

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC142507.1 - phase: 0
(542 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
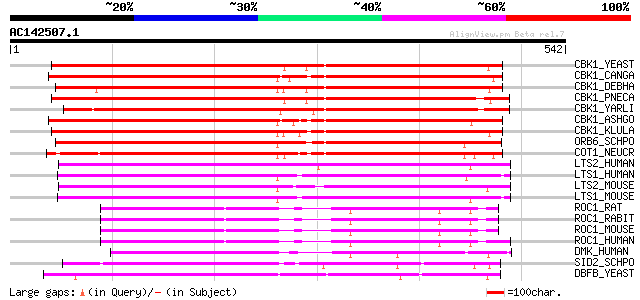
Score E
Sequences producing significant alignments: (bits) Value
CBK1_YEAST (P53894) Serine/threonine-protein kinase CBK1 (EC 2.7... 400 e-111
CBK1_CANGA (Q6FP74) Serine/threonine-protein kinase CBK1 (EC 2.7... 400 e-111
CBK1_DEBHA (Q6BLJ9) Serine/threonine-protein kinase CBK1 (EC 2.7... 394 e-109
CBK1_PNECA (Q6TGC6) Serine/threonine-protein kinase CBK1 (EC 2.7... 393 e-109
CBK1_YARLI (Q6CFS5) Serine/threonine-protein kinase CBK1 (EC 2.7... 392 e-108
CBK1_ASHGO (Q754N7) Serine/threonine-protein kinase CBK1 (EC 2.7... 392 e-108
CBK1_KLULA (P31034) Serine/threonine-protein kinase CBK1 (EC 2.7... 390 e-108
ORB6_SCHPO (O13310) Serine/threonine-protein kinase orb6 (EC 2.7... 387 e-107
COT1_NEUCR (P38679) Serine/threonine-protein kinase cot-1 (EC 2.... 356 1e-97
LTS2_HUMAN (Q9NRM7) Serine/threonine protein kinase LATS2 (EC 2.... 346 8e-95
LTS1_HUMAN (O95835) Serine/threonine protein kinase LATS1 (EC 2.... 340 7e-93
LTS2_MOUSE (Q7TSJ6) Serine/threonine protein kinase LATS2 (EC 2.... 339 9e-93
LTS1_MOUSE (Q8BYR2) Serine/threonine protein kinase LATS1 (EC 2.... 337 5e-92
ROC1_RAT (Q63644) Rho-associated protein kinase 1 (EC 2.7.1.37) ... 291 2e-78
ROC1_RABIT (O77819) Rho-associated protein kinase 1 (EC 2.7.1.37... 291 2e-78
ROC1_MOUSE (P70335) Rho-associated protein kinase 1 (EC 2.7.1.37... 291 2e-78
ROC1_HUMAN (Q13464) Rho-associated protein kinase 1 (EC 2.7.1.37... 290 5e-78
DMK_HUMAN (Q09013) Myotonin-protein kinase (EC 2.7.1.-) (Myotoni... 267 5e-71
SID2_SCHPO (Q09898) Serine/threonine-protein kinase sid2 (EC 2.7... 246 8e-65
DBFB_YEAST (P32328) Serine/threonine-protein kinase DBF20 (EC 2.... 233 1e-60
>CBK1_YEAST (P53894) Serine/threonine-protein kinase CBK1 (EC
2.7.1.37) (Cell wall biosynthesis kinase)
Length = 756
Score = 400 bits (1027), Expect = e-111
Identities = 212/471 (45%), Positives = 312/471 (66%), Gaps = 32/471 (6%)
Query: 42 TKLKVEAAKHFVESYYKNQKQNQQERKERRNILENKLADAEVSKEDKKNLLKNFEEMETE 101
T+ K A K +E++Y++ + ER ERR LE +L S+E K L + + E++
Sbjct: 279 TQDKAAAVKLKIENFYQSSVKYAIERNERRVELETELTSHNWSEERKSRQLSSLGKKESQ 338
Query: 102 IMRRQRLKMGADDFEPLTMIGKGAFGEVRICREKTTGQVYAMKKLQKSEMLRRGQVEYVK 161
+R +R ++ +DF + +IGKGAFGEVR+ ++K TG++YAMK L KSEM ++ Q+ +VK
Sbjct: 339 FLRLRRTRLSLEDFHTVKVIGKGAFGEVRLVQKKDTGKIYAMKTLLKSEMYKKDQLAHVK 398
Query: 162 SERNLLAEVDSNCIVKLYCSFQDDEYLYLIMEYLPGGDMMTLLMRREVLTENEAKFYIGE 221
+ER++LA DS +V LY SFQD +YLYLIME+LPGGD+MT+L+R ++ TE+ +FY+ E
Sbjct: 399 AERDVLAGSDSPWVVSLYYSFQDAQYLYLIMEFLPGGDLMTMLIRWQLFTEDVTRFYMAE 458
Query: 222 TVLAIESIHKHNYIHRDIKPDNLLLDRNGHVKLSDFGLCKPLDCSN--------LQEKDL 273
+LAIE+IHK +IHRDIKPDN+L+D GH+KLSDFGL ++ LQ+ +
Sbjct: 459 CILAIETIHKLGFIHRDIKPDNILIDIRGHIKLSDFGLSTGFHKTHDSNYYKKLLQQDEA 518
Query: 274 SDGVNRSGVLQSDEL------------TLSPNQSQQQQLEHWRKNHSRMLAYSTVGTPDY 321
++G+++ G ++ ++S S +QQ++ WRK+ R++AYSTVGTPDY
Sbjct: 519 TNGISKPGTYNANTTDTANKRQTMVVDSISLTMSNRQQIQTWRKSR-RLMAYSTVGTPDY 577
Query: 322 IAPEVILKRGYGVECDWWSLGAIMYEMLVGYPPFHSDDPRTTCRKIAHWKTYLKFPKD-K 380
IAPE+ L +GYG ECDWWSLGAIMYE L+G+PPF S+ P+ T RKI +++ L+FP D
Sbjct: 578 IAPEIFLYQGYGQECDWWSLGAIMYECLIGWPPFCSETPQETYRKIMNFEQTLQFPDDIH 637
Query: 381 LSPEAKDLICRLLCNVKQRLGAK-GADEIKAHPWFKVVEWEKLYQMRAPFIPKVNDELDT 439
+S EA+DLI RLL + QRLG GADEIK+HP+F+ V+W + Q+ AP+IPK++ DT
Sbjct: 638 ISYEAEDLIRRLLTHADQRLGRHGGADEIKSHPFFRGVDWNTIRQVEAPYIPKLSSITDT 697
Query: 440 RNFDKFEEEDQKTEPS-PKAGPWRKMLQ--------SKDINFVGYTYKNYE 481
R F E E+ P+ +A R+ + +D+ F+GYTY ++
Sbjct: 698 RFFPTDELENVPDSPAMAQAAKQREQMTKQGGSAPVKEDLPFIGYTYSRFD 748
>CBK1_CANGA (Q6FP74) Serine/threonine-protein kinase CBK1 (EC
2.7.1.37)
Length = 773
Score = 400 bits (1027), Expect = e-111
Identities = 220/476 (46%), Positives = 309/476 (64%), Gaps = 37/476 (7%)
Query: 39 SNETKLKVEAAKHFVESYYKNQKQNQQERKERRNILENKLADAEVSKEDKKNLLKNFEEM 98
S T+ K A K VE+YY+ + ER ERR LE +L S+E L + +
Sbjct: 294 SKSTQDKAAAVKLKVENYYQQSVKYAIERNERRVELETELGSHNWSEERNARQLASLGKK 353
Query: 99 ETEIMRRQRLKMGADDFEPLTMIGKGAFGEVRICREKTTGQVYAMKKLQKSEMLRRGQVE 158
E++ +R +R ++ +DF + +IGKGAFGEVR+ ++K TG++YAMK L KSEM ++ Q+
Sbjct: 354 ESQFLRLRRTRLSLEDFHTVQVIGKGAFGEVRLVQKKDTGKIYAMKTLLKSEMYKKDQLA 413
Query: 159 YVKSERNLLAEVDSNCIVKLYCSFQDDEYLYLIMEYLPGGDMMTLLMRREVLTENEAKFY 218
+VK+ER++LA DS IV LY SFQD +YLYLIME+LPGGD+MT+L+R ++ TE+ +FY
Sbjct: 414 HVKAERDVLAGTDSPWIVSLYYSFQDAQYLYLIMEFLPGGDLMTMLIRWQLFTEDVTRFY 473
Query: 219 IGETVLAIESIHKHNYIHRDIKPDNLLLDRNGHVKLSDFGLC----KPLDCSNLQEK--- 271
+ E +LAIE+IHK +IHRDIKPDN+L+D GH+KLSDFGL K D SN +K
Sbjct: 474 MAECILAIETIHKLGFIHRDIKPDNILIDIRGHIKLSDFGLSTGFHKTHD-SNYYKKLLQ 532
Query: 272 ------------DLSDGVNRSGVLQSDELTLSPNQSQQQQLEHWRKNHSRMLAYSTVGTP 319
