

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC142094.8 - phase: 0 /pseudo
(82 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
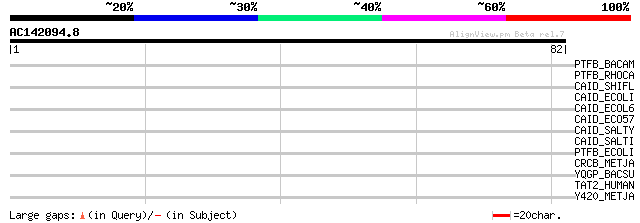
Score E
Sequences producing significant alignments: (bits) Value
PTFB_BACAM (P41029) PTS system, fructose-specific IIBC component... 39 0.002
PTFB_RHOCA (P23387) PTS system, fructose-specific IIBC component... 38 0.004
CAID_SHIFL (P59395) Carnitinyl-CoA dehydratase (EC 4.2.1.-) (Cro... 31 0.65
CAID_ECOLI (P31551) Carnitinyl-CoA dehydratase (EC 4.2.1.-) (Cro... 31 0.65
CAID_ECOL6 (Q8FLA6) Carnitinyl-CoA dehydratase (EC 4.2.1.-) (Cro... 31 0.65
CAID_ECO57 (Q8XA35) Carnitinyl-CoA dehydratase (EC 4.2.1.-) (Cro... 31 0.65
CAID_SALTY (Q8ZRX5) Carnitinyl-CoA dehydratase (EC 4.2.1.-) (Cro... 30 1.4
CAID_SALTI (Q8Z9L5) Carnitinyl-CoA dehydratase (EC 4.2.1.-) (Cro... 30 1.4
PTFB_ECOLI (P20966) PTS system, fructose-specific IIBC component... 29 1.9
CRCB_METJA (Q58918) Protein crcB homolog 28 4.2
YQGP_BACSU (P54493) Hypothetical protein yqgP 27 7.2
TAT2_HUMAN (Q93075) TatD DNase domain containing deoxyribonuclea... 27 7.2
Y420_METJA (Q57863) Hypothetical protein MJ0420 27 9.3
>PTFB_BACAM (P41029) PTS system, fructose-specific IIBC component
(EIIBC-Fru) (Fructose-permease IIBC component)
(Phosphotransferase enzyme II, BC component) (EC
2.7.1.69) (EII-Fru) (Fragment)
Length = 304
Score = 38.9 bits (89), Expect = 0.002
Identities = 24/60 (40%), Positives = 32/60 (53%), Gaps = 1/60 (1%)
Query: 17 GIVPIVNNFGLIGGLIPGFLLGFVLLCKKDPFVLPDQKLHKRCLPIICFILLSTGWTGVL 76
G + N G +GGLI GFL G+V++ K FV Q L P++ + LL TGVL
Sbjct: 55 GFMATQANAGFLGGLIAGFLAGYVVILLKKLFVFIPQSL-DGLKPVLIYPLLGIFITGVL 113
>PTFB_RHOCA (P23387) PTS system, fructose-specific IIBC component
(EIIBC-Fru) (Fructose-permease IIBC component)
(Phosphotransferase enzyme II, BC component) (EC
2.7.1.69) (EII-Fru)
Length = 578
Score = 38.1 bits (87), Expect = 0.004
Identities = 23/70 (32%), Positives = 38/70 (53%), Gaps = 4/70 (5%)
Query: 7 GMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCKKDPFVLPDQKLHKRCLPIICFI 66
G+ P L G++ + N G +GG++ GFL G+V +D LP + + P++
Sbjct: 313 GLTP--GLIGGMLAVNLNAGFLGGIVAGFLAGYVARWLRDAIKLP--RTLEGLKPVLILP 368
Query: 67 LLSTGWTGVL 76
LLST TG++
Sbjct: 369 LLSTAITGLI 378
>CAID_SHIFL (P59395) Carnitinyl-CoA dehydratase (EC 4.2.1.-)
(Crotonobetainyl-CoA hydratase)
Length = 260
Score = 30.8 bits (68), Expect = 0.65
Identities = 14/49 (28%), Positives = 22/49 (44%)
Query: 7 GMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCKKDPFVLPDQKL 55
G+ +FNL ++ VN + GG F++ F LP+ KL
Sbjct: 85 GLTEIFNLDKPVIAAVNGYAFGGGFELALAADFIVCADNASFALPEAKL 133
>CAID_ECOLI (P31551) Carnitinyl-CoA dehydratase (EC 4.2.1.-)
(Crotonobetainyl-CoA hydratase)
Length = 260
Score = 30.8 bits (68), Expect = 0.65
Identities = 14/49 (28%), Positives = 22/49 (44%)
Query: 7 GMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCKKDPFVLPDQKL 55
G+ +FNL ++ VN + GG F++ F LP+ KL
Sbjct: 85 GLTEIFNLDKPVIAAVNGYAFGGGFELALAADFIVCADNASFALPEAKL 133
>CAID_ECOL6 (Q8FLA6) Carnitinyl-CoA dehydratase (EC 4.2.1.-)
(Crotonobetainyl-CoA hydratase)
Length = 260
Score = 30.8 bits (68), Expect = 0.65
Identities = 14/49 (28%), Positives = 22/49 (44%)
Query: 7 GMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCKKDPFVLPDQKL 55
G+ +FNL ++ VN + GG F++ F LP+ KL
Sbjct: 85 GLTEIFNLDKPVIAAVNGYAFGGGFELALAADFIVCADNASFALPEAKL 133
>CAID_ECO57 (Q8XA35) Carnitinyl-CoA dehydratase (EC 4.2.1.-)
(Crotonobetainyl-CoA hydratase)
Length = 260
Score = 30.8 bits (68), Expect = 0.65
Identities = 14/49 (28%), Positives = 22/49 (44%)
Query: 7 GMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCKKDPFVLPDQKL 55
