

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144930.1 - phase: 0 /pseudo
(1322 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
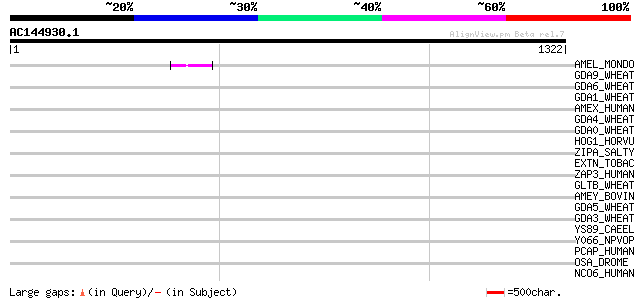
Score E
Sequences producing significant alignments: (bits) Value
AMEL_MONDO (Q28462) Amelogenin 46 6e-04
GDA9_WHEAT (P18573) Alpha/beta-gliadin MM1 precursor (Prolamin) 45 0.001
GDA6_WHEAT (P04726) Alpha/beta-gliadin clone PW1215 precursor (P... 44 0.002
GDA1_WHEAT (P04721) Alpha/beta-gliadin A-I precursor (Prolamin) 43 0.007
AMEX_HUMAN (Q99217) Amelogenin, X isoform precursor 43 0.007
GDA4_WHEAT (P04724) Alpha/beta-gliadin A-IV precursor (Prolamin) 41 0.019
GDA0_WHEAT (P02863) Alpha/beta-gliadin precursor (Prolamin) 41 0.019
HOG1_HORVU (P17990) Gamma-hordein 1 precursor 41 0.025
ZIPA_SALTY (P55894) Cell division protein zipA 40 0.042
EXTN_TOBAC (P13983) Extensin precursor (Cell wall hydroxyproline... 40 0.042
ZAP3_HUMAN (P49750) Nuclear protein ZAP3 (ZAP113) 40 0.055
GLTB_WHEAT (P10386) Glutenin, low molecular weight subunit 1D1 p... 40 0.055
AMEY_BOVIN (Q99004) Amelogenin, class II precursor 40 0.055
GDA5_WHEAT (P04725) Alpha/beta-gliadin A-V precursor (Prolamin) 39 0.072
GDA3_WHEAT (P04723) Alpha/beta-gliadin A-III precursor (Prolamin) 39 0.094
YS89_CAEEL (Q09624) Hypothetical protein ZK945.9 in chromosome II 39 0.12
Y066_NPVOP (Q83949) Hypothetical 98.6 kDa protein (ORF71) 38 0.16
PCAP_HUMAN (Q96RN5) Positive cofactor 2 glutamine/Q-rich-associa... 38 0.16
OSA_DROME (Q8IN94) Trithorax group protein OSA (Eyelid protein) 38 0.16
NCO6_HUMAN (Q14686) Nuclear receptor coactivator 6 (Amplified in... 38 0.16
>AMEL_MONDO (Q28462) Amelogenin
Length = 202
Score = 46.2 bits (108), Expect = 6e-04
Identities = 40/107 (37%), Positives = 50/107 (46%), Gaps = 8/107 (7%)
Query: 383 PINAAQMP--PSYPYVPY-SQHPFFPPFYHQYPLP-PG-QPQVPVNAIAQQMKQQLPVQQ 437
PI AAQ P P P +P QHP P +HQ LP PG QP P AQQ + P+Q
Sbjct: 71 PIMAAQQPAPPQQPVMPVPGQHPMAPTQHHQPNLPQPGQQPYQP--QPAQQPQPHQPIQP 128
Query: 438 -QQNQQARPTFPPIPMLYAELLPTLLHRGHCTTRQGKPPPDPLPPRF 483
Q Q +P P PM + + + + + PP PLPP F
Sbjct: 129 IQPIQPIQPMQPMQPMQPMQPMQPMQPQTPVHAVRPLPPQPPLPPMF 175
>GDA9_WHEAT (P18573) Alpha/beta-gliadin MM1 precursor (Prolamin)
Length = 307
Score = 45.4 bits (106), Expect = 0.001