D DG N + D + L+ S +QQ++ WRK+ R++AYSTVGTP
Sbjct: 533 QDEATTKNGAPNDAGDGSNNRQTMIVDSINLT--MSNRQQIQTWRKSR-RLMAYSTVGTP 589
Query: 320 DYIAPEVILKRGYGVECDWWSLGAIMYEMLVGYPPFHSDDPRTTCRKIAHWKTYLKFPKD 379
DYIAPE+ L +GYG +CDWWSLGAIMYE L+G+PPF S+ P+ T RKI +++ L+FP+D
Sbjct: 590 DYIAPEIFLYQGYGQDCDWWSLGAIMYECLIGWPPFCSETPQETYRKIMNFEQTLQFPED 649
Query: 380 -KLSPEAKDLICRLLCNVKQRLGAK-GADEIKAHPWFKVVEWEKLYQMRAPFIPKVNDEL 437
+S EA+DLI RLL + QRLG + GADEIK+HP+F+ V+W + Q+ AP+IPK++
Sbjct: 650 VHISYEAEDLIRRLLTHSNQRLGRQGGADEIKSHPFFRGVDWNTIRQVEAPYIPKLSSIT 709
Query: 438 DTRNFDKFEEEDQKTEPS-PKAGPWRKMLQSKDIN-----------FVGYTYKNYE 481
DTR F E E+ P+ +A R+ + +N F+GYTY ++
Sbjct: 710 DTRFFPTDELENVPDSPAMAQAAKQREQMMKNGVNPNQNQVKEDLPFIGYTYSRFD 765
>CBK1_DEBHA (Q6BLJ9) Serine/threonine-protein kinase CBK1 (EC
2.7.1.37)
Length = 716
Score = 394 bits (1013), Expect = e-109
Identities = 218/467 (46%), Positives = 312/467 (66%), Gaps = 31/467 (6%)
Query: 45 KVEAAKHFVESYYKNQKQNQQERKERRNILENKLADAEV--SKEDKKNLLKNFEEMETEI 102
K + K +E+YY+ + ER +RR LE+KL + E S+E K L+N + E++
Sbjct: 243 KASSIKLKLENYYQMSVAHAIERNQRRLDLEHKLLNEESGSSEERKNRQLQNLGKKESQF 302
Query: 103 MRRQRLKMGADDFEPLTMIGKGAFGEVRICREKTTGQVYAMKKLQKSEMLRRGQVEYVKS 162
+R +R K+ +DF + +IGKGAFGEVR+ +++ TG++YAMK L KSEM ++ Q+ +VK+
Sbjct: 303 LRLRRTKLSLEDFNTVKVIGKGAFGEVRLVQKRDTGKIYAMKTLLKSEMYKKDQLAHVKA 362
Query: 163 ERNLLAEVDSNCIVKLYCSFQDDEYLYLIMEYLPGGDMMTLLMRREVLTENEAKFYIGET 222
ER++LA DS +V LY SFQD +YLYLIME+LPGGD+MT+L+R ++ TE+ +FY+ E
Sbjct: 363 ERDVLAGSDSPWVVSLYYSFQDAQYLYLIMEFLPGGDLMTMLIRWQIFTEDITRFYMAEC 422
Query: 223 VLAIESIHKHNYIHRDIKPDNLLLDRNGHVKLSDFGLC----KPLDCS---NLQEKDLSD 275
VLAIE+IHK +IHRDIKPDN+L+D GH+KLSDFGL K D + L EK+
Sbjct: 423 VLAIEAIHKLGFIHRDIKPDNILIDIRGHIKLSDFGLSTGFHKTHDSNYYKKLLEKENPH 482
Query: 276 GVN-RSGVLQSDEL-----------TLSPNQSQQQQLEHWRKNHSRMLAYSTVGTPDYIA 323
N ++G LQ+ + + S +QQ++ WRK+ R++AYSTVGTPDYIA
Sbjct: 483 HTNPQNGNLQAPSMATNNRNSMMVDAIHLTMSNRQQMQTWRKSR-RLMAYSTVGTPDYIA 541
Query: 324 PEVILKRGYGVECDWWSLGAIMYEMLVGYPPFHSDDPRTTCRKIAHWKTYLKFPKD-KLS 382
PE+ + +GYG ECDWWSLGAIM+E L+G+PPF S+ P T RKI +W+ L+ P D LS
Sbjct: 542 PEIFVHQGYGQECDWWSLGAIMFECLIGWPPFCSETPHETYRKILNWQETLQIPDDIHLS 601
Query: 383 PEAKDLICRLLCNVKQRLGA-KGADEIKAHPWFKVVEWEKLYQMRAPFIPKVNDELDTRN 441
PE++DLI +LL N + RLG GADE+K HP+F+ V+W+ + ++ APFIPK+ DTR
Sbjct: 602 PESEDLIRKLLTNAENRLGRYNGADELKLHPFFRGVDWDTIRKVDAPFIPKLRSITDTRF 661
Query: 442 FDKFEEEDQKTEPS-PKAGPWR-KMLQS-----KDINFVGYTYKNYE 481
F E E+ P+ KA R ++LQ+ +D+ F+GYTY ++
Sbjct: 662 FPTDELENVPDSPALSKAMEQRDQVLQNGGNVKEDLPFIGYTYSRFD 708
>CBK1_PNECA (Q6TGC6) Serine/threonine-protein kinase CBK1 (EC
2.7.1.37)
Length = 507
Score = 393 bits (1010), Expect = e-109
Identities = 207/461 (44%), Positives = 300/461 (64%), Gaps = 21/461 (4%)
Query: 42 TKLKVEAAKHFVESYYKNQKQNQQERKERRNILENKLADAEVSKEDKKNLLKNFEEMETE 101
T + + K +E YK + ER +RR E KLA S+E KK L + + E++
Sbjct: 53 THERAASVKLKLEHLYKVTVEQTVERNQRRMDFEAKLAQDRGSEERKKRQLNSLGQKESQ 112
Query: 102 IMRRQRLKMGADDFEPLTMIGKGAFGEVRICREKTTGQVYAMKKLQKSEMLRRGQVEYVK 161
+R +R K+ +DF + +IGKGAFGEVR+ ++ TG++YAMK L KSEM ++ Q+ +VK
Sbjct: 113 FLRLRRTKLSLNDFHTVKVIGKGAFGEVRLVQKIDTGKIYAMKTLLKSEMFKKDQLAHVK 172
Query: 162 SERNLLAEVDSNCIVKLYCSFQDDEYLYLIMEYLPGGDMMTLLMRREVLTENEAKFYIGE 221
+ER++LAE DS +V LY SFQD +YLYLIME+LPGGD+MT+L++ + +E+ +FYI E
Sbjct: 173 AERDVLAESDSPWVVSLYYSFQDSQYLYLIMEFLPGGDLMTMLIKYDTFSEDVTRFYIAE 232
Query: 222 TVLAIESIHKHNYIHRDIKPDNLLLDRNGHVKLSDFGLCKPLDCSN---LQEKDLSDGVN 278
+LAIE++HK +IHRDIKPDN+L+D+ GH+KLSDFGL ++ ++ +N
Sbjct: 233 CILAIEAVHKLGFIHRDIKPDNILIDKTGHIKLSDFGLSMGFHKTHDNAYYQRLFESKIN 292
Query: 279 RSGVLQSDEL---TLSPNQSQQQQLEHWRKNHSRMLAYSTVGTPDYIAPEVILKRGYGVE 335
S + L T+S S + ++ W+KN R++AYSTVGTPDYIAPE+ + GYG E
Sbjct: 293 TSTSSTQNSLMVDTISLTMSSKDKIATWKKNR-RIMAYSTVGTPDYIAPEIFTQHGYGQE 351
Query: 336 CDWWSLGAIMYEMLVGYPPFHSDDPRTTCRKIAHWKTYLKFPKD-KLSPEAKDLICRLLC 394
CDWWSLGAIM+E L+G+PPF S++ T RKI +W+ L FP+D LS EA+DLI +LL
Sbjct: 352 CDWWSLGAIMFECLIGWPPFCSENAHETYRKIINWRENLYFPEDLHLSAEAEDLIRKLLT 411
Query: 395 NVKQRLGAKGADEIKAHPWFKVVEWEKLYQMRAPFIPKVNDELDTRNFDKFEEEDQKTEP 454
+ QRLG GA++IK HP+F+ V W+ + ++ APFIP++ DT F++ + T
Sbjct: 412 SADQRLGRYGANDIKLHPFFRGVNWDTIREINAPFIPQLKSITDTSYFEEIDTIPNITMN 471
Query: 455 SPKAGPWRKMLQSK-------DINFVGYTYKNYEMVDANAI 488
SP +LQ+K ++ FVGYTYK ++M+ I
Sbjct: 472 SP------PVLQNKIPSDVDQNLAFVGYTYKRFDMMTQKGI 506
>CBK1_YARLI (Q6CFS5) Serine/threonine-protein kinase CBK1 (EC
2.7.1.37)
Length = 588
Score = 392 bits (1006), Expect = e-108
Identities = 211/449 (46%), Positives = 295/449 (64%), Gaps = 19/449 (4%)
Query: 53 VESYYKNQKQNQQERKERRNILENKLADAEVSKEDKKNLLKNFEEMETEIMRRQRLKMGA 112
+ +YYK Q+ ER +RR+ E + A S+E + +L N+ + ET +R +R +M
Sbjct: 146 ISTYYKGIVQHAIERHQRRSAAEASVEQA-TSEERRNRVLTNYGKKETAYLRMRRTRMAL 204
Query: 113 DDFEPLTMIGKGAFGEVRICREKTTGQVYAMKKLQKSEMLRRGQVEYVKSERNLLAEVDS 172
