

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144723.3 + phase: 0
(127 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
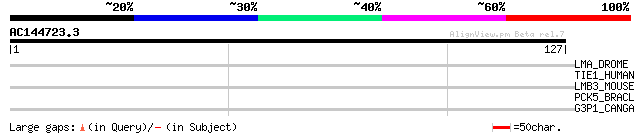
Score E
Sequences producing significant alignments: (bits) Value
LMA_DROME (Q00174) Laminin alpha chain precursor 28 2.7
TIE1_HUMAN (P35590) Tyrosine-protein kinase receptor Tie-1 precu... 28 4.7
LMB3_MOUSE (Q61087) Laminin beta-3 chain precursor (Laminin 5 be... 28 4.7
PCK5_BRACL (Q9NJ15) Proprotein convertase subtilisin/kexin type ... 27 8.0
G3P1_CANGA (Q6FPW3) Glyceraldehyde-3-phosphate dehydrogenase 1 (... 27 8.0
>LMA_DROME (Q00174) Laminin alpha chain precursor
Length = 3712
Score = 28.5 bits (62), Expect = 2.7
Identities = 25/83 (30%), Positives = 32/83 (38%), Gaps = 18/83 (21%)
Query: 27 CKGPKYPPKECCG-SFKEFACPYADVINDLTNDCASTMFSYINLYGRY-----PPGLF-- 78
CK K P C + E CP +VI DL CA + + + G P G+
Sbjct: 1418 CKPCKCPNSAMCEPTTGECMCP-PNVIGDLCEKCAPNTYGFHQVIGCEECACNPMGIANG 1476
Query: 79 ---------ASECREGKEGLACD 92
ECR+ EG ACD
Sbjct: 1477 NSQCDLFNGTCECRQNIEGRACD 1499
>TIE1_HUMAN (P35590) Tyrosine-protein kinase receptor Tie-1
precursor (EC 2.7.1.112)
Length = 1138
Score = 27.7 bits (60), Expect = 4.7
Identities = 22/87 (25%), Positives = 33/87 (37%), Gaps = 25/87 (28%)
Query: 20 YTIITSKCKGPKYPP---KECCGSFKEFACPYADVINDLTNDCASTMFSYINLYGRYPPG 76
+ +I C ++ P KEC G C + V +D +C PPG
Sbjct: 208 FRLIVRGCGAGRWGPGCTKECPG------CLHGGVCHDHDGECVC------------PPG 249
Query: 77 LFASEC----REGKEGLACDALPPSVS 99
+ C REG+ G +C P +S
Sbjct: 250 FTGTRCEQACREGRFGQSCQEQCPGIS 276
>LMB3_MOUSE (Q61087) Laminin beta-3 chain precursor (Laminin 5 beta
3) (Kalinin B1 chain)
Length = 1168
Score = 27.7 bits (60), Expect = 4.7
Identities = 17/48 (35%), Positives = 20/48 (41%), Gaps = 1/48 (2%)
Query: 46 CPYADVINDLTNDCASTMFSYINLYGRYPPGLFASECR-EGKEGLACD 92
CP + LT A+ YG P G A +C G EG ACD
Sbjct: 497 CPCREGFGGLTCSSAAIRQCPDQTYGHVPTGCRACDCDFRGTEGPACD 544
>PCK5_BRACL (Q9NJ15) Proprotein convertase subtilisin/kexin type 5
precursor (EC 3.4.21.-) (Proprotein convertase PC6-like)
(aPC6)
Length = 1696
Score = 26.9 bits (58), Expect = 8.0
Identities = 24/79 (30%), Positives = 35/79 (43%), Gaps = 8/79 (10%)
Query: 8 PAGCSVNFEFLNYTIITSKCKGPKYPPKECCGSFKEFACPYADVINDLTNDCASTMFSYI 67
PAG N E ++ + S C+ +Y E G ++ CPY D DCA +YI
Sbjct: 1152 PAGFIGNAE--SHECVESSCEQDQYYSSET-GRCED--CPYNCRACDNDGDCAECAPTYI 1206
Query: 68 NLYGRYPPGLFASECREGK 86
+ GR P C +G+
Sbjct: 1207 VVDGRCRP---EETCEDGE 1222
>G3P1_CANGA (Q6FPW3) Glyceraldehyde-3-phosphate dehydrogenase 1
(EC 1.2.1.12) (GAPDH 1)
Length = 332
Score = 26.9 bits (58), Expect = 8.0
Identities = 13/24 (54%), Positives = 17/24 (70%), Gaps = 2/24 (8%)
Query: 52 IND--LTNDCASTMFSYINLYGRY 73
IND +TND A+ MF Y + +GRY
Sbjct: 31 INDPFITNDYAAYMFKYDSTHGRY 54
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.325 0.141 0.454
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,961,033
Number of Sequences: 164201
Number of extensions: 621189
Number of successful extensions: 1576
Number of sequences better than 10.0: 5
Number of HSP's better than 10.0 without gapping: 0
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 1575
Number of HSP's gapped (non-prelim): 6
length of query: 127
length of database: 59,974,054
effective HSP length: 103
effective length of query: 24
effective length of database: 43,061,351
effective search space: 1033472424
effective search space used: 1033472424
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 58 (26.9 bits)
Medicago: description of AC144723.3