Identities = 31/90 (34%), Positives = 42/90 (46%), Gaps = 6/90 (6%)
Query: 376 QSMATVAPINAAQMPPSYPYVPYSQ----HPFFPPFYHQYPLPPGQPQV--PVNAIAQQM 429
Q + P Q+P P +PY Q +P PF Q P P QPQ P I+QQ
Sbjct: 73 QPYLQLQPFPQPQLPYPQPQLPYPQPQLPYPQPQPFRPQQPYPQSQPQYSQPQQPISQQQ 132
Query: 430 KQQLPVQQQQNQQARPTFPPIPMLYAELLP 459
+QQ QQQ+ QQ + +L +L+P
Sbjct: 133 QQQQQQQQQKQQQQQQQQILQQILQQQLIP 162
>GDA6_WHEAT (P04726) Alpha/beta-gliadin clone PW1215 precursor
(Prolamin)
Length = 296
Score = 44.3 bits (103), Expect = 0.002
Identities = 25/55 (45%), Positives = 28/55 (50%), Gaps = 2/55 (3%)
Query: 390 PPSYPYVPYSQHPFFPPFYHQYPLPPGQPQV--PVNAIAQQMKQQLPVQQQQNQQ 442
P P+ P +P PPF Q P P QPQ P I+QQ QQ QQQQ QQ
Sbjct: 82 PQPQPFPPQLPYPQPPPFSPQQPYPQPQPQYPQPQQPISQQQAQQQQQQQQQQQQ 136
Score = 36.2 bits (82), Expect = 0.61
Identities = 35/106 (33%), Positives = 41/106 (38%), Gaps = 31/106 (29%)
Query: 383 PINAAQMPPSYPYV---------PYSQHPFFP---PFYHQ--YPLPP------------- 415
P Q PP PY PY Q FP PF Q YP PP
Sbjct: 51 PGQQQQFPPQQPYPQPQPFPSQQPYLQLQPFPQPQPFPPQLPYPQPPPFSPQQPYPQPQP 110
Query: 416 --GQPQVPVNAIAQQMKQQLPVQQQQNQQARPTFPPIPMLYAELLP 459
QPQ P++ Q +QQ QQQQ QQ + I L +L+P
Sbjct: 111 QYPQPQQPISQQQAQQQQQQQQQQQQQQQQQQILQQI--LQQQLIP 154
>GDA1_WHEAT (P04721) Alpha/beta-gliadin A-I precursor (Prolamin)
Length = 262
Score = 42.7 bits (99), Expect = 0.007
Identities = 29/70 (41%), Positives = 36/70 (51%), Gaps = 7/70 (10%)
Query: 394 PYVPYSQHPFFPPFYHQYPLPPGQPQV--PVNAIAQQMKQQLPVQQQQNQQARPTFPPIP 451
P +PYSQ PF Q P P QPQ P I+QQ +QQ QQQQ QQ + I
Sbjct: 84 PQLPYSQPQ---PFRPQQPYPQPQPQYSQPQQPISQQQQQQQQQQQQQQQQQQQIIQQI- 139
Query: 452 MLYAELLPTL 461
L +L+P +
Sbjct: 140 -LQQQLIPCM 148
>AMEX_HUMAN (Q99217) Amelogenin, X isoform precursor
Length = 191
Score = 42.7 bits (99), Expect = 0.007
Identities = 35/119 (29%), Positives = 45/119 (37%), Gaps = 37/119 (31%)
Query: 381 VAPINAAQMPPSYPYVPYSQHPFFPPFYHQYPLPPGQPQVPVNA---------------- 424
+ P+ + Q PP++ P+ P P Q P+ P QP +PV
Sbjct: 65 IIPVLSQQHPPTHTLQPHHHIPVVPA---QQPVIPQQPMMPVPGQHSMTPIQHHQPNLPP 121
Query: 425 IAQQMKQQLPVQQQQNQQARPTFPPIPMLYAELLPTLLHRGHCTTRQGKPPPDPLPPRF 483
AQQ Q PVQ Q +Q +P P PM Q PP PLPP F
Sbjct: 122 PAQQPYQPQPVQPQPHQPMQPQPPVHPM------------------QPLPPQPPLPPMF 162
>GDA4_WHEAT (P04724) Alpha/beta-gliadin A-IV precursor (Prolamin)
Length = 297
Score = 41.2 bits (95), Expect = 0.019