+DF + +IGKGAFGEVR+ +++ G++YAMK + K EM + Q +VK+ER++LA+ DS
Sbjct: 205 EDFVTVKVIGKGAFGEVRLVQKRDNGKIYAMKTMLKKEMDMKEQWAHVKAERDVLADSDS 264
Query: 173 NCIVKLYCSFQDDEYLYLIMEYLPGGDMMTLLMRREVLTENEAKFYIGETVLAIESIHKH 232
IV LY SFQDD YLYLIME+LPGGD+MT+L++ +V +E+ +FYI E VLAIE+IHK
Sbjct: 265 PWIVSLYFSFQDDLYLYLIMEFLPGGDLMTMLIKYDVFSEDITRFYIAECVLAIEAIHKL 324
Query: 233 NYIHRDIKPDNLLLDRNGHVKLSDFGLCKPL--DCSNLQEKDLSDGVNRSGVLQSDELTL 290
+IHRDIKPDN+L+D+ GH+KLSDFGL S+ K L DG + + Q
Sbjct: 325 GFIHRDIKPDNILIDKTGHIKLSDFGLSTGFHKTHSSAYWKKLKDGTSSNPATQMGPPQN 384
Query: 291 SPNQS--------QQQQLEHWRKNHSRMLAYSTVGTPDYIAPEVILKRGYGVECDWWSLG 342
+ QS +QQ+ WRKN R++AYSTVGTPDYIAPE+ + +GYG ECDWWSLG
Sbjct: 385 TNRQSTYDSIHLTMRQQISTWRKNR-RLMAYSTVGTPDYIAPEIFVHQGYGQECDWWSLG 443
Query: 343 AIMYEMLVGYPPFHSDDPRTTCRKIAHWKTYLKFPKD-KLSPEAKDLICRLLCNVKQRLG 401
AIM+E LVG+PPF S+ PR T KI +W+ L+FP D LSPE++DLI RLL + + RLG
Sbjct: 444 AIMFECLVGWPPFCSEQPRETYHKIINWRETLQFPDDVHLSPESEDLIRRLLTSSENRLG 503
Query: 402 -AKGADEIKAHPWFKVVEWEKLYQMRAPFIPKVNDELDTRNFDKFEEEDQKTEPSPKAGP 460
GA+EIK+HP+F+ V+W + + APF+PK++ DT F E D P +
Sbjct: 504 RIGGANEIKSHPFFRGVDWSSIREFNAPFVPKLSSITDTSYFPTDELGDVSEYPQQSS-- 561
Query: 461 WRKMLQSKDINFVGYTYKNYE-MVDANAI 488
+ +S D+ F+GYT+ ++ M NAI
Sbjct: 562 --RSDRSSDLPFIGYTFSRFDNMTRRNAI 588
>CBK1_ASHGO (Q754N7) Serine/threonine-protein kinase CBK1 (EC
2.7.1.37)
Length = 719
Score = 392 bits (1006), Expect = e-108
Identities = 219/483 (45%), Positives = 313/483 (64%), Gaps = 45/483 (9%)
Query: 39 SNETKLKVEAAKHFVESYYKNQKQNQQERKERRNILENKLADAEVSKEDKKNLLKNFEEM 98
S T+ K A K VE++Y++ + ER +RR LE++L S+E K L + +
Sbjct: 234 SKSTQEKAAAVKLKVENFYQSSVNHAIERNQRRVELESQLLSHGWSEERKNRQLSSLGKK 293
Query: 99 ETEIMRRQRLKMGADDFEPLTMIGKGAFGEVRICREKTTGQVYAMKKLQKSEMLRRGQVE 158
E++ +R +R ++ +DF + +IGKGAFGEVR+ ++K TG++YAMK L KSEM ++ Q+
Sbjct: 294 ESQFLRLRRTRLSLEDFHTVKVIGKGAFGEVRLVQKKDTGKIYAMKTLLKSEMYKKDQLA 353
Query: 159 YVKSERNLLAEVDSNCIVKLYCSFQDDEYLYLIMEYLPGGDMMTLLMRREVLTENEAKFY 218
+VK+ER++LA DS +V LY SFQD +YLYLIME+LPGGD+MT+L+R ++ TE+ +FY
Sbjct: 354 HVKAERDVLAGSDSPWVVSLYYSFQDAQYLYLIMEFLPGGDLMTMLIRWQIFTEDVTRFY 413
Query: 219 IGETVLAIESIHKHNYIHRDIKPDNLLLDRNGHVKLSDFGLC----KPLDCSNLQEKDLS 274
+ E +LAIE+IHK +IHRDIKPDN+L+D GH+KLSDFGL K D SN +K L
Sbjct: 414 MAECILAIEAIHKLGFIHRDIKPDNILIDIRGHIKLSDFGLSTGFHKTHD-SNYYKKLLQ 472
Query: 275 D----------------------GVNRSGVLQSDELTLSPNQSQQQQLEHWRKNHSRMLA 312
+ G NR+ +L D + L+ + +QQ++ WRK+ R++A
Sbjct: 473 EDEQQQNGGNMGKYPASGGGGNGGGNRNTML-VDAIHLT--MTNRQQMQTWRKSR-RLMA 528
Query: 313 YSTVGTPDYIAPEVILKRGYGVECDWWSLGAIMYEMLVGYPPFHSDDPRTTCRKIAHWKT 372
YSTVGTPDYIAPE+ L +GYG ECDWWSLGAIMYE L+G+PPF S+ P+ T RKI +++
Sbjct: 529 YSTVGTPDYIAPEIFLYQGYGQECDWWSLGAIMYECLIGWPPFCSETPQETYRKIMNFEQ 588
Query: 373 YLKFPKD-KLSPEAKDLICRLLCNVKQRLGAKGADEIKAHPWFKVVEWEKLYQMRAPFIP 431
L FP D +S EA+DLI RLL + +RLG GA+EIK HP+F+ V+WE + Q+ AP+IP
Sbjct: 589 TLVFPDDIHISYEAEDLIRRLLSHADERLGRHGANEIKNHPFFRGVDWETIRQVGAPYIP 648
Query: 432 KVNDELDTRNFDKFEEED------------QKTEPSPKAGPWRKMLQSK-DINFVGYTYK 478
K++ DTR F E E+ Q+ + + G Q+K D+ F+GYTY
Sbjct: 649 KLSSVTDTRFFPTDELENVPDSPAMAQAAKQREQMLKQGGSAANTAQAKEDLPFIGYTYS 708
Query: 479 NYE 481
++
Sbjct: 709 RFD 711
>CBK1_KLULA (P31034) Serine/threonine-protein kinase CBK1 (EC
2.7.1.37)
Length = 718
Score = 390 bits (1002), Expect = e-108
Identities = 217/484 (44%), Positives = 309/484 (63%), Gaps = 47/484 (9%)
Query: 42 TKLKVEAAKHFVESYYKNQKQNQQERKERRNILENKLADAEVSKEDKKNLLKNFEEMETE 101
T+ K A K +E++Y++ ER +RR LE++LA + S+E K L + + E++
Sbjct: 230 TQDKAAAVKLKIENFYQSSVGYAIERNQRRLELESELASQDWSEERKNRQLASLGKKESQ 289
Query: 102 IMRRQRLKMGADDFEPLTMIGKGAFGEVRICREKTTGQVYAMKKLQKSEMLRRGQVEYVK 161
+R +R ++ DDF + +IGKGAFGEVR+ ++K TG++YAMK L KSEM + Q+ +VK
Sbjct: 290 FLRLRRTRLSLDDFNSVKVIGKGAFGEVRLVQKKDTGKIYAMKTLLKSEMYNKDQLAHVK 349
Query: 162 SERNLLAEVDSNCIVKLYCSFQDDEYLYLIMEYLPGGDMMTLLMRREVLTENEAKFYIGE 221
+ER++LA DS +V LY SFQD +YLYLIME+LPGGD+MT+L+R ++ TE+ +FY+ E
Sbjct: 350 AERDVLAGSDSPWVVSLYYSFQDSQYLYLIMEFLPGGDLMTMLIRWQIFTEDVTRFYMAE 409
Query: 222 TVLAIESIHKHNYIHRDIKPDNLLLDRNGHVKLSDFGLC----KPLDCS----------- 266
+LAIE IHK +IHRDIKPDN+L+D GH+KLSDFGL K D S
Sbjct: 410 CILAIEVIHKLGFIHRDIKPDNILIDIRGHIKLSDFGLSTGFHKTHDSSYYKKLLQEDEA 469
Query: 267 -------------NLQEKDLSDGVNRSG--VLQSDELTLSPNQSQQQQLEHWRKNHSRML 311
NLQ+ L + N + D + L+ + +QQ++ WRK+ R++
Sbjct: 470 KKQQQQQQQQQQLNLQKPQLPNETNNGNRNTMLVDAIHLT--MTNRQQMQTWRKSR-RLM 526
Query: 312 AYSTVGTPDYIAPEVILKRGYGVECDWWSLGAIMYEMLVGYPPFHSDDPRTTCRKIAHWK 371
AYSTVGTPDYIAPE+ L +GYG ECDWWSLGAIMYE L+G+PPF S+ P+ T RKI +++
Sbjct: 527 AYSTVGTPDYIAPEIFLYQGYGQECDWWSLGAIMYECLIGWPPFCSETPQETYRKIMNFE 586
Query: 372 TYLKFPKD-KLSPEAKDLICRLLCNVKQRLGAK-GADEIKAHPWFKVVEWEKLYQMRAPF 429
L+FP D +S EA+DLI RLL + + RLG GADEIKAHP+F V+W + Q+ AP+
