

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139344.4 - phase: 0 /pseudo
(826 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
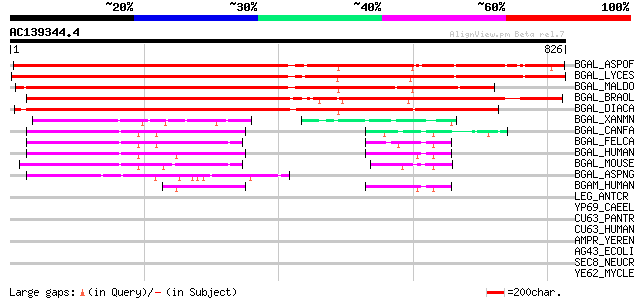
Score E
Sequences producing significant alignments: (bits) Value
BGAL_ASPOF (P45582) Beta-galactosidase precursor (EC 3.2.1.23) (... 945 0.0
BGAL_LYCES (P48980) Beta-galactosidase precursor (EC 3.2.1.23) (... 926 0.0
BGAL_MALDO (P48981) Beta-galactosidase precursor (EC 3.2.1.23) (... 825 0.0
BGAL_BRAOL (P49676) Beta-galactosidase precursor (EC 3.2.1.23) (... 820 0.0
BGAL_DIACA (Q00662) Putative beta-galactosidase precursor (EC 3.... 772 0.0
BGAL_XANMN (P48982) Beta-galactosidase precursor (EC 3.2.1.23) (... 167 8e-41
BGAL_CANFA (Q9TRY9) Beta-galactosidase precursor (EC 3.2.1.23) (... 158 5e-38
BGAL_FELCA (O19015) Beta-galactosidase precursor (EC 3.2.1.23) (... 156 2e-37
BGAL_HUMAN (P16278) Beta-galactosidase precursor (EC 3.2.1.23) (... 149 2e-35
BGAL_MOUSE (P23780) Beta-galactosidase precursor (EC 3.2.1.23) (... 145 3e-34
BGAL_ASPNG (P29853) Beta-galactosidase precursor (EC 3.2.1.23) (... 116 2e-25
BGAM_HUMAN (P16279) Beta-galactosidase-related protein precursor... 52 8e-06
LEG_ANTCR (P22031) D-galactoside-specific lectin (Sea urchin egg... 37 0.28
YP69_CAEEL (Q09218) Hypothetical protein B0495.9 in chromosome II 33 2.4
CU63_PANTR (Q68US5) Protein C21orf63 homolog precursor 33 4.0
CU63_HUMAN (P58658) Protein C21orf63 precursor (Protein PRED34) ... 33 4.0
AMPR_YEREN (P45461) HTH-type transcriptional activator ampR 33 4.0
AG43_ECOLI (P39180) Antigen 43 precursor (AG43) (Fluffing protein) 33 4.0
SEC8_NEUCR (Q9HE88) Probable exocyst complex component sec8 32 5.2
YE62_MYCLE (Q49682) Hypothetical UPF0051 protein ML0594 32 6.9
>BGAL_ASPOF (P45582) Beta-galactosidase precursor (EC 3.2.1.23)
(Lactase)
Length = 832
Score = 945 bits (2442), Expect = 0.0
Identities = 466/836 (55%), Positives = 580/836 (68%), Gaps = 40/836 (4%)
Query: 6 IVFVLLWFLGVYVPASFCSNVTYDHRALVIDGKRRVLMSGSIHYPRSTPQMWPDLIQKSK 65
++ ++ V+ P + ++VTYDH++++I+G+RR+L+SGSIHYPRSTP+MWPDLIQK+K
Sbjct: 7 LMLMVALLAAVWSPPAVTASVTYDHKSVIINGQRRILISGSIHYPRSTPEMWPDLIQKAK 66
Query: 66 DGGIDVIETYVFWNLHEPVRGQYNFEGRGDLVGFVKAVAAAGLYVHLRIGPYVCAEWNYG 125
DGG+DVI+TYVFWN HEP GQY F GR DLV F+K V AGLY HLRIGPYVCAEWN+G
