

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144928.9 - phase: 2 /pseudo
(1036 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
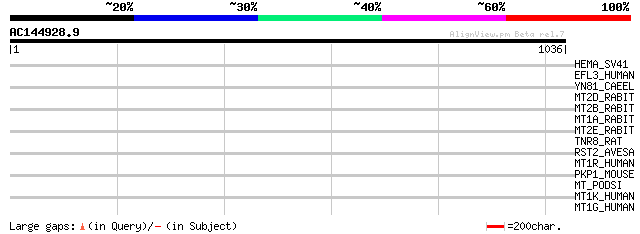
HEMA_SV41 (P25180) Hemagglutinin-neuraminidase (EC 3.2.1.18) 40 0.032
EFL3_HUMAN (O75095) Multiple EGF-like-domain protein 3 (Multiple... 34 1.8
YN81_CAEEL (Q03610) Hypothetical protein ZC84.1 in chromosome III 33 3.0
MT2D_RABIT (P80291) Metallothionein-IID (MT-2D) 33 4.0
MT2B_RABIT (P80289) Metallothionein-IIB (MT-2B) 33 4.0
MT1A_RABIT (P11957) Metallothionein-IA (MT-1A) 33 4.0
MT2E_RABIT (P80292) Metallothionein-IIE (MT-2E) 33 5.2
TNR8_RAT (P97525) Tumor necrosis factor receptor superfamily mem... 32 6.8
RST2_AVESA (P50696) Thaumatin-like pathogenesis-related protein ... 32 6.8
MT1R_HUMAN (Q93083) Metallothionein-IR (MT-1R) 32 6.8
PKP1_MOUSE (P97350) Plakophilin 1 32 8.8
MT_PODSI (Q708T3) Metallothionein (MT) 32 8.8
MT1K_HUMAN (P80296) Metallothionein-IK (MT-1K) 32 8.8
MT1G_HUMAN (P13640) Metallothionein-IG (MT-1G) 32 8.8
>HEMA_SV41 (P25180) Hemagglutinin-neuraminidase (EC 3.2.1.18)
Length = 568
Score = 40.0 bits (92), Expect = 0.032
Identities = 26/77 (33%), Positives = 36/77 (45%), Gaps = 10/77 (12%)
Query: 117 PCPLGSYCPIATLNKTTGVC-------EPYLYQLPPMQPNHTCGGANVWADFSSSSETFC 169
PC G++CP N TGV P+ + PN+ GGA +WAD + + TF
Sbjct: 448 PCSAGNFCPA---NCITGVYADVWPMNNPFPAGSSGVNPNYLFGGAFLWADVARVNPTFY 504
Query: 170 SAGSYCPTTTTKFPCSS 186
A + TT FP S+
Sbjct: 505 MASATQYKNTTGFPNSN 521
>EFL3_HUMAN (O75095) Multiple EGF-like-domain protein 3 (Multiple
epidermal growth factor-like domains 6) (Fragment)
Length = 1246
Score = 34.3 bits (77), Expect = 1.8
Identities = 39/138 (28%), Positives = 52/138 (37%), Gaps = 18/138 (13%)
Query: 59 RNCNLTS--WV--SGCEPGWACSVPSGQKIDLKD---SKEMPARTSNCRACCE-GFFCPH 110
R C+LT W GC C P Q D +D S + R C+A CE G+F P
Sbjct: 547 RFCHLTCPPWAFGPGCSEECQCVQPHTQSCDKRDGSCSCKAGFRGERCQAECELGYFGP- 605
Query: 111 GITCMIPCPLGSYCPIATLNKTTGVCEPYLYQLPPMQPNHTCGGANVWADFSSSSETFCS 170
G CP+G C + +G C + P CG F + + CS
Sbjct: 606 GCWQACTCPVGVAC-----DSVSGECGK---RCPAGFQGEDCGQECPVGTFGVNCSSSCS 657
Query: 171 -AGSYCPTTTTKFPCSSG 187
G+ C T + C G
Sbjct: 658 CGGAPCHGVTGQCRCPPG 675
>YN81_CAEEL (Q03610) Hypothetical protein ZC84.1 in chromosome III
Length = 1416
Score = 33.5 bits (75), Expect = 3.0
Identities = 37/171 (21%), Positives = 59/171 (33%), Gaps = 20/171 (11%)
Query: 33 CTAAEVKFYLNSLMEKSSSANYLKPNRNCNLTSWVSGCEPGWACSVPSGQKIDLKDSKEM 92
CT +++ N+ + Y N N + C+ ++P + I + K+M
