

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144928.8 + phase: 0 /pseudo
(359 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
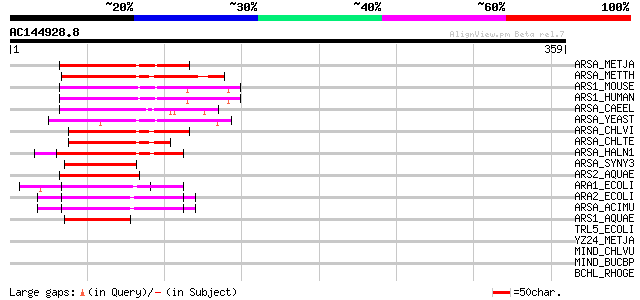
Score E
Sequences producing significant alignments: (bits) Value
ARSA_METJA (Q58542) Putative arsenical pump-driving ATPase (EC 3... 74 4e-13
ARSA_METTH (O27555) Putative arsenical pump-driving ATPase (EC 3... 74 6e-13
ARS1_MOUSE (O54984) Arsenical pump-driving ATPase (EC 3.6.3.16) ... 69 2e-11
ARS1_HUMAN (O43681) Arsenical pump-driving ATPase (EC 3.6.3.16) ... 69 2e-11
ARSA_CAEEL (P30632) Putative arsenical pump-driving ATPase (EC 3... 68 3e-11
ARSA_YEAST (Q12154) Putative arsenical pump-driving ATPase (EC 3... 67 9e-11
ARSA_CHLVI (Q46465) Putative arsenical pump-driving ATPase (EC 3... 64 6e-10
ARSA_CHLTE (Q46366) Putative arsenical pump-driving ATPase (EC 3... 63 1e-09
ARSA_HALN1 (O52027) Putative arsenical pump-driving ATPase (EC 3... 63 1e-09
ARSA_SYNY3 (Q55794) Putative arsenical pump-driving ATPase (EC 3... 61 5e-09
ARS2_AQUAE (O66674) Putative arsenical pump-driving ATPase 2 (EC... 60 7e-09
ARA1_ECOLI (P08690) Arsenical pump-driving ATPase (EC 3.6.3.16) ... 59 1e-08
ARA2_ECOLI (P52145) Arsenical pump-driving ATPase (EC 3.6.3.16) ... 56 2e-07
ARSA_ACIMU (O50593) Arsenical pump-driving ATPase (EC 3.6.3.16) ... 54 5e-07
ARS1_AQUAE (O66908) Putative arsenical pump-driving ATPase 1 (EC... 50 7e-06
TRL5_ECOLI (Q00187) Protein traL 40 0.009
YZ24_METJA (Q60283) Hypothetical protein MJECL24 39 0.016
MIND_CHLVU (P56346) Putative septum site-determining protein minD 38 0.035
MIND_BUCBP (Q89AI3) Septum site-determining protein minD (Cell d... 38 0.045
BCHL_RHOGE (Q9JPA5) Light-independent protochlorophyllide reduct... 37 0.059
>ARSA_METJA (Q58542) Putative arsenical pump-driving ATPase (EC
3.6.3.16) (Arsenite-translocating ATPase) (Arsenical
resistance ATPase) (Arsenite-transporting ATPase)
Length = 349
Score = 74.3 bits (181), Expect = 4e-13
Identities = 40/84 (47%), Positives = 55/84 (64%), Gaps = 2/84 (2%)
Query: 33 KYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSDSFAQDLTGGALVQVDG 92
KY M GGKGGVGKT+ +A+ V A G ++VSTDPAHSL D F Q+ G +V G
Sbjct: 27 KYIMFGGKGGVGKTTMSAATGVYLAEKGLKVVIVSTDPAHSLRDIFEQEF-GHEPTKVKG 85
Query: 93 PDYPLFALEINPEKAREDFRDVAK 116
D L+ +EI+P+KA E++++ K
Sbjct: 86 YD-NLYVVEIDPQKAMEEYKEKLK 108
>ARSA_METTH (O27555) Putative arsenical pump-driving ATPase (EC
3.6.3.16) (Arsenite-translocating ATPase) (Arsenical
resistance ATPase) (Arsenite-transporting ATPase)
Length = 324