Sbjct: 587 QTLQFPDDIHISYEAEDLIRRLLTHSENRLGRHGGADEIKAHPFFSGVDWNTIRQVEAPY 646
Query: 430 IPKVNDELDTRNFDKFEEEDQKTEPS-PKAGPWRKMLQS-----------KDINFVGYTY 477
IPK++ DTR F E E+ P+ +A R+ + + +D+ F+GYTY
Sbjct: 647 IPKLSSVTDTRFFPTDELENVPDSPAMAQAARQREQMMTQQQGVPQSNAKEDLPFIGYTY 706
Query: 478 KNYE 481
++
Sbjct: 707 SRFD 710
>ORB6_SCHPO (O13310) Serine/threonine-protein kinase orb6 (EC
2.7.1.37)
Length = 469
Score = 387 bits (994), Expect = e-107
Identities = 199/443 (44%), Positives = 291/443 (64%), Gaps = 12/443 (2%)
Query: 45 KVEAAKHFVESYYKNQKQNQQERKERRNILENKLADAEVSKEDKKNLLKNFEEMETEIMR 104
KV+ K ++E YYK + ER +RR LE +LA S+E K L+ E E++ +R
Sbjct: 23 KVQKTKKYIEHYYKVAVDHAVERNQRRINLEQRLATERGSEERKNRQLRASGEKESQFLR 82
Query: 105 RQRLKMGADDFEPLTMIGKGAFGEVRICREKTTGQVYAMKKLQKSEMLRRGQVEYVKSER 164
+R ++ +DF + +IGKGAFGEVR+ ++ TG++YAMK L K+EM +R Q+ +VK+ER
Sbjct: 83 FRRTRLSLEDFSTIKVIGKGAFGEVRLVQKLDTGKIYAMKSLLKTEMFKRDQLAHVKAER 142
Query: 165 NLLAEVDSNCIVKLYCSFQDDEYLYLIMEYLPGGDMMTLLMRREVLTENEAKFYIGETVL 224
+LL E DS +V LY +FQD YLYLIME+LPGGD+MT+L+ + +E+ +FY+ E VL
Sbjct: 143 DLLVESDSPWVVSLYYAFQDSLYLYLIMEFLPGGDLMTMLINYDTFSEDVTRFYMAECVL 202
Query: 225 AIESIHKHNYIHRDIKPDNLLLDRNGHVKLSDFGLC----KPLDCSNLQEKDLSDGVNRS 280
AI +H+ YIHRDIKPDN+L+DR+GH+KLSDFGL K ++ + + V R
Sbjct: 203 AIADVHRMGYIHRDIKPDNILIDRDGHIKLSDFGLSTGFYKQDQSASYMKPRTGNTVKRG 262
Query: 281 GVLQSDELTLSPNQSQQQQLEHWRKNHSRMLAYSTVGTPDYIAPEVILKRGYGVECDWWS 340
++ + LT+ S + ++ W+KN R++AYSTVGTPDYIAPE+ L++GYG +CDWWS
Sbjct: 263 QMVDAIWLTM----SSKDKMATWKKNR-RVMAYSTVGTPDYIAPEIFLQQGYGQDCDWWS 317
Query: 341 LGAIMYEMLVGYPPFHSDDPRTTCRKIAHWKTYLKFPKD-KLSPEAKDLICRLLCNVKQR 399
LGAIM+E L+G+PPF S++ T RKI +W+ L FP D LS EA+DL+ RL+ + + R
Sbjct: 318 LGAIMFECLIGWPPFCSENSHETYRKIINWRETLTFPNDIHLSIEARDLMDRLMTDSEHR 377
Query: 400 LGAKGADEIKAHPWFKVVEWEKLYQMRAPFIPKVNDELDTRNF--DKFEEEDQKTEPSPK 457
LG GA EI HP+F ++W+ + + APFIP + DT F D+ E+ ++
Sbjct: 378 LGRGGAIEIMQHPFFTGIDWDHIRETAAPFIPNLKSITDTHYFPVDELEQVPEQPVTQQP 437
Query: 458 AGPWRKMLQSKDINFVGYTYKNY 480
A + L+ ++ F+GYTYK +
Sbjct: 438 ASVDPQTLEQTNLAFLGYTYKKF 460
>COT1_NEUCR (P38679) Serine/threonine-protein kinase cot-1 (EC
2.7.1.37) (Colonial temperature-sensitive 1)
Length = 598
Score = 356 bits (913), Expect = 1e-97
Identities = 191/463 (41%), Positives = 299/463 (64%), Gaps = 26/463 (5%)
Query: 37 PPSNETKLKVEAAKHFVESYYKNQKQNQQERKERRNILENKLADAEVSKEDKKNLLKNFE 96
P +N + K ++K+ + +ER +R++ +E KL + ++ ++++
Sbjct: 140 PNANNNQKK---CSQLASDFFKDSVKRARERNQRQSEMEQKLGETNDARR-RESIWSTAG 195
Query: 97 EMETEIMRRQRLKMGADDFEPLTMIGKGAFGEVRICREKTTGQVYAMKKLQKSEMLRRGQ 156
E + +R R K ++++ + +IGKGAFGEV++ ++K G+VYAMK L K+EM ++ Q
Sbjct: 196 RKEGQYLRFLRTKDKPENYQTIKIIGKGAFGEVKLVQKKADGKVYAMKSLIKTEMFKKDQ 255
Query: 157 VEYVKSERNLLAEVDSNCIVKLYCSFQDDEYLYLIMEYLPGGDMMTLLMRREVLTENEAK 216
+ +V++ER++LAE DS +VKLY +FQD +LY++ME+LPGGD+MT+L++ E+ +E+ +
Sbjct: 256 LAHVRAERDILAESDSPWVVKLYTTFQDANFLYMLMEFLPGGDLMTMLIKYEIFSEDITR 315
Query: 217 FYIGETVLAIESIHKHNYIHRDIKPDNLLLDRNGHVKLSDFGLC---KPLDCSN-----L 268
FYI E VLAI+++HK +IHRDIKPDN+LLDR GHVKL+DFGL L +N L
Sbjct: 316 FYIAEIVLAIDAVHKLGFIHRDIKPDNILLDRGGHVKLTDFGLSTGFHKLHDNNYYTQLL 375
Query: 269 QEKDLSDGVNRSGVLQSDELTLSPNQSQQQQLEHWRKNHSRMLAYSTVGTPDYIAPEVIL 328
Q K NR+ V D++ L+ S + Q+ WR++ R++AYSTVGTPDYIAPE+
Sbjct: 376 QGKSNKPRDNRNSV-AIDQINLT--VSNRAQINDWRRSR-RLMAYSTVGTPDYIAPEIFT 431
Query: 329 KRGYGVECDWWSLGAIMYEMLVGYPPFHSDDPRTTCRKIAHWKTYLKFPKD-KLSPEAKD 387
GY +CDWWSLG IM+E LVG+PPF ++D T RKI +W+ L FP D L +A++
Sbjct: 432 GHGYSFDCDWWSLGTIMFECLVGWPPFCAEDSHDTYRKIVNWRHSLYFPDDITLGVDAEN 491
Query: 388 LICRLLCNVKQRLGAKGADEIKAHPWFKVVEWEKLYQMRAPFIPKVNDELDTRNF--DKF 445
LI L+CN + RLG GA EIK+H +F+ VE++ L ++RAPF P++ +DT F D+
Sbjct: 492 LIRSLICNTENRLGRGGAHEIKSHAFFRGVEFDSLRRIRAPFEPRLTSAIDTTYFPTDEI 551
Query: 446 EEEDQKT-----EPSPKAGPWRKMLQSKDIN--FVGYTYKNYE 481
++ D T + + A + +S +++ F+GYT+K ++
Sbjct: 552 DQTDNATLLKAQQAARGAAAPAQQEESPELSLPFIGYTFKRFD 594
>LTS2_HUMAN (Q9NRM7) Serine/threonine protein kinase LATS2 (EC
2.7.1.37) (Large tumor suppressor homolog 2)
(Serine/threonine kinase kpm) (Kinase phosphorylated
during mitosis protein) (Warts-like kinase)
Length = 1088
Score = 346 bits (888), Expect = 8e-95
Identities = 189/452 (41%), Positives = 274/452 (59%), Gaps = 10/452 (2%)
Query: 48 AAKHFVESYYKNQKQNQQERKERRNILENKLADAEVSKEDKKNLLKNFEEMETEIMRRQR 107
A K F+E + +N + Q++ RR LE ++A A + + +++ + K + E+ R +R
Sbjct: 601 AFKFFMEQHVENVIKTYQQKVNRRLQLEQEMAKAGLCEAEQEQMRKILYQKESNYNRLKR 660
Query: 108 LKMGADDFEPLTMIGKGAFGEVRICREKTTGQVYAMKKLQKSEMLRRGQVEYVKSERNLL 167
KM F + +G GAFGEV + + T +YAMK L+K ++L R QV +VK+ER++L
Sbjct: 661 AKMDKSMFVKIKTLGIGAFGEVCLACKVDTHALYAMKTLRKKDVLNRNQVVHVKAERDIL 720
Query: 168 AEVDSNCIVKLYCSFQDDEYLYLIMEYLPGGDMMTLLMRREVLTENEAKFYIGETVLAIE 227
AE D+ +VKLY SFQD + LY +M+Y+PGGDMM+LL+R EV E+ A+FYI E LAIE
