

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145221.10 - phase: 0
(68 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
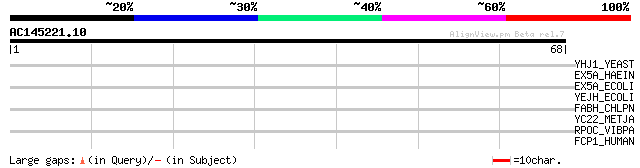
Score E
Sequences producing significant alignments: (bits) Value
YHJ1_YEAST (P38766) Hypothetical helicase in SLT2-PUT2 intergeni... 31 0.52
EX5A_HAEIN (P45158) Exodeoxyribonuclease V alpha chain (EC 3.1.1... 29 2.6
EX5A_ECOLI (P04993) Exodeoxyribonuclease V alpha chain (EC 3.1.1... 28 4.4
YEJH_ECOLI (P33919) Hypothetical protein yejH 27 7.5
FABH_CHLPN (Q9Z8P0) 3-oxoacyl-[acyl-carrier-protein] synthase II... 27 7.5
YC22_METJA (Q58619) Hypothetical protein MJ1222 27 9.8
RPOC_VIBPA (Q87KQ5) DNA-directed RNA polymerase beta' chain (EC ... 27 9.8
FCP1_HUMAN (Q9Y5B0) RNA polymerase II subunit A C-terminal domai... 27 9.8
>YHJ1_YEAST (P38766) Hypothetical helicase in SLT2-PUT2 intergenic
region
Length = 723
Score = 31.2 bits (69), Expect = 0.52
Identities = 18/64 (28%), Positives = 37/64 (57%), Gaps = 1/64 (1%)
Query: 1 MTINKSQGQSLKHDGVYLPTPVFSHGQLHVVVSKVTSREWLKILIIDEDGEDTDETSNIV 60
++I+K+QGQ+++ V L +F GQ++V +S+ + + L++L D T+E
Sbjct: 657 LSIHKAQGQTIQRLKVDLRR-IFEAGQVYVALSRAVTMDTLQVLNFDPGKIRTNERVKDF 715
Query: 61 YEEV 64
Y+ +
Sbjct: 716 YKRL 719
>EX5A_HAEIN (P45158) Exodeoxyribonuclease V alpha chain (EC
3.1.11.5)
Length = 640
Score = 28.9 bits (63), Expect = 2.6
Identities = 13/22 (59%), Positives = 15/22 (68%)
Query: 1 MTINKSQGQSLKHDGVYLPTPV 22
MTI+KSQG KH + LPT V
Sbjct: 568 MTIHKSQGSEFKHTVMVLPTEV 589
>EX5A_ECOLI (P04993) Exodeoxyribonuclease V alpha chain (EC
3.1.11.5) (Exodeoxyribonuclease V 67 kDa polypeptide)
Length = 608
Score = 28.1 bits (61), Expect = 4.4
Identities = 14/41 (34%), Positives = 22/41 (53%), Gaps = 3/41 (7%)
Query: 1 MTINKSQGQSLKHDGVYLP---TPVFSHGQLHVVVSKVTSR 38
MT++KSQG H + LP TPV + ++ V++ R
Sbjct: 536 MTVHKSQGSEFDHAALILPSQRTPVVTRELVYTAVTRARRR 576
>YEJH_ECOLI (P33919) Hypothetical protein yejH
Length = 586
Score = 27.3 bits (59), Expect = 7.5
Identities = 11/17 (64%), Positives = 11/17 (64%)
Query: 39 EWLKILIIDEDGEDTDE 55
EWLKI DEDG D E
Sbjct: 490 EWLKITYYDEDGADVSE 506
>FABH_CHLPN (Q9Z8P0) 3-oxoacyl-[acyl-carrier-protein] synthase III
(EC 2.3.1.41) (Beta-ketoacyl-ACP synthase III) (KAS
III)
Length = 335
Score = 27.3 bits (59), Expect = 7.5
Identities = 14/40 (35%), Positives = 23/40 (57%), Gaps = 2/40 (5%)
Query: 2 TINKSQGQSLKHDGVYLPTPVFSHGQLHVVVSKVTSREWL 41
++NK++ ++ G YLP V S+ L +V TS EW+
Sbjct: 4 SVNKNKKAAIWATGSYLPEKVLSNADLEKMVD--TSDEWI 41
>YC22_METJA (Q58619) Hypothetical protein MJ1222
Length = 243
Score = 26.9 bits (58), Expect = 9.8
Identities = 16/63 (25%), Positives = 30/63 (47%), Gaps = 1/63 (1%)
Query: 1 MTINKSQGQSLKHDGVYLPTPVFSHGQLHVVVSKVTSREWLKILIIDEDGEDTDETSNIV 60
M SQ + D +++ P F+ ++ K +E K +++ +DG D+TS I
Sbjct: 4 MKKQNSQINEINKDEIFVVVPAFNEEKMIGETLKNLKKEGYKNIVVVDDG-SMDKTSEIA 62
Query: 61 YEE 63
+E
Sbjct: 63 KKE 65
>RPOC_VIBPA (Q87KQ5) DNA-directed RNA polymerase beta' chain (EC
2.7.7.6) (RNAP beta' subunit) (Transcriptase beta'
chain) (RNA polymerase beta' subunit)
Length = 1400
Score = 26.9 bits (58), Expect = 9.8
Identities = 14/52 (26%), Positives = 28/52 (52%), Gaps = 2/52 (3%)
Query: 10 SLKHDGVYLPTPVFSHGQLHVVVSKVTSREWLKIL--IIDEDGEDTDETSNI 59
++K +G+YL P + +++ +R ++I ++DEDG T ET +
Sbjct: 519 NVKGEGMYLSGPAEAEKAYRTKQAELHARVKVRITETVVDEDGNSTTETKMV 570
>FCP1_HUMAN (Q9Y5B0) RNA polymerase II subunit A C-terminal domain
phosphatase (EC 3.1.3.16) (TFIIF-associating CTD
phosphatase)
Length = 961
Score = 26.9 bits (58), Expect = 9.8
Identities = 15/38 (39%), Positives = 24/38 (62%), Gaps = 1/38 (2%)
Query: 32 VSKVTSREWLKILIIDEDGEDTDETSNIVY-EEVFQNV 68
+S VT+ E L + +E+ EDTDE +++Y EE+ V
Sbjct: 563 LSGVTAGESLDQSMEEEEEEDTDEDDHLIYLEEILVRV 600
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.314 0.133 0.377
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,111,629
Number of Sequences: 164201
Number of extensions: 267821
Number of successful extensions: 474
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 472
Number of HSP's gapped (non-prelim): 8
length of query: 68
length of database: 59,974,054
effective HSP length: 44
effective length of query: 24
effective length of database: 52,749,210
effective search space: 1265981040
effective search space used: 1265981040
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 58 (26.9 bits)
Medicago: description of AC145221.10