Sbjct: 67 DGGLDVIQTYVFWNGHEPSPGQYYFGGRYDLVRFLKLVKQAGLYAHLRIGPYVCAEWNFG 126
Query: 126 GFPLWLHFIAGIKFRTNNEPFKAEMKRFTAKIVDMMKQENLYASQGGPIILSQIENEYGN 185
GFP+WL ++ GI FRT+N PFKA M +FT KIV MMK E LY +QGGPIILSQIENEYG
Sbjct: 127 GFPVWLKYVPGIHFRTDNGPFKAAMGKFTEKIVSMMKAEGLYETQGGPIILSQIENEYGP 186
Query: 186 IDTHDARAAKSYIDWAASMATSLDTGVPWIMCQQANAPDPIINTCNSFYCDQFTPNSDNK 245
++ +D A KSY +WAA MA L+TGVPW+MC+Q +APDP+INTCN FYCD F+PN DNK
Sbjct: 187 VEYYDGAAGKSYTNWAAKMAVGLNTGVPWVMCKQDDAPDPVINTCNGFYCDYFSPNKDNK 246
Query: 246 PKMWTENWSGWFLAFGGAVPYRPVEDLAFAVARFFQRGGTFQNYYMYHGGTNFGRTTGGP 305
PKMWTE W+GWF FGGAVP RP ED+AFAVARF Q+GG+F NYYMYHGGTNFGRT GGP
Sbjct: 247 PKMWTEAWTGWFTGFGGAVPQRPAEDMAFAVARFIQKGGSFINYYMYHGGTNFGRTAGGP 306
Query: 306 FISTSYDYDAPIDEYGDIRQPKWGHLKDLHKAIKLCEEALIASDPTITSPGPNLETAVYK 365
FISTSYDYDAPIDEYG +RQPKWGHL+DLHKAIKLCE AL++ +PTITS G N E+ VY+
Sbjct: 307 FISTSYDYDAPIDEYGLLRQPKWGHLRDLHKAIKLCEPALVSGEPTITSLGQNQESYVYR 366
Query: 366 TGAVCSAFLANIGMS-DATVTFNGNSYHLPGWSVSILPDCKNVVLNTAKVNTASMISSFA 424
+ + C+AFLAN ATVTFNG Y+LP WSVSILPDCK V NTA+V
Sbjct: 367 SKSSCAAFLANFNSRYYATVTFNGMHYNLPPWSVSILPDCKTTVFNTARV---------G 417
Query: 425 TESLKEKVDSLDSSSSGWSWISEPVGISTPDAFTKSGLLEQINTTADRSDYLWYSLSIVY 484
++ K+ L S W +E + FTK GL+EQ++TT DRSDYLWY+ +
Sbjct: 418 AQTTTMKMQYLGGFS--WKAYTEDTDALNDNTFTKDGLVEQLSTTWDRSDYLWYTTYVDI 475
Query: 485 EDN-----AGDQPVLHIESLGHALHAFVNGKLAGSKAGSSGNAKVNVDIPITLVTGKNTI 539
N G P L + S GHA+H F+NG+L+G+ GS N K+ L G N I
Sbjct: 476 AKNEEFLKTGKYPYLTVMSAGHAVHVFINGQLSGTAYGSLDNPKLTYSGSAKLWAGSNKI 535
Query: 540 DLLSLTVGLQNYGAFYDTVGAGITGPVILKGLKNGSSVDLTSQQWTYQVGLQGEFVGL-- 597
+LS++VGL N G ++T G+ GPV L GL G DL+ Q+WTYQ+GL GE + L
Sbjct: 536 SILSVSVGLPNVGNHFETWNTGVLGPVTLTGLNEGKR-DLSLQKWTYQIGLHGETLSLHS 594
Query: 598 --SSGNVGQWNSQSNLPANQPLTWYKTNFVAPSGSNPVAIDFTGMGKGEAWVNGQSIGRY 655
S NV +W S QPLTWYKT F AP G+ P+A+D MGKG+ W+NGQSIGRY
Sbjct: 595 LTGSSNV-EWGEASQ---KQPLTWYKTFFNAPPGNEPLALDMNTMGKGQIWINGQSIGRY 650
Query: 656 WPTYISPNSGCTDSCNYRGTYSASKCLKNCGKPSQTLYHVPRAWLKPDSNTFVLFEESGG 715
WP Y + SG SC+YRGTY+ KCL NCG+ SQ YHVPR+WL P N V+ EE GG
Sbjct: 651 WPAYKA--SGSCGSCDYRGTYNEKKCLSNCGEASQRWYHVPRSWLIPTGNFLVVLEEWGG 708
Query: 716 DPTKISFGTKQIESVCSHVTESHPPPVDTWNSNAESERKVGPVLSLECPYPNQAISSIKF 775
DPT IS + + SVC+ V E P +D W + A KV L C P Q +S IKF