Sbjct: 721 AEADNEWVVKLYYSFQDKDSLYFVMDYIPGGDMMSLLIRMEVFPEHLARFYIAELTLAIE 780
Query: 228 SIHKHNYIHRDIKPDNLLLDRNGHVKLSDFGLCKPLDCS-NLQEKDLSDGVNRSGVLQSD 286
S+HK +IHRDIKPDN+L+D +GH+KL+DFGLC + N + V + + SD
Sbjct: 781 SVHKMGFIHRDIKPDNILIDLDGHIKLTDFGLCTGFRWTHNSKYYQKGSHVRQDSMEPSD 840
Query: 287 ELTLSPNQSQQQQL----EHWRKNHSRMLAYSTVGTPDYIAPEVILKRGYGVECDWWSLG 342
N +L + RK H R LA+S VGTP+YIAPEV+L++GY CDWWS+G
Sbjct: 841 LWDDVSNCRCGDRLKTLEQRARKQHQRCLAHSLVGTPNYIAPEVLLRKGYTQLCDWWSVG 900
Query: 343 AIMYEMLVGYPPFHSDDPRTTCRKIAHWKTYLKFPKD-KLSPEAKDLICRLLCNVKQRLG 401
I++EMLVG PPF + P T K+ +W+ L P KLSPEA+DLI +L C+ RLG
Sbjct: 901 VILFEMLVGQPPFLAPTPTETQLKVINWENTLHIPAQVKLSPEARDLITKLCCSADHRLG 960
Query: 402 AKGADEIKAHPWFKVVEWEK-LYQMRAPFIPKVNDELDTRNFDKFEEE---DQKTEPSPK 457
GAD++KAHP+F +++ + + AP++P ++ +DT NFD +EE + +E S K
Sbjct: 961 RNGADDLKAHPFFSAIDFSSDIRKQPAPYVPTISHPMDTSNFDPVDEESPWNDASEGSTK 1020
Query: 458 AGPWRKMLQSKDINFVGYTYKNYEMVDANAIP 489
A +K Y + D N P
Sbjct: 1021 AWDTLTSPNNKHPEHAFYEFTFRRFFDDNGYP 1052
>LTS1_HUMAN (O95835) Serine/threonine protein kinase LATS1 (EC
2.7.1.37) (Large tumor suppressor homolog 1) (WARTS
protein kinase) (h-warts)
Length = 1130
Score = 340 bits (871), Expect = 7e-93
Identities = 185/460 (40%), Positives = 274/460 (59%), Gaps = 23/460 (5%)
Query: 47 EAAKHFVESYYKNQKQNQQERKERRNILENKLADAEVSKEDKKNLLKNFEEMETEIMRRQ 106
+A K F+E + +N ++ Q+R R+ LEN++ +S++ + + K + E+ +R +
Sbjct: 637 QAFKFFMEQHVENVLKSHQQRLHRKKQLENEMMRVGLSQDAQDQMRKMLCQKESNYIRLK 696
Query: 107 RLKMGADDFEPLTMIGKGAFGEVRICREKTTGQVYAMKKLQKSEMLRRGQVEYVKSERNL 166
R KM F + +G GAFGEV + R+ T +YA K L+K ++L R QV +VK+ER++
Sbjct: 697 RAKMDKSMFVKIKTLGIGAFGEVCLARKVDTKALYATKTLRKKDVLLRNQVAHVKAERDI 756
Query: 167 LAEVDSNCIVKLYCSFQDDEYLYLIMEYLPGGDMMTLLMRREVLTENEAKFYIGETVLAI 226
LAE D+ +V+LY SFQD + LY +M+Y+PGGDMM+LL+R + E+ A+FYI E A+
Sbjct: 757 LAEADNEWVVRLYYSFQDKDNLYFVMDYIPGGDMMSLLIRMGIFPESLARFYIAELTCAV 816
Query: 227 ESIHKHNYIHRDIKPDNLLLDRNGHVKLSDFGLC---------KPLDCSNLQEKDLSDGV 277
ES+HK +IHRDIKPDN+L+DR+GH+KL+DFGLC K + +D D
Sbjct: 817 ESVHKMGFIHRDIKPDNILIDRDGHIKLTDFGLCTGFRWTHDSKYYQSGDHPRQDSMDFS 876
Query: 278 NRSGVLQSDELTLSPNQSQQQQLEHWRKNHSRMLAYSTVGTPDYIAPEVILKRGYGVECD 337
N G D + + + H R LA+S VGTP+YIAPEV+L+ GY CD
Sbjct: 877 NEWG----DPSSCRCGDRLKPLERRAARQHQRCLAHSLVGTPNYIAPEVLLRTGYTQLCD 932
Query: 338 WWSLGAIMYEMLVGYPPFHSDDPRTTCRKIAHWKTYLKF-PKDKLSPEAKDLICRLLCNV 396
WWS+G I++EMLVG PPF + P T K+ +W+T L P+ KLSPEA DLI +L
Sbjct: 933 WWSVGVILFEMLVGQPPFLAQTPLETQMKVINWQTSLHIPPQAKLSPEASDLIIKLCRGP 992
Query: 397 KQRLGAKGADEIKAHPWFKVVEWEK-LYQMRAPFIPKVNDELDTRNFDK------FEEED 449
+ RLG GADEIKAHP+FK +++ L Q A +IPK+ DT NFD + +++
Sbjct: 993 EDRLGKNGADEIKAHPFFKTIDFSSDLRQQSASYIPKITHPTDTSNFDPVDPDKLWSDDN 1052
Query: 450 QKTEPSPKAGPWRKMLQSKDINFVGYTYKNYEMVDANAIP 489
++ + W K + + F +T++ + D N P
Sbjct: 1053 EEENVNDTLNGWYKNGKHPEHAFYEFTFRRF--FDDNGYP 1090
>LTS2_MOUSE (Q7TSJ6) Serine/threonine protein kinase LATS2 (EC
2.7.1.37) (Large tumor suppressor homolog 2)
(Serine/threonine kinase kpm) (Kinase phosphorylated
during mitosis protein)
Length = 1042
Score = 339 bits (870), Expect = 9e-93
Identities = 189/462 (40%), Positives = 275/462 (58%), Gaps = 30/462 (6%)
Query: 48 AAKHFVESYYKNQKQNQQERKERRNILENKLADAEVSKEDKKNLLKNFEEMETEIMRRQR 107
A K F+E + +N + Q++ RR LE ++A A + + +++ + K + E+ R +R
Sbjct: 559 AFKFFMEQHVENVIKTYQQKVSRRLQLEQEMAKAGLCEAEQEQMRKILYQKESNYNRLKR 618
Query: 108 LKMGADDFEPLTMIGKGAFGEVRICREKTTGQVYAMKKLQKSEMLRRGQVEYVKSERNLL 167
KM F + +G GAFGEV + + T +YAMK L+K ++L R QV +VK+ER++L
Sbjct: 619 AKMDKSMFVKIKTLGIGAFGEVCLACKLDTHALYAMKTLRKKDVLNRNQVAHVKAERDIL 678
Query: 168 AEVDSNCIVKLYCSFQDDEYLYLIMEYLPGGDMMTLLMRREVLTENEAKFYIGETVLAIE 227
AE D+ +VKLY SFQD + LY +M+Y+PGGDMM+LL+R EV E+ A+FYI E LAIE
Sbjct: 679 AEADNEWVVKLYYSFQDKDSLYFVMDYIPGGDMMSLLIRMEVFPEHLARFYIAELTLAIE 738
Query: 228 SIHKHNYIHRDIKPDNLLLDRNGHVKLSDFGLC---------------KPLDCSNLQEKD 272
S+HK +IHRDIKPDN+L+D +GH+KL+DFGLC + +++ D
Sbjct: 739 SVHKMGFIHRDIKPDNILIDLDGHIKLTDFGLCTGFRWTHNSKYYQKGNHMRQDSMEPGD 798
Query: 273 LSDGVNRSGVLQSDELTLSPNQSQQQQLEHWRKNHSRMLAYSTVGTPDYIAPEVILKRGY 332
L D V S D L ++Q+Q H R LA+S VGTP+YIAPEV+L++GY
Sbjct: 799 LWDDV--SNCRCGDRLKTLEQRAQKQ--------HQRCLAHSLVGTPNYIAPEVLLRKGY 848
Query: 333 GVECDWWSLGAIMYEMLVGYPPFHSDDPRTTCRKIAHWKTYLKFPKD-KLSPEAKDLICR 391
CDWWS+G I++EMLVG PPF + P T K+ +W++ L P +LS EA+DLI +
Sbjct: 849 TQLCDWWSVGVILFEMLVGQPPFLAPTPTETQLKVINWESTLHIPTQVRLSAEARDLITK 908
Query: 392 LLCNVKQRLGAKGADEIKAHPWFKVVEWEK-LYQMRAPFIPKVNDELDTRNFDKFEEEDQ 450
L C RLG GAD++KAHP+F +++ + + + AP++P ++ +DT NFD +EE
Sbjct: 909 LCCAADCRLGRDGADDLKAHPFFNTIDFSRDIRKQPAPYVPTISHPMDTSNFDPVDEESP 968
Query: 451 KTEPS-PKAGPWRKML--QSKDINFVGYTYKNYEMVDANAIP 489
E S A W + SK Y + D N P
Sbjct: 969 WHEASGESAKAWDTLASPSSKHPEHAFYEFTFRRFFDDNGYP 1010
>LTS1_MOUSE (Q8BYR2) Serine/threonine protein kinase LATS1 (EC