Score = 73.9 bits (180), Expect = 6e-13
Identities = 43/106 (40%), Positives = 65/106 (60%), Gaps = 10/106 (9%)
Query: 34 YYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSDSFAQDLTGGALVQVDGP 93
+ +GGKGGVGKT+ +A+ A+ A +G TLV+STDPAHSLSDS +++ G ++
Sbjct: 16 FVFIGGKGGVGKTTISAATALWMARSGKKTLVISTDPAHSLSDSLEREI-GHTPTKI--- 71
Query: 94 DYPLFALEINPEKAREDFRDVAKQNGGSTGVKDFMDGMGLGMIVDQ 139
L+A+EI+PE A E+++ ++ GMGL M+ DQ
Sbjct: 72 TENLYAVEIDPEVAMEEYQAKLQEQAAMN------PGMGLDMLQDQ 111
>ARS1_MOUSE (O54984) Arsenical pump-driving ATPase (EC 3.6.3.16)
(Arsenite-translocating ATPase) (Arsenical resistance
ATPase) (Arsenite-transporting ATPase) (ARSA)
Length = 348
Score = 68.9 bits (167), Expect = 2e-11
Identities = 46/122 (37%), Positives = 67/122 (54%), Gaps = 7/122 (5%)
Query: 33 KYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSDSFAQDLTGGALVQVDG 92
K+ +GGKGGVGKT+C+ SLAV+ + L++STDPAH++SD+F Q + +V G
Sbjct: 38 KWIFVGGKGGVGKTTCSCSLAVQLSKGRESVLIISTDPAHNISDAFDQKFS-KVPTKVKG 96
Query: 93 PDYPLFALEINPEKAREDFRD--VAKQNGGSTGVKDFMDGMGLGMIVDQT---AKVRRAK 147
D LFA+EI+P + D + N S G K + M +D+ A+V R
Sbjct: 97 YD-NLFAMEIDPSLGVAELPDEFFEEDNMLSMGKKMMQEAMSAFPGIDEAMSYAEVMRLV 155
Query: 148 TG 149
G
Sbjct: 156 KG 157
>ARS1_HUMAN (O43681) Arsenical pump-driving ATPase (EC 3.6.3.16)
(Arsenite-translocating ATPase) (Arsenical resistance
ATPase) (Arsenite-transporting ATPase) (ARSA) (ASNA-I)
Length = 348
Score = 68.9 bits (167), Expect = 2e-11
Identities = 46/122 (37%), Positives = 67/122 (54%), Gaps = 7/122 (5%)
Query: 33 KYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSDSFAQDLTGGALVQVDG 92
K+ +GGKGGVGKT+C+ SLAV+ + L++STDPAH++SD+F Q + +V G
Sbjct: 38 KWIFVGGKGGVGKTTCSCSLAVQLSKGRESVLIISTDPAHNISDAFDQKFS-KVPTKVKG 96
Query: 93 PDYPLFALEINPEKAREDFRD--VAKQNGGSTGVKDFMDGMGLGMIVDQT---AKVRRAK 147
D LFA+EI+P + D + N S G K + M +D+ A+V R
Sbjct: 97 YD-NLFAMEIDPSLGVAELPDEFFEEDNMLSMGKKMMQEAMSAFPGIDEAMSYAEVMRLV 155
Query: 148 TG 149
G
Sbjct: 156 KG 157
>ARSA_CAEEL (P30632) Putative arsenical pump-driving ATPase (EC
3.6.3.16) (Arsenite-translocating ATPase) (Arsenical
resistance ATPase) (Arsenite-transporting ATPase)
Length = 342
Score = 68.2 bits (165), Expect = 3e-11
Identities = 45/115 (39%), Positives = 65/115 (56%), Gaps = 14/115 (12%)
Query: 33 KYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSDSFAQDLTGGALVQVDG 92
K+ +GGKGGVGKT+C+ SLA + + L++STDPAH++SD+F+Q T + V+G
Sbjct: 19 KWIFVGGKGGVGKTTCSCSLAAQLSKVRERVLLISTDPAHNISDAFSQKFTKTPTL-VEG 77
Query: 93 PDYPLFALEI--NPE-------KAREDFRDVAKQNGGSTGV---KDFMDGMGLGM 135
