

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144514.8 - phase: 0 /pseudo
(555 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
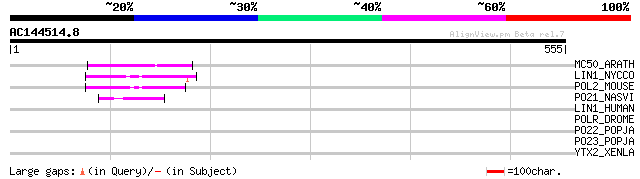
Score E
Sequences producing significant alignments: (bits) Value
MC50_ARATH (P92555) Hypothetical mitochondrial protein AtMg01250... 78 5e-14
LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog 50 1e-05
POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains... 46 2e-04
PO21_NASVI (Q03278) Retrovirus-related Pol polyprotein from type... 46 3e-04
LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog 40 0.012
POLR_DROME (P16423) Retrovirus-related Pol polyprotein from type... 39 0.046
PO22_POPJA (Q03274) Retrovirus-related Pol polyprotein from type... 35 0.67
PO23_POPJA (Q05118) Retrovirus-related Pol polyprotein from type... 34 1.1
YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein ... 33 2.5
>MC50_ARATH (P92555) Hypothetical mitochondrial protein AtMg01250
(ORF102)
Length = 122
Score = 78.2 bits (191), Expect = 5e-14
Identities = 42/105 (40%), Positives = 59/105 (56%), Gaps = 1/105 (0%)
Query: 78 LVNNDAVGPFIPGRGLRQGDPLSPYMFILCVEGLSSLIRDAEEHRIISGISICRGAPSIS 137
++N G P RGLRQGDPLSPY+FILC E LS L R A+E + GI + +P I+
Sbjct: 13 IINGAPQGLVTPSRGLRQGDPLSPYLFILCTEVLSGLCRRAQEQGRLPGIRVSNNSPRIN 72
Query: 138 HLLFADDCFLFFKADEGQTQVMKNILITYKAASGQAISLPISEIY 182
HLLFADD + Q+ ++ + G ++ P+S +Y
Sbjct: 73 HLLFADDT-SSARWIPLAAQIWPIFFLSMRLFQGNPVNHPMSNLY 116
>LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog
Length = 1260
Score = 50.4 bits (119), Expect = 1e-05
Identities = 36/113 (31%), Positives = 56/113 (48%), Gaps = 7/113 (6%)
Query: 76 SVLVNNDAVGPFIPGRGLRQGDPLSPYMFILCVEGLSSLIRDAEEHRIISGISICRGAPS 135
++++N + F G RQG PLSP +F + +E L+ IR E + I GI I G+
Sbjct: 639 NIILNGVKLKSFPLRSGTRQGCPLSPLLFNIVMEVLAIAIR---EEKAIKGIHI--GSEE 693
Query: 136 ISHLLFADDCFLFFKADEGQTQVMKNILITYKAASGQAISL--PISEIYCNQN 186
I LFADD ++ + T + ++ Y SG I+ ++ IY N N
Sbjct: 694 IKLSLFADDMIVYLENTRDSTTKLLEVIKEYSNVSGYKINTHKSVAFIYTNNN 746
>POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains:
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1300
Score = 46.2 bits (108), Expect = 2e-04
Identities = 30/100 (30%), Positives = 50/100 (50%), Gaps = 5/100 (5%)
Query: 76 SVLVNNDAVGPFIPGRGLRQGDPLSPYMFILCVEGLSSLIRDAEEHRIISGISICRGAPS 135
++ VN + + G RQG PLSPY+F + +E L+ IR +E I GI I G
Sbjct: 666 NIKVNGEKLEAIPLKSGTRQGCPLSPYLFNIVLEVLARAIRQQKE---IKGIQI--GKEE 720
Query: 136 ISHLLFADDCFLFFKADEGQTQVMKNILITYKAASGQAIS 175
+ L ADD ++ + T+ + N++ ++ G I+
Sbjct: 721 VKISLLADDMIVYISDPKNSTRELLNLINSFGEVVGYKIN 760
>PO21_NASVI (Q03278) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 1025
Score = 45.8 bits (107), Expect = 3e-04
Identities = 26/66 (39%), Positives = 33/66 (49%), Gaps = 8/66 (12%)
Query: 89 PGRGLRQGDPLSPYMFILCVEGLSSLIRDAEEHRIISGISICRGAPSISHLLFADDCFLF 148
P RG+RQGDPLSP +F + + DA R+ GA I L+FADD L
Sbjct: 509 PARGVRQGDPLSPLLF--------NCVMDAVLRRLPENTGFLMGAEKIGALVFADDLVLL 560
Query: 149 FKADEG 154