2.7.1.37) (Large tumor suppressor homolog 1) (WARTS
protein kinase)
Length = 1129
Score = 337 bits (864), Expect = 5e-92
Identities = 187/460 (40%), Positives = 269/460 (57%), Gaps = 23/460 (5%)
Query: 47 EAAKHFVESYYKNQKQNQQERKERRNILENKLADAEVSKEDKKNLLKNFEEMETEIMRRQ 106
+A K F+E + +N ++ Q+R R+ LEN++ +S++ + + K + E+ +R +
Sbjct: 636 QAFKFFMEQHVENVLKSHQQRLHRKKQLENEMMRVGLSQDAQDQMRKMLCQKESNYIRLK 695
Query: 107 RLKMGADDFEPLTMIGKGAFGEVRICREKTTGQVYAMKKLQKSEMLRRGQVEYVKSERNL 166
R KM F + +G GAFGEV + R+ T +YA K L+K ++L R QV +VK+ER++
Sbjct: 696 RAKMDKSMFVKIKTLGIGAFGEVCLARKVDTKALYATKTLRKKDVLLRNQVAHVKAERDI 755
Query: 167 LAEVDSNCIVKLYCSFQDDEYLYLIMEYLPGGDMMTLLMRREVLTENEAKFYIGETVLAI 226
LAE D+ +V+LY SFQD + LY +M+Y+PGGDMM+LL+R + EN A+FYI E A+
Sbjct: 756 LAEADNEWVVRLYYSFQDKDNLYFVMDYIPGGDMMSLLIRMGIFPENLARFYIAELTCAV 815
Query: 227 ESIHKHNYIHRDIKPDNLLLDRNGHVKLSDFGLC---------KPLDCSNLQEKDLSDGV 277
ES+HK +IHRDIKPDN+L+DR+GH+KL+DFGLC K + +D D
Sbjct: 816 ESVHKMGFIHRDIKPDNILIDRDGHIKLTDFGLCTGFRWTHDSKYYQSGDHPRQDSMDFS 875
Query: 278 NRSGVLQSDELTLSPNQSQQQQLEHWRKNHSRMLAYSTVGTPDYIAPEVILKRGYGVECD 337
N G D + + H R LA+S VGTP+YIAPEV+L+ GY CD
Sbjct: 876 NEWG----DPSNCRCGDRLKPLERRAARQHQRCLAHSLVGTPNYIAPEVLLRTGYTQLCD 931
Query: 338 WWSLGAIMYEMLVGYPPFHSDDPRTTCRKIAHWKTYLKF-PKDKLSPEAKDLICRLLCNV 396
WWS+G I+ EMLVG PPF + P T K+ W+T L P+ KLSPEA DLI +L
Sbjct: 932 WWSVGVILCEMLVGQPPFLAQTPLETQMKVIIWQTSLHIPPQAKLSPEASDLIIKLCRGP 991
Query: 397 KQRLGAKGADEIKAHPWFKVVEWEK-LYQMRAPFIPKVNDELDTRNFDKFEEE------D 449
+ RLG GADEIKAHP+FK +++ L Q A +IPK+ DT NFD + +
Sbjct: 992 EDRLGKNGADEIKAHPFFKTIDFSSDLRQQSASYIPKITHPTDTSNFDPVDPDKLWSDGS 1051
Query: 450 QKTEPSPKAGPWRKMLQSKDINFVGYTYKNYEMVDANAIP 489
++ S W K + + F +T++ + D N P
Sbjct: 1052 EEENISDTLSGWYKNGKHPEHAFYEFTFRRF--FDDNGYP 1089
>ROC1_RAT (Q63644) Rho-associated protein kinase 1 (EC 2.7.1.37)
(Rho-associated, coiled-coil containing protein kinase
1) (p160 ROCK-1) (p160ROCK) (p150 RhoA-binding kinase
ROK beta)
Length = 1369
Score = 291 bits (746), Expect = 2e-78
Identities = 162/398 (40%), Positives = 236/398 (58%), Gaps = 57/398 (14%)
Query: 89 KNLLKNFEEMETEIMRRQRLKMGADDFEPLTMIGKGAFGEVRICREKTTGQVYAMKKLQK 148
KN+ + I + + L+M A+D+E + +IG+GAFGEV++ R K+T +VYAMK L K
Sbjct: 50 KNIDNFLSRYKDTINKIRDLRMKAEDYEVVKVIGRGAFGEVQLVRHKSTRKVYAMKLLSK 109
Query: 149 SEMLRRGQVEYVKSERNLLAEVDSNCIVKLYCSFQDDEYLYLIMEYLPGGDMMTLLMRRE 208
EM++R + ER+++A +S +V+L+ +FQDD YLY++MEY+PGGD++ L+ +
Sbjct: 110 FEMIKRSDSAFFWEERDIMAFANSPWVVQLFYAFQDDRYLYMVMEYMPGGDLVNLMSNYD 169
Query: 209 VLTENEAKFYIGETVLAIESIHKHNYIHRDIKPDNLLLDRNGHVKLSDFGLCKPLDCSNL 268
V E A+FY E VLA+++IH +IHRD+KPDN+LLD++GH+KL+DFG C +
Sbjct: 170 V-PEKWARFYTAEVVLALDAIHSMGFIHRDVKPDNMLLDKSGHLKLADFGTCMKM----- 223
Query: 269 QEKDLSDGVNRSGVLQSDELTLSPNQSQQQQLEHWRKNHSRMLAYSTVGTPDYIAPEVIL 328
N+ G+++ D + VGTPDYI+PEV+
Sbjct: 224 ---------NKEGMVRCD---------------------------TAVGTPDYISPEVLK 247
Query: 329 KRG----YGVECDWWSLGAIMYEMLVGYPPFHSDDPRTTCRKIAHWKTYLKFPKDK-LSP 383
+G YG ECDWWS+G +YEMLVG PF++D T KI + K L FP D +S
Sbjct: 248 SQGGDGYYGRECDWWSVGVFLYEMLVGDTPFYADSLVGTYSKIMNHKNSLTFPDDNDISK 307
Query: 384 EAKDLICRLLCNVKQRLGAKGADEIKAHPWFKVVE--WEKLYQMRAPFIPKVNDELDTRN 441
EAK+LIC L + + RLG G +EIK H +FK + WE L AP +P ++ ++DT N
Sbjct: 308 EAKNLICAFLTDREVRLGRNGVEEIKRHLFFKNDQWAWETLRDTVAPVVPDLSSDIDTSN 367
Query: 442 FDKFEEE--DQKTEPSPKAGPWRKMLQSKDINFVGYTY 477
FD EE+ D++T P PKA + FVG+TY
Sbjct: 368 FDDLEEDKGDEETFPIPKA------FVGNQLPFVGFTY 399
>ROC1_RABIT (O77819) Rho-associated protein kinase 1 (EC 2.7.1.37)
(Rho-associated, coiled-coil containing protein kinase
1) (p160 ROCK-1) (p160ROCK) (cAMP dependent protein
kinase ROCK-I) (CePKA) (Corneal epithelial
Rho-associated-Ser/Thr kinase 1) (HEBM
Length = 1354
Score = 291 bits (746), Expect = 2e-78
Identities = 162/398 (40%), Positives = 236/398 (58%), Gaps = 57/398 (14%)
Query: 89 KNLLKNFEEMETEIMRRQRLKMGADDFEPLTMIGKGAFGEVRICREKTTGQVYAMKKLQK 148
KN+ + I + + L+M A+D+E + +IG+GAFGEV++ R K+T +VYAMK L K
Sbjct: 50 KNIDNFLSRYKDTINKIRDLRMKAEDYEVVKVIGRGAFGEVQLVRHKSTRKVYAMKLLSK 109
Query: 149 SEMLRRGQVEYVKSERNLLAEVDSNCIVKLYCSFQDDEYLYLIMEYLPGGDMMTLLMRRE 208
EM++R + ER+++A +S +V+L+ +FQDD YLY++MEY+PGGD++ L+ +
Sbjct: 110 FEMIKRSDSAFFWEERDIMAFANSPWVVQLFYAFQDDRYLYMVMEYMPGGDLVNLMSNYD 169
Query: 209 VLTENEAKFYIGETVLAIESIHKHNYIHRDIKPDNLLLDRNGHVKLSDFGLCKPLDCSNL 268
V E A+FY E VLA+++IH +IHRD+KPDN+LLD++GH+KL+DFG C +
Sbjct: 170 V-PEKWARFYTAEVVLALDAIHSMGFIHRDVKPDNMLLDKSGHLKLADFGTCMKM----- 223
Query: 269 QEKDLSDGVNRSGVLQSDELTLSPNQSQQQQLEHWRKNHSRMLAYSTVGTPDYIAPEVIL 328
N+ G+++ D + VGTPDYI+PEV+
Sbjct: 224 ---------NKEGMVRCD---------------------------TAVGTPDYISPEVLK 247
Query: 329 KRG----YGVECDWWSLGAIMYEMLVGYPPFHSDDPRTTCRKIAHWKTYLKFPKDK-LSP 383
+G YG ECDWWS+G +YEMLVG PF++D T KI + K L FP D +S