Identities = 25/66 (37%), Positives = 30/66 (44%), Gaps = 1/66 (1%)
Query: 376 QSMATVAPINAAQMPPSYPYVPYSQHPFFPPFYHQYPLPPGQPQVPVNAIAQQMKQQLPV 435
Q + P Q+P P +PY Q F P YP P Q P I+QQ +QQ
Sbjct: 73 QPYMQLQPFPQPQLPYPQPQLPYPQPQPFRP-QQSYPQPQPQYSQPQQPISQQQQQQQQQ 131
Query: 436 QQQQNQ 441
QQQQ Q
Sbjct: 132 QQQQQQ 137
>GDA0_WHEAT (P02863) Alpha/beta-gliadin precursor (Prolamin)
Length = 286
Score = 41.2 bits (95), Expect = 0.019
Identities = 26/55 (47%), Positives = 29/55 (52%), Gaps = 5/55 (9%)
Query: 390 PPSYPYVPYSQHPFFPPFYHQYPLPPGQPQV--PVNAIAQQMKQQLPVQQQQNQQ 442
P P +PYSQ PF Q P P QPQ P I+QQ +QQ QQQQ QQ
Sbjct: 80 PFPQPQLPYSQPQ---PFRPQQPYPQPQPQYSQPQQPISQQQQQQQQQQQQQQQQ 131
>HOG1_HORVU (P17990) Gamma-hordein 1 precursor
Length = 305
Score = 40.8 bits (94), Expect = 0.025
Identities = 43/128 (33%), Positives = 53/128 (40%), Gaps = 30/128 (23%)
Query: 377 SMATVAPINAAQMPPSY--------PYVPYSQHPFFPPFYHQYPLP----PGQPQVPVNA 424
+MAT + Q+ PS PY P SQ PF Q+P P P QPQ P
Sbjct: 11 AMATTFATSEMQVNPSVQVQPTQQQPY-PESQQPFISQSQQQFPQPQQPFPQQPQQPF-- 67
Query: 425 IAQQMKQQLPVQQQQNQQARPT--FPPIPML---------YAELLPTLLHRGHCTTRQGK 473
QQ +QQ Q+Q +PT FP P+L +LLP H+ Q
Sbjct: 68 ---PQSQQQCLQQPQHQFPQPTQQFPQRPLLPFTHPFLTFPDQLLPQPPHQSFPQPPQSY 124
Query: 474 PPPDPLPP 481
P P PL P
Sbjct: 125 PQP-PLQP 131
Score = 33.1 bits (74), Expect = 5.2
Identities = 26/78 (33%), Positives = 36/78 (45%), Gaps = 13/78 (16%)
Query: 388 QMPPSYPYVPYSQHPF--FP------PFYHQYPLPP-GQPQVPVNAIAQQMKQQLPVQQQ 438
Q P P +P++ HPF FP P + +P PP PQ P+ Q +Q+ P +
Sbjct: 87 QQFPQRPLLPFT-HPFLTFPDQLLPQPPHQSFPQPPQSYPQPPLQPFPQPPQQKYP---E 142
Query: 439 QNQQARPTFPPIPMLYAE 456
Q QQ P P LY +
Sbjct: 143 QPQQPFPWQQPTIQLYLQ 160
>ZIPA_SALTY (P55894) Cell division protein zipA
Length = 328
Score = 40.0 bits (92), Expect = 0.042
Identities = 30/88 (34%), Positives = 35/88 (39%), Gaps = 14/88 (15%)
Query: 364 ETEIGMVSAGAGQSMATVAPINAAQMPPSYPYVPYSQHPFFPPFYHQYPLP--PGQPQVP 421
E + V+ GQS AP Q P QH + PP+ P P P QPQ P
Sbjct: 65 EVRVHRVNHAPGQSQEHDAP---RQSP---------QHQYQPPYASAQPRPAAPPQPQAP 112
Query: 422 VNAIAQQMKQQLPVQQQQNQQARPTFPP 449
+ QQ Q P QQ A P PP
Sbjct: 113 MQQPVQQPVQPAPQPQQVQPSAPPVQPP 140
>EXTN_TOBAC (P13983) Extensin precursor (Cell wall
hydroxyproline-rich glycoprotein)
Length = 620
Score = 40.0 bits (92), Expect = 0.042
