

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
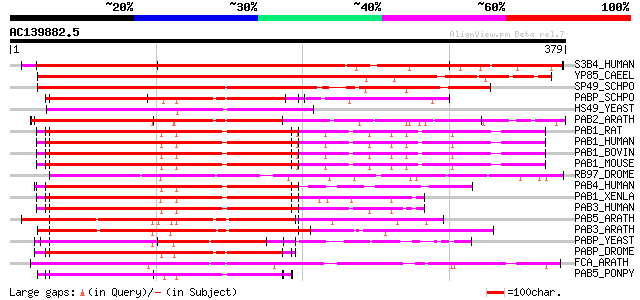
Query= AC139882.5 - phase: 0
(379 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
S3B4_HUMAN (Q15427) Splicing factor 3B subunit 4 (Spliceosome as... 400 e-111
YP85_CAEEL (Q09442) Hypothetical RNA-binding protein C08B11.5 in... 388 e-107
SP49_SCHPO (O14102) Spliceosome-associated protein 49 264 2e-70
PABP_SCHPO (P31209) Polyadenylate-binding protein (Poly(A)-bindi... 132 2e-30
HS49_YEAST (Q99181) HSH49 protein 131 3e-30
PAB2_ARATH (P42731) Polyadenylate-binding protein 2 (Poly(A)-bin... 129 1e-29
PAB1_RAT (Q9EPH8) Polyadenylate-binding protein 1 (Poly(A)-bindi... 127 4e-29
PAB1_HUMAN (P11940) Polyadenylate-binding protein 1 (Poly(A)-bin... 127 4e-29
PAB1_BOVIN (P61286) Polyadenylate-binding protein 1 (Poly(A)-bin... 127 4e-29
PAB1_MOUSE (P29341) Polyadenylate-binding protein 1 (Poly(A)-bin... 127 5e-29
RB97_DROME (Q02926) Ribonucleoprotein RB97D 126 1e-28
PAB4_HUMAN (Q13310) Polyadenylate-binding protein 4 (Poly(A)-bin... 124 3e-28
PAB1_XENLA (P20965) Polyadenylate-binding protein 1 (Poly(A)-bin... 122 1e-27
PAB3_HUMAN (Q9H361) Polyadenylate-binding protein 3 (Poly(A)-bin... 120 7e-27
PAB5_ARATH (Q05196) Polyadenylate-binding protein 5 (Poly(A)-bin... 119 2e-26
PAB3_ARATH (O64380) Polyadenylate-binding protein 3 (Poly(A)-bin... 117 4e-26
PABP_YEAST (P04147) Polyadenylate-binding protein, cytoplasmic a... 117 5e-26
PABP_DROME (P21187) Polyadenylate-binding protein (Poly(A)-bindi... 114 3e-25
FCA_ARATH (O04425) Flowering time control protein FCA 114 3e-25
PAB5_PONPY (P60050) Polyadenylate-binding protein 5 (Poly(A)-bin... 113 7e-25
>S3B4_HUMAN (Q15427) Splicing factor 3B subunit 4 (Spliceosome
associated protein 49) (SAP 49) (SF3b50) (Pre-mRNA
splicing factor SF3b 49 kDa subunit)
Length = 424
Score = 400 bits (1027), Expect = e-111
Identities = 221/389 (56%), Positives = 251/389 (63%), Gaps = 36/389 (9%)
Query: 19 AERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEED 78
+ERNQDAT YVG LD ++SE LLWELF+QAGPVVN ++PKDRVT QHQGYGFVEF SEED
Sbjct: 7 SERNQDATVYVGGLDEKVSEPLLWELFLQAGPVVNTHMPKDRVTGQHQGYGFVEFLSEED 66
Query: 79 ADYAIKVLNMIKLYGKPIRVNKASQDKKSLDVGANLFIGNLDPDVDEKLLYDTFSAFGVI 138
ADYAIK++NMIKLYGKPIRVNKAS K+LDVGAN+FIGNLDP++DEKLLYDTFSAFGVI
Sbjct: 67 ADYAIKIMNMIKLYGKPIRVNKASAHNKNLDVGANIFIGNLDPEIDEKLLYDTFSAFGVI 126
Query: 139 VTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAYKKDTKGER 198
+ PKIMRDPDTGNS+G+ FI++ SF+ASD+AIEAMNGQYLCNR ITVSYA+KKD+KGER
Sbjct: 127 LQTPKIMRDPDTGNSKGYAFINFASFDASDAAIEAMNGQYLCNRPITVSYAFKKDSKGER 186
Query: 199 HGTPAERVLAASNPTAQKSRPHTLFASGPPSLPNAPQ----------------ANGTIPA 242
HG+ AER+LAA NP +Q RPH LFA PP P+AP G+ P
Sbjct: 187 HGSAAERLLAAQNPLSQADRPHQLFADAPPP-PSAPNPVVSSLGSGLPPPGMPPPGSFPP 245
Query: 243 PVPPRPFANGVAPPPIHVIQPPPPQAPAF----QPMQMPPPGQQVWHQQQQGQPMMQQGM 298
PVPP G PP I PPPP P P P H P GM
Sbjct: 246 PVPP----PGALPPGIPPAMPPPPMPPGAAGHGPPSAGTPGAGHPGHGHSHPHPFPPGGM 301
Query: 299 P-PPMQQFR----PPHSMQMP-PPPPPQGMQGPPRPLPPPSVMAGQQQVWRPP--PPPQQ 350
P P M Q + PH + P PP G Q PPRP PP G + PP PP
Sbjct: 302 PHPGMSQMQLAHHGPHGLGHPHAGPPGSGGQPPPRP-PPGMPHPGPPPMGMPPRGPPFGS 360
Query: 351 QGGHPMGYPQNSMPPPPP--PNHHGMRPP 377
GHP P + M PPP P H PP
Sbjct: 361 PMGHPGPMPPHGMRGPPPLMPPHGYTGPP 389
Score = 56.6 bits (135), Expect = 1e-07
Identities = 30/94 (31%), Positives = 51/94 (53%), Gaps = 1/94 (1%)
Query: 9 VGANLLGQHSAERNQDATAYVGNLDPQISEELLWELFVQAGPVVNV-YVPKDRVTNQHQG 67
+ N H+ + A ++GNLDP+I E+LL++ F G ++ + +D T +G
Sbjct: 84 IRVNKASAHNKNLDVGANIFIGNLDPEIDEKLLYDTFSAFGVILQTPKIMRDPDTGNSKG 143
Query: 68 YGFVEFRSEEDADYAIKVLNMIKLYGKPIRVNKA 101
Y F+ F S + +D AI+ +N L +PI V+ A
Sbjct: 144 YAFINFASFDASDAAIEAMNGQYLCNRPITVSYA 177
Score = 49.3 bits (116), Expect = 2e-05
Identities = 30/87 (34%), Positives = 33/87 (37%), Gaps = 18/87 (20%)
Query: 301 PMQQFRPPHSMQMPPPPPPQ---------GMQGPPRPLPPPSVMAGQQQVWRPPPPPQQQ 351
P+ Q PH + PPPP G PP +PPP PPP
Sbjct: 200 PLSQADRPHQLFADAPPPPSAPNPVVSSLGSGLPPPGMPPPGSF---------PPPVPPP 250
Query: 352 GGHPMGYPQNSMPPPPPPNHHGMRPPS 378
G P G P PPP PP G PPS
Sbjct: 251 GALPPGIPPAMPPPPMPPGAAGHGPPS 277
>YP85_CAEEL (Q09442) Hypothetical RNA-binding protein C08B11.5 in
chromosome II
Length = 388
Score = 388 bits (996), Expect = e-107
Identities = 211/391 (53%), Positives = 252/391 (63%), Gaps = 50/391 (12%)
Query: 20 ERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDA 79
ERNQDAT YVG LD ++SE +LWEL VQAGPVV+V +PKDRVT HQG+GFVEF EEDA
Sbjct: 8 ERNQDATIYVGGLDEKVSESILWELMVQAGPVVSVNMPKDRVTANHQGFGFVEFMGEEDA 67
Query: 80 DYAIKVLNMIKLYGKPIRVNKASQDKKSLDVGANLFIGNLDPDVDEKLLYDTFSAFGVIV 139
DYAIK+LNMIKLYGKPI+VNKAS +K++DVGAN+F+GNLDP+VDEKLLYDTFSAFGVI+
Sbjct: 68 DYAIKILNMIKLYGKPIKVNKASAHEKNMDVGANIFVGNLDPEVDEKLLYDTFSAFGVIL 127
Query: 140 TNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAYKKDTKGERH 199
PKIMRD D+G S+GF FI++ SFEASD+A+EAMNGQ+LCNR ITVSYA+K+D+KGERH
Sbjct: 128 QVPKIMRDVDSGTSKGFAFINFASFEASDTALEAMNGQFLCNRAITVSYAFKRDSKGERH 187
Query: 200 GTPAERVLAASNPTAQKSRPHTLFASGPPSLP-NAPQANGTIPA---------------- 242
GT AER+LAA NP K RPH +F+ P +P N P A + A
Sbjct: 188 GTAAERMLAAQNPLFPKDRPHQVFSDVPLGVPANTPLAMPGVHAAIAAHATGRPGYQPPP 247
Query: 243 --------------PVPPRPFANGVAPPPIHVI------QPPPPQAPAFQPMQMPPPGQQ 282
PVPP P + PPP+ PPPP + + P PPPG+
Sbjct: 248 LMGMAQSGYQGQYPPVPPPPPSVTPMPPPMPPTPGMTPRPPPPPSSGMWPPPPPPPPGRT 307
Query: 283 VWHQQQQGQPMMQQGMPPPMQQFRPPHSMQMPPPPPPQGMQ---GPPRPLPPPSVMAGQQ 339
G P PPP +F PP MPPPPPP GM+ G P P PP AG
Sbjct: 308 PGPPGMPGMP-----PPPPPSRFGPPGMGGMPPPPPP-GMRYPGGMPPPPPPRYPSAGPG 361
Query: 340 QVWRPPPPPQQQGGHPMGYPQNSMPPPPPPN 370
PPPPP + P G+ +PPPPPP+
Sbjct: 362 MY--PPPPPSRPPAPPSGH--GMIPPPPPPS 388
>SP49_SCHPO (O14102) Spliceosome-associated protein 49
Length = 335
Score = 264 bits (675), Expect = 2e-70
Identities = 148/320 (46%), Positives = 202/320 (62%), Gaps = 33/320 (10%)
Query: 20 ERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDA 79
+RNQDAT Y+GNLD ++++ +L+EL +QAGPVVN+++P+DRV N H G+GF EF E+D
Sbjct: 6 DRNQDATIYLGNLDEKVTDSILFELCLQAGPVVNIHIPRDRVRNSHNGFGFCEFLHEQDV 65
Query: 80 DYAIKVLNMIKLYGKPIRVNKASQDKK-SLDVGANLFIGNLDPDVDEKLLYDTFSAFGVI 138
+YA ++LN +KL+GKPIRVN+ASQD+ + +GANLF+GNLDP VDE++LYDTFSA G +
Sbjct: 66 EYACQILNQVKLFGKPIRVNRASQDRGVNTLIGANLFVGNLDPLVDERVLYDTFSALGQL 125
Query: 139 VTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAYKKDTKGER 198
V P++ RD + G S+G+GF+SYDSFE +D+AIEAMN Q+L N+ ITVSYA+K++ KGER
Sbjct: 126 VKAPQVARD-ENGRSKGYGFVSYDSFETADAAIEAMNNQFLMNKPITVSYAFKREGKGER 184
Query: 199 HGTPAERVLAASNPTAQKSRPHTLFASGPPSLPNAPQANGTIP---APVPPRPFANGVAP 255
HG AER LAA+ A+K++ + PQ+ T+P +P P P + P
Sbjct: 185 HGDIAERKLAAA---AKKNK-----------VAVTPQS--TLPPGFSPATPAPTSAANTP 228