+ EG
Sbjct: 561 AETREG 566
>LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog
Length = 1259
Score = 40.4 bits (93), Expect = 0.012
Identities = 28/95 (29%), Positives = 46/95 (47%), Gaps = 5/95 (5%)
Query: 92 GLRQGDPLSPYMFILCVEGLSSLIRDAEEHRIISGISICRGAPSISHLLFADDCFLFFKA 151
G RQG PLSP + + +E L+ IR +E I GI + G + LFADD ++ +
Sbjct: 655 GTRQGCPLSPLLPNIVLEVLARAIRQEKE---IKGIQL--GKEEVKLSLFADDMIVYLEN 709
Query: 152 DEGQTQVMKNILITYKAASGQAISLPISEIYCNQN 186
Q + ++ + SG I++ S+ + N
Sbjct: 710 PIVSAQNLLKLISNFSKVSGYKINVQKSQAFLYTN 744
>POLR_DROME (P16423) Retrovirus-related Pol polyprotein from type II
retrotransposable element R2DM [Contains: Protease (EC
3.4.23.-); Reverse transcriptase (EC 2.7.7.49);
Endonuclease]
Length = 1057
Score = 38.5 bits (88), Expect = 0.046
Identities = 24/68 (35%), Positives = 33/68 (48%), Gaps = 8/68 (11%)
Query: 87 FIPGRGLRQGDPLSPYMFILCVEGLSSLIRDAEEHRIISGISICRGAPSISHLLFADDCF 146
F+P RG++QGDPLSP +F +L+ D + S I G + FADD
Sbjct: 533 FVPARGVKQGDPLSPILF--------NLVMDRLLRTLPSEIGAKVGNAITNAAAFADDLV 584
Query: 147 LFFKADEG 154
LF + G
Sbjct: 585 LFAETRMG 592
>PO22_POPJA (Q03274) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 711
Score = 34.7 bits (78), Expect = 0.67
Identities = 21/57 (36%), Positives = 29/57 (50%), Gaps = 6/57 (10%)
Query: 91 RGLRQGDPLSPYMFILCVEGLSSLIRDAEEHRIISGISICRGAPSISHLLFADDCFL 147
RG++QGDPLSP++F ++ L ++ GI G I L FADD L
Sbjct: 199 RGVKQGDPLSPFLFNAVLDELLCSLQST------PGIGGTIGEEKIPVLAFADDLLL 249
>PO23_POPJA (Q05118) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 606
Score = 33.9 bits (76), Expect = 1.1
Identities = 22/61 (36%), Positives = 33/61 (54%), Gaps = 6/61 (9%)
Query: 92 GLRQGDPLSPYMFILCVEGLSSLIRDAEEHRIISGISICRGAPSISHLLFADDCFLFFKA 151
G++QGDPLSP +F + ++ L + + D + G S+ A I+ L FADD L
Sbjct: 85 GVKQGDPLSPVLFNIVLDELVTRLNDEQ-----PGASM-TPACKIASLAFADDLLLLEDR 138
Query: 152 D 152
D
Sbjct: 139 D 139
>YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein
(ORF 2)
Length = 1308
Score = 32.7 bits (73), Expect = 2.5
Identities = 21/71 (29%), Positives = 34/71 (47%), Gaps = 5/71 (7%)
Query: 77 VLVNNDAVGPFIPGRGLRQGDPLSPYMFILCVEGLSSLIRDAEEHRIISGISICRGAPSI 136
V +N P GRG+RQG PLS ++ L +E L+R + ++G+ + +
Sbjct: 635 VKINWSLTAPLAFGRGVRQGCPLSGQLYSLAIEPFLCLLR-----KRLTGLVLKEPDMRV 689
Query: 137 SHLLFADDCFL 147
+ADD L
Sbjct: 690 VLSAYADDVIL 700
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.343 0.152 0.517
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 60,240,388
Number of Sequences: 164201
Number of extensions: 2398650
Number of successful extensions: 7452
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 7444
Number of HSP's gapped (non-prelim): 9
length of query: 555
length of database: 59,974,054
effective HSP length: 115
effective length of query: 440
effective length of database: 41,090,939
effective search space: 18080013160
effective search space used: 18080013160
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.5 bits)
S2: 68 (30.8 bits)
Medicago: description of AC144514.8