Identities = 26/91 (28%), Positives = 37/91 (40%), Gaps = 9/91 (9%)
Query: 390 PPSYPYVPYSQHPFFPPFYHQYPLPPGQPQVPVNAIAQQMKQQLPVQQQQNQQARPTFPP 449
PP++ + P + P P H P P QPQ P + P Q+ Q PT+ P
Sbjct: 234 PPTHRHAPPTHQP--SPLRHLPPSPRRQPQPPTYSPP-------PPAYAQSPQPSPTYSP 284
Query: 450 IPMLYAELLPTLLHRGHCTTRQGKPPPDPLP 480
P Y+ P+ ++ PPP P P
Sbjct: 285 PPPTYSPPPPSPIYSPPPPAYSPSPPPTPTP 315
Score = 38.5 bits (88), Expect = 0.12
Identities = 31/106 (29%), Positives = 40/106 (37%), Gaps = 12/106 (11%)
Query: 382 APINAAQMPPSYPYVP--YSQHPFFPPFYHQYPLPPGQPQVPVNAIAQQMKQQLPVQQQQ 439
AP + PP+Y P Y+Q P PP Y P P P + P Q
Sbjct: 405 APPTYSPPPPTYSPPPPTYAQPPPLPPTYSPPPPAYSPPPPPTYSPPPPTYSPPPPAYAQ 464
Query: 440 NQQARPTFPPIPMLYAELLPTLLHRGHCTTRQGKPPP--DPLPPRF 483
PT+ P P Y+ P+ ++ PPP PLPP F
Sbjct: 465 PPPPPPTYSPPPPAYSPPPPSPIY--------SPPPPQVQPLPPTF 502
Score = 37.0 bits (84), Expect = 0.36
Identities = 28/94 (29%), Positives = 37/94 (38%), Gaps = 14/94 (14%)
Query: 382 APINAAQMPPSYPYVPYSQHPFFPPFYHQYPLPPGQPQVPVNAIAQQMKQQLPVQQQQNQ 441
AP + PP + P + P PP H +P PP Q P I Q P
Sbjct: 149 APPSGGHTPPRGQHPPSHRRPS-PPSRHGHPPPPTYAQPPPTPIYSPSPQVQP------- 200
Query: 442 QARPTFPPIPMLYAELLPTLLHRGHCTTRQGKPP 475
PT+ P P + + P+ RGH Q +PP
Sbjct: 201 --PPTYSPPPPTHVQPTPSPPSRGH----QPQPP 228
>ZAP3_HUMAN (P49750) Nuclear protein ZAP3 (ZAP113)
Length = 1822
Score = 39.7 bits (91), Expect = 0.055
Identities = 27/97 (27%), Positives = 42/97 (42%), Gaps = 4/97 (4%)
Query: 389 MPPSYPYVPYSQHP--FFPPFYHQYPLPPGQPQVPVNAIAQQMKQQLP--VQQQQNQQAR 444
MPPS Y+P Q P ++PP Q LPP QP + +Q P + Q+
Sbjct: 158 MPPSQSYMPPPQPPPSYYPPTSSQPYLPPAQPSPSQSPPSQSYLAPTPSYSSSSSSSQSY 217
Query: 445 PTFPPIPMLYAELLPTLLHRGHCTTRQGKPPPDPLPP 481
+ + ++ P+ +GH ++ PPP PP
Sbjct: 218 LSHSQSYLPSSQASPSRPSQGHSKSQLLAPPPPSAPP 254
>GLTB_WHEAT (P10386) Glutenin, low molecular weight subunit 1D1
precursor
Length = 307
Score = 39.7 bits (91), Expect = 0.055
Identities = 24/65 (36%), Positives = 31/65 (46%), Gaps = 8/65 (12%)
Query: 388 QMPPSYPYVPYSQHPFF---PPFYHQYPLPPGQPQVPVNAIAQQMKQQLPVQQQQNQQAR 444
Q+ P P Q PF+ PPF Q P+ P QP +QQ + LP Q +QQ +
Sbjct: 57 QLFPQQPSFSQQQPPFWQQQPPFSQQQPILPQQPP-----FSQQQQLVLPQQPPFSQQQQ 111
Query: 445 PTFPP 449
P PP
Sbjct: 112 PVLPP 116
>AMEY_BOVIN (Q99004) Amelogenin, class II precursor
Length = 192