Query: 256 PPIHVIQ-PPPPQAP------AFQPMQMPPPGQQVWHQQQQGQPMMQQGMPPPMQQFRPP 308
I PP P P A P+ +P Q G P M P M P
Sbjct: 229 ATIAATSIPPVPNVPLVGATTAVPPLSIPNVLPFTAAQHFPGMPAM-----PMMNVPMGP 283
Query: 309 HSMQMPPPPPPQGMQGPPRP 328
+ PPPPP + P P
Sbjct: 284 GGAPLVPPPPPGMVMASPSP 303
>PABP_SCHPO (P31209) Polyadenylate-binding protein (Poly(A)-binding
protein) (PABP)
Length = 653
Score = 132 bits (331), Expect = 2e-30
Identities = 68/165 (41%), Positives = 109/165 (65%), Gaps = 3/165 (1%)
Query: 25 ATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIK 84
A+ YVG LDP ++E +L+ELF GPV ++ V +D VT + GY +V F + ED + A+
Sbjct: 80 ASLYVGELDPSVTEAMLFELFNSIGPVASIRVCRDAVTRRSLGYAYVNFHNMEDGEKALD 139
Query: 85 VLNMIKLYGKPIRVNKASQDKKSLDVG-ANLFIGNLDPDVDEKLLYDTFSAFGVIVTNPK 143
LN + G+P R+ + +D +G N+FI NLDP +D K L+DTFSAFG I+ + K
Sbjct: 140 ELNYTLIKGRPCRIMWSQRDPSLRKMGTGNVFIKNLDPAIDNKALHDTFSAFGKIL-SCK 198
Query: 144 IMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSY 188
+ D + GN++G+GF+ +DS E++++AIE +NG L ++++ V +
Sbjct: 199 VAVD-ELGNAKGYGFVHFDSVESANAAIEHVNGMLLNDKKVYVGH 242
Score = 107 bits (268), Expect = 4e-23
Identities = 89/298 (29%), Positives = 142/298 (46%), Gaps = 34/298 (11%)
Query: 28 YVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIKVLN 87
Y+ NLD +I+E+ +LF Q G + ++ + KD+ ++ +G+GFV + + E A A+ LN
Sbjct: 264 YIKNLDTEITEQEFSDLFGQFGEITSLSLVKDQ-NDKPRGFGFVNYANHECAQKAVDELN 322
Query: 88 MIKLYGKPIRVNKASQ-----------------DKKSLDVGANLFIGNLDPDVDEKLLYD 130
+ GK + V +A + +K + G NLFI NL +VD++ L
Sbjct: 323 DKEYKGKKLYVGRAQKKHEREEELRKRYEQMKLEKMNKYQGVNLFIKNLQDEVDDERLKA 382
Query: 131 TFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAY 190
FSAFG I T+ KIM D + G S+GFGF+ Y + E ++ A+ MN + L + + V+ A
Sbjct: 383 EFSAFGTI-TSAKIMTD-EQGKSKGFGFVCYTTPEEANKAVTEMNQRMLAGKPLYVALAQ 440
Query: 191 KKDTKGERHGTPAERVLAASNPTAQKSRPHTLFASGPPSL---PNAPQANGTIPAPVPPR 247
+K+ + + E + A N + + A+G P++ P G P+P
Sbjct: 441 RKEVRRSQ----LEAQIQARNQF--RLQQQVAAAAGIPAVQYGATGPLIYGPGGYPIPAA 494
Query: 248 PFANGVAPPPIH-VIQPPPPQAPAFQPMQMPPPGQQVWHQQ----QQGQPMMQQGMPP 300
G+ P H P P P P P PG + + QG+PMM G P
Sbjct: 495 VNGRGMPMVPGHNGPMPMYPGMPTQFPAGGPAPGYPGMNARGPVPAQGRPMMMPGSVP 552
Score = 82.4 bits (202), Expect = 2e-15
Identities = 56/177 (31%), Positives = 95/177 (53%), Gaps = 10/177 (5%)
Query: 28 YVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIKVLN 87
++ NLDP I + L + F G +++ V D + N +GYGFV F S E A+ AI+ +N
Sbjct: 171 FIKNLDPAIDNKALHDTFSAFGKILSCKVAVDELGNA-KGYGFVHFDSVESANAAIEHVN 229
Query: 88 MIKLYGKPIRV-NKASQDKKSLDVGA------NLFIGNLDPDVDEKLLYDTFSAFGVIVT 140
+ L K + V + S+ ++ V A N++I NLD ++ E+ D F FG I T
Sbjct: 230 GMLLNDKKVYVGHHVSRRERQSKVEALKANFTNVYIKNLDTEITEQEFSDLFGQFGEI-T 288
Query: 141 NPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAYKKDTKGE 197
+ +++D RGFGF++Y + E + A++ +N + +++ V A KK + E
Sbjct: 289 SLSLVKD-QNDKPRGFGFVNYANHECAQKAVDELNDKEYKGKKLYVGRAQKKHEREE 344
Score = 50.4 bits (119), Expect = 7e-06
Identities = 25/107 (23%), Positives = 56/107 (51%), Gaps = 2/107 (1%)
Query: 95 PIRVNKASQDKKSLDVGANLFIGNLDPDVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSR 154
P + + + A+L++G LDP V E +L++ F++ G V + ++ RD T S
Sbjct: 63 PTNASSVATPSGTAPTSASLYVGELDPSVTEAMLFELFNSIGP-VASIRVCRDAVTRRSL 121
Query: 155 GFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAYKKDTKGERHGT 201
G+ ++++ + E + A++ +N + R + ++ ++D + GT
Sbjct: 122 GYAYVNFHNMEDGEKALDELNYTLIKGRPCRIMWS-QRDPSLRKMGT 167
>HS49_YEAST (Q99181) HSH49 protein
Length = 213
Score = 131 bits (329), Expect = 3e-30
Identities = 69/195 (35%), Positives = 117/195 (59%), Gaps = 15/195 (7%)
Query: 26 TAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIKV 85
T YVGN+DP+I++E L+ELF+Q PV+ + PKD+V +QGY F+EF ++ DA YAIK+
Sbjct: 10 TVYVGNIDPRITKEQLYELFIQINPVLRIKYPKDKVLQAYQGYAFIEFYNQGDAQYAIKI 69
Query: 86 L-NMIKLYGKPIRVNKASQDKKSLDVGAN-----------LFIGNLDPDVDEKLLYDTFS 133
+ N ++LY + I+V + + + ++ +N LFI NL +D L F+
Sbjct: 70 MNNTVRLYDRLIKVRQVTNSTGTTNLPSNISKDMILPIAKLFIKNLADSIDSDQLVKIFN 129
Query: 134 AFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAYKKD 193
FG ++ P+I + ++ ++ FE +D AI+++N Q + N +ITV YA+K++
Sbjct: 130 KFGKLIREPEIFY--LSNGKLKCAYVYFEDFEKADLAIKSLNNQLVANNRITVDYAFKEN 187
Query: 194 TKGE-RHGTPAERVL 207
KG ++G +R+L
Sbjct: 188 GKGNAKYGDDVDRLL 202
Score = 34.7 bits (78), Expect = 0.41
Identities = 19/69 (27%), Positives = 36/69 (51%), Gaps = 1/69 (1%)
Query: 107 SLDVGANLFIGNLDPDVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEA 166
S D G +++GN+DP + ++ LY+ F ++ K +D +G+ FI + +
Sbjct: 4 SADSGNTVYVGNIDPRITKEQLYELFIQINPVL-RIKYPKDKVLQAYQGYAFIEFYNQGD 62
Query: 167 SDSAIEAMN 175
+ AI+ MN
Sbjct: 63 AQYAIKIMN 71
>PAB2_ARATH (P42731) Polyadenylate-binding protein 2
(Poly(A)-binding protein 2) (PABP 2)
Length = 629
Score = 129 bits (324), Expect = 1e-29
Identities = 75/173 (43%), Positives = 110/173 (63%), Gaps = 3/173 (1%)
Query: 15 GQHSAERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFR 74
G +A + + + YVG+LD +++ L++ F Q G VV V V +D VT + GYG+V F
Sbjct: 26 GGATATQFGNTSLYVGDLDFNVTDSQLFDAFGQMGTVVTVRVCRDLVTRRSLGYGYVNFT 85
Query: 75 SEEDADYAIKVLNMIKLYGKPIRVNKASQDKKSLDVGA-NLFIGNLDPDVDEKLLYDTFS 133
+ +DA AI+ LN I LYGKPIRV + +D GA N+FI NLD +D K L+DTFS
Sbjct: 86 NPQDAARAIQELNYIPLYGKPIRVMYSHRDPSVRRSGAGNIFIKNLDESIDHKALHDTFS 145
Query: 134 AFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITV 186
+FG IV + K+ D +G S+G+GF+ Y + E++ AIE +NG L ++Q+ V
Sbjct: 146 SFGNIV-SCKVAVD-SSGQSKGYGFVQYANEESAQKAIEKLNGMLLNDKQVYV 196
Score = 113 bits (283), Expect = 7e-25
Identities = 95/332 (28%), Positives = 155/332 (46%), Gaps = 34/332 (10%)
Query: 18 SAERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEE 77
+A + + YV NL +++ L F + G + + V KD + +G+GFV F + +
Sbjct: 208 TANKTKFTNVYVKNLAESTTDDDLKNAFGEYGKITSAVVMKDG-EGKSKGFGFVNFENAD 266
Query: 78 DADYAIKVLNMIKLYGKPI---RVNKASQDKKSLDV--------------GANLFIGNLD 120
DA A++ LN K K R K S+ + L V +NL++ NLD
Sbjct: 267 DAARAVESLNGHKFDDKEWYVGRAQKKSERETELRVRYEQNLKEAADKFQSSNLYVKNLD 326
Query: 121 PDVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLC 180
P + ++ L + FS FG VT+ K+MRDP+ G S+G GF+++ + E + A+ ++G+ +
Sbjct: 327 PSISDEKLKEIFSPFGT-VTSSKVMRDPN-GTSKGSGFVAFATPEEATEAMSQLSGKMIE 384
Query: 181 NRQITVSYAYKKDTKGERHGTPAERVL-AASNPTAQKSRPHTLFASGPPSLPNAPQANGT 239
++ + V+ A +K+ + R +V A P+ P ++ G P +
Sbjct: 385 SKPLYVAIAQRKEDRRVRLQAQFSQVRPVAMQPSVGPRMP--VYPPGGPGIGQQMFYGQA 442
Query: 240 IPAPVPPRPFANGVAPPPIHVIQPPPPQAPAF-----QPMQMPPPGQQ----VWHQQQQG 290
PA +PP+P G + ++P P+F QP Q P G + + H QQQ
Sbjct: 443 PPAMIPPQP-GYGYQQQLVPGMRPGGGPVPSFFMPMVQPQQQRPGGGRRPGGIQHSQQQ- 500
Query: 291 QPMMQQGMPPPMQQFRPPHSMQMPPPPPPQGM 322
PMMQQ M P + FR P PP M
