = 2e-53
Identities = 114/295 (38%), Positives = 187/295 (62%), Gaps = 19/295 (6%)
Query: 141 EEGNVIGEGGYGVVYRGVLQDGCVVAVKNL-HNNKGQAEKEFKVEVEAIGKVRHKNLVRL 199
+E N+IG+GG G+VYRG + + VA+K L G+++ F E++ +G++RH+++VRL
Sbjct: 693 KEENIIGKGGAGIVYRGSMPNNVDVAIKRLVGRGTGRSDHGFTAEIQTLGRIRHRHIVRL 752
Query: 200 VGYCAEGARRMLVYEYVENGNLEQWLHGNVGPTSPLTWDIRMKIAIGTAKGLTYLHEGLE 259
+GY A +L+YEY+ NG+L + LHG+ G L W+ R ++A+ AKGL YLH
Sbjct: 753 LGYVANKDTNLLLYEYMPNGSLGELLHGSKG--GHLQWETRHRVAVEAAKGLCYLHHDCS 810
Query: 260 PKVVHRDIKSSNILLDKNWNAKVSDFGLAK-LLGSEKTHVTTRVMGTFGYVSPEYASTGM 318
P ++HRD+KS+NILLD ++ A V+DFGLAK L+ + + + G++GY++PEYA T
Sbjct: 811 PLILHRDVKSNNILLDSDFEAHVADFGLAKFLVDGAASECMSSIAGSYGYIAPEYAYTLK 870
Query: 319 LNERSDVYSFGVLLMEIITGRSPIDYSRPPGE-MNLVDWFKAMVS--SRRSD-----ELV 370
++E+SDVYSFGV+L+E+I G+ P+ GE +++V W + ++ SD +V
Sbjct: 871 VDEKSDVYSFGVVLLELIAGKKPVGEF---GEGVDIVRWVRNTEEEITQPSDAAIVVAIV 927
Query: 371 DPLIETPPSPRALKRVLLICLRCIDLDVIKRPKMGQIVHMLESDDFPFRSVSSML 425
DP + P + V I + C++ + RP M ++VHML + P +SV++++
Sbjct: 928 DPRLTGYPLTSVI-HVFKIAMMCVEEEAAARPTMREVVHMLTN---PPKSVANLI 978
>RLK5_ARATH (P47735) Receptor-like protein kinase 5 precursor (EC
2.7.1.37)
Length = 999
Score = 200 bits (508), Expect = 7e-51
Identities = 112/285 (39%), Positives = 170/285 (59%), Gaps = 16/285 (5%)
Query: 141 EEGNVIGEGGYGVVYRGVLQDGCVVAVKNLHNNKGQAEKE----------FKVEVEAIGK 190
+E NVIG G G VY+ L+ G VVAVK L+ + + E F EVE +G
Sbjct: 684 DEKNVIGFGSSGKVYKVELRGGEVVAVKKLNKSVKGGDDEYSSDSLNRDVFAAEVETLGT 743
Query: 191 VRHKNLVRLVGYCAEGARRMLVYEYVENGNLEQWLHGNVGPTSPLTWDIRMKIAIGTAKG 250
+RHK++VRL C+ G ++LVYEY+ NG+L LHG+ L W R++IA+ A+G
Sbjct: 744 IRHKSIVRLWCCCSSGDCKLLVYEYMPNGSLADVLHGDRKGGVVLGWPERLRIALDAAEG 803
Query: 251 LTYLHEGLEPKVVHRDIKSSNILLDKNWNAKVSDFGLAK---LLGSEKTHVTTRVMGTFG 307
L+YLH P +VHRD+KSSNILLD ++ AKV+DFG+AK + GS+ + + G+ G
Sbjct: 804 LSYLHHDCVPPIVHRDVKSSNILLDSDYGAKVADFGIAKVGQMSGSKTPEAMSGIAGSCG 863
Query: 308 YVSPEYASTGMLNERSDVYSFGVLLMEIITGRSPIDYSRPPGEMNLVDWFKAMVSSRRSD 367
Y++PEY T +NE+SD+YSFGV+L+E++TG+ P D G+ ++ W + +
Sbjct: 864 YIAPEYVYTLRVNEKSDIYSFGVVLLELVTGKQPTDSEL--GDKDMAKWVCTALDKCGLE 921
Query: 368 ELVDPLIETPPSPRALKRVLLICLRCIDLDVIKRPKMGQIVHMLE 412
++DP ++ + +V+ I L C + RP M ++V ML+
Sbjct: 922 PVIDPKLDL-KFKEEISKVIHIGLLCTSPLPLNRPSMRKVVIMLQ 965
>SIRK_ARATH (O64483) Senescence-induced receptor-like
serine/threonine kinase precursor (FLG22-induced
receptor-like kinase 1)
Length = 876
Score = 199 bits (505), Expect = 2e-50
Identities = 114/288 (39%), Positives = 176/288 (60%), Gaps = 7/288 (2%)
Query: 126 RWYSLKEVEMATRGFEEGNVIGEGGYGVVYRGVLQDGCVVAVKNLHNNKGQAEKEFKVEV 185
R++ EV T FE VIG+GG+G VY GV+ +G VAVK L Q KEF+ EV
Sbjct: 562 RYFKYSEVVNITNNFER--VIGKGGFGKVYHGVI-NGEQVAVKVLSEESAQGYKEFRAEV 618
Query: 186 EAIGKVRHKNLVRLVGYCAEGARRMLVYEYVENGNLEQWLHGNVGPTSPLTWDIRMKIAI 245