Score = 39.7 bits (91), Expect = 0.055
Identities = 32/119 (26%), Positives = 45/119 (36%), Gaps = 37/119 (31%)
Query: 381 VAPINAAQMPPSYPYVPYSQHPFFPPFYHQYPLPPGQPQVPVNAI--------------- 425
+ P+ + Q P ++ P+ +P P Q P+ P QP +PV
Sbjct: 66 IIPVVSQQSPQNHALQPHHHNPMVPA---QQPVVPQQPMMPVPGQHSMTPIQHHQPNLPL 122
Query: 426 -AQQMKQQLPVQQQQNQQARPTFPPIPMLYAELLPTLLHRGHCTTRQGKPPPDPLPPRF 483
AQQ Q P+Q Q +Q +P P P+ Q PP PLPP F
Sbjct: 123 PAQQSFQPQPIQPQPHQPLQPQPPVHPI------------------QRLPPQPPLPPIF 163
>GDA5_WHEAT (P04725) Alpha/beta-gliadin A-V precursor (Prolamin)
Length = 319
Score = 39.3 bits (90), Expect = 0.072
Identities = 25/60 (41%), Positives = 29/60 (47%), Gaps = 8/60 (13%)
Query: 388 QMPPSYPYVPYSQHPFFPPFYHQYPLPPGQPQV-----PVNAIAQQMKQQLPVQQQQNQQ 442
Q P P +PY Q FPP Q P P QPQ P++ Q +QQ QQQQ QQ
Sbjct: 83 QPQPFPPQLPYPQPQSFPP---QQPYPQQQPQYLQPQQPISQQQAQQQQQQQQQQQQQQQ 139
Score = 34.3 bits (77), Expect = 2.3
Identities = 28/76 (36%), Positives = 29/76 (37%), Gaps = 16/76 (21%)
Query: 383 PINAAQMPPSYPYV---------PYSQHPFFP---PFYHQYPLPPGQPQVPVNAIAQQMK 430
P Q PP PY PY Q FP PF Q P P Q P QQ
Sbjct: 51 PGQQQQFPPQQPYPQPQPFPSQQPYLQLQPFPQPQPFPPQLPYPQPQSFPPQQPYPQQQP 110
Query: 431 QQL----PVQQQQNQQ 442
Q L P+ QQQ QQ
Sbjct: 111 QYLQPQQPISQQQAQQ 126
>GDA3_WHEAT (P04723) Alpha/beta-gliadin A-III precursor (Prolamin)
Length = 282
Score = 38.9 bits (89), Expect = 0.094
Identities = 26/57 (45%), Positives = 28/57 (48%), Gaps = 5/57 (8%)
Query: 388 QMPPSYPYVPYSQHPFFPPFYHQYPLPPGQPQV--PVNAIAQQMKQQLPVQQQQNQQ 442
Q P P +PY Q FPP Q P P QPQ P I+QQ QQ QQQ QQ
Sbjct: 78 QPQPFPPQLPYPQTQPFPP---QQPYPQPQPQYPQPQQPISQQQAQQQQQQQQTLQQ 131
Score = 34.3 bits (77), Expect = 2.3
Identities = 25/73 (34%), Positives = 36/73 (49%), Gaps = 13/73 (17%)
Query: 412 PLPPGQPQVPVNAIAQQMKQQLPVQQQQNQ--QARPTFPPIPMLYAELLPTLLHRGHCTT 469
P+P QPQ P QQ ++Q+P+ QQQ Q + FPP + P H+ +
Sbjct: 24 PVPQLQPQNPSQ---QQPQEQVPLMQQQQQFPGQQEQFPP-----QQPYP---HQQPFPS 72
Query: 470 RQGKPPPDPLPPR 482
+Q P P P PP+
Sbjct: 73 QQPYPQPQPFPPQ 85
>YS89_CAEEL (Q09624) Hypothetical protein ZK945.9 in chromosome II
Length = 3178
Score = 38.5 bits (88), Expect = 0.12
Identities = 34/130 (26%), Positives = 54/130 (41%), Gaps = 18/130 (13%)
Query: 10 TLREELAKANDVMTALLAAQEQSATAIPIATTIPMT---TSMLPTASTDARFAMPAGFPY 66
TLR A D ++T P TT+ T T+ +PT+++ AM