Sbjct: 709 DPTGISMVKRSVASVCAEVEELQ-PTMDNWRTKAYGRPKV----HLSCD-PGQKMSKIKF 762
Query: 776 ASFGTPRGTCGNYNHGSCSSNRALSIVQKA-----CIGSSSCNIGVSINTF-GNPC 825
ASFGTP+GTCG+++ GSC ++++ ++ C+G C++ V+ F G+PC
Sbjct: 763 ASFGTPQGTCGSFSEGSCHAHKSYDAFEQEGLMQNCVGQEFCSVNVAPEVFGGDPC 818
>BGAL_LYCES (P48980) Beta-galactosidase precursor (EC 3.2.1.23)
(Lactase) (Acid beta-galactosidase)
(Exo-(1-->4)-beta-D-galactanase)
Length = 835
Score = 926 bits (2392), Expect = 0.0
Identities = 464/839 (55%), Positives = 573/839 (67%), Gaps = 33/839 (3%)
Query: 3 GTQIVFVLLWFLGVYVPASFCSNVTYDHRALVIDGKRRVLMSGSIHYPRSTPQMWPDLIQ 62
G + +L+ L ++V S V+YDH+A++++G+R++L+SGSIHYPRSTP+MWPDLIQ
Sbjct: 2 GFWMAMLLMLLLCLWVSCGIAS-VSYDHKAIIVNGQRKILISGSIHYPRSTPEMWPDLIQ 60
Query: 63 KSKDGGIDVIETYVFWNLHEPVRGQYNFEGRGDLVGFVKAVAAAGLYVHLRIGPYVCAEW 122
K+K+GG+DVI+TYVFWN HEP G+Y FE R DLV F+K V AGLYVHLRIGPY CAEW
Sbjct: 61 KAKEGGVDVIQTYVFWNGHEPEEGKYYFEERYDLVKFIKVVQEAGLYVHLRIGPYACAEW 120
Query: 123 NYGGFPLWLHFIAGIKFRTNNEPFKAEMKRFTAKIVDMMKQENLYASQGGPIILSQIENE 182
N+GGFP+WL ++ GI FRTNNEPFKA M++FT KIVDMMK E LY +QGGPIILSQIENE
Sbjct: 121 NFGGFPVWLKYVPGISFRTNNEPFKAAMQKFTTKIVDMMKAEKLYETQGGPIILSQIENE 180
Query: 183 YGNIDTHDARAAKSYIDWAASMATSLDTGVPWIMCQQANAPDPIINTCNSFYCDQFTPNS 242
YG ++ K Y +WAA MA L TGVPWIMC+Q + PDPIINTCN FYCD FTPN
Sbjct: 181 YGPMEWELGEPGKVYSEWAAKMAVDLGTGVPWIMCKQDDVPDPIINTCNGFYCDYFTPNK 240
Query: 243 DNKPKMWTENWSGWFLAFGGAVPYRPVEDLAFAVARFFQRGGTFQNYYMYHGGTNFGRTT 302
NKPKMWTE W+ WF FGG VPYRP ED+AFAVARF Q GG+F NYYMYHGGTNFGRT+
Sbjct: 241 ANKPKMWTEAWTAWFTEFGGPVPYRPAEDMAFAVARFIQTGGSFINYYMYHGGTNFGRTS 300
Query: 303 GGPFISTSYDYDAPIDEYGDIRQPKWGHLKDLHKAIKLCEEALIASDPTITSPGPNLETA 362
GGPFI+TSYDYDAP+DE+G +RQPKWGHLKDLH+AIKLCE AL++ DPT+TS G E
Sbjct: 301 GGPFIATSYDYDAPLDEFGSLRQPKWGHLKDLHRAIKLCEPALVSVDPTVTSLGNYQEAR 360
Query: 363 VYKT-GAVCSAFLANIGM-SDATVTFNGNSYHLPGWSVSILPDCKNVVLNTAKVNTASMI 420
V+K+ C+AFLAN S A V F Y+LP WS+SILPDCKN V NTA+V
Sbjct: 361 VFKSESGACAAFLANYNQHSFAKVAFGNMHYNLPPWSISILPDCKNTVYNTARV------ 414
Query: 421 SSFATESLKEKVDSLDSSSSGWSW--ISEPVGISTPDAFTKSGLLEQINTTADRSDYLWY 478
+S + K+ + S G+SW +E D FT GLLEQIN T D SDYLWY
Sbjct: 415 ---GAQSAQMKMTPV---SRGFSWESFNEDAASHEDDTFTVVGLLEQINITRDVSDYLWY 468
Query: 479 SLSIVYED-----NAGDQPVLHIESLGHALHAFVNGKLAGSKAGSSGNAKVNVDIPITLV 533