+ + +V H NL LVGYC E +L+YEY+ N NL +L G + L+W+ R+KI++
Sbjct: 619 DLLMRVHHTNLTSLVGYCNEINHMVLIYEYMANENLGDYLAGK--RSFILSWEERLKISL 676
Query: 246 GTAKGLTYLHEGLEPKVVHRDIKSSNILLDKNWNAKVSDFGLAKLLGSEKT-HVTTRVMG 304
A+GL YLH G +P +VHRD+K +NILL++ AK++DFGL++ E + ++T V G
Sbjct: 677 DAAQGLEYLHNGCKPPIVHRDVKPTNILLNEKLQAKMADFGLSRSFSVEGSGQISTVVAG 736
Query: 305 TFGYVSPEYASTGMLNERSDVYSFGVLLMEIITGRSPIDYSRPPGEMNLVDWFKAMVSSR 364
+ GY+ PEY ST +NE+SDVYS GV+L+E+ITG+ I S+ ++++ D ++++++
Sbjct: 737 SIGYLDPEYYSTRQMNEKSDVYSLGVVLLEVITGQPAIASSKTE-KVHISDHVRSILANG 795
Query: 365 RSDELVDPLIETPPSPRALKRVLLICLRCIDLDVIKRPKMGQIVHMLE 412
+VD + + ++ I L C + +RP M Q+V L+
Sbjct: 796 DIRGIVDQRLRERYDVGSAWKMSEIALACTEHTSAQRPTMSQVVMELK 843
>RPK1_IPONI (P93194) Receptor-like protein kinase precursor (EC
2.7.1.37)
Length = 1109
Score = 191 bits (486), Expect = 2e-48
Identities = 118/307 (38%), Positives = 173/307 (55%), Gaps = 11/307 (3%)
Query: 130 LKEVEMATRGFEEGNVIGEGGYGVVYRGVLQDGCVVAVKNL-HNNKGQAEKEFKVEVEAI 188
L +V AT + VIG+G +G +Y+ L V AVK L E+E I
Sbjct: 806 LNKVLEATENLNDKYVIGKGAHGTIYKATLSPDKVYAVKKLVFTGIKNGSVSMVREIETI 865
Query: 189 GKVRHKNLVRLVGYCAEGARRMLVYEYVENGNLEQWLHGNVGPTSPLTWDIRMKIAIGTA 248
GKVRH+NL++L + +++Y Y+ENG+L LH P PL W R IA+GTA
Sbjct: 866 GKVRHRNLIKLEEFWLRKEYGLILYTYMENGSLHDILH-ETNPPKPLDWSTRHNIAVGTA 924
Query: 249 KGLTYLHEGLEPKVVHRDIKSSNILLDKNWNAKVSDFGLAKLLGSEKTHV-TTRVMGTFG 307
GL YLH +P +VHRDIK NILLD + +SDFG+AKLL T + + V GT G
Sbjct: 925 HGLAYLHFDCDPAIVHRDIKPMNILLDSDLEPHISDFGIAKLLDQSATSIPSNTVQGTIG 984
Query: 308 YVSPEYASTGMLNERSDVYSFGVLLMEIITGRSPIDYSRPPGEMNLVDWFKAM-VSSRRS 366
Y++PE A T + + SDVYS+GV+L+E+IT + +D S GE ++V W +++ +
Sbjct: 985 YMAPENAFTTVKSRESDVYSYGVVLLELITRKKALDPSF-NGETDIVGWVRSVWTQTGEI 1043
Query: 367 DELVDP-LIETPPSPRALKRV---LLICLRCIDLDVIKRPKMGQIVHMLESDDFPFRSVS 422
++VDP L++ +++V L + LRC + +V KRP M +V L + RS S
Sbjct: 1044 QKIVDPSLLDELIDSSVMEQVTEALSLALRCAEKEVDKRPTMRDVVKQLTR--WSIRSYS 1101
Query: 423 SMLQVQS 429
S ++ +S
Sbjct: 1102 SSVRNKS 1108
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.319 0.138 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 50,764,561
Number of Sequences: 164201
Number of extensions: 2268132
Number of successful extensions: 9858
Number of sequences better than 10.0: 1632
Number of HSP's better than 10.0 without gapping: 1211
Number of HSP's successfully gapped in prelim test: 421
Number of HSP's that attempted gapping in prelim test: 6086
Number of HSP's gapped (non-prelim): 1879
length of query: 433
length of database: 59,974,054
effective HSP length: 113
effective length of query: 320
effective length of database: 41,419,341
effective search space: 13254189120
effective search space used: 13254189120
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 67 (30.4 bits)
Medicago: description of AC146682.5