Sbjct: 501 NPMMQQQMHPRGRMFRYPQGRGGSGDVPPYDM 532
Score = 86.3 bits (212), Expect = 1e-16
Identities = 100/390 (25%), Positives = 162/390 (40%), Gaps = 41/390 (10%)
Query: 18 SAERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEE 77
S R+ ++ NLD I + L + F G +V+ V D + Q +GYGFV++ +EE
Sbjct: 117 SVRRSGAGNIFIKNLDESIDHKALHDTFSSFGNIVSCKVAVDS-SGQSKGYGFVQYANEE 175
Query: 78 DADYAIKVLNMIKLYGKPIRVNKASQDKKSLDVG-----ANLFIGNLDPDVDEKLLYDTF 132
A AI+ LN + L K + V + ++ N+++ NL + L + F
Sbjct: 176 SAQKAIEKLNGMLLNDKQVYVGPFLRRQERDSTANKTKFTNVYVKNLAESTTDDDLKNAF 235
Query: 133 SAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAYKK 192
+G I T+ +M+D + G S+GFGF+++++ + + A+E++NG +++ V A K
Sbjct: 236 GEYGKI-TSAVVMKDGE-GKSKGFGFVNFENADDAARAVESLNGHKFDDKEWYVGRAQK- 292
Query: 193 DTKGERHGTPAERVLAASNPTAQKSRPHTLFASG-PPSLPNAPQAN-----GTIPAPVPP 246
K ER R A K + L+ PS+ + GT+ +
Sbjct: 293 --KSERETELRVRYEQNLKEAADKFQSSNLYVKNLDPSISDEKLKEIFSPFGTVTSSKVM 350
Query: 247 RPFANGVAPPPIHVIQPPPPQAP------AFQPMQMPP----PGQQVWHQQQQGQPMMQQ 296
R NG + V P +A + + ++ P Q+ ++ + Q Q
Sbjct: 351 RD-PNGTSKGSGFVAFATPEEATEAMSQLSGKMIESKPLYVAIAQRKEDRRVRLQAQFSQ 409
Query: 297 GMPPPMQQFRPPHSMQMPPPPPPQGM-----QGPPRPLPPPSVMAGQQQVWRPPPPPQQQ 351
P MQ P PP P G Q PP +PP QQQ+ P +
Sbjct: 410 VRPVAMQPSVGPRMPVYPPGGPGIGQQMFYGQAPPAMIPPQPGYGYQQQL----VPGMRP 465
Query: 352 GGHPMGYPQNSMPPPPPPNHH--GMRPPSG 379
GG P+ P MP P G R P G
Sbjct: 466 GGGPV--PSFFMPMVQPQQQRPGGGRRPGG 493
Score = 49.3 bits (116), Expect = 2e-05
Identities = 30/83 (36%), Positives = 46/83 (55%), Gaps = 1/83 (1%)
Query: 16 QHSAERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRS 75
+ +A++ Q + YV NLDP IS+E L E+F G V + V +D +G GFV F +
Sbjct: 309 KEAADKFQSSNLYVKNLDPSISDEKLKEIFSPFGTVTSSKVMRD-PNGTSKGSGFVAFAT 367
Query: 76 EEDADYAIKVLNMIKLYGKPIRV 98
E+A A+ L+ + KP+ V
Sbjct: 368 PEEATEAMSQLSGKMIESKPLYV 390
>PAB1_RAT (Q9EPH8) Polyadenylate-binding protein 1 (Poly(A)-binding
protein 1) (PABP 1)
Length = 636
Score = 127 bits (320), Expect = 4e-29
Identities = 66/175 (37%), Positives = 109/175 (61%), Gaps = 5/175 (2%)
Query: 25 ATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIK 84
A+ YVG+L P ++E +L+E F AGP++++ V +D +T + GY +V F+ DA+ A+
Sbjct: 11 ASLYVGDLHPDVTEAMLYEKFSPAGPILSIRVCRDMITRRSLGYAYVNFQQPADAERALD 70
Query: 85 VLNMIKLYGKPIRVNKASQDKKSLDVG-ANLFIGNLDPDVDEKLLYDTFSAFGVIVTNPK 143
+N + GKP+R+ + +D G N+FI NLD +D K LYDTFSAFG I++
Sbjct: 71 TMNFDVIKGKPVRIMWSQRDPSLRKSGVGNIFIKNLDKSIDNKALYDTFSAFGNILSCKV 130
Query: 144 IMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVS-YAYKKDTKGE 197
+ D S+G+GF+ +++ EA++ AIE MNG L +R++ V + +K+ + E
Sbjct: 131 VC---DENGSKGYGFVHFETQEAAERAIEKMNGMLLNDRKVFVGRFKSRKEREAE 182
Score = 105 bits (261), Expect = 2e-22
Identities = 104/385 (27%), Positives = 160/385 (41%), Gaps = 51/385 (13%)
Query: 19 AERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEED 78
A + Y+ N + +E L ELF + GP ++V V D + + +G+GFV F ED
Sbjct: 185 ARAKEFTNVYIKNFGEDMDDERLKELFGKFGPALSVKVMTDE-SGKSKGFGFVSFERHED 243
Query: 79 ADYAIKVLNMIKLYGKPIRVNKAS-----------------QDKKSLDVGANLFIGNLDP 121
A A+ +N +L GK I V +A QD+ + G NL++ NLD
Sbjct: 244 AQKAVDEMNGKELNGKQIYVGRAQKKVERQTELKRKFEQMKQDRITRYQGVNLYVKNLDD 303
Query: 122 DVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCN 181
+D++ L FS FG I T+ K+M + G S+GFGF+ + S E + A+ MNG+ +
Sbjct: 304 GIDDERLRKEFSPFGTI-TSAKVMMEG--GRSKGFGFVCFSSPEEATKAVTEMNGRIVAT 360
Query: 182 RQITVSYAYKKDTK----GERHGTPAERVLAASNPTA---QKSRPHTLFASGPPSLPNAP 234
+ + V+ A +K+ + ++ V A NP Q + P F + P N
Sbjct: 361 KPLYVALAQRKEERQAHLTNQYMQRMASVRAVPNPVINPYQPAPPSGYFMAAIPQTQN-- 418
Query: 235 QANGTIPAPV-----PPRPFANGVAPPPIH----VIQPPPPQAP--AFQPMQMPPPGQQV 283
+A P+ + PR A G P P I+P P+ P +P P +V
Sbjct: 419 RAAYYPPSQIAQLRPSPRWTAQGARPHPFQNMPGAIRPAAPRPPFSTMRPASSQVP--RV 476
Query: 284 WHQQQQGQPMMQQGMPPP--MQQFRPPHSMQMPPPPPPQGMQGPPRPLPPPSVMAGQQQV 341
Q+ Q P P P +P G++ P + L Q QV
Sbjct: 477 MSTQRVANTSTQTMGPRPAAAATAATPAVRTVPQYKYAAGVRNPQQHL------NAQPQV 530
Query: 342 WRPPPPPQQQGGHPMGYPQNSMPPP 366
P QG P+ + PP
Sbjct: 531 TMQQPAVHVQGQEPLTASMLASAPP 555
Score = 92.8 bits (229), Expect = 1e-18
Identities = 57/172 (33%), Positives = 100/172 (58%), Gaps = 11/172 (6%)
Query: 28 YVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIKVLN 87
++ NLD I + L++ F G +++ V D N +GYGFV F ++E A+ AI+ +N
Sbjct: 102 FIKNLDKSIDNKALYDTFSAFGNILSCKVVCDE--NGSKGYGFVHFETQEAAERAIEKMN 159
Query: 88 MIKLYGKPIRVNK-ASQDKKSLDVGA------NLFIGNLDPDVDEKLLYDTFSAFGVIVT 140
+ L + + V + S+ ++ ++GA N++I N D+D++ L + F FG ++
Sbjct: 160 GMLLNDRKVFVGRFKSRKEREAELGARAKEFTNVYIKNFGEDMDDERLKELFGKFGPALS 219
Query: 141 NPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAYKK 192
K+M D ++G S+GFGF+S++ E + A++ MNG+ L +QI V A KK
Sbjct: 220 -VKVMTD-ESGKSKGFGFVSFERHEDAQKAVDEMNGKELNGKQIYVGRAQKK 269
>PAB1_HUMAN (P11940) Polyadenylate-binding protein 1
(Poly(A)-binding protein 1) (PABP 1)
Length = 636
Score = 127 bits (320), Expect = 4e-29
Identities = 66/175 (37%), Positives = 109/175 (61%), Gaps = 5/175 (2%)
Query: 25 ATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIK 84
A+ YVG+L P ++E +L+E F AGP++++ V +D +T + GY +V F+ DA+ A+
Sbjct: 11 ASLYVGDLHPDVTEAMLYEKFSPAGPILSIRVCRDMITRRSLGYAYVNFQQPADAERALD 70
Query: 85 VLNMIKLYGKPIRVNKASQDKKSLDVG-ANLFIGNLDPDVDEKLLYDTFSAFGVIVTNPK 143
+N + GKP+R+ + +D G N+FI NLD +D K LYDTFSAFG I++
Sbjct: 71 TMNFDVIKGKPVRIMWSQRDPSLRKSGVGNIFIKNLDKSIDNKALYDTFSAFGNILSCKV 130
Query: 144 IMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVS-YAYKKDTKGE 197
+ D S+G+GF+ +++ EA++ AIE MNG L +R++ V + +K+ + E
Sbjct: 131 VC---DENGSKGYGFVHFETQEAAERAIEKMNGMLLNDRKVFVGRFKSRKEREAE 182
Score = 103 bits (258), Expect = 6e-22
Identities = 103/385 (26%), Positives = 160/385 (40%), Gaps = 51/385 (13%)
Query: 19 AERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEED 78
A + Y+ N + +E L +LF + GP ++V V D + + +G+GFV F ED
Sbjct: 185 ARAKEFTNVYIKNFGEDMDDERLKDLFGKFGPALSVKVMTDE-SGKSKGFGFVSFERHED 243
Query: 79 ADYAIKVLNMIKLYGKPIRVNKAS-----------------QDKKSLDVGANLFIGNLDP 121
A A+ +N +L GK I V +A QD+ + G NL++ NLD
Sbjct: 244 AQKAVDEMNGKELNGKQIYVGRAQKKVERQTELKRKFEQMKQDRITRYQGVNLYVKNLDD 303
Query: 122 DVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCN 181
+D++ L FS FG I T+ K+M + G S+GFGF+ + S E + A+ MNG+ +
Sbjct: 304 GIDDERLRKEFSPFGTI-TSAKVMMEG--GRSKGFGFVCFSSPEEATKAVTEMNGRIVAT 360
Query: 182 RQITVSYAYKKDTK----GERHGTPAERVLAASNPTA---QKSRPHTLFASGPPSLPNAP 234
+ + V+ A +K+ + ++ V A NP Q + P F + P N
Sbjct: 361 KPLYVALAQRKEERQAHLTNQYMQRMASVRAVPNPVINPYQPAPPSGYFMAAIPQTQN-- 418
Query: 235 QANGTIPAPV-----PPRPFANGVAPPPIH----VIQPPPPQAP--AFQPMQMPPPGQQV 283
+A P+ + PR A G P P I+P P+ P +P P +V
Sbjct: 419 RAAYYPPSQIAQLRPSPRWTAQGARPHPFQNMPGAIRPAAPRPPFSTMRPASSQVP--RV 476