I + N+G+ P L + S GHALH FVNG+LAG+ GS N K+ I L
Sbjct: 469 MTDIEIDPTEGFLNSGNWPWLTVFSAGHALHVFVNGQLAGTVYGSLENPKLTFSNGINLR 528
Query: 534 TGKNTIDLLSLTVGLQNYGAFYDTVGAGITGPVILKGLKNGSSVDLTSQQWTYQVGLQGE 593
G N I LLS+ VGL N G ++T AG+ GPV L GL G+ DLT Q+W Y+VGL+GE
Sbjct: 529 AGVNKISLLSIAVGLPNVGPHFETWNAGVLGPVSLNGLNEGTR-DLTWQKWFYKVGLKGE 587
Query: 594 FV---GLSSGNVGQWNSQSNLPANQPLTWYKTNFVAPSGSNPVAIDFTGMGKGEAWVNGQ 650
+ LS +W S + QPL+WYKT F AP G+ P+A+D MGKG+ W+NGQ
Sbjct: 588 ALSLHSLSGSPSVEWVEGSLVAQKQPLSWYKTTFNAPDGNEPLALDMNTMGKGQVWINGQ 647
Query: 651 SIGRYWPTYISPNSGCTDSCNYRGTYSASKCLKNCGKPSQTLYHVPRAWLKPDSNTFVLF 710
S+GR+WP Y S SG CNY G + KCL NCG+ SQ YHVPR+WL P N V+F
Sbjct: 648 SLGRHWPAYKS--SGSCSVCNYTGWFDEKKCLTNCGEGSQRWYHVPRSWLYPTGNLLVVF 705
Query: 711 EESGGDPTKISFGTKQIESVCSHVTESHPPPVDTWNS--NAESERKVGPVLSLECPYPNQ 768
EE GGDP I+ ++I SVC+ + E P ++ W + + +R + P L+C P Q
Sbjct: 706 EEWGGDPYGITLVKREIGSVCADIYEWQPQLLN-WQRLVSGKFDRPLRPKAHLKCA-PGQ 763
Query: 769 AISSIKFASFGTPRGTCGNYNHGSCSSNRALSIVQKACIGSSSCNIGVSINTF-GNPCR 826
ISSIKFASFGTP G CGN+ GSC + R+ +K C+G SC++ V+ F G+PCR
Sbjct: 764 KISSIKFASFGTPEGVCGNFQQGSCHAPRSYDAFKKNCVGKESCSVQVTPENFGGDPCR 822
>BGAL_MALDO (P48981) Beta-galactosidase precursor (EC 3.2.1.23)
(Lactase) (Acid beta-galactosidase)
(Exo-(1-->4)-beta-D-galactanase)
Length = 731
Score = 825 bits (2130), Expect = 0.0
Identities = 398/722 (55%), Positives = 511/722 (70%), Gaps = 23/722 (3%)
Query: 9 VLLWFLGVYVPASFCSNVTYDHRALVIDGKRRVLMSGSIHYPRSTPQMWPDLIQKSKDGG 68
+LL F ++ AS ++V+YDH+A++I+G++R+L+SGSIHYPRSTP+MWPDLIQK+KDGG
Sbjct: 11 ILLLFSCIFSAAS--ASVSYDHKAIIINGQKRILISGSIHYPRSTPEMWPDLIQKAKDGG 68
Query: 69 IDVIETYVFWNLHEPVRGQYNFEGRGDLVGFVKAVAAAGLYVHLRIGPYVCAEWNYGGFP 128
+DVI+TYVFWN HEP G Y FE R DLV F+K V GL+V+LRIGPYVCAEWN+GGFP
Sbjct: 69 LDVIQTYVFWNGHEPSPGNYYFEERYDLVKFIKLVQQEGLFVNLRIGPYVCAEWNFGGFP 128
Query: 129 LWLHFIAGIKFRTNNEPFKAEMKRFTAKIVDMMKQENLYASQGGPIILSQIENEYGNIDT 188
+WL ++ GI FRT+NEPFKA M++FT KIV MMK E L+ +QGGPIILSQIENE+G ++
Sbjct: 129 VWLKYVPGIAFRTDNEPFKAAMQKFTEKIVSMMKAEKLFQTQGGPIILSQIENEFGPVEW 188
Query: 189 HDARAAKSYIDWAASMATSLDTGVPWIMCQQANAPDPIINTCNSFYCDQFTPNSDNKPKM 248
K+Y WAA MA LDTGVPWIMC+Q +APDP+I+TCN FYC+ F PN D KPKM
Sbjct: 189 EIGAPGKAYTKWAAQMAVGLDTGVPWIMCKQEDAPDPVIDTCNGFYCENFKPNKDYKPKM 248
Query: 249 WTENWSGWFLAFGGAVPYRPVEDLAFAVARFFQRGGTFQNYYMYHGGTNFGRTTGGPFIS 308