Sbjct: 443 CTENVKRYWYNARTRQCQMFEYTGCQGNDNNFDSIMDCQNFCKNAIPEPKCIQGQAYKDM 502
Query: 93 PARTSNCRACCEGFFCPHGITCMIPCPLGSYCP----IATLNKTTGV-----------CE 137
N C G CP C CP +LN +G+
Sbjct: 503 ---FGNFVTCSNGMGCPANYECYFDGSQWGCCPTKAFTCSLNTDSGIQCGAGSTFKYYYN 559
Query: 138 PYLYQLPPMQPNHTCGGANVWADFSSSSETFCSAGSYCPTTTTKFPCSSGL 188
P Q N G +N +A+ + E++CS G CP T SG+
Sbjct: 560 PQTQNCESFQYNGCDGNSNNFAN-RDACESYCSVGG-CPNGGTPLRDHSGM 608
>MT2D_RABIT (P80291) Metallothionein-IID (MT-2D)
Length = 61
Score = 33.1 bits (74), Expect = 4.0
Identities = 19/41 (46%), Positives = 21/41 (50%), Gaps = 1/41 (2%)
Query: 243 SS*KCKENCKCTPKMESCKRCC*KRCNWAASSIITQVLS*K 283
SS KCKE CKCT +SC CC C A I + S K
Sbjct: 17 SSCKCKE-CKCTSCKKSCCSCCPAGCTKCAQGCICKGASDK 56
>MT2B_RABIT (P80289) Metallothionein-IIB (MT-2B)
Length = 61
Score = 33.1 bits (74), Expect = 4.0
Identities = 19/41 (46%), Positives = 21/41 (50%), Gaps = 1/41 (2%)
Query: 243 SS*KCKENCKCTPKMESCKRCC*KRCNWAASSIITQVLS*K 283
SS KCKE CKCT +SC CC C A I + S K
Sbjct: 17 SSCKCKE-CKCTSCKKSCCSCCPAGCTKCAQGCICKGASDK 56
>MT1A_RABIT (P11957) Metallothionein-IA (MT-1A)
Length = 61
Score = 33.1 bits (74), Expect = 4.0
Identities = 19/41 (46%), Positives = 21/41 (50%), Gaps = 1/41 (2%)
Query: 243 SS*KCKENCKCTPKMESCKRCC*KRCNWAASSIITQVLS*K 283
SS KCKE CKCT +SC CC C A I + S K
Sbjct: 17 SSCKCKE-CKCTSCKKSCCSCCPAGCTKCAQGCICKGASDK 56
>MT2E_RABIT (P80292) Metallothionein-IIE (MT-2E)
Length = 61
Score = 32.7 bits (73), Expect = 5.2
Identities = 17/34 (50%), Positives = 18/34 (52%), Gaps = 1/34 (2%)
Query: 243 SS*KCKENCKCTPKMESCKRCC*KRCNWAASSII 276
SS KCKE CKCT +SC CC C A I
Sbjct: 17 SSCKCKE-CKCTSCKKSCCSCCPAGCTKCAQGCI 49
>TNR8_RAT (P97525) Tumor necrosis factor receptor superfamily member
8 precursor (CD30L receptor) (Lymphocyte activation
antigen CD30)
Length = 493
Score = 32.3 bits (72), Expect = 6.8
Identities = 23/83 (27%), Positives = 32/83 (37%), Gaps = 14/83 (16%)
Query: 100 RACCEGFFCPHGITCMIPCPLG-----SYCPIATLNKTTGVCEPYLYQLPPMQPNHTCGG 154
R CC + CP G++ PCP G + + T + G C + LP + C G
Sbjct: 42 RNCC--YQCPSGLSPTQPCPQGPATARNSVILTTTSNEDGKCTACVTCLPGLVEKAPCSG 99
Query: 155 ANVWADFSSSSETFCSAGSYCPT 177
+S C G YC T
Sbjct: 100 -------NSPRICECQPGMYCST 115
>RST2_AVESA (P50696) Thaumatin-like pathogenesis-related protein 2
precursor
Length = 169
Score = 32.3 bits (72), Expect = 6.8
Identities = 26/108 (24%), Positives = 42/108 (38%), Gaps = 20/108 (18%)
Query: 32 LCTAAEVKFYLNSLMEKSSSANYLKPNRNCNLTSWVSGCEPGWACSVPSGQKIDLKDSKE 91
+ T++ V F+L ++ +SA + NC T W +G G + S Q ++