Query: 284 WHQQQQGQPMMQQGMPPP--MQQFRPPHSMQMPPPPPPQGMQGPPRPLPPPSVMAGQQQV 341
Q+ Q P P P +P G++ P + L Q QV
Sbjct: 477 MSTQRVANTSTQTMGPRPAAAAAAATPAVRTVPQYKYAAGVRNPQQHL------NAQPQV 530
Query: 342 WRPPPPPQQQGGHPMGYPQNSMPPP 366
P QG P+ + PP
Sbjct: 531 TMQQPAVHVQGQEPLTASMLASAPP 555
Score = 94.4 bits (233), Expect = 4e-19
Identities = 58/172 (33%), Positives = 100/172 (57%), Gaps = 11/172 (6%)
Query: 28 YVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIKVLN 87
++ NLD I + L++ F G +++ V D N +GYGFV F ++E A+ AI+ +N
Sbjct: 102 FIKNLDKSIDNKALYDTFSAFGNILSCKVVCDE--NGSKGYGFVHFETQEAAERAIEKMN 159
Query: 88 MIKLYGKPIRVNK-ASQDKKSLDVGA------NLFIGNLDPDVDEKLLYDTFSAFGVIVT 140
+ L + + V + S+ ++ ++GA N++I N D+D++ L D F FG ++
Sbjct: 160 GMLLNDRKVFVGRFKSRKEREAELGARAKEFTNVYIKNFGEDMDDERLKDLFGKFGPALS 219
Query: 141 NPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAYKK 192
K+M D ++G S+GFGF+S++ E + A++ MNG+ L +QI V A KK
Sbjct: 220 -VKVMTD-ESGKSKGFGFVSFERHEDAQKAVDEMNGKELNGKQIYVGRAQKK 269
>PAB1_BOVIN (P61286) Polyadenylate-binding protein 1
(Poly(A)-binding protein 1) (PABP 1)
Length = 636
Score = 127 bits (320), Expect = 4e-29
Identities = 66/175 (37%), Positives = 109/175 (61%), Gaps = 5/175 (2%)
Query: 25 ATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIK 84
A+ YVG+L P ++E +L+E F AGP++++ V +D +T + GY +V F+ DA+ A+
Sbjct: 11 ASLYVGDLHPDVTEAMLYEKFSPAGPILSIRVCRDMITRRSLGYAYVNFQQPADAERALD 70
Query: 85 VLNMIKLYGKPIRVNKASQDKKSLDVG-ANLFIGNLDPDVDEKLLYDTFSAFGVIVTNPK 143
+N + GKP+R+ + +D G N+FI NLD +D K LYDTFSAFG I++
Sbjct: 71 TMNFDVIKGKPVRIMWSQRDPSLRKSGVGNIFIKNLDKSIDNKALYDTFSAFGNILSCKV 130
Query: 144 IMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVS-YAYKKDTKGE 197
+ D S+G+GF+ +++ EA++ AIE MNG L +R++ V + +K+ + E
Sbjct: 131 VC---DENGSKGYGFVHFETQEAAERAIEKMNGMLLNDRKVFVGRFKSRKEREAE 182
Score = 103 bits (258), Expect = 6e-22
Identities = 103/385 (26%), Positives = 160/385 (40%), Gaps = 51/385 (13%)
Query: 19 AERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEED 78
A + Y+ N + +E L +LF + GP ++V V D + + +G+GFV F ED
Sbjct: 185 ARAKEFTNVYIKNFGEDMDDERLKDLFGKFGPALSVKVMTDE-SGKSKGFGFVSFERHED 243
Query: 79 ADYAIKVLNMIKLYGKPIRVNKAS-----------------QDKKSLDVGANLFIGNLDP 121
A A+ +N +L GK I V +A QD+ + G NL++ NLD
Sbjct: 244 AQKAVDEMNGKELNGKQIYVGRAQKKVERQTELKRKFEQMKQDRITRYQGVNLYVKNLDD 303
Query: 122 DVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCN 181
+D++ L FS FG I T+ K+M + G S+GFGF+ + S E + A+ MNG+ +
Sbjct: 304 GIDDERLRKEFSPFGTI-TSAKVMMEG--GRSKGFGFVCFSSPEEATKAVTEMNGRIVAT 360
Query: 182 RQITVSYAYKKDTK----GERHGTPAERVLAASNPTA---QKSRPHTLFASGPPSLPNAP 234
+ + V+ A +K+ + ++ V A NP Q + P F + P N
Sbjct: 361 KPLYVALAQRKEERQAHLTNQYMQRMASVRAVPNPVINPYQPAPPSGYFMAAIPQTQN-- 418
Query: 235 QANGTIPAPV-----PPRPFANGVAPPPIH----VIQPPPPQAP--AFQPMQMPPPGQQV 283
+A P+ + PR A G P P I+P P+ P +P P +V
Sbjct: 419 RAAYYPPSQIAQLRPSPRWTAQGARPHPFQNMPGAIRPAAPRPPFSTMRPASSQVP--RV 476
Query: 284 WHQQQQGQPMMQQGMPPP--MQQFRPPHSMQMPPPPPPQGMQGPPRPLPPPSVMAGQQQV 341
Q+ Q P P P +P G++ P + L Q QV
Sbjct: 477 MSTQRVANTSTQTMGPRPAAAAAAATPAVRTVPQYKYAAGVRNPQQHL------NAQPQV 530
Query: 342 WRPPPPPQQQGGHPMGYPQNSMPPP 366
P QG P+ + PP
Sbjct: 531 TMQQPAVHVQGQEPLTASMLASAPP 555
Score = 94.4 bits (233), Expect = 4e-19
Identities = 58/172 (33%), Positives = 100/172 (57%), Gaps = 11/172 (6%)
Query: 28 YVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIKVLN 87
++ NLD I + L++ F G +++ V D N +GYGFV F ++E A+ AI+ +N
Sbjct: 102 FIKNLDKSIDNKALYDTFSAFGNILSCKVVCDE--NGSKGYGFVHFETQEAAERAIEKMN 159
Query: 88 MIKLYGKPIRVNK-ASQDKKSLDVGA------NLFIGNLDPDVDEKLLYDTFSAFGVIVT 140
+ L + + V + S+ ++ ++GA N++I N D+D++ L D F FG ++
Sbjct: 160 GMLLNDRKVFVGRFKSRKEREAELGARAKEFTNVYIKNFGEDMDDERLKDLFGKFGPALS 219
Query: 141 NPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAYKK 192
K+M D ++G S+GFGF+S++ E + A++ MNG+ L +QI V A KK
Sbjct: 220 -VKVMTD-ESGKSKGFGFVSFERHEDAQKAVDEMNGKELNGKQIYVGRAQKK 269
>PAB1_MOUSE (P29341) Polyadenylate-binding protein 1
(Poly(A)-binding protein 1) (PABP 1)
Length = 636
Score = 127 bits (319), Expect = 5e-29
Identities = 66/175 (37%), Positives = 109/175 (61%), Gaps = 5/175 (2%)
Query: 25 ATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIK 84
A+ YVG+L P ++E +L+E F AGP++++ V +D +T + GY +V F+ DA+ A+
Sbjct: 11 ASLYVGDLHPDVTEAMLYEKFSPAGPILSIRVCRDMITRRSLGYAYVNFQQPADAERALD 70
Query: 85 VLNMIKLYGKPIRVNKASQDKKSLDVG-ANLFIGNLDPDVDEKLLYDTFSAFGVIVTNPK 143
+N + GKP+R+ + +D G N+FI NLD +D K LYDTFSAFG I++
Sbjct: 71 TMNFDVIKGKPVRIMWSQRDPSLRKSGVGNIFIKNLDKSIDNKALYDTFSAFGNILSCKV 130
Query: 144 IMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVS-YAYKKDTKGE 197
+ D S+G+GF+ +++ EA++ AIE MNG L +R++ V + +K+ + E
Sbjct: 131 VC---DENGSKGYGFVHFETQEAAERAIEKMNGMLLNDRKVFVGRFKSQKEREAE 182
Score = 105 bits (261), Expect = 2e-22
Identities = 104/385 (27%), Positives = 160/385 (41%), Gaps = 51/385 (13%)
Query: 19 AERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEED 78
A + Y+ N + +E L ELF + GP ++V V D + + +G+GFV F ED
Sbjct: 185 ARAKEFTNVYIKNFGEDMDDERLKELFGKFGPALSVKVMTDE-SGKSKGFGFVSFERHED 243
Query: 79 ADYAIKVLNMIKLYGKPIRVNKAS-----------------QDKKSLDVGANLFIGNLDP 121
A A+ +N +L GK I V +A QD+ + G NL++ NLD
Sbjct: 244 AQKAVDEMNGKELNGKQIYVGRAQKKVERQTELKRKFEQMKQDRITRYQGVNLYVKNLDD 303
Query: 122 DVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCN 181
+D++ L FS FG I T+ K+M + G S+GFGF+ + S E + A+ MNG+ +
Sbjct: 304 GIDDERLRKEFSPFGTI-TSAKVMMEG--GRSKGFGFVCFSSPEEATKAVTEMNGRIVAT 360
Query: 182 RQITVSYAYKKDTK----GERHGTPAERVLAASNPTA---QKSRPHTLFASGPPSLPNAP 234
+ + V+ A +K+ + ++ V A NP Q + P F + P N
Sbjct: 361 KPLYVALAQRKEERQAHLTNQYMQRMASVRAVPNPVINPYQPAPPSGYFMAAIPQTQN-- 418
Query: 235 QANGTIPAPV-----PPRPFANGVAPPPIH----VIQPPPPQAP--AFQPMQMPPPGQQV 283
+A P+ + PR A G P P I+P P+ P +P P +V
Sbjct: 419 RAAYYPPSQIAQLRPSPRWTAQGARPHPFQNMPGAIRPAAPRPPFSTMRPASSQVP--RV 476
Query: 284 WHQQQQGQPMMQQGMPPP--MQQFRPPHSMQMPPPPPPQGMQGPPRPLPPPSVMAGQQQV 341
Q+ Q P P P +P G++ P + L Q QV
Sbjct: 477 MSTQRVANTSTQTMGPRPAAAAAAATPAVRTVPQYKYAAGVRNPQQHL------NAQPQV 530
Query: 342 WRPPPPPQQQGGHPMGYPQNSMPPP 366
P QG P+ + PP
Sbjct: 531 TMQQPAVHVQGQEPLTASMLASAPP 555
Score = 94.4 bits (233), Expect = 4e-19
Identities = 58/172 (33%), Positives = 100/172 (57%), Gaps = 11/172 (6%)
Query: 28 YVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIKVLN 87
++ NLD I + L++ F G +++ V D N +GYGFV F ++E A+ AI+ +N
Sbjct: 102 FIKNLDKSIDNKALYDTFSAFGNILSCKVVCDE--NGSKGYGFVHFETQEAAERAIEKMN 159
Query: 88 MIKLYGKPIRVNK-ASQDKKSLDVGA------NLFIGNLDPDVDEKLLYDTFSAFGVIVT 140
+ L + + V + SQ ++ ++GA N++I N D+D++ L + F FG ++
Sbjct: 160 GMLLNDRKVFVGRFKSQKEREAELGARAKEFTNVYIKNFGEDMDDERLKELFGKFGPALS 219
Query: 141 NPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAYKK 192