WTE W+GW+ FGGAVP RP ED+AF+VARF Q GG+F NYYMYHGGTNFGRT GGPF++
Sbjct: 249 WTEVWTGWYTEFGGAVPTRPAEDVAFSVARFIQSGGSFLNYYMYHGGTNFGRTAGGPFMA 308
Query: 309 TSYDYDAPIDEYGDIRQPKWGHLKDLHKAIKLCEEALIASDPTITSPGPNLETAVYKTGA 368
TSYDYDAP+DEYG R+PKWGHL+DLHKAIK CE AL++ DP++T G N E V+K+ +
Sbjct: 309 TSYDYDAPLDEYGLPREPKWGHLRDLHKAIKSCESALVSVDPSVTKLGSNQEAHVFKSES 368
Query: 369 VCSAFLANIGMS-DATVTFNGNSYHLPGWSVSILPDCKNVVLNTAKVNTASMISSFATES 427
C+AFLAN V+F G Y LP WS+SILPDCK V NTAKV ++S
Sbjct: 369 DCAAFLANYDAKYSVKVSFGGGQYDLPPWSISILPDCKTEVYNTAKV---------GSQS 419
Query: 428 LKEKVDSLDSSSSGWSWISEPVGISTPDAFTKSGLLEQINTTADRSDYLWYSLSI-VYED 486
+ ++ + S S+I E D T GL EQIN T D +DYLWY I + D
Sbjct: 420 SQVQMTPVHSGFPWQSFIEETTSSDETDTTTLDGLYEQINITRDTTDYLWYMTDITIGSD 479
Query: 487 NA----GDQPVLHIESLGHALHAFVNGKLAGSKAGSSGNAKVNVDIPITLVTGKNTIDLL 542
A G P+L I S GHAL+ F+NG+L+G+ GS N K++ + L +G N + LL
Sbjct: 480 EAFLKNGKSPLLTIFSAGHALNVFINGQLSGTVYGSLENPKLSFSQNVNLRSGINKLALL 539
Query: 543 SLTVGLQNYGAFYDTVGAGITGPVILKGLKNGSSVDLTSQQWTYQVGLQGEFVGL---SS 599
S++VGL N G ++T AG+ GP+ LKGL +G + D++ +WTY+ GL+GE +GL +
Sbjct: 540 SISVGLPNVGTHFETWNAGVLGPITLKGLNSG-TWDMSGWKWTYKTGLKGEALGLHTVTG 598
Query: 600 GNVGQWNSQSNLPANQPLTWYKTNFVAPSGSNPVAIDFTGMGKGEAWVNGQSIGRYWPTY 659
+ +W ++ QPLTWYK F AP G P+A+D MGKG+ W+NGQS+GR+WP Y
Sbjct: 599 SSSVEWVEGPSMAEKQPLTWYKATFNAPPGDAPLALDMGSMGKGQIWINGQSVGRHWPGY 658
Query: 660 ISPNSGCTDSCNYRGTYSASKCLKNCGKPSQTLYHVPRAWLKPDSNTFVLFEESGGDPTK 719
I+ S C D C+Y GTY KC +CG+PSQ YH+PR+WL P N V+FEE GGDP++
Sbjct: 659 IARGS-CGD-CSYAGTYDDKKCRTHCGEPSQRWYHIPRSWLTPTGNLLVVFEEWGGDPSR 716
Query: 720 IS 721
IS
Sbjct: 717 IS 718
>BGAL_BRAOL (P49676) Beta-galactosidase precursor (EC 3.2.1.23)
(Lactase)
Length = 828
Score = 820 bits (2119), Expect = 0.0
Identities = 411/820 (50%), Positives = 530/820 (64%), Gaps = 57/820 (6%)
Query: 26 VTYDHRALVIDGKRRVLMSGSIHYPRSTPQMWPDLIQKSKDGGIDVIETYVFWNLHEPVR 85
V++D RA+ IDG+RR+L+SGSIHYPRST MWPDLI K+KDGG+D IETYVFWN HEP R
Sbjct: 27 VSHDERAITIDGQRRILLSGSIHYPRSTSDMWPDLISKAKDGGLDTIETYVFWNAHEPSR 86
Query: 86 GQYNFEGRGDLVGFVKAVAAAGLYVHLRIGPYVCAEWNYGGFPLWLHFIAGIKFRTNNEP 145
QY+F G DLV F+K + +AGLY LRIGPYVCAEWNYGGFP+WLH + +KFRT N
Sbjct: 87 RQYDFSGNLDLVRFIKTIQSAGLYSVLRIGPYVCAEWNYGGFPVWLHNMPDMKFRTINPG 146
