

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144929.16 + phase: 0
(143 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
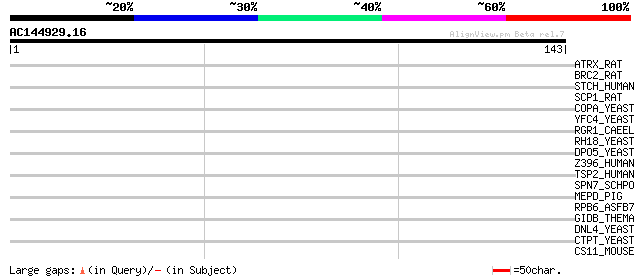
Score E
Sequences producing significant alignments: (bits) Value
ATRX_RAT (P70486) Transcriptional regulator ATRX (X-linked nucle... 32 0.35
BRC2_RAT (O35923) Breast cancer type 2 susceptibility protein ho... 32 0.60
STCH_HUMAN (P48723) Microsomal stress 70 protein ATPase core pre... 31 1.0
SCP1_RAT (Q03410) Synaptonemal complex protein 1 (SCP-1 protein) 30 1.8
COPA_YEAST (P53622) Coatomer alpha subunit (Alpha-coat protein) ... 30 2.3
YFC4_YEAST (P43572) Hypothetical 96.7 kDa protein in STE2-FRS2 i... 29 3.0
RGR1_CAEEL (Q03570) Mediator complex subunit rgr-1 29 3.0
RH18_YEAST (Q12749) DNA repair protein RHC18 (Rad18 homolog) 28 5.1
DPO5_YEAST (P39985) DNA polymerase V (EC 2.7.7.7) (POL V) 28 5.1
Z396_HUMAN (Q96N95) Zinc finger protein 396 (Zinc finger and SCA... 28 6.7
TSP2_HUMAN (P35442) Thrombospondin 2 precursor 28 6.7
SPN7_SCHPO (O60165) Septin homolog spn7 28 6.7
MEPD_PIG (P47788) Thimet oligopeptidase (EC 3.4.24.15) (Endopept... 28 6.7
RPB6_ASFB7 (P42484) DNA-directed RNA polymerase, subunit 6 homol... 28 8.7
GIDB_THEMA (Q9WZG6) Methyltransferase gidB (EC 2.1.-.-) (Glucose... 28 8.7
DNL4_YEAST (Q08387) DNA ligase II (EC 6.5.1.1) (Polydeoxyribonuc... 28 8.7
CTPT_YEAST (P13259) Choline-phosphate cytidylyltransferase (EC 2... 28 8.7
CS11_MOUSE (Q9D269) Cystatin 11 precursor 28 8.7
>ATRX_RAT (P70486) Transcriptional regulator ATRX (X-linked nuclear
protein) (pABP-2) (Fragment)
Length = 527
Score = 32.3 bits (72), Expect = 0.35
Identities = 27/123 (21%), Positives = 51/123 (40%), Gaps = 10/123 (8%)
Query: 14 ISYELEKFVEADEECFYESSGRISFRSNVTLSRKKTDGLETEDLGNKVICPLQEYLLGSS 73
+S +++ E C +SS R+ V+L KK L + G + C +
Sbjct: 161 LSDSVDRLPVKGESC--DSSEDKKTRNRVSLREKKQFSLPAKSSGKRPECSSSDTERSVK 218
Query: 74 FEIREKTEVRIERASVRETQVKQGRRSALHIIKKMSNMVLSSSKSCNTYGNTADHATSTN 133
E + T+ R++R +RE + +RS + V S S S + G++ D
Sbjct: 219 GECCDSTDKRVKRIDLRERRSSNSKRS--------TKEVKSGSSSSDAEGSSEDAKKQKK 270
Query: 134 EKL 136
+++
Sbjct: 271 QRM 273
>BRC2_RAT (O35923) Breast cancer type 2 susceptibility protein homolog
Length = 3343
Score = 31.6 bits (70), Expect = 0.60
Identities = 19/85 (22%), Positives = 43/85 (50%), Gaps = 5/85 (5%)
Query: 38 FRSNVTLSRKKTDGLETEDLGNKVICPLQEYLLGSSFEIREKTEVRIERASVRETQ---- 93
F+SN +L K + D ++ + PL + LG SF E+++ +V++++
Sbjct: 955 FKSNSSLYLKSDGNNDYLDKWSEFLDPLMNHKLGGSFRTASNKEIKLSEDNVKKSKMFFK 1014
Query: 94 -VKQGRRSALHIIKKMSNMVLSSSK 117
+++ ++L I +S + L++ K
Sbjct: 1015 DIEEQYPTSLDCIDTVSTLQLANKK 1039