K+M D ++G S+GFGF+S++ E + A++ MNG+ L +QI V A KK
Sbjct: 220 -VKVMTD-ESGKSKGFGFVSFERHEDAQKAVDEMNGKELNGKQIYVGRAQKK 269
>RB97_DROME (Q02926) Ribonucleoprotein RB97D
Length = 471
Score = 126 bits (316), Expect = 1e-28
Identities = 118/409 (28%), Positives = 172/409 (41%), Gaps = 73/409 (17%)
Query: 28 YVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIKVLN 87
++G L P +EE L + Q G VV+V V +D T + +G+GF+ + D A +
Sbjct: 35 FIGGLAPYTTEENLKLFYGQWGKVVDVVVMRDAATKRSRGFGFITYTKSLMVDRAQENRP 94
Query: 88 MIKLYGKPIRVNKA----SQDKKSLDVGAN-LFIGNLDPDVDEKLLYDTFSAFGVIVTNP 142
I + GK + +A ++ + ++ LF+G L + DE+ L + F FG +V+
Sbjct: 95 HI-IDGKTVEAKRALPRPERESRETNISVKKLFVGGLKDNHDEECLREYFLQFGNVVS-V 152
Query: 143 KIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYA--------YKKDT 194
K++ D TG RGF F+ +D ++A D AI +Q + Y Y D
Sbjct: 153 KLLTDKTTGKRRGFAFVEFDDYDAVDKAI--------LKKQHAIKYVHVDVKKSIYNLDK 204
Query: 195 KGERH-GTPAERVLAASNPTAQKS----RPHTLFASGPPSLPNAP--QANGTIPAPVPPR 247
K ++ G A + + N Q+ +P P P P QA G PP
Sbjct: 205 KEKQQPGGLANAIKPSLNQQQQQQGGGQQPPNGNMQAPSFRPPVPPQQAMGPYQQQPPPA 264
Query: 248 PFANGVAPPP-IHVIQPPPPQAPAFQ----PMQM-------------------PPPGQQV 283
P + APPP + PPPP P + P QM PPPG Q
Sbjct: 265 PMS---APPPNFNYWGPPPPAMPPYYQQPPPQQMNGWGPYPPPQQNGWNAPPPPPPGAQQ 321
Query: 284 WHQQQQGQPMMQQGMPPPMQQFRPPHSMQMPPPPPPQGMQGPPRPLPPPSVMAGQQQVWR 343
WH Q G P Q + PP+ PP PPP P PP P+P + + QQQ +
Sbjct: 322 WHANQWGCPPPVQQV-PPVGAVPPPMGHNGPPPTAPGNWNMPP-PVPGAAPPSHQQQSSQ 379
Query: 344 PPPP--------PQQQGGHPMGYPQ---NSMPP---PPPPNHHGMRPPS 378
P P Q GG P + N M P PP ++H +PP+
Sbjct: 380 QPTPQPNFGTGYQQNYGGGPSKHNNMNANRMNPYSAGPPNSYHTPQPPA 428
>PAB4_HUMAN (Q13310) Polyadenylate-binding protein 4
(Poly(A)-binding protein 4) (PABP 4) (Inducible
poly(A)-binding protein) (iPABP) (Activated-platelet
protein-1) (APP-1)
Length = 644
Score = 124 bits (312), Expect = 3e-28
Identities = 67/175 (38%), Positives = 107/175 (60%), Gaps = 5/175 (2%)
Query: 25 ATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIK 84
A+ YVG+L ++E +L+E F AGPV+++ V +D +T + GY +V F+ DA+ A+
Sbjct: 11 ASLYVGDLHSDVTEAMLYEKFSPAGPVLSIRVCRDMITRRSLGYAYVNFQQPADAERALD 70
Query: 85 VLNMIKLYGKPIRVNKASQDKKSLDVG-ANLFIGNLDPDVDEKLLYDTFSAFGVIVTNPK 143
+N + GKPIR+ + +D G N+FI NLD +D K LYDTFSAFG I++
Sbjct: 71 TMNFDVIKGKPIRIMWSQRDPSLRKSGVGNVFIKNLDKSIDNKALYDTFSAFGNILSCKV 130
Query: 144 IMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVS-YAYKKDTKGE 197
+ D S+G+ F+ +++ EA+D AIE MNG L +R++ V + +K+ + E
Sbjct: 131 VC---DENGSKGYAFVHFETQEAADKAIEKMNGMLLNDRKVFVGRFKSRKEREAE 182
Score = 108 bits (271), Expect = 2e-23
Identities = 93/315 (29%), Positives = 147/315 (46%), Gaps = 49/315 (15%)
Query: 19 AERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEED 78
A+ + Y+ N ++ +E L ELF Q G ++V V +D + +G+GFV + ED
Sbjct: 185 AKAKEFTNVYIKNFGEEVDDESLKELFSQFGKTLSVKVMRDP-NGKSKGFGFVSYEKHED 243
Query: 79 ADYAIKVLNMIKLYGKPIRVNKAS-----------------QDKKSLDVGANLFIGNLDP 121
A+ A++ +N ++ GK I V +A Q++ S G NL+I NLD
Sbjct: 244 ANKAVEEMNGKEISGKIIFVGRAQKKVERQAELKRKFEQLKQERISRYQGVNLYIKNLDD 303
Query: 122 DVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCN 181
+D++ L FS FG I T+ K+M + G S+GFGF+ + S E + A+ MNG+ + +
Sbjct: 304 TIDDEKLRKEFSPFGSI-TSAKVMLED--GRSKGFGFVCFSSPEEATKAVTEMNGRIVGS 360
Query: 182 RQITVSYAYKKDTKGERHGTPAERVLAASNPTAQKSRPHTLFASGPPSLPNAPQANGTIP 241
+ + V+ A +K+ ER +N Q+ +G +LP N P
Sbjct: 361 KPLYVALAQRKE----------ERKAHLTNQYMQR-------VAGMRALPANAILNQFQP 403
Query: 242 APVPPRPFANGVAPPPIHVIQPPPPQAPAFQPMQMPPPGQQVWHQQQQGQPMMQQGMPPP 301
A A G P + Q PP Q QM P + W QQ G+P QGMP
Sbjct: 404 A-------AGGYFVPAVPQAQGRPPYYTPNQLAQMRPNPR--W--QQGGRPQGFQGMPSA 452
Query: 302 MQQFRPPHSMQMPPP 316
++Q P +++ P
Sbjct: 453 IRQSGPRPTLRHLAP 467
Score = 98.2 bits (243), Expect = 3e-20
Identities = 61/182 (33%), Positives = 104/182 (56%), Gaps = 11/182 (6%)
Query: 18 SAERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEE 77
S ++ ++ NLD I + L++ F G +++ V D N +GY FV F ++E
Sbjct: 92 SLRKSGVGNVFIKNLDKSIDNKALYDTFSAFGNILSCKVVCDE--NGSKGYAFVHFETQE 149
Query: 78 DADYAIKVLNMIKLYGKPIRVNK-ASQDKKSLDVGA------NLFIGNLDPDVDEKLLYD 130
AD AI+ +N + L + + V + S+ ++ ++GA N++I N +VD++ L +
Sbjct: 150 AADKAIEKMNGMLLNDRKVFVGRFKSRKEREAELGAKAKEFTNVYIKNFGEEVDDESLKE 209
Query: 131 TFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAY 190
FS FG ++ K+MRDP+ G S+GFGF+SY+ E ++ A+E MNG+ + + I V A
Sbjct: 210 LFSQFGKTLS-VKVMRDPN-GKSKGFGFVSYEKHEDANKAVEEMNGKEISGKIIFVGRAQ 267
Query: 191 KK 192
KK
Sbjct: 268 KK 269
>PAB1_XENLA (P20965) Polyadenylate-binding protein 1
(Poly(A)-binding protein 1) (PABP 1)
Length = 633
Score = 122 bits (306), Expect = 1e-27
Identities = 63/175 (36%), Positives = 108/175 (61%), Gaps = 5/175 (2%)
Query: 25 ATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIK 84
A+ YVG+L ++E +L+E F AGP++++ V +D +T + GY +V F+ DA+ A+
Sbjct: 11 ASLYVGDLHQDVTEAMLYEKFSPAGPILSIRVCRDMITRRSLGYAYVNFQQPADAERALD 70
Query: 85 VLNMIKLYGKPIRVNKASQDKKSLDVG-ANLFIGNLDPDVDEKLLYDTFSAFGVIVTNPK 143
+N + G+P+R+ + +D G N+FI NLD +D K LYDTFSAFG I++
Sbjct: 71 TMNFDVIKGRPVRIMWSQRDPSLRKSGVGNIFIKNLDKSIDNKALYDTFSAFGNILSCKV 130
Query: 144 IMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVS-YAYKKDTKGE 197
+ D S+G+GF+ +++ EA++ AI+ MNG L +R++ V + +K+ + E
Sbjct: 131 VC---DENGSKGYGFVHFETQEAAERAIDKMNGMLLNDRKVFVGRFKSRKEREAE 182
Score = 98.2 bits (243), Expect = 3e-20
Identities = 86/295 (29%), Positives = 134/295 (45%), Gaps = 37/295 (12%)
Query: 19 AERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEED 78
A + Y+ N +++E L E+F + GP ++V V D + +G+GFV F ED
Sbjct: 185 ARAKEFTNVYIKNFGDDMNDERLKEMFGKYGPALSVKVMTDD-NGKSKGFGFVSFERHED 243
Query: 79 ADYAIKVLNMIKLYGKPIRVNKA-----------------SQDKKSLDVGANLFIGNLDP 121
A A+ + + GK + V +A +QD+ + G NL++ NLD
Sbjct: 244 AQKAVDEMYGKDMNGKSMFVGRAQKKVERQTELKRKFEQMNQDRITRYQGVNLYVKNLDD 303
Query: 122 DVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCN 181
+D++ L F FG I T+ K+M + G S+GFGF+ + S E + A+ MNG+ +
Sbjct: 304 GIDDERLRKEFLPFGTI-TSAKVMMEG--GRSKGFGFVCFSSPEEATKAVTEMNGRIVAT 360
Query: 182 RQITVSYAYKKDTKGERHGTPAERVLAAS----NPTAQ--KSRPHTLFASGPPSLPN--A 233
+ + V+ A +K+ + + H T AS NP + P + F + P N A
Sbjct: 361 KPLYVALAQRKEER-QAHLTNQYMQRMASVRVPNPVINPYQPPPSSYFMAAIPPAQNRAA 419
Query: 234 PQANGTIPAPVP-PRPFANGVAPPPIH----VIQPPPPQAPAFQPMQMPPPGQQV 283
G I P PR A G P P I+P P+ P F M+ P QV
Sbjct: 420 YYPPGQIAQLRPSPRWTAQGARPHPFQNMPGAIRPTAPRPPTFSTMR--PASNQV 472
Score = 82.8 bits (203), Expect = 1e-15
Identities = 52/172 (30%), Positives = 96/172 (55%), Gaps = 11/172 (6%)
Query: 28 YVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIKVLN 87
++ NLD I + L++ F G +++ V D N +GYGFV F ++E A+ AI +N