Sbjct: 1 MATSSAVLFFLLAVFAAGASAATFRITNNCGFTVWPAGIPVGGGFQLNSKQSSNI----N 56
Query: 92 MPARTS-------------NCRACCEGFFCPHGITCMI---PCPLGSY 123
+PA TS N R C C ++C + P L Y
Sbjct: 57 VPAGTSAGRIWGRTGCSFNNGRGSCATGDCAGALSCTLSGQPATLAEY 104
>MT1R_HUMAN (Q93083) Metallothionein-IR (MT-1R)
Length = 61
Score = 32.3 bits (72), Expect = 6.8
Identities = 19/47 (40%), Positives = 22/47 (46%), Gaps = 1/47 (2%)
Query: 237 GKIKRISS*KCKENCKCTPKMESCKRCC*KRCNWAASSIITQVLS*K 283
G SS KCKE CKCT +SC CC C A + + S K
Sbjct: 11 GSCSCASSCKCKE-CKCTSCKKSCCSCCPMGCAKCAQGCVCKGASEK 56
>PKP1_MOUSE (P97350) Plakophilin 1
Length = 728
Score = 32.0 bits (71), Expect = 8.8
Identities = 17/53 (32%), Positives = 26/53 (48%), Gaps = 1/53 (1%)
Query: 421 DSYTESSKQTYIEKCNRENQTWPYHRYHGPVRGWQDNISFSS-SWKSTWMLGY 472
++Y +S+ +Y K N +W Y Y+G ++ DN FSS S W Y
Sbjct: 79 NNYGTTSRSSYFSKFQAGNGSWGYPIYNGTLKREPDNRRFSSYSQMENWSRHY 131
>MT_PODSI (Q708T3) Metallothionein (MT)
Length = 63
Score = 32.0 bits (71), Expect = 8.8
Identities = 15/31 (48%), Positives = 17/31 (54%), Gaps = 1/31 (3%)
Query: 246 KCKENCKCTPKMESCKRCC*KRCNWAASSII 276
KCK NCKCT +SC CC C A S +
Sbjct: 21 KCK-NCKCTSCKKSCCSCCPAGCAKCAKSCV 50
>MT1K_HUMAN (P80296) Metallothionein-IK (MT-1K)
Length = 62
Score = 32.0 bits (71), Expect = 8.8
Identities = 19/41 (46%), Positives = 21/41 (50%), Gaps = 1/41 (2%)
Query: 243 SS*KCKENCKCTPKMESCKRCC*KRCNWAASSIITQVLS*K 283
SS KCKE CKCT +SC CC C A I + S K
Sbjct: 18 SSCKCKE-CKCTSCKKSCCSCCPVGCAKCAQGCICKGASEK 57
>MT1G_HUMAN (P13640) Metallothionein-IG (MT-1G)
Length = 61
Score = 32.0 bits (71), Expect = 8.8
Identities = 19/41 (46%), Positives = 21/41 (50%), Gaps = 1/41 (2%)
Query: 243 SS*KCKENCKCTPKMESCKRCC*KRCNWAASSIITQVLS*K 283
SS KCKE CKCT +SC CC C A I + S K
Sbjct: 17 SSCKCKE-CKCTSCKKSCCSCCPVGCAKCAQGCICKGASEK 56
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.354 0.153 0.599
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 107,828,539
Number of Sequences: 164201
Number of extensions: 4045761
Number of successful extensions: 15663
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 13
Number of HSP's that attempted gapping in prelim test: 15648
Number of HSP's gapped (non-prelim): 30
length of query: 1036
length of database: 59,974,054
effective HSP length: 120
effective length of query: 916
effective length of database: 40,269,934
effective search space: 36887259544
effective search space used: 36887259544
T: 11
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.6 bits)
S2: 71 (32.0 bits)
Medicago: description of AC144928.9