Query: 146 FKAEMKRFTAKIVDMMKQENLYASQGGPIILSQIENEYGNIDTHDARAAKSYIDWAASMA 205
F EM+ FT KIV+MMK+E+L+ASQGGPIIL+QIENEYGN+ + K+YIDW A+MA
Sbjct: 147 FMNEMQNFTTKIVNMMKEESLFASQGGPIILAQIENEYGNVISSYGAEGKAYIDWCANMA 206
Query: 206 TSLDTGVPWIMCQQANAPDPIINTCNSFYCDQFTPNSDNKPKMWTENWSGWFLAFGGAVP 265
SLD GVPWIMCQQ +AP P+I TCN FYCDQ+ P++ + PKMWTENW+GWF +GG P
Sbjct: 207 NSLDIGVPWIMCQQPHAPQPMIETCNGFYCDQYKPSNPSSPKMWTENWTGWFKNWGGKHP 266
Query: 266 YRPVEDLAFAVARFFQRGGTFQNYYMYHGGTNFGRTTGGPFISTSYDYDAPIDEYGDIRQ 325
YR EDLAF+VARFFQ GGTFQNYYMYHGGTNFGR GGP+I+TSYDYDAP+DEYG++ Q
Sbjct: 267 YRTAEDLAFSVARFFQTGGTFQNYYMYHGGTNFGRVAGGPYITTSYDYDAPLDEYGNLNQ 326
Query: 326 PKWGHLKDLHKAIKLCEEALIASDPTITSPGPNLETAVYKTGAVCSAFLANI-GMSDATV 384
PKWGHLK LH +K E+ L + + G ++ VY T S F+ N+ +DA V
Sbjct: 327 PKWGHLKQLHTLLKSMEKPLTYGNISTIDLGNSVTATVYSTNEKSSCFIGNVNATADALV 386
Query: 385 TFNGNSYHLPGWSVSILPDCKNVVLNTAKVNTASMISSFATESLKEKVDSLDSSSSGWSW 444
F G Y++P WSVS+LPDC NTA+VNT +S TE ++ + L W+W
Sbjct: 387 NFKGKDYNVPAWSVSVLPDCDKEAYNTARVNTQ---TSIITEDSCDEPEKLK-----WTW 438
Query: 445 ISEPVGISTPDAFTK-------SGLLEQINTTADRSDYLWYSLSIVYEDNAGDQPV---- 493
E +T K GL++Q + T D SDYLWY ++ V+ D P+
Sbjct: 439 RPE---FTTQKTILKGSGDLIAKGLVDQKDVTNDASDYLWY-MTRVHLDK--KDPIWSRN 492
Query: 494 --LHIESLGHALHAFVNGKLAGSKAGSSGNAKVNVDIPITLVTGKNTIDLLSLTVGLQNY 551
L + S H LHA+VNGK G++ + + LV G N + LLS++VGLQNY
Sbjct: 493 MSLRVHSNAHVLHAYVNGKYVGNQIVRDNKFDYRFEKKVNLVHGTNHLALLSVSVGLQNY 552
Query: 552 GAFYDTVGAGITGPVILKGLKNGSSV--DLTSQQWTYQVGLQG----EFVGLSSGNVGQW 605
G F+++ GI GPV L G K ++ DL+ QW Y++GL G F S+G+ +
Sbjct: 553 GPFFESGPTGINGPVKLVGYKGDETIEKDLSKHQWDYKIGLNGFNHKLFSMKSAGHHHRK 612
Query: 606 NSQSNLPANQPLTWYKTNFVAPSGSNPVAIDFTGMGKGEAWVNGQSIGRYWPTYISPNSG 665
S LPA++ L+WYK NF AP G +PV +D G+GKGE W+NGQSIGRYWP++ S + G
Sbjct: 613 WSTEKLPADRMLSWYKANFKAPLGKDPVIVDLNGLGKGEVWINGQSIGRYWPSFNSSDEG 672
Query: 666 CTDSCNYRGTYSASKCLKNCGKPSQTLYHVPRAWLKPDS-NTFVLFEESGGDPTKISFGT 724
CT+ C+YRG Y + KC CGKP+Q YHVPR++L NT LFEE GGDP+ + F T
Sbjct: 673 CTEECDYRGEYGSDKCAFMCGKPTQRWYHVPRSFLNDKGHNTITLFEEMGGDPSMVKFKT 732
Query: 725 KQIESVCSHVTESHPPPVDTWNSNAESERKVGPVLSLECPYPNQAISSIKFASFGTPRGT 784
VC+ E + +E N+ IS++KFASFG P G
Sbjct: 733 VVTGRVCAKAHEHN---------------------KVELSCNNRPISAVKFASFGNPSGQ 771
Query: 785 CGNYNHGSCSSNR-ALSIVQKACIGSSSCNIGVSINTFGN 823