Sbjct: 102 FIKNLDKSIDNKALYDTFSAFGNILSCKVVCDE--NGSKGYGFVHFETQEAAERAIDKMN 159
Query: 88 MIKLYGKPIRVNK-ASQDKKSLDVGA------NLFIGNLDPDVDEKLLYDTFSAFGVIVT 140
+ L + + V + S+ ++ ++GA N++I N D++++ L + F +G ++
Sbjct: 160 GMLLNDRKVFVGRFKSRKEREAELGARAKEFTNVYIKNFGDDMNDERLKEMFGKYGPALS 219
Query: 141 NPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAYKK 192
K+M D D G S+GFGF+S++ E + A++ M G+ + + + V A KK
Sbjct: 220 -VKVMTD-DNGKSKGFGFVSFERHEDAQKAVDEMYGKDMNGKSMFVGRAQKK 269
>PAB3_HUMAN (Q9H361) Polyadenylate-binding protein 3
(Poly(A)-binding protein 3) (PABP 3) (Testis-specific
poly(A)-binding protein)
Length = 631
Score = 120 bits (300), Expect = 7e-27
Identities = 60/175 (34%), Positives = 108/175 (61%), Gaps = 5/175 (2%)
Query: 25 ATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIK 84
A+ YVG+L P ++E +L+E F AGP++++ + +D +T+ Y +V F+ +DA++A+
Sbjct: 11 ASLYVGDLHPDVTEAMLYEKFSPAGPILSIRICRDLITSGSSNYAYVNFQHTKDAEHALD 70
Query: 85 VLNMIKLYGKPIRVNKASQDKKSLDVG-ANLFIGNLDPDVDEKLLYDTFSAFGVIVTNPK 143
+N + GKP+R+ + +D G N+F+ NLD ++ K LYDT SAFG I++
Sbjct: 71 TMNFDVIKGKPVRIMWSQRDPSLRKSGVGNIFVKNLDKSINNKALYDTVSAFGNILSCNV 130
Query: 144 IMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITV-SYAYKKDTKGE 197
+ D S+G+GF+ +++ EA++ AI+ MNG L R++ V + +K+ + E
Sbjct: 131 VC---DENGSKGYGFVHFETHEAAERAIKKMNGMLLNGRKVFVGQFKSRKEREAE 182
Score = 98.2 bits (243), Expect = 3e-20
Identities = 83/292 (28%), Positives = 133/292 (45%), Gaps = 36/292 (12%)
Query: 19 AERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEED 78
A + Y+ N + +E L +LF + GP ++V V D + + +G+GFV F ED
Sbjct: 185 ARAKEFPNVYIKNFGEDMDDERLKDLFGKFGPALSVKVMTDE-SGKSKGFGFVSFERHED 243
Query: 79 ADYAIKVLNMIKLYGKPIRVNKAS-----------------QDKKSLDVGANLFIGNLDP 121
A A+ +N +L GK I V +A QD+ + NL++ NLD
Sbjct: 244 AQKAVDEMNGKELNGKQIYVGRAQKKVERQTELKRTFEQMKQDRITRYQVVNLYVKNLDD 303
Query: 122 DVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCN 181
+D++ L FS FG I T+ K+M + G S+GFGF+ + S E + A+ MNG+ +
Sbjct: 304 GIDDERLRKAFSPFGTI-TSAKVMMEG--GRSKGFGFVCFSSPEEATKAVTEMNGRIVAT 360
Query: 182 RQITVSYAYKKDTK-GERHGTPAERVLAASNPTAQKSRPHTLFASGPPSLPNAPQANGTI 240
+ + V+ A +K+ + +R+ + Q++ P F + P N A
Sbjct: 361 KPLYVALAQRKEERQAYLTNEYMQRMASVRAVPNQRAPPSGYFMTAVPQTQN--HAAYYP 418
Query: 241 PAPV-----PPRPFANGVAPPPIH----VIQPPPPQAPAFQPMQMPPPGQQV 283
P+ + PR A G P P I+P P+ P F M+ P QV
Sbjct: 419 PSQIARLRPSPRWTAQGARPHPFQNKPSAIRPGAPRVP-FSTMR--PASSQV 467
Score = 95.5 bits (236), Expect = 2e-19
Identities = 60/172 (34%), Positives = 100/172 (57%), Gaps = 11/172 (6%)
Query: 28 YVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIKVLN 87
+V NLD I+ + L++ G +++ V D N +GYGFV F + E A+ AIK +N
Sbjct: 102 FVKNLDKSINNKALYDTVSAFGNILSCNVVCDE--NGSKGYGFVHFETHEAAERAIKKMN 159
Query: 88 MIKLYGKPIRVNK-ASQDKKSLDVGA------NLFIGNLDPDVDEKLLYDTFSAFGVIVT 140
+ L G+ + V + S+ ++ ++GA N++I N D+D++ L D F FG ++
Sbjct: 160 GMLLNGRKVFVGQFKSRKEREAELGARAKEFPNVYIKNFGEDMDDERLKDLFGKFGPALS 219
Query: 141 NPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAYKK 192
K+M D ++G S+GFGF+S++ E + A++ MNG+ L +QI V A KK
Sbjct: 220 -VKVMTD-ESGKSKGFGFVSFERHEDAQKAVDEMNGKELNGKQIYVGRAQKK 269
>PAB5_ARATH (Q05196) Polyadenylate-binding protein 5
(Poly(A)-binding protein 5) (PABP 5)
Length = 668
Score = 119 bits (297), Expect = 2e-26
Identities = 65/188 (34%), Positives = 114/188 (60%), Gaps = 4/188 (2%)
Query: 9 VGANLLGQHSAERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGY 68
V A + + + +++ YVG+LDP ++E L +LF Q PV N+ V +D +T++ GY
Sbjct: 29 VAAAAAAAEALQTHPNSSLYVGDLDPSVNESHLLDLFNQVAPVHNLRVCRD-LTHRSLGY 87
Query: 69 GFVEFRSEEDADYAIKVLNMIKLYGKPIRVNKASQDKKS-LDVGANLFIGNLDPDVDEKL 127
+V F + EDA A++ LN + +PIR+ +++D + L N+FI NLD +D K
Sbjct: 88 AYVNFANPEDASRAMESLNYAPIRDRPIRIMLSNRDPSTRLSGKGNVFIKNLDASIDNKA 147
Query: 128 LYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVS 187
LY+TFS+FG I+ + K+ D G S+G+GF+ ++ E + +AI+ +NG L ++Q+ V
Sbjct: 148 LYETFSSFGTIL-SCKVAMDV-VGRSKGYGFVQFEKEETAQAAIDKLNGMLLNDKQVFVG 205
Query: 188 YAYKKDTK 195
+ ++ +
Sbjct: 206 HFVRRQDR 213
Score = 89.4 bits (220), Expect = 1e-17
Identities = 62/177 (35%), Positives = 95/177 (53%), Gaps = 10/177 (5%)
Query: 28 YVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIKVLN 87
++ NLD I + L+E F G +++ V D V + +GYGFV+F EE A AI LN
Sbjct: 135 FIKNLDASIDNKALYETFSSFGTILSCKVAMD-VVGRSKGYGFVQFEKEETAQAAIDKLN 193
Query: 88 MIKLYGKPIRVNK--ASQDKKSLDVGA-----NLFIGNLDPDVDEKLLYDTFSAFGVIVT 140
+ L K + V QD+ + GA N+++ NL ++ + L TF +G I +
Sbjct: 194 GMLLNDKQVFVGHFVRRQDRARSESGAVPSFTNVYVKNLPKEITDDELKKTFGKYGDI-S 252
Query: 141 NPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAYKKDTKGE 197
+ +M+D +GNSR FGF+++ S EA+ A+E MNG L + V A KK + E
Sbjct: 253 SAVVMKD-QSGNSRSFGFVNFVSPEAAAVAVEKMNGISLGEDVLYVGRAQKKSDREE 308
Score = 84.7 bits (208), Expect = 3e-16
Identities = 88/305 (28%), Positives = 139/305 (44%), Gaps = 43/305 (14%)
Query: 28 YVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIKVLN 87
YV NL +I+++ L + F + G + + V KD+ N + +GFV F S E A A++ +N
Sbjct: 228 YVKNLPKEITDDELKKTFGKYGDISSAVVMKDQSGNS-RSFGFVNFVSPEAAAVAVEKMN 286
Query: 88 MIKLYGKPI----RVNKASQDKKSLD--------------VGANLFIGNLDPDVDEKLLY 129
I L G+ + R K S ++ L G+NL++ NLD V+++ L
Sbjct: 287 GISL-GEDVLYVGRAQKKSDREEELRRKFEQERISRFEKLQGSNLYLKNLDDSVNDEKLK 345
Query: 130 DTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYA 189
+ FS +G VT+ K+M + G SRGFGF++Y + E + A++ MNG+ + + + V+ A
Sbjct: 346 EMFSEYGN-VTSCKVMMNSQ-GLSRGFGFVAYSNPEEALLAMKEMNGKMIGRKPLYVALA 403
Query: 190 YKKDTKGERHGTPAERVLAASNPTAQKSRP------HTLFASGPPSLPNAPQANG-TIPA 242
+K+ ER +P P H GP S P+ P G
Sbjct: 404 QRKE---ERQAHLQSLFTQIRSPGTMSPVPSPMSGFHHHPPGGPMSGPHHPMFIGHNGQG 460
Query: 243 PVPPRPFANGVAPPPIHVIQP---PPPQAPAFQPMQMPPPGQQV--------WHQQQQGQ 291
VP +P G + ++P PP F + PG +V QQ Q Q
Sbjct: 461 LVPSQPMGYGYQVQFMPGMRPGAGPPNFMMPFPLQRQTQPGPRVGFRRGANNMQQQFQQQ 520
Query: 292 PMMQQ 296
M+QQ
Sbjct: 521 QMLQQ 525
Score = 42.4 bits (98), Expect = 0.002
Identities = 26/87 (29%), Positives = 48/87 (54%), Gaps = 1/87 (1%)
Query: 20 ERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDA 79
E+ Q + Y+ NLD +++E L E+F + G V + V + +G+GFV + + E+A
Sbjct: 323 EKLQGSNLYLKNLDDSVNDEKLKEMFSEYGNVTSCKVMMNS-QGLSRGFGFVAYSNPEEA 381
Query: 80 DYAIKVLNMIKLYGKPIRVNKASQDKK 106
A+K +N + KP+ V A + ++
Sbjct: 382 LLAMKEMNGKMIGRKPLYVALAQRKEE 408
>PAB3_ARATH (O64380) Polyadenylate-binding protein 3
(Poly(A)-binding protein 3) (PABP 3)
Length = 660
Score = 117 bits (294), Expect = 4e-26
Identities = 61/187 (32%), Positives = 114/187 (60%), Gaps = 4/187 (2%)
Query: 20 ERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDA 79
+ + +++ Y G+LDP+++E L++LF VV+V V +D+ + GY ++ F + DA
Sbjct: 44 QTHPNSSLYAGDLDPKVTEAHLFDLFKHVANVVSVRVCRDQ-NRRSLGYAYINFSNPNDA 102