LFA+EI NP E ++ A+ GGS G KDF+ G+
Sbjct: 78 -FKNLFAMEIDSNPNGEGVEMGNIEEMLQNAAQNEGGSGGFSMGKDFLQSFAGGL 131
>ARSA_YEAST (Q12154) Putative arsenical pump-driving ATPase (EC
3.6.3.16) (Arsenite-translocating ATPase) (Arsenical
resistance ATPase) (Arsenite-transporting ATPase)
Length = 354
Score = 66.6 bits (161), Expect = 9e-11
Identities = 42/124 (33%), Positives = 66/124 (52%), Gaps = 8/124 (6%)
Query: 26 MVAGTERKYYMLGGKGGVGKTSCAASLAVKFA--NNGHPTLVVSTDPAHSLSDSFAQDLT 83
++ T K+ +GGKGGVGKT+ + S+A++ A L++STDPAH+LSD+F +
Sbjct: 12 LITSTTHKWIFVGGKGGVGKTTSSCSIAIQMALSQPNKQFLLISTDPAHNLSDAFGEKF- 70
Query: 84 GGALVQVDGPDYPLFALEINPEKAREDFRDVAKQNGGSTGVKDFMDGMGL----GMIVDQ 139
G +V G + L +EI+P A +D D+A + G D +G G + D
Sbjct: 71 GKDARKVTGMN-NLSCMEIDPSAALKDMNDMAVSRANNNGSDGQGDDLGSLLQGGALADL 129
Query: 140 TAKV 143
T +
Sbjct: 130 TGSI 133
>ARSA_CHLVI (Q46465) Putative arsenical pump-driving ATPase (EC
3.6.3.16) (Arsenite-translocating ATPase) (Arsenical
resistance ATPase) (Arsenite-transporting ATPase)
Length = 405
Score = 63.9 bits (154), Expect = 6e-10
Identities = 39/79 (49%), Positives = 51/79 (64%), Gaps = 5/79 (6%)
Query: 39 GKGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSDSFAQDLTGGALVQVDGPDYPLF 98
GKGGVGKTS +A+ AV+ + GH TLV+STDPAHSLSDSF L G ++ L
Sbjct: 8 GKGGVGKTSVSAATAVRLSEMGHRTLVLSTDPAHSLSDSFNLQL-GAEPTKI---KENLH 63
Query: 99 ALEINP-EKAREDFRDVAK 116
A+E+NP +E++ V K
Sbjct: 64 AIEVNPYVDLKENWHSVQK 82
>ARSA_CHLTE (Q46366) Putative arsenical pump-driving ATPase (EC
3.6.3.16) (Arsenite-translocating ATPase) (Arsenical
resistance ATPase) (Arsenite-transporting ATPase)
Length = 405
Score = 63.2 bits (152), Expect = 1e-09
Identities = 36/66 (54%), Positives = 45/66 (67%), Gaps = 4/66 (6%)
Query: 39 GKGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSDSFAQDLTGGALVQVDGPDYPLF 98
GKGGVGKTS +A+ AV+ + GH TLV+STDPAHSLSDSF L G ++ L
Sbjct: 8 GKGGVGKTSVSAATAVRLSEMGHRTLVLSTDPAHSLSDSFNIQL-GAEPTKI---KENLH 63
Query: 99 ALEINP 104
A+E+NP
Sbjct: 64 AIEVNP 69
>ARSA_HALN1 (O52027) Putative arsenical pump-driving ATPase (EC
3.6.3.16) (Arsenite-translocating ATPase) (Arsenical
resistance ATPase) (Arsenite-transporting ATPase)
Length = 644
Score = 62.8 bits (151), Expect = 1e-09
Identities = 35/83 (42%), Positives = 55/83 (66%), Gaps = 4/83 (4%)
Query: 31 ERKYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSDSFAQDLTGGALVQV 90
+ ++ GKGGVGK++ + + A A+N + TL+V+TDPA +LSD F QD+ G + +
Sbjct: 18 DTEFVFFSGKGGVGKSTVSCATATWLADNDYDTLLVTTDPAPNLSDIFNQDI-GHEVTAI 76
Query: 91 DGPDYP-LFALEINPEKAREDFR 112
D D P L A+EI+P+ A E++R