>STCH_HUMAN (P48723) Microsomal stress 70 protein ATPase core
precursor
Length = 471
Score = 30.8 bits (68), Expect = 1.0
Identities = 21/66 (31%), Positives = 31/66 (46%), Gaps = 1/66 (1%)
Query: 31 ESSGRISFRSNVTLSRKKTDGLETEDLGNKVICPLQEYLLGSSFEIREKTEVRIERASVR 90
+ SG +SRK D L EDL K++ P+Q+ L E E EV + S R
Sbjct: 352 KKSGESQVLFETEISRKLFDTLN-EDLFQKILVPIQQVLKEGHLEKTEIDEVVLVGGSTR 410
Query: 91 ETQVKQ 96
+++Q
Sbjct: 411 IPRIRQ 416
>SCP1_RAT (Q03410) Synaptonemal complex protein 1 (SCP-1 protein)
Length = 997
Score = 30.0 bits (66), Expect = 1.8
Identities = 33/108 (30%), Positives = 46/108 (42%), Gaps = 11/108 (10%)
Query: 8 ETQLKLISYELEKFVEADEECFYESSGRISFRSN--VTLSRKKTDGLETEDL--GNKVIC 63
E QLKLI+ EL+K EE F++N V L KT E + L K +
Sbjct: 403 EDQLKLITMELQKKSSELEE-------MTKFKNNKEVELEELKTILAEDQKLLDEKKQVE 455
Query: 64 PLQEYLLGSSFEIREKTEVRIERASVRETQVKQGRRSALHIIKKMSNM 111
L E L G E+ + R + E QV + S H +K++ M
Sbjct: 456 KLAEELQGKEQELTFLLQTREKEIHDLEVQVTVTKTSEEHYLKQVEEM 503
>COPA_YEAST (P53622) Coatomer alpha subunit (Alpha-coat protein)
(Alpha-COP)
Length = 1201
Score = 29.6 bits (65), Expect = 2.3
Identities = 17/54 (31%), Positives = 27/54 (49%), Gaps = 8/54 (14%)
Query: 27 ECFYESSGRISFRSNVTLSRKKTDGLETEDLGNKVICPLQEYLLGSSFEIREKT 80
ECF E+ RI+ D E E L +K++ +EY+LG S E+ ++
Sbjct: 1011 ECFREAIYRITLLM--------VDDAEDEKLAHKILETAREYILGLSIELERRS 1056
>YFC4_YEAST (P43572) Hypothetical 96.7 kDa protein in STE2-FRS2
intergenic region
Length = 832
Score = 29.3 bits (64), Expect = 3.0
Identities = 28/117 (23%), Positives = 47/117 (39%), Gaps = 20/117 (17%)
Query: 24 ADEECFYESSGRISFRSNVTLSRKKTDGLETEDLGNKVICPLQEYLLGSSFEIREKTEVR 83
+D+ Y+S+GR + N L K+ E D P QE L S E K R
Sbjct: 589 SDDIRIYDSNGRSRNKDNYNLDTKRIKKTELYD-------PFQENLEIHSREYPIKFRKR 641
Query: 84 IERASVRETQVKQGRRSALHIIKKMSNMVLSSSKSCNTYGNTADHATSTNEKLCKVG 140
+ R++++ + +M N SS+KS + + D + E+ + G
Sbjct: 642 VGRSNIK-------------YVDRMPNFTTSSTKSACSLMDFVDFDSIEKEQYSREG 685
>RGR1_CAEEL (Q03570) Mediator complex subunit rgr-1
Length = 1374
Score = 29.3 bits (64), Expect = 3.0
Identities = 21/76 (27%), Positives = 37/76 (48%), Gaps = 4/76 (5%)
Query: 19 EKFVEADEECFYESSGRISFRSNVTLSRKKT--DGLETEDLGNKVICPLQEYLLGSSFEI 76
E+F+++ +E E R++ S L T + +D ++ CPL EYL S+
Sbjct: 926 EQFMDSLDERPEEIDPRMAITSQPILMSHDTIIKACDFKDTEGRITCPLDEYLCSISY-- 983
Query: 77 REKTEVRIERASVRET 92
++ + +ER S R T
Sbjct: 984 LQRALLTLERMSPRAT 999
>RH18_YEAST (Q12749) DNA repair protein RHC18 (Rad18 homolog)
Length = 1114
Score = 28.5 bits (62), Expect = 5.1
Identities = 23/80 (28%), Positives = 39/80 (48%), Gaps = 11/80 (13%)
Query: 31 ESSGRISFRSNVTLSRKKTDGLETEDLGNKVICPLQEYLL----GSSFEIREKTEVRIER 86