Query: 80 DYAIKVLNMIKLYGKPIRVNKASQDKKS-LDVGANLFIGNLDPDVDEKLLYDTFSAFGVI 138
A++ LN L+ +PIR+ +++D + L N+FI NLD +D K L++TFS+FG I
Sbjct: 103 YRAMEALNYTPLFDRPIRIMLSNRDPSTRLSGKGNIFIKNLDASIDNKALFETFSSFGTI 162
Query: 139 VTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAYKKDTKGER 198
++ K+ D TG S+G+GF+ ++ E++ +AI+ +NG + ++Q+ V + ++ +
Sbjct: 163 LSC-KVAMDV-TGRSKGYGFVQFEKEESAQAAIDKLNGMLMNDKQVFVGHFIRRQERARD 220
Query: 199 HGTPAER 205
TP R
Sbjct: 221 ENTPTPR 227
Score = 96.7 bits (239), Expect = 9e-20
Identities = 63/177 (35%), Positives = 96/177 (53%), Gaps = 10/177 (5%)
Query: 28 YVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIKVLN 87
++ NLD I + L+E F G +++ V D VT + +GYGFV+F EE A AI LN
Sbjct: 139 FIKNLDASIDNKALFETFSSFGTILSCKVAMD-VTGRSKGYGFVQFEKEESAQAAIDKLN 197
Query: 88 MIKLYGKPI-------RVNKASQDKKSLDVGANLFIGNLDPDVDEKLLYDTFSAFGVIVT 140
+ + K + R +A + N+++ NL ++ E L TF FGVI +
Sbjct: 198 GMLMNDKQVFVGHFIRRQERARDENTPTPRFTNVYVKNLPKEIGEDELRKTFGKFGVI-S 256
Query: 141 NPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAYKKDTKGE 197
+ +MRD +GNSR FGF++++ EA+ SA+E MNG L + + V A KK + E
Sbjct: 257 SAVVMRD-QSGNSRCFGFVNFECTEAAASAVEKMNGISLGDDVLYVGRAQKKSEREE 312
Score = 92.4 bits (228), Expect = 2e-18
Identities = 92/332 (27%), Positives = 144/332 (42%), Gaps = 36/332 (10%)
Query: 28 YVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIKVLN 87
YV NL +I E+ L + F + G + + V +D+ N + +GFV F E A A++ +N
Sbjct: 232 YVKNLPKEIGEDELRKTFGKFGVISSAVVMRDQSGNS-RCFGFVNFECTEAAASAVEKMN 290
Query: 88 MIKLYGKPI---RVNKASQDKKSL--------------DVGANLFIGNLDPDVDEKLLYD 130
I L + R K S+ ++ L GANL++ NLD VD++ L +
Sbjct: 291 GISLGDDVLYVGRAQKKSEREEELRRKFEQERINRFEKSQGANLYLKNLDDSVDDEKLKE 350
Query: 131 TFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAY 190
FS +G VT+ K+M +P G SRGFGF++Y + E + A+ MNG+ + + + ++ A
Sbjct: 351 MFSEYGN-VTSSKVMLNPQ-GMSRGFGFVAYSNPEEALRALSEMNGKMIGRKPLYIALAQ 408
Query: 191 KKDTKGERHGTPAERVLAASNPTAQKSRPHTLFASGPPSLPNAPQANGTIPAPVPPRPFA 250
+K+ + H + A P + P GPP Q NG + VP +P
Sbjct: 409 RKEDR-RAHLQALFSQIRAPGPMSGFHHPPGGPMPGPPQHMYVGQ-NGA--SMVPSQPIG 464
Query: 251 NGVAP-----------PPIHVIQPPPPQAPAFQPMQMPPPGQQVWHQQQQGQPMMQQGMP 299
G P P ++ P + P P G Q Q Q +M +
Sbjct: 465 YGFQPQFMPGMRPGSGPGNFIVPYPLQRQPQTGPRMGFRRGATNVQQHIQQQQLMHRNPS 524
Query: 300 PPMQQFR-PPHSMQMPPPPPPQGMQGPPRPLP 330
P M+ + PQG+ P PLP
Sbjct: 525 PGMRYMNGASNGRNGMDSSVPQGILPPIIPLP 556
Score = 41.6 bits (96), Expect = 0.003
Identities = 25/84 (29%), Positives = 45/84 (52%), Gaps = 1/84 (1%)
Query: 20 ERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDA 79
E++Q A Y+ NLD + +E L E+F + G V + V + +G+GFV + + E+A
Sbjct: 327 EKSQGANLYLKNLDDSVDDEKLKEMFSEYGNVTSSKVMLN-PQGMSRGFGFVAYSNPEEA 385
Query: 80 DYAIKVLNMIKLYGKPIRVNKASQ 103
A+ +N + KP+ + A +
Sbjct: 386 LRALSEMNGKMIGRKPLYIALAQR 409
>PABP_YEAST (P04147) Polyadenylate-binding protein, cytoplasmic and
nuclear (Poly(A)-binding protein) (PABP) (ARS consensus
binding protein ACBP-67) (Polyadenylate tail-binding
protein)
Length = 576
Score = 117 bits (293), Expect = 5e-26
Identities = 65/167 (38%), Positives = 100/167 (58%), Gaps = 3/167 (1%)
Query: 22 NQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADY 81
N A+ YVG+L+P +SE L+++F G V ++ V +D +T GY +V F E
Sbjct: 34 NSSASLYVGDLEPSVSEAHLYDIFSPIGSVSSIRVCRDAITKTSLGYAYVNFNDHEAGRK 93
Query: 82 AIKVLNMIKLYGKPIRVNKASQDKKSLDVGA-NLFIGNLDPDVDEKLLYDTFSAFGVIVT 140
AI+ LN + G+ R+ + +D G+ N+FI NL PD+D K LYDTFS FG I++
Sbjct: 94 AIEQLNYTPIKGRLCRIMWSQRDPSLRKKGSGNIFIKNLHPDIDNKALYDTFSVFGDILS 153
Query: 141 NPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVS 187
+ KI D + G S+GFGF+ ++ A+ AI+A+NG L ++I V+
Sbjct: 154 S-KIATD-ENGKSKGFGFVHFEEEGAAKEAIDALNGMLLNGQEIYVA 198
Score = 106 bits (265), Expect = 9e-23
Identities = 90/305 (29%), Positives = 140/305 (45%), Gaps = 57/305 (18%)
Query: 28 YVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIKVLN 87
YV N++ + ++E ELF + GP+V+ + KD + +G+GFV + EDA A++ LN
Sbjct: 221 YVKNINSETTDEQFQELFAKFGPIVSASLEKD-ADGKLKGFGFVNYEKHEDAVKAVEALN 279
Query: 88 MIKLYGKPIRVNKASQDKKSLDV-----------------GANLFIGNLDPDVDEKLLYD 130
+L G+ + V +A + + + V G NLF+ NLD VD++ L +
Sbjct: 280 DSELNGEKLYVGRAQKKNERMHVLKKQYEAYRLEKMAKYQGVNLFVKNLDDSVDDEKLEE 339
Query: 131 TFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAY 190
F+ +G I T+ K+MR + G S+GFGF+ + + E + AI N Q + + + V+ A
Sbjct: 340 EFAPYGTI-TSAKVMRT-ENGKSKGFGFVCFSTPEEATKAITEKNQQIVAGKPLYVAIAQ 397
Query: 191 KKDTKGERHGTPAERVLAASNPTAQKSRPHTLFASGPPSLPNAPQANGTIPAPVPPRPFA 250
+KD R A+++ A + Q+ + A A +P P P
Sbjct: 398 RKDV---RRSQLAQQIQARNQMRYQQ------------ATAAAAAAAAGMPGQFMP-PMF 441
Query: 251 NGVAPPPIHVIQPPPPQAPAFQPMQMPPPGQQVWHQQQQGQPMMQQGMPPPMQQFRPPHS 310
GV PP P PQ QM P G G P + GMPP QFR
Sbjct: 442 YGVMPPRGVPFNGPNPQ-------QMNPMG---------GMP--KNGMPP---QFRNGPV 480
Query: 311 MQMPP 315
+PP
Sbjct: 481 YGVPP 485
Score = 80.1 bits (196), Expect = 9e-15
Identities = 52/185 (28%), Positives = 94/185 (50%), Gaps = 10/185 (5%)
Query: 18 SAERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEE 77
S + ++ NL P I + L++ F G +++ + D + +G+GFV F E
Sbjct: 118 SLRKKGSGNIFIKNLHPDIDNKALYDTFSVFGDILSSKIATDE-NGKSKGFGFVHFEEEG 176
Query: 78 DADYAIKVLNMIKLYGKPI-------RVNKASQDKKSLDVGANLFIGNLDPDVDEKLLYD 130
A AI LN + L G+ I R + SQ +++ NL++ N++ + ++ +
Sbjct: 177 AAKEAIDALNGMLLNGQEIYVAPHLSRKERDSQLEETKAHYTNLYVKNINSETTDEQFQE 236
Query: 131 TFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAY 190
F+ FG IV+ + +D D G +GFGF++Y+ E + A+EA+N L ++ V A
Sbjct: 237 LFAKFGPIVS-ASLEKDAD-GKLKGFGFVNYEKHEDAVKAVEALNDSELNGEKLYVGRAQ 294
Query: 191 KKDTK 195
KK+ +
Sbjct: 295 KKNER 299
Score = 52.0 bits (123), Expect = 2e-06
Identities = 26/74 (35%), Positives = 45/74 (60%), Gaps = 1/74 (1%)
Query: 102 SQDKKSLDVGANLFIGNLDPDVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISY 161
S+ + + A+L++G+L+P V E LYD FS G V++ ++ RD T S G+ ++++
Sbjct: 27 SESQSVENSSASLYVGDLEPSVSEAHLYDIFSPIG-SVSSIRVCRDAITKTSLGYAYVNF 85
Query: 162 DSFEASDSAIEAMN 175
+ EA AIE +N
Sbjct: 86 NDHEAGRKAIEQLN 99
>PABP_DROME (P21187) Polyadenylate-binding protein (Poly(A)-binding
protein) (PABP)
Length = 632
Score = 114 bits (286), Expect = 3e-25
Identities = 61/163 (37%), Positives = 104/163 (63%), Gaps = 3/163 (1%)
Query: 25 ATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIK 84
A+ YVG+L ++E L++ F AGPV+++ V +D +T + GY +V F+ DA+ A+
Sbjct: 2 ASLYVGDLPQDVNESGLFDKFSSAGPVLSIRVCRDVITRRSLGYAYVNFQQPADAERALD 61
Query: 85 VLNMIKLYGKPIRVNKASQDKKSLDVG-ANLFIGNLDPDVDEKLLYDTFSAFGVIVTNPK 143
+N + KPIR+ + +D G N+FI NLD +D K +YDTFSAFG I+ + K
Sbjct: 62 TMNFDLVRNKPIRIMWSQRDPSLRRSGVGNVFIKNLDRAIDNKAIYDTFSAFGNIL-SCK 120
Query: 144 IMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITV 186