CG++ GSC + A+ +V K C+G +C + VS + FG+
Sbjct: 772 CGSFAAGSCEGAKDAVKVVAKECVGKLNCTMNVSSHKFGS 811
>BGAL_DIACA (Q00662) Putative beta-galactosidase precursor (EC
3.2.1.23) (Lactase) (SR12 protein)
Length = 731
Score = 772 bits (1994), Expect = 0.0
Identities = 385/731 (52%), Positives = 494/731 (66%), Gaps = 27/731 (3%)
Query: 7 VFVLLWFLG-VYVPASFCSNVTYDHRALVIDGKRRVLMSGSIHYPRSTPQMWPDLIQKSK 65
VFVL+ + VY NV YD+RA+ I+ +RR+L+SGSIHYPRSTP+MWPD+I+K+K
Sbjct: 17 VFVLITLISCVY------GNVWYDYRAIKINDQRRILLSGSIHYPRSTPEMWPDIIEKAK 70
Query: 66 DGGIDVIETYVFWNLHEPVRGQYNFEGRGDLVGFVKAVAAAGLYVHLRIGPYVCAEWNYG 125
D +DVI+TYVFWN HEP G+Y FEGR DLV F+K + AGL+VHLRIGP+ CAEWN+G
Sbjct: 71 DSQLDVIQTYVFWNGHEPSEGKYYFEGRYDLVKFIKLIHQAGLFVHLRIGPFACAEWNFG 130
Query: 126 GFPLWLHFIAGIKFRTNNEPFKAEMKRFTAKIVDMMKQENLYASQGGPIILSQIENEYGN 185
GFP+WL ++ GI+FRT+N PFK +M+ FT KIVDMMK E L+ QGGPIIL+QIENEYG
Sbjct: 131 GFPVWLKYVPGIEFRTDNGPFKEKMQVFTTKIVDMMKAEKLFHWQGGPIILNQIENEYGP 190
Query: 186 IDTHDARAAKSYIDWAASMATSLDTGVPWIMCQQ-ANAPDPIINTCNSFYCDQFTPNSDN 244
++ K+Y WAA MA SL+ GVPWIMC+Q ++ PD +I+TCN FYC+ F P +
Sbjct: 191 VEWEIGAPGKAYTHWAAQMAQSLNAGVPWIMCKQDSDVPDNVIDTCNGFYCEGFVPKDKS 250
Query: 245 KPKMWTENWSGWFLAFGGAVPYRPVEDLAFAVARFFQRGGTFQNYYMYHGGTNFGRTTGG 304
KPKMWTENW+GW+ +G VPYRP ED+AF+VARF Q GG+F NYYM+HGGTNF TT G
Sbjct: 251 KPKMWTENWTGWYTEYGKPVPYRPAEDVAFSVARFIQNGGSFMNYYMFHGGTNF-ETTAG 309
Query: 305 PFISTSYDYDAPIDEYGDIRQPKWGHLKDLHKAIKLCEEALIASDPTITSPGPNLETAVY 364
F+STSYDYDAP+DEYG R+PK+ HLK+LHKAIK+CE AL++SD +T+ G N E VY
Sbjct: 310 RFVSTSYDYDAPLDEYGLPREPKYTHLKNLHKAIKMCEPALVSSDAKVTNLGSNQEAHVY 369
Query: 365 KTGA-VCSAFLANIGMS-DATVTFNGNSYHLPGWSVSILPDCKNVVLNTAKVNTASMISS 422
+ + C+AFLAN VTF+G + LP WS+SILPDCK V NTA+VN S
Sbjct: 370 SSNSGSCAAFLANYDPKWSVKVTFSGMEFELPAWSISILPDCKKEVYNTARVNEPS---- 425
Query: 423 FATESLKEKVDSLDSSSSGWSWISEPVGISTPDAFTKSGLLEQINTTADRSDYLWYSLSI 482
L K+ + S+ + S+ E +P F + L EQIN T D+SDYLWY +
Sbjct: 426 ---PKLHSKMTPVISNLNWQSYSDEVPTADSPGTFREKKLYEQINMTWDKSDYLWYMTDV 482
Query: 483 VYEDN-----AGDQPVLHIESLGHALHAFVNGKLAGSKAGSSGNAKVNVDIPITLVTGKN 537
V + N GD+P L + S GH LH FVNG+L G GS ++ + + G N
Sbjct: 483 VLDGNEGFLKKGDEPWLTVNSAGHVLHVFVNGQLQGHAYGSLAKPQLTFSQKVKMTAGVN 542
Query: 538 TIDLLSLTVGLQNYGAFYDTVGAGITGPVILKGLKNGSSVDLTSQQWTYQVGLQGEFVGL 597
I LLS VGL N G ++ G+ GPV L GL G+ DLT Q W+Y++G +GE +