Sbjct: 77 D--DVPNLSAIEIDPDVAAEEYR 97
Score = 61.6 bits (148), Expect = 3e-09
Identities = 38/96 (39%), Positives = 58/96 (59%), Gaps = 3/96 (3%)
Query: 17 TESISVFDDMVAGTERKYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSD 76
T++ +V +++V E +Y GKGGVGK++ A++ AV A G+ TLVV+TDPA L+D
Sbjct: 328 TDADAVAEELVPVEETRYLFFTGKGGVGKSTIASTTAVSLAEAGYETLVVTTDPAAHLAD 387
Query: 77 SFAQDLTGGALVQVDGPDYPLFALEINPEKAREDFR 112
F Q + G V + L A I+ E+A E++R
Sbjct: 388 IFEQPV-GHEPTSVGQAN--LDAARIDQERALEEYR 420
>ARSA_SYNY3 (Q55794) Putative arsenical pump-driving ATPase (EC
3.6.3.16) (Arsenite-translocating ATPase) (Arsenical
resistance ATPase) (Arsenite-transporting ATPase)
Length = 396
Score = 60.8 bits (146), Expect = 5e-09
Identities = 29/47 (61%), Positives = 37/47 (78%)
Query: 36 MLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSDSFAQDL 82
++ GKGGVGKTS AA+ ++ A GH TLV+STDPAHSL+DSF +L
Sbjct: 5 LMTGKGGVGKTSVAAATGLRCAELGHKTLVLSTDPAHSLADSFDLEL 51
>ARS2_AQUAE (O66674) Putative arsenical pump-driving ATPase 2 (EC
3.6.3.16) (Arsenite-translocating ATPase 2) (Arsenical
resistance ATPase 2) (Arsenite-transporting ATPase 2)
Length = 299
Score = 60.5 bits (145), Expect = 7e-09
Identities = 27/52 (51%), Positives = 37/52 (70%)
Query: 33 KYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSDSFAQDLTG 84
+ + GGKGGVGKT+ +++ AVK + G L++STDPAHSLSD F +L G
Sbjct: 2 RVFFFGGKGGVGKTTASSAFAVKLSEQGKKVLLLSTDPAHSLSDVFNTELQG 53
>ARA1_ECOLI (P08690) Arsenical pump-driving ATPase (EC 3.6.3.16)
(Arsenite-translocating ATPase) (Arsenical resistance
ATPase) (Arsenite-transporting ATPase)
Length = 583
Score = 59.3 bits (142), Expect = 1e-08
Identities = 33/79 (41%), Positives = 46/79 (57%), Gaps = 2/79 (2%)
Query: 34 YYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSDSFAQDLTGGALVQVDGP 93
Y GKGGVGKTS + + A++ A G L+VSTDPA ++ F+Q T G +Q
Sbjct: 10 YLFFTGKGGVGKTSISCATAIRLAEQGKRVLLVSTDPASNVGQVFSQ--TIGITIQAIAS 67
Query: 94 DYPLFALEINPEKAREDFR 112
L ALEI+P+ A + +R
Sbjct: 68 VPGLSALEIDPQAAAQQYR 86
Score = 52.4 bits (124), Expect = 2e-06
Identities = 34/96 (35%), Positives = 51/96 (52%), Gaps = 11/96 (11%)
Query: 7 LLSVKSVAAPTE-----------SISVFDDMVAGTERKYYMLGGKGGVGKTSCAASLAVK 55
LLS + VA+P+ S+S D +A E ML GKGGVGKT+ AA++AV+
Sbjct: 291 LLSTQPVASPSSDEYLQQRPDIPSLSALVDDIARNEHGLIMLMGKGGVGKTTMAAAIAVR 350
Query: 56 FANNGHPTLVVSTDPAHSLSDSFAQDLTGGALVQVD 91
A+ G + ++DPA LS + L + ++D
Sbjct: 351 LADMGFDVHLTTSDPAAHLSMTLNGSLNNLQVSRID 386
>ARA2_ECOLI (P52145) Arsenical pump-driving ATPase (EC 3.6.3.16)
(Arsenite-translocating ATPase) (Arsenical resistance