Sbjct: 254 TLRRMKRDAGDNTCDYTIESTSTSTTTPTTTTVTSTVTSTTTVPTSTSTVTTAMSTSTS- 312
Query: 67 GLPPFFTPSTAAGTSGTANNVLIPATNAASINATLPQTTAAVTEPLVHTMPQSVNINTQH 126
TPST+ T+ A+ + S +T Q+++ +T + P S ++T
Sbjct: 313 ------TPSTSTTIESTSTTFTSTASTSTSSTSTTQQSSSTIT-----SSPSSTTLST-- 359
Query: 127 GSIPVTKTME 136
SIP T T E
Sbjct: 360 -SIPTTTTPE 368
Score = 33.5 bits (75), Expect = 4.0
Identities = 25/97 (25%), Positives = 43/97 (43%), Gaps = 4/97 (4%)
Query: 30 EQSATAIPIATTIPMTTSMLPTASTDARFAMPAGFPYGLPPFFTPSTAAGTSGTANNVLI 89
E ++T+ + TT P TT TAST P+ P +P T+ TS ++++ +
Sbjct: 415 EVTSTSSTVTTTEPTTTLTTSTASTST--TEPSTSTVTTSPSTSPVTSTVTSSSSSSTTV 472
Query: 90 PATNAASINATLPQT--TAAVTEPLVHTMPQSVNINT 124
+ +T P + T + T P T S + +T
Sbjct: 473 TTPTSTESTSTSPSSTVTTSTTAPSTSTTGPSSSSST 509
>Y066_NPVOP (Q83949) Hypothetical 98.6 kDa protein (ORF71)
Length = 875
Score = 38.1 bits (87), Expect = 0.16
Identities = 29/65 (44%), Positives = 31/65 (47%), Gaps = 9/65 (13%)
Query: 388 QMPPSYPYVPYSQH-PFFPPFYHQYPLPPGQPQVPVNAIAQQMKQQLPVQQQQNQQARPT 446
Q PP P+ PY Q+ P PP P P QPQ P QQ QQ P QQ Q Q P
Sbjct: 85 QPPPPQPFYPYGQYWPQQPP--QPPPDQPQQPQPP-----QQPPQQPPQQQPQPPQP-PQ 136
Query: 447 FPPIP 451
PP P
Sbjct: 137 QPPQP 141
>PCAP_HUMAN (Q96RN5) Positive cofactor 2 glutamine/Q-rich-associated
protein (PC2 glutamine/Q-rich-associated protein)
(TPA-inducible gene-1) (TIG-1) (Activator-recruited
cofactor 105 kDa component) (ARC105) (CTG repeat protein
7a)
Length = 788
Score = 38.1 bits (87), Expect = 0.16
Identities = 32/86 (37%), Positives = 40/86 (46%), Gaps = 13/86 (15%)
Query: 408 YHQYPLPPGQPQVPVNAIAQ----QMKQQLPVQQQQNQQARPTFPPI---PMLY-----A 455
+HQ Q Q + IAQ Q +QQ QQQQ QQA PPI PM +
Sbjct: 224 HHQNQQQIQQQQQQLQRIAQLQLQQQQQQQQQQQQQQQQALQAQPPIQQPPMQQPQPPPS 283
Query: 456 ELLPTLLHRGHCTTRQGKPPPDPLPP 481
+ LP L + H T+ +PPP P P
Sbjct: 284 QALPQQLQQMH-HTQHHQPPPQPQQP 308
>OSA_DROME (Q8IN94) Trithorax group protein OSA (Eyelid protein)
Length = 2716
Score = 38.1 bits (87), Expect = 0.16
Identities = 31/102 (30%), Positives = 38/102 (36%), Gaps = 7/102 (6%)
Query: 382 APINAAQMPPSYPYVPYSQHPFFPPFY--HQYPLPPGQPQVPVNAIAQQMKQQLPVQQQQ 439
AP A PP P+ P Q P + H+YP G P P Q +QQ P QQ
Sbjct: 1530 APPRGAAPPPGAPHGPPIQQPAGVAQWDQHRYPPQQGPPPPPQQQQQPQQQQQQPPYQQV 1589
Query: 440 NQQARPTFPPIPMLYAELLPTLLHRGHCTTRQGKPPPDPLPP 481