E S + +SN+ RKK D L NK I L+E L G ++R++ E ++E+
Sbjct: 396 EKSNQAEAQSNIDQGRKKVDAL------NKTIAHLEEELTKEMGGDKDQMRQELE-QLEK 448
Query: 87 ASVRETQVKQGRRSALHIIK 106
A+ + +V +L +K
Sbjct: 449 ANEKLREVNNSLVVSLQDVK 468
>DPO5_YEAST (P39985) DNA polymerase V (EC 2.7.7.7) (POL V)
Length = 1022
Score = 28.5 bits (62), Expect = 5.1
Identities = 22/98 (22%), Positives = 45/98 (45%), Gaps = 3/98 (3%)
Query: 44 LSRKKTDGLETEDLGNKVICPLQEYLLGSSFEIREKTEVRIERASVRETQVKQGRRSALH 103
LS D + E + ++ + L + L F+ R++ I + R+ +VKQ R + +
Sbjct: 788 LSDDGGDDEDEESMDDEKMMELDDQL-SEIFKRRKEALSSISTGNQRKFEVKQSRENVIS 846
Query: 104 IIKKMSNMVLSSSKSCN--TYGNTADHATSTNEKLCKV 139
++ +M+ K C T N ++H+ + L K+
Sbjct: 847 FKHRVVDMLAVYVKYCEKLTLANKSEHSNNLGGSLSKL 884
>Z396_HUMAN (Q96N95) Zinc finger protein 396 (Zinc finger and SCAN
domain containing protein 14)
Length = 335
Score = 28.1 bits (61), Expect = 6.7
Identities = 18/76 (23%), Positives = 38/76 (49%), Gaps = 7/76 (9%)
Query: 54 TEDLGNKVICPLQEYLLGSSFEIREKTEVRIERASVRETQVKQGRRS----ALHIIKKMS 109
TE+L + + P+++ L G+S+E++ +R ++ T VK R + + +S
Sbjct: 159 TEELPSSQLMPVKKQLQGASWELQ---SLRPHDEDIKTTNVKSASRQKTSLGIELHCNVS 215
Query: 110 NMVLSSSKSCNTYGNT 125
N++ + +TY T
Sbjct: 216 NILHMNGSQSSTYRGT 231
>TSP2_HUMAN (P35442) Thrombospondin 2 precursor
Length = 1172
Score = 28.1 bits (61), Expect = 6.7
Identities = 16/65 (24%), Positives = 32/65 (48%), Gaps = 1/65 (1%)
Query: 78 EKTEVRIERASVRETQVKQGRRSALHIIKKMSNMVLSSSKSCNTYGNTADHATSTNEKLC 137
EK+ + + + S RE+ + G +H++ + S + S K C +A S N +
Sbjct: 186 EKSRMYVAKGSARESHFR-GLLQNVHLVFENSVEDILSKKGCQQGQGAEINAISENTETL 244
Query: 138 KVGPY 142
++GP+
Sbjct: 245 RLGPH 249
>SPN7_SCHPO (O60165) Septin homolog spn7
Length = 428
Score = 28.1 bits (61), Expect = 6.7
Identities = 22/65 (33%), Positives = 39/65 (59%), Gaps = 10/65 (15%)
Query: 78 EKTEVR--IERASVRETQVKQGRRSALHIIKKMSNMVL--SSSKSCNTYGNTADHATSTN 133
E+TE R I+++ +R T V +S +KK++++ + +SS C++YGNT T TN
Sbjct: 319 ERTETRSSIDQSEMR-TNVSDSTKS--EELKKINSIKVDNTSSLKCDSYGNT---KTKTN 372
Query: 134 EKLCK 138
+ C+
Sbjct: 373 QLNCE 377
>MEPD_PIG (P47788) Thimet oligopeptidase (EC 3.4.24.15)
(Endopeptidase 24.15)
Length = 686
Score = 28.1 bits (61), Expect = 6.7
Identities = 14/40 (35%), Positives = 21/40 (52%)
Query: 78 EKTEVRIERASVRETQVKQGRRSALHIIKKMSNMVLSSSK 117
+K +R E A E +K GRR+ LH+ K+ + S K
Sbjct: 129 QKDSLRPEAARYLERLIKLGRRNGLHLPKETQEKIKSIKK 168
>RPB6_ASFB7 (P42484) DNA-directed RNA polymerase, subunit 6 homolog
(EC 2.7.7.6)
Length = 147
Score = 27.7 bits (60), Expect = 8.7
Identities = 22/94 (23%), Positives = 41/94 (43%), Gaps = 4/94 (4%)
Query: 18 LEKFVEADEECFYESSGRISFRSNVTLSRKKTDGLETEDLGNKVICPLQEYLLG---SSF 74
+E++VE +EE F +S + + S +G VI P E + ++F