ATPase) (Arsenite-transporting ATPase)
Length = 583
Score = 55.8 bits (133), Expect = 2e-07
Identities = 33/79 (41%), Positives = 45/79 (56%), Gaps = 2/79 (2%)
Query: 34 YYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSDSFAQDLTGGALVQVDGP 93
Y GKGGVGKTS + + A++ A G L+VSTDPA ++ F D T G +Q
Sbjct: 10 YLFFTGKGGVGKTSISCATAIRLAELGKRVLLVSTDPASNVGQVF--DQTIGNTIQPVTA 67
Query: 94 DYPLFALEINPEKAREDFR 112
L ALEI+P+ A + +R
Sbjct: 68 VSGLSALEIDPQDAAQQYR 86
Score = 50.8 bits (120), Expect = 5e-06
Identities = 35/102 (34%), Positives = 50/102 (48%), Gaps = 12/102 (11%)
Query: 19 SISVFDDMVAGTERKYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSDSF 78
S+SV D +A +E ML GKGGVGKT+ AA++AV A+ G + ++DPA LS +
Sbjct: 314 SLSVLVDDIARSEHGLIMLMGKGGVGKTTMAAAIAVSLADKGFNVHLTTSDPAAHLSTT- 372
Query: 79 AQDLTGGALVQVDGPDYPLFALEINPEKAREDFRDVAKQNGG 120
++G L INP E +R + G
Sbjct: 373 -----------LNGSLKNLQVSRINPHDETERYRQHVLETKG 403
>ARSA_ACIMU (O50593) Arsenical pump-driving ATPase (EC 3.6.3.16)
(Arsenite-translocating ATPase) (Arsenical resistance
ATPase) (Arsenite-transporting ATPase)
Length = 583
Score = 54.3 bits (129), Expect = 5e-07
Identities = 32/79 (40%), Positives = 44/79 (55%), Gaps = 2/79 (2%)
Query: 34 YYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSDSFAQDLTGGALVQVDGP 93
Y GKGGVGKTS + + A+ A G L+VSTDPA ++ F DL G ++
Sbjct: 10 YLFFTGKGGVGKTSISCATAIHLAEQGKRVLLVSTDPASNVGQVF--DLAIGNTIRPVTA 67
Query: 94 DYPLFALEINPEKAREDFR 112
L ALEI+P++A +R
Sbjct: 68 VPGLSALEIDPQEAARQYR 86
Score = 48.5 bits (114), Expect = 3e-05
Identities = 34/102 (33%), Positives = 50/102 (48%), Gaps = 12/102 (11%)
Query: 19 SISVFDDMVAGTERKYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSDSF 78
S+S D +A +E ML GKGGVGKT+ AA++AV+ A+ G + ++DPA LS +
Sbjct: 314 SLSGLVDDIARSEHGLIMLMGKGGVGKTTMAAAIAVRLADMGFDVHLTTSDPAAHLSTT- 372
Query: 79 AQDLTGGALVQVDGPDYPLFALEINPEKAREDFRDVAKQNGG 120
++G L INP E +R + G
Sbjct: 373 -----------LNGSLKNLQVSRINPHDETERYRQHVLETKG 403
>ARS1_AQUAE (O66908) Putative arsenical pump-driving ATPase 1 (EC
3.6.3.16) (Arsenite-translocating ATPase 1) (Arsenical
resistance ATPase 1) (Arsenite-transporting ATPase 1)
Length = 396
Score = 50.4 bits (119), Expect = 7e-06
Identities = 23/43 (53%), Positives = 30/43 (69%)
Query: 36 MLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSDSF 78
+ GKGGVGKT+ +A+ K + G +VVS DPAHSL+DSF
Sbjct: 5 LFSGKGGVGKTTISAATGYKLSQLGKKVIVVSLDPAHSLADSF 47
>TRL5_ECOLI (Q00187) Protein traL
Length = 241
Score = 40.0 bits (92), Expect = 0.009