P P +A++ P G PP PL P
Sbjct: 1590 AGPPGQQPPQAPPQWAQMNP-----GQTAQSGIAPPGSPLRP 1626
Score = 34.7 bits (78), Expect = 1.8
Identities = 29/96 (30%), Positives = 35/96 (36%), Gaps = 8/96 (8%)
Query: 383 PINAAQMPPSYPYVPYSQHPFFPPFYHQYPLPPGQPQVPVNAIAQQMKQQLPVQQQQNQQ 442
P Q PP P SQ + P+ +YP PPG P N Q P + N+
Sbjct: 357 PPGTGQQPPQQNTPPTSQ---YSPYPQRYPTPPGLPAGGSNHRTAYSTHQYP---EPNRP 410
Query: 443 ARPTFPPIPMLYAELLPTLLHRGHCTTRQGKPPPDP 478
P P L P H H Q +PPP P
Sbjct: 411 WPGGSSPSPGSGHPLPPASPH--HVPPLQQQPPPPP 444
Score = 33.9 bits (76), Expect = 3.0
Identities = 27/98 (27%), Positives = 39/98 (39%), Gaps = 17/98 (17%)
Query: 389 MPPSYPYVPYSQH---PFFPPFYHQYPLPPGQPQVPVNAIAQQMKQQLPVQQQQNQQARP 445
+PP +P+ Y ++ P P+ + PLP G+P +QQ QQQ QQ P
Sbjct: 140 LPPPHPHPAYGRYHADPNMDPYRYGQPLPGGKP---------PQQQQPHPQQQPPQQPGP 190
Query: 446 TFPPIPMLYAELLPTLLHRGHCTT-----RQGKPPPDP 478
P +P +G T + PPP P
Sbjct: 191 GGSPNRPPQQRYIPGQPPQGPTPTLNSLLQSSNPPPPP 228
>NCO6_HUMAN (Q14686) Nuclear receptor coactivator 6 (Amplified in
breast cancer-3 protein) (Cancer-amplified
transcriptional coactivator ASC-2) (Activating signal
cointegrator-2) (ASC-2) (Peroxisome
proliferator-activated receptor-interacting protein) (PP
Length = 2063
Score = 38.1 bits (87), Expect = 0.16
Identities = 23/58 (39%), Positives = 27/58 (45%), Gaps = 8/58 (13%)
Query: 390 PPSYPYVPYSQHPF--FPPFYHQYPLPPGQPQVPVNAIAQQMKQQLPVQQQQNQQARP 445
PP YP P Q P PP Q+ PP QP QQ + QLP QQQ ++P
Sbjct: 970 PPGYPSQPVEQRPLQQMPPQLMQHVAPPPQPP------QQQPQPQLPQQQQPPPPSQP 1021
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.332 0.143 0.464
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 153,512,144
Number of Sequences: 164201
Number of extensions: 6631712
Number of successful extensions: 24190
Number of sequences better than 10.0: 153
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 155
Number of HSP's that attempted gapping in prelim test: 23164
Number of HSP's gapped (non-prelim): 663
length of query: 1322
length of database: 59,974,054
effective HSP length: 122
effective length of query: 1200
effective length of database: 39,941,532
effective search space: 47929838400
effective search space used: 47929838400
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 72 (32.3 bits)
Medicago: description of AC144930.1