G+ +FNL ++ VN + GG F++ F LP+ KL
Sbjct: 85 GLTEIFNLDKPVIAAVNGYAFGGGFELALAADFIVCADNASFALPEAKL 133
>CAID_SALTY (Q8ZRX5) Carnitinyl-CoA dehydratase (EC 4.2.1.-)
(Crotonobetainyl-CoA hydratase)
Length = 260
Score = 29.6 bits (65), Expect = 1.4
Identities = 13/49 (26%), Positives = 23/49 (46%)
Query: 7 GMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCKKDPFVLPDQKL 55
G+ +F+L ++ VN + GG F++ + F LP+ KL
Sbjct: 85 GLTEIFDLDKPVIAAVNGYAFGGGFELALAADFIVCAENASFALPEAKL 133
>CAID_SALTI (Q8Z9L5) Carnitinyl-CoA dehydratase (EC 4.2.1.-)
(Crotonobetainyl-CoA hydratase)
Length = 260
Score = 29.6 bits (65), Expect = 1.4
Identities = 13/49 (26%), Positives = 23/49 (46%)
Query: 7 GMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCKKDPFVLPDQKL 55
G+ +F+L ++ VN + GG F++ + F LP+ KL
Sbjct: 85 GLTEIFDLDKPVIAAVNGYAFGGGFELALAADFIVCAENASFALPEAKL 133
>PTFB_ECOLI (P20966) PTS system, fructose-specific IIBC component
(EIIBC-Fru) (Fructose-permease IIBC component)
(Phosphotransferase enzyme II, BC component) (EC
2.7.1.69) (EII-Fru)
Length = 563
Score = 29.3 bits (64), Expect = 1.9
Identities = 19/64 (29%), Positives = 30/64 (46%), Gaps = 4/64 (6%)
Query: 7 GMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCKKDPFVLPDQKLHKRCLPIICFI 66
G+ P L G++ + G IGG+I GFL G++ LP + PI+
Sbjct: 298 GLTP--GLIGGMLAVSTGSGFIGGIIAGFLAGYIAKLISTQLKLPQSM--EALKPILIIP 353
Query: 67 LLST 70
L+S+
Sbjct: 354 LISS 357
>CRCB_METJA (Q58918) Protein crcB homolog
Length = 124
Score = 28.1 bits (61), Expect = 4.2
Identities = 24/69 (34%), Positives = 31/69 (44%), Gaps = 18/69 (26%)
Query: 14 LTIGIVPIVNNFGLIGG-----LIPGFLLGFVLLCKKDPFVLPDQKLHKRCLPIICFILL 68
L GIVP+ FGL G LI F+LGF+L C + + KL +
Sbjct: 21 LISGIVPV--KFGLPTGTLAVNLIGSFILGFLLYCSLFAPIPTEYKL-----------FI 67
Query: 69 STGWTGVLT 77
TG+ G LT
Sbjct: 68 GTGFCGALT 76
>YQGP_BACSU (P54493) Hypothetical protein yqgP
Length = 507
Score = 27.3 bits (59), Expect = 7.2
Identities = 27/79 (34%), Positives = 37/79 (46%), Gaps = 13/79 (16%)
Query: 1 MFNLSIG----MVPVFNLTIGI-VPIVNNFGLIGGLIPGFLLGFVL-LCKKDPFVLPDQK 54
MF +IG ++ + NL G V ++N G IGGLI GF L L K F
Sbjct: 308 MFLRTIGTNIIVIIIINLGFGFAVSNIDNSGHIGGLIGGFFAAAALGLPKAGAF------ 361
Query: 55 LHKRCLPIICFILLSTGWT 73
KR L + I L+ G++
Sbjct: 362 -GKRLLSAVFLIALAVGFS 379
>TAT2_HUMAN (Q93075) TatD DNase domain containing deoxyribonuclease
2 (EC 3.1.21.-)
Length = 761
Score = 27.3 bits (59), Expect = 7.2
Identities = 17/42 (40%), Positives = 22/42 (51%), Gaps = 9/42 (21%)
Query: 48 FVLPDQKLHKRCL----PII-----CFILLSTGWTGVLTKGS 80
FV PD K+H+ C P+I F +S G+T VLT S
Sbjct: 645 FVPPDYKIHRHCFTGSYPVIEPLLKYFPNMSVGFTAVLTYSS 686
>Y420_METJA (Q57863) Hypothetical protein MJ0420
Length = 380
Score = 26.9 bits (58), Expect = 9.3
Identities = 22/69 (31%), Positives = 35/69 (49%), Gaps = 13/69 (18%)
Query: 6 IGMVPVFNLTIGIVPIVNNFGL-----IGGLIPGFLLGFVLLCKKDPFVLPDQKLHKRCL 60
IG + +F L P +++ GL I G++ F++GF L PF+ P Q R +
Sbjct: 24 IGHLMIFILAF---PYLDDIGLYSLFKILGVLSIFIVGFFL-----PFIFPMQIKFNRHI 75
Query: 61 PIICFILLS 69
+ F+LLS
Sbjct: 76 LYLSFLLLS 84
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.331 0.154 0.501
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,410,157
Number of Sequences: 164201
Number of extensions: 408916
Number of successful extensions: 1022
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 9
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 1013
Number of HSP's gapped (non-prelim): 13
length of query: 82
length of database: 59,974,054
effective HSP length: 58
effective length of query: 24
effective length of database: 50,450,396
effective search space: 1210809504
effective search space used: 1210809504
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC142094.8