Identities = 29/91 (31%), Positives = 43/91 (46%), Gaps = 1/91 (1%)
Query: 34 YYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSDSFAQDLTGGALVQVDGP 93
+ +L GKGGVGK+ AA++A A G L + TDP ++ + + + L L +DG
Sbjct: 5 HMVLQGKGGVGKSMIAATIAQYKAGKGQKPLCIDTDPVNATFEGY-KALNVRRLNIMDGD 63
Query: 94 DYPLFALEINPEKAREDFRDVAKQNGGSTGV 124
D + E DV NG S+ V
Sbjct: 64 DINTRNFDALVEMIASAEGDVIIDNGASSFV 94
>YZ24_METJA (Q60283) Hypothetical protein MJECL24
Length = 259
Score = 39.3 bits (90), Expect = 0.016
Identities = 18/36 (50%), Positives = 24/36 (66%)
Query: 40 KGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLS 75
KGGVGKT+ A +L+ A G+ TLV+ DP +LS
Sbjct: 10 KGGVGKTTIALNLSFTLAEKGYDTLVIDLDPQFNLS 45
>MIND_CHLVU (P56346) Putative septum site-determining protein minD
Length = 282
Score = 38.1 bits (87), Expect = 0.035
Identities = 18/46 (39%), Positives = 25/46 (54%)
Query: 24 DDMVAGTERKYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTD 69
DD G ER + GKGGVGKT+ A+L + A G+ ++ D
Sbjct: 9 DDNSKGLERVIVITSGKGGVGKTTTTANLGMSIARLGYRVALIDAD 54
>MIND_BUCBP (Q89AI3) Septum site-determining protein minD (Cell
division inhibitor minD)
Length = 269
Score = 37.7 bits (86), Expect = 0.045
Identities = 17/31 (54%), Positives = 22/31 (70%)
Query: 39 GKGGVGKTSCAASLAVKFANNGHPTLVVSTD 69
GKGGVGKT+ +A+LA FA G T+V+ D
Sbjct: 9 GKGGVGKTTSSAALATGFAKKGKKTVVIDFD 39
>BCHL_RHOGE (Q9JPA5) Light-independent protochlorophyllide
reductase iron-sulfur ATP-binding protein (EC 1.18.-.-)
(LI-POR subunit L) (DPOR subunit L)
Length = 302
Score = 37.4 bits (85), Expect = 0.059
Identities = 15/40 (37%), Positives = 26/40 (64%)
Query: 33 KYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTDPAH 72
K + + GKGG+GK++ +++L+V F+ G L + DP H
Sbjct: 37 KVFAIYGKGGIGKSTTSSNLSVAFSKLGKRVLQIGCDPKH 76
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.347 0.152 0.531
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 37,738,596
Number of Sequences: 164201
Number of extensions: 1466440
Number of successful extensions: 6505
Number of sequences better than 10.0: 123
Number of HSP's better than 10.0 without gapping: 114
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 6379
Number of HSP's gapped (non-prelim): 127
length of query: 359
length of database: 59,974,054
effective HSP length: 111
effective length of query: 248
effective length of database: 41,747,743
effective search space: 10353440264
effective search space used: 10353440264
T: 11
A: 40
X1: 14 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.7 bits)
S2: 66 (30.0 bits)
Medicago: description of AC144928.8