Sbjct: 15 VEEYVETEEENFVDSEEESEDKDEIVESPSICEGFVQASSQTLVIIPDNERITSNVLTTF 74
Query: 75 EIREKTEVRIERASVR-ETQVKQGRRSALHIIKK 107
E VR ++ ++ T +K+ S + I K+
Sbjct: 75 EATRLVAVRAQQLAINGSTMLKKKYSSPIDIAKQ 108
>GIDB_THEMA (Q9WZG6) Methyltransferase gidB (EC 2.1.-.-) (Glucose
inhibited division protein B)
Length = 238
Score = 27.7 bits (60), Expect = 8.7
Identities = 13/31 (41%), Positives = 19/31 (60%)
Query: 77 REKTEVRIERASVRETQVKQGRRSALHIIKK 107
R E+++E VRE +K G R AL I++K
Sbjct: 187 RAMKELKVELEKVREYSLKTGERRALLILRK 217
>DNL4_YEAST (Q08387) DNA ligase II (EC 6.5.1.1)
(Polydeoxyribonucleotide synthase [ATP]) (DNA ligase IV
homolog)
Length = 944
Score = 27.7 bits (60), Expect = 8.7
Identities = 25/90 (27%), Positives = 39/90 (42%), Gaps = 7/90 (7%)
Query: 39 RSNVTLSRKKTDGLETEDL--GNKVICPLQEYLLGSSFEI-----REKTEVRIERASVRE 91
RS + L RKK + D N+ P+ G F + E T +RI RA + +
Sbjct: 654 RSQLGLIRKKRKRVLISDSFHQNRKQLPISNIFAGLLFYVLSDYVTEDTGIRITRAELEK 713
Query: 92 TQVKQGRRSALHIIKKMSNMVLSSSKSCNT 121
T V+ G + ++I K ++ SC T
Sbjct: 714 TIVEHGGKLIYNVILKRHSIGDVRLISCKT 743
>CTPT_YEAST (P13259) Choline-phosphate cytidylyltransferase (EC
2.7.7.15) (Phosphorylcholine transferase)
(CTP:phosphocholine cytidylyltransferase) (CT) (CCT)
Length = 424
Score = 27.7 bits (60), Expect = 8.7
Identities = 20/110 (18%), Positives = 49/110 (44%), Gaps = 4/110 (3%)
Query: 8 ETQLKLISYELEKFVEADEECFYESSGRISFRSNVTLSRKKTDGLETEDLGNKVICPLQE 67
E ++ ++++ +V AD + Y+ + L+ ++T+G+ T D+ K+I +
Sbjct: 187 EHKIDYVAHDDIPYVSADSDDIYKPIKEMG----KFLTTQRTNGVSTSDIITKIIRDYDK 242
Query: 68 YLLGSSFEIREKTEVRIERASVRETQVKQGRRSALHIIKKMSNMVLSSSK 117
YL+ + + E+ + E + K+ KK + ++S+
Sbjct: 243 YLMRNFARGATRQELNVSWLKKNELEFKKHINEFRSYFKKNQTNLNNASR 292
>CS11_MOUSE (Q9D269) Cystatin 11 precursor
Length = 139
Score = 27.7 bits (60), Expect = 8.7
Identities = 11/28 (39%), Positives = 19/28 (67%)
Query: 69 LLGSSFEIREKTEVRIERASVRETQVKQ 96
L+ S++++ KT +RIE S E+ VK+
Sbjct: 19 LVAFSYQVKRKTFIRIEEVSALESSVKE 46
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.312 0.128 0.345
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,569,625
Number of Sequences: 164201
Number of extensions: 497142
Number of successful extensions: 1010
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 13
Number of HSP's that attempted gapping in prelim test: 1005
Number of HSP's gapped (non-prelim): 18
length of query: 143
length of database: 59,974,054
effective HSP length: 99
effective length of query: 44
effective length of database: 43,718,155
effective search space: 1923598820
effective search space used: 1923598820
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC144929.16