+ D + GNS+G+GF+ +++ EA++++I+ +NG L +++ V
Sbjct: 121 VATD-EKGNSKGYGFVHFETEEAANTSIDKVNGMLLNGKKVYV 162
Score = 88.2 bits (217), Expect = 3e-17
Identities = 57/182 (31%), Positives = 103/182 (56%), Gaps = 11/182 (6%)
Query: 18 SAERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEE 77
S R+ ++ NLD I + +++ F G +++ V D N +GYGFV F +EE
Sbjct: 83 SLRRSGVGNVFIKNLDRAIDNKAIYDTFSAFGNILSCKVATDEKGNS-KGYGFVHFETEE 141
Query: 78 DADYAIKVLNMIKLYGKPIRVNKASQDKKSLDVG------ANLFIGNLDPDVDEKLLYDT 131
A+ +I +N + L GK + V K +K ++G N+++ N D D++ L +
Sbjct: 142 AANTSIDKVNGMLLNGKKVYVGKFIP-RKEQELGEKAKLFTNVYVKNFTEDFDDEKLKEF 200
Query: 132 FSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLC-NRQITVSYAY 190
F +G I T+ K+M D G S+GFGF+++++ EA+++A++A+NG+ + + + V+ A
Sbjct: 201 FEPYGKI-TSYKVMSKED-GKSKGFGFVAFETTEAAEAAVQALNGKDMGEGKSLYVARAQ 258
Query: 191 KK 192
KK
Sbjct: 259 KK 260
Score = 74.3 bits (181), Expect = 5e-13
Identities = 52/186 (27%), Positives = 91/186 (47%), Gaps = 21/186 (11%)
Query: 28 YVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIKVLN 87
YV N +E L E F G + + Y + + +G+GFV F + E A+ A++ LN
Sbjct: 184 YVKNFTEDFDDEKLKEFFEPYGKITS-YKVMSKEDGKSKGFGFVAFETTEAAEAAVQALN 242
Query: 88 MIKL-YGKPIRVNKAS-----------------QDKKSLDVGANLFIGNLDPDVDEKLLY 129
+ GK + V +A Q + G NL++ NLD +D+ L
Sbjct: 243 GKDMGEGKSLYVARAQKKAERQQELKRKFEELKQKRHESVFGVNLYVKNLDDTIDDDRLR 302
Query: 130 DTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYA 189
FS +G I T+ K+M D + G S+GFGF+ ++ + A+ +NG+ + ++ + V+ A
Sbjct: 303 IAFSPYGNI-TSAKVMTDEE-GRSKGFGFVCFNPESEATCAVTELNGRVVGSKPLYVALA 360
Query: 190 YKKDTK 195
+K+ +
Sbjct: 361 QRKEER 366
>FCA_ARATH (O04425) Flowering time control protein FCA
Length = 747
Score = 114 bits (286), Expect = 3e-25
Identities = 116/393 (29%), Positives = 169/393 (42%), Gaps = 55/393 (13%)
Query: 15 GQHSAERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFR 74
G ++R+ +VG++ +EE + F Q G V+ V + KD+ T Q QG FV++
Sbjct: 110 GTDVSDRSSTVKLFVGSVPRTATEEEIRPYFEQHGNVLEVALIKDKRTGQQQGCCFVKYA 169
Query: 75 SEEDADYAIKVL-NMIKLYG--KPIRVNKASQDKKSL-DVGANLFIGNLDPDVDEKLLYD 130
+ +DAD AI+ L N I L G P++V A +++ + + LF+G+L+ EK + +
Sbjct: 170 TSKDADRAIRALHNQITLPGGTGPVQVRYADGERERIGTLEFKLFVGSLNKQATEKEVEE 229
Query: 131 TFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYL---CNRQITVS 187
F FG V + +MRD + SRG GF+ Y S E + +AI+ +NG Y CN+ + V
Sbjct: 230 IFLQFG-HVEDVYLMRD-EYRQSRGCGFVKYSSKETAMAAIDGLNGTYTMRGCNQPLIVR 287
Query: 188 YAYKKDTKGERHGTPAERVLAASNPTAQKSRPHTLFASGPPSLPNAPQANGTIPAPVPPR 247
+A K K A V S P Q ASGP N ++G + P R
Sbjct: 288 FAEPKRPKPGESREMAPPVGLGSGPRFQ--------ASGPRPTSNFGDSSGDVSHTNPWR 339
Query: 248 P-FANGVAPPPIHVIQPPPPQAPAFQPMQMPPPGQQVWHQQQQGQPMMQQGMPP--PM-- 302
P + V PP I+ P P PGQ QG P+ G+PP P+
Sbjct: 340 PATSRNVGPPSNTGIRGAGSDFP-------PKPGQATL-PSNQGGPLGGYGVPPLNPLPV 391
Query: 303 ----------QQFRPPHSMQMPPPPPPQGMQGPPRPLPPPSVMAGQQQVWRPPPPPQQQG 352
QQ R P P QG P PL P + G Q P Q
Sbjct: 392 PGVSSSATLQQQNRAAGQHITPLKKPLHSPQGLPLPLRPQTNFPGAQ------APLQNPY 445
Query: 353 GHPMGYPQNSMPP---------PPPPNHHGMRP 376
+ P + +PP P P + +RP
Sbjct: 446 AYSSQLPTSQLPPQQNISRATAPQTPLNINLRP 478
>PAB5_PONPY (P60050) Polyadenylate-binding protein 5
(Poly(A)-binding protein 5) (PABP 5)
Length = 382
Score = 113 bits (283), Expect = 7e-25
Identities = 62/163 (38%), Positives = 96/163 (58%), Gaps = 4/163 (2%)
Query: 25 ATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIK 84
A YVG+LDP ++E++L++ F AGP+ + +D VT GYG+V FR DA++A+
Sbjct: 18 AALYVGDLDPDVTEDMLYKKFRPAGPLRFTRICRDPVTRSPLGYGYVNFRFPADAEWALN 77
Query: 85 VLNMIKLYGKPIRVNKASQDKKSLDVG-ANLFIGNLDPDVDEKLLYDTFSAFGVIVTNPK 143
+N + GKP R+ + D + G N+FI NLD +D + L+ FSAFG I++
Sbjct: 78 TMNFDLINGKPFRLMWSQPDDRLRKSGVGNIFIKNLDKSIDNRALFYLFSAFGNILSCKV 137
Query: 144 IMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITV 186
+ D S+G+ ++ +DS A++ AI MNG L NRQ+ V
Sbjct: 138 VC---DDNGSKGYAYVHFDSLAAANRAIWHMNGVRLNNRQVYV 177
Score = 81.6 bits (200), Expect = 3e-15
Identities = 53/185 (28%), Positives = 96/185 (51%), Gaps = 21/185 (11%)
Query: 20 ERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDA 79
+R +V N+ I +E L ELF + GP +V V +D + + +G+GFV + + E A
Sbjct: 194 DRATFTNVFVKNIGDDIDDEKLKELFCEYGPTESVKVIRD-ASGKSKGFGFVRYETHEAA 252
Query: 80 DYAIKVLNMIKLYGKPIRVNKASQD-----------------KKSLDVGANLFIGNLDPD 122
A+ L+ + GK + V +A + +KS G ++I NLD
Sbjct: 253 QKAVLDLHGKSIDGKVLYVGRAQKKIERLAELRRRFERLRLKEKSRPPGVPIYIKNLDET 312
Query: 123 VDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNR 182
++++ L + FS+FG I + K+M + G +GFG + + SFE + A++ MNG+ + ++
Sbjct: 313 INDEKLKEEFSSFGSI-SRAKVMME--VGQGKGFGVVCFSSFEEATKAVDEMNGRIVGSK 369
Query: 183 QITVS 187
+ V+
Sbjct: 370 PLHVT 374
Score = 73.6 bits (179), Expect = 8e-13
Identities = 49/173 (28%), Positives = 93/173 (53%), Gaps = 12/173 (6%)
Query: 28 YVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIKVLN 87
++ NLD I L+ LF G +++ V D N +GY +V F S A+ AI +N
Sbjct: 109 FIKNLDKSIDNRALFYLFSAFGNILSCKVVCD--DNGSKGYAYVHFDSLAAANRAIWHMN 166
Query: 88 MIKLYGKPIRVNKAS-QDKKSLDVGA-------NLFIGNLDPDVDEKLLYDTFSAFGVIV 139
++L + + V + ++++ +V N+F+ N+ D+D++ L + F +G
Sbjct: 167 GVRLNNRQVYVGRFKFPEERAAEVRTRDRATFTNVFVKNIGDDIDDEKLKELFCEYGP-T 225
Query: 140 TNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAYKK 192
+ K++RD +G S+GFGF+ Y++ EA+ A+ ++G+ + + + V A KK
Sbjct: 226 ESVKVIRDA-SGKSKGFGFVRYETHEAAQKAVLDLHGKSIDGKVLYVGRAQKK 277
Score = 47.4 bits (111), Expect = 6e-05
Identities = 29/95 (30%), Positives = 50/95 (52%), Gaps = 3/95 (3%)
Query: 99 NKASQDKKSLDVGANLFIGNLDPDVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGF 158
N A + KK L A L++G+LDPDV E +LY F G + +I RDP T + G+G+
Sbjct: 7 NPAGKKKKYLK--AALYVGDLDPDVTEDMLYKKFRPAGPL-RFTRICRDPVTRSPLGYGY 63
Query: 159 ISYDSFEASDSAIEAMNGQYLCNRQITVSYAYKKD 193
+++ ++ A+ MN + + + ++ D
Sbjct: 64 VNFRFPADAEWALNTMNFDLINGKPFRLMWSQPDD 98
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.315 0.136 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 58,939,142
Number of Sequences: 164201
Number of extensions: 3499904
Number of successful extensions: 49336
Number of sequences better than 10.0: 2062
Number of HSP's better than 10.0 without gapping: 826
Number of HSP's successfully gapped in prelim test: 1278
Number of HSP's that attempted gapping in prelim test: 14733
Number of HSP's gapped (non-prelim): 10968
length of query: 379
length of database: 59,974,054
effective HSP length: 112
effective length of query: 267
effective length of database: 41,583,542
effective search space: 11102805714
effective search space used: 11102805714
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 67 (30.4 bits)
Medicago: description of AC139882.5