Sbjct: 248 SQGGDGYYGRECDWWSVGVFLYEMLVGDTPFYADSLVGTYSKIMNHKNSLTFPDDNDISK 307
Query: 384 EAKDLICRLLCNVKQRLGAKGADEIKAHPWFKVVE--WEKLYQMRAPFIPKVNDELDTRN 441
EAK+LIC L + + RLG G +EIK H +FK + WE L AP +P ++ ++DT N
Sbjct: 308 EAKNLICAFLTDREVRLGRNGVEEIKRHLFFKNDQWAWETLRDTVAPVVPDLSSDIDTSN 367
Query: 442 FDKFEEE--DQKTEPSPKAGPWRKMLQSKDINFVGYTY 477
FD EE+ D++T P PKA + FVG+TY
Sbjct: 368 FDDLEEDKGDEETFPIPKA------FVGNQLPFVGFTY 399
>ROC1_MOUSE (P70335) Rho-associated protein kinase 1 (EC 2.7.1.37)
(Rho-associated, coiled-coil containing protein kinase
1) (p160 ROCK-1) (p160ROCK)
Length = 1354
Score = 291 bits (746), Expect = 2e-78
Identities = 162/398 (40%), Positives = 236/398 (58%), Gaps = 57/398 (14%)
Query: 89 KNLLKNFEEMETEIMRRQRLKMGADDFEPLTMIGKGAFGEVRICREKTTGQVYAMKKLQK 148
KN+ + I + + L+M A+D+E + +IG+GAFGEV++ R K+T +VYAMK L K
Sbjct: 50 KNIDNFLSRYKDTINKIRDLRMKAEDYEVVKVIGRGAFGEVQLVRHKSTRKVYAMKLLSK 109
Query: 149 SEMLRRGQVEYVKSERNLLAEVDSNCIVKLYCSFQDDEYLYLIMEYLPGGDMMTLLMRRE 208
EM++R + ER+++A +S +V+L+ +FQDD YLY++MEY+PGGD++ L+ +
Sbjct: 110 FEMIKRSDSAFFWEERDIMAFANSPWVVQLFYAFQDDRYLYMVMEYMPGGDLVNLMSNYD 169
Query: 209 VLTENEAKFYIGETVLAIESIHKHNYIHRDIKPDNLLLDRNGHVKLSDFGLCKPLDCSNL 268
V E A+FY E VLA+++IH +IHRD+KPDN+LLD++GH+KL+DFG C +
Sbjct: 170 V-PEKWARFYTAEVVLALDAIHSMGFIHRDVKPDNMLLDKSGHLKLADFGTCMKM----- 223
Query: 269 QEKDLSDGVNRSGVLQSDELTLSPNQSQQQQLEHWRKNHSRMLAYSTVGTPDYIAPEVIL 328
N+ G+++ D + VGTPDYI+PEV+
Sbjct: 224 ---------NKEGMVRCD---------------------------TAVGTPDYISPEVLK 247
Query: 329 KRG----YGVECDWWSLGAIMYEMLVGYPPFHSDDPRTTCRKIAHWKTYLKFPKDK-LSP 383
+G YG ECDWWS+G +YEMLVG PF++D T KI + K L FP D +S
Sbjct: 248 SQGGDGYYGRECDWWSVGVFLYEMLVGDTPFYADSLVGTYSKIMNHKNSLTFPDDNDISK 307
Query: 384 EAKDLICRLLCNVKQRLGAKGADEIKAHPWFKVVE--WEKLYQMRAPFIPKVNDELDTRN 441
EAK+LIC L + + RLG G +EIK H +FK + WE L AP +P ++ ++DT N
Sbjct: 308 EAKNLICAFLTDREVRLGRNGVEEIKRHLFFKNDQWAWETLRDTVAPVVPDLSSDIDTSN 367
Query: 442 FDKFEEE--DQKTEPSPKAGPWRKMLQSKDINFVGYTY 477
FD EE+ D++T P PKA + FVG+TY
Sbjct: 368 FDDLEEDKGDEETFPIPKA------FVGNQLPFVGFTY 399
>ROC1_HUMAN (Q13464) Rho-associated protein kinase 1 (EC 2.7.1.37)
(Rho-associated, coiled-coil containing protein kinase
1) (p160 ROCK-1) (p160ROCK)
Length = 1354
Score = 290 bits (743), Expect = 5e-78
Identities = 163/410 (39%), Positives = 241/410 (58%), Gaps = 57/410 (13%)
Query: 89 KNLLKNFEEMETEIMRRQRLKMGADDFEPLTMIGKGAFGEVRICREKTTGQVYAMKKLQK 148
KN+ + I + + L+M A+D+E + +IG+GAFGEV++ R K+T +VYAMK L K
Sbjct: 50 KNIDNFLSRYKDTINKIRDLRMKAEDYEVVKVIGRGAFGEVQLVRHKSTRKVYAMKLLSK 109
Query: 149 SEMLRRGQVEYVKSERNLLAEVDSNCIVKLYCSFQDDEYLYLIMEYLPGGDMMTLLMRRE 208
EM++R + ER+++A +S +V+L+ +FQDD YLY++MEY+PGGD++ L+ +
Sbjct: 110 FEMIKRSDSAFFWEERDIMAFANSPWVVQLFYAFQDDRYLYMVMEYMPGGDLVNLMSNYD 169
Query: 209 VLTENEAKFYIGETVLAIESIHKHNYIHRDIKPDNLLLDRNGHVKLSDFGLCKPLDCSNL 268
V E A+FY E VLA+++IH +IHRD+KPDN+LLD++GH+KL+DFG C +
Sbjct: 170 V-PEKWARFYTAEVVLALDAIHSMGFIHRDVKPDNMLLDKSGHLKLADFGTCMKM----- 223
Query: 269 QEKDLSDGVNRSGVLQSDELTLSPNQSQQQQLEHWRKNHSRMLAYSTVGTPDYIAPEVIL 328
N+ G+++ D + VGTPDYI+PEV+
Sbjct: 224 ---------NKEGMVRCD---------------------------TAVGTPDYISPEVLK 247
Query: 329 KRG----YGVECDWWSLGAIMYEMLVGYPPFHSDDPRTTCRKIAHWKTYLKFPKDK-LSP 383
+G YG ECDWWS+G +YEMLVG PF++D T KI + K L FP D +S
Sbjct: 248 SQGGDGYYGRECDWWSVGVFLYEMLVGDTPFYADSLVGTYSKIMNHKNSLTFPDDNDISK 307
Query: 384 EAKDLICRLLCNVKQRLGAKGADEIKAHPWFKVVE--WEKLYQMRAPFIPKVNDELDTRN 441
EAK+LIC L + + RLG G +EIK H +FK + WE L AP +P ++ ++DT N
Sbjct: 308 EAKNLICAFLTDREVRLGRNGVEEIKRHLFFKNDQWAWETLRDTVAPVVPDLSSDIDTSN 367
Query: 442 FDKFEEE--DQKTEPSPKAGPWRKMLQSKDINFVGYTYKNYEMVDANAIP 489
FD EE+ +++T P PKA + FVG+TY + ++A P
Sbjct: 368 FDDLEEDKGEEETFPIPKA------FVGNQLPFVGFTYYSNRRYLSSANP 411
>DMK_HUMAN (Q09013) Myotonin-protein kinase (EC 2.7.1.-) (Myotonic
dystrophy protein kinase) (MDPK) (DM-kinase) (DMK)
(DMPK) (MT-PK)
Length = 639
Score = 267 bits (683), Expect = 5e-71
Identities = 157/407 (38%), Positives = 221/407 (53%), Gaps = 59/407 (14%)
Query: 99 ETEIMRRQRLKMGADDFEPLTMIGKGAFGEVRICREKTTGQVYAMKKLQKSEMLRRGQVE 158
E ++R + +++ DDFE L +IG+GAF EV + + K TGQVYAMK + K +ML+RG+V
Sbjct: 65 EPIVVRLKEVRLQRDDFEILKVIGRGAFSEVAVVKMKQTGQVYAMKIMNKWDMLKRGEVS 124
Query: 159 YVKSERNLLAEVDSNCIVKLYCSFQDDEYLYLIMEYLPGGDMMTLLMR-REVLTENEAKF 217
+ ER++L D I +L+ +FQD+ YLYL+MEY GGD++TLL + E + A+F
Sbjct: 125 CFREERDVLVNGDRRWITQLHFAFQDENYLYLVMEYYVGGDLLTLLSKFGERIPAEMARF 184
Query: 218 YIGETVLAIESIHKHNYIHRDIKPDNLLLDRNGHVKLSDFGLCKPLDCSNLQEKDLSDGV 277
Y+ E V+AI+S+H+ Y+HRDIKPDN+LLDR GH++L+DFG C L +DG
Sbjct: 185 YLAEIVMAIDSVHRLGYVHRDIKPDNILLDRCGHIRLADFGSCLKL---------RADGT 235
Query: 278 NRSGVLQSDELTLSPNQSQQQQLEHWRKNHSRMLAYSTVGTPDYIAPEVILKRG------ 331
RS V VGTPDY++PE++ G
Sbjct: 236 VRSLV--------------------------------AVGTPDYLSPEILQAVGGGPGTG 263
Query: 332 -YGVECDWWSLGAIMYEMLVGYPPFHSDDPRTTCRKIAHWKTYLKFP--KDKLSPEAKDL 388
YG ECDWW+LG YEM G PF++D T KI H+K +L P + + EA+D
Sbjct: 264 SYGPECDWWALGVFAYEMFYGQTPFYADSTAETYGKIVHYKEHLSLPLVDEGVPEEARDF 323
Query: 389 ICRLLCNVKQRLGAKGADEIKAHPWFKVVEWEKLYQMRAPFIPKVNDELDTRNFDKFEEE 448
I RLLC + RLG GA + + HP+F ++W+ L PF P DT NFD E
Sbjct: 324 IQRLLCPPETRLGRGGAGDFRTHPFFFGLDWDGLRDSVPPFTPDFEGATDTCNFDLV--E 381
Query: 449 DQKTEPSPKAGPWRKMLQ-----SKDINFVGYTYKNYEMVDANAIPG 490
D T G ++ + FVGY+Y + D+ +PG
Sbjct: 382 DGLTAMVSGGGETLSDIREGAPLGVHLPFVGYSYSCMALRDSE-VPG 427
>SID2_SCHPO (Q09898) Serine/threonine-protein kinase sid2 (EC
2.7.1.37)
Length = 607
Score = 246 bits (629), Expect = 8e-65
Identities = 140/446 (31%), Positives = 241/446 (53%), Gaps = 29/446 (6%)
Query: 52 FVESYYKNQKQNQQERKERRNILENKLADAEVSKEDKKNLLKNFEEMETEIMRRQRLKMG 111
F +Y Q RK+R + E +L S+ D+ L+K + E +R++R ++
Sbjct: 147 FFLDHYFEQLHYLYTRKQRARLFEEQLLKEPDSRRDE--LVKRYNGRERVYLRKRRTRIS 204
Query: 112 ADDFEPLTMIGKGAFGEVRICREKTTGQVYAMKKLQKSEMLRRGQVEYVKSERNLLAEVD 171
DF+ +T +G+G +G V + R++ T ++ A+K + KS + + ++ +V +ER++L +
Sbjct: 205 HGDFQTITQVGQGGYGSVWLARKRDTKEIVALKIMNKSVLHKMDEIRHVLTERDILTTAN 264
Query: 172 SNCIVKLYCSFQDDEYLYLIMEYLPGGDMMTLLMRREVLTENEAKFYIGETVLAIESIHK 231
S +V+L +FQD +YL ME++PGGD TLL VL ++ AKFY E LAI+++H+
Sbjct: 265 SEWLVRLLYAFQDTSNIYLAMEFVPGGDFRTLLSNSGVLRDHHAKFYATEMFLAIDALHQ 324
Query: 232 HNYIHRDIKPDNLLLDRNGHVKLSDFGLCKPLDCSNLQEKDLSDGVNRSGVLQSDELTLS 291
YIHRD+KP+N L+ +GH+KL+DFGL + + +K + R LQ +
Sbjct: 325 LGYIHRDLKPENFLVGASGHIKLTDFGLSSGI----ISKKKIESMKIR---LQEVNNVVV 377
Query: 292 PNQSQQQQLEHWRK--NHSRMLAYSTVGTPDYIAPEVILKRGYGVECDWWSLGAIMYEML 349
P +S +++ + +R + + A+S VG+PDY+APEV+ Y D+WSLG IMYE L
Sbjct: 378 PERSMRERRQVFRTLLSQDPVYAHSVVGSPDYMAPEVLRGENYNHSVDYWSLGCIMYECL 437
Query: 350 VGYPPFHSDDPRTTCRKIAHWKTYLKFP--------KDKLSPEAKDLICRLLCNVKQRLG 401
G+PPF + T + +W+ + P + +A D +C + + K R
Sbjct: 438 SGFPPFSGSNVNETWSNLKNWRKCFQRPHYDDPRDLEFNWRDDAWDFVCHCITDPKDRFC 497
Query: 402 AKGADEIKAHPWFKVVEWEKL-YQMRAPFIPKVNDELDTRNFDKFEEEDQKT---EPSPK 457
+ ++ HP+F ++W+ + R PF+P +N E+D FD F E+ + E K
Sbjct: 498 S--LKQVMQHPYFSKIDWKNVRTAYRPPFVPDLNSEIDAGYFDDFTNENDMSKYKEVHEK 555
Query: 458 AGPWRKMLQS----KDINFVGYTYKN 479
M+ + K F+G+T+++
Sbjct: 556 QAAIANMVNTFNKPKRNAFIGFTFRH 581
>DBFB_YEAST (P32328) Serine/threonine-protein kinase DBF20 (EC
2.7.1.37)
Length = 564
Score = 233 bits (594), Expect = 1e-60
Identities = 139/460 (30%), Positives = 239/460 (51%), Gaps = 18/460 (3%)
Query: 34 NEEPPSNETKLKVEAAKHFVESYYKNQKQ----NQQERKERRNILENKLADAEVSKEDKK 89
+E ++T+ V + + YY + +Q K+ LE + + VS +
Sbjct: 84 HERASQSKTQRVVNVCQLYFLDYYCDMFDYVISRRQRTKQVLRYLEQQRSVKNVSNKVLN 143
Query: 90 NLLKNFEEMETEIMRRQRLKMGADDFEPLTMIGKGAFGEVRICREKTTGQVYAMKKLQKS 149
+ + E E++R++RLK DF+ LT +G+G +G+V + ++K + ++ A+K L K
Sbjct: 144 EEWALYLQREHEVLRKRRLKPKHKDFQILTQVGQGGYGQVYLAKKKDSDEICALKILNKK 203
Query: 150 EMLRRGQVEYVKSERNLLAEVDSNCIVKLYCSFQDDEYLYLIMEYLPGGDMMTLLMRREV 209
+ + + +V +ER++L S+ +VKL +FQD E LYL ME++PGGD TLL+ +
Sbjct: 204 LLFKLNETNHVLTERDILTTTRSDWLVKLLYAFQDPESLYLAMEFVPGGDFRTLLINTRI 263
Query: 210 LTENEAKFYIGETVLAIESIHKHNYIHRDIKPDNLLLDRNGHVKLSDFGLCKPLDCSNLQ 269
L A+FYI E A+ ++H+ Y HRD+KP+N L+D GH+KL+DFGL SN +
Sbjct: 264 LKSGHARFYISEMFCAVNALHELGYTHRDLKPENFLIDATGHIKLTDFGLAAG-TVSNER 322
Query: 270 EKDLSDGVNRSGVLQSDELTLSPNQSQQQQLEHWRKNHSRMLAYSTVGTPDYIAPEVILK 329
+ + + L+ T + +++ + RK A S VG+PDY+A EV+
Sbjct: 323 IESMKIRLEEVKNLEFPAFTERSIEDRRKIYHNMRKTEIN-YANSMVGSPDYMALEVLEG 381
Query: 330 RGYGVECDWWSLGAIMYEMLVGYPPFHSDDPRTTCRKIAHWKTYLKFPKDK-----LSPE 384
+ Y D+WSLG +++E LVGY PF T + +WK L+ P+ + S
Sbjct: 382 KKYDFTVDYWSLGCMLFESLVGYTPFSGSSTNETYENLRYWKKTLRRPRTEDRRAAFSDR 441
Query: 385 AKDLICRLLCNVKQRLGAKGADEIKAHPWFKVVEWEKLYQMRAPFIPKVNDELDTRNFDK 444
DLI RL+ + R+ + ++++ +F + +E L PFIP+++DE D FD
Sbjct: 442 TWDLITRLIADPINRV--RSFEQVRKMSYFAEINFETLRTSSPPFIPQLDDETDAGYFDD 499
Query: 445 FEEEDQKTEPSPKAGPWRKML-----QSKDINFVGYTYKN 479
F E+ + + K+ + D VG+T+++
Sbjct: 500 FTNEEDMAKYADVFKRQNKLSAMVDDSAVDSKLVGFTFRH 539
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.319 0.135 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 64,953,400
Number of Sequences: 164201
Number of extensions: 2856207
Number of successful extensions: 14784
Number of sequences better than 10.0: 1702
Number of HSP's better than 10.0 without gapping: 1456
Number of HSP's successfully gapped in prelim test: 250
Number of HSP's that attempted gapping in prelim test: 9862
Number of HSP's gapped (non-prelim): 3364
length of query: 542
length of database: 59,974,054
effective HSP length: 115
effective length of query: 427
effective length of database: 41,090,939
effective search space: 17545830953
effective search space used: 17545830953
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 68 (30.8 bits)
Medicago: description of AC142507.1