Sbjct: 543 RISLLSAVVGLANVGWHFERYNQGVLGPVTLSGLNEGTR-DLTWQYWSYKIGTKGEEQQV 601
Query: 598 SSGNVGQWNSQSNLPA-NQPLTWYKTNFVAPSGSNPVAIDFTGMGKGEAWVNGQSIGRYW 656
+ G + Q PA QPL WYKT F AP G++P+A+D MGKG+AW+NGQSIGR+W
Sbjct: 602 YNSG-GSSHVQWGPPAWKQPLVWYKTTFDAPGGNDPLALDLGSMGKGQAWINGQSIGRHW 660
Query: 657 PTYISPNSGCTDSCNYRGTYSASKCLKNCGKPSQTLYHVPRAWLKPDSNTFVLFEESGGD 716
I+ S C D+CNY GTY+ +KCL +CGK SQ YHVPR+WL+P N V+FEE GGD
Sbjct: 661 SNNIAKGS-CNDNCNYAGTYTETKCLSDCGKSSQKWYHVPRSWLQPRGNLLVVFEEWGGD 719
Query: 717 PTKISFGTKQI 727
+S + I
Sbjct: 720 TKWVSLVKRTI 730
>BGAL_XANMN (P48982) Beta-galactosidase precursor (EC 3.2.1.23)
(Lactase)
Length = 598
Score = 167 bits (424), Expect = 8e-41
Identities = 122/348 (35%), Positives = 171/348 (49%), Gaps = 30/348 (8%)
Query: 34 VIDGKRRVLMSGSIHYPRSTPQMWPDLIQKSKDGGIDVIETYVFWNLHEPVRGQYNFEGR 93
V DGK L+SG+IH+ R W D +QK++ G++ +ETYVFWNL EP +GQ++F G
Sbjct: 38 VRDGKPYQLLSGAIHFQRIPRAYWKDRLQKARALGLNTVETYVFWNLVEPQQGQFDFSGN 97
Query: 94 GDLVGFVKAVAAAGLYVHLRIGPYVCAEWNYGGFPLWLHFIAGIKFRTNNEPFKAEMKRF 153
D+ FVK AA GL V LR GPY CAEW GG+P WL I+ R+ + F A + +
Sbjct: 98 NDVAAFVKEAAAQGLNVILRPGPYACAEWEAGGYPAWLFGKGNIRVRSRDPRFLAASQAY 157
Query: 154 TAKIVDMMKQENLYASQGGPIILSQIENEYGNIDTHDARAAKS---YIDWAASMATSLDT 210
+ + + L GGPII Q+ENEYG+ A A + Y+ A L T
Sbjct: 158 LDALAKQV--QPLLNHNGGPIIAVQVENEYGSYADDHAYMADNRAMYVKAGFDKAL-LFT 214
Query: 211 GVPWIMCQQANAPD--PIINTC-----NSFYCDQFTPNSDNKPKMWTENWSGWFLAFGGA 263
M PD ++N ++F D+ ++P+M E W+GWF +G
Sbjct: 215 SDGADMLANGTLPDTLAVVNFAPGEAKSAF--DKLIKFRPDQPRMVGEYWAGWFDHWG-- 270
Query: 264 VPYRPVEDLAFAVA-RFFQRGGTFQNYYMYHGGTNFGRTTGGPF----------ISTSYD 312
P+ + A + R G N YM+ GGT+FG G F +TSYD
Sbjct: 271 KPHAATDARQQAEEFEWILRQGHSANLYMFIGGTSFGFMNGANFQNNPSDHYAPQTTSYD 330
Query: 313 YDAPIDEYGDIRQPKWGHLKD-LHKAIKLCEEALIASDPTITSPGPNL 359
YDA +DE G PK+ ++D + + + AL A T T P L
Sbjct: 331 YDAILDEAGH-PTPKFALMRDAIARVTGVQPPALPAPITTTTLPATPL 377
Score = 45.8 bits (107), Expect = 5e-04
Identities = 56/238 (23%), Positives = 87/238 (36%), Gaps = 45/238 (18%)
Query: 435 LDSSSSGWSWISEPVGISTPDAFTKSGLLEQINTTADRSDYLWYSLSIVYEDNAGDQPVL 494
L S+S W + P+ I TP + G Y+ Y +I + L
Sbjct: 377 LRESASLWDNLPTPIAIDTPQPMEQFG---------QDYGYILYRTTIT----GPRKGPL 423
Query: 495 HIESLGHALHAFVNGKLAGSKAGSSGNAKVNVDIPITLVTGKNTIDLLSLTVGLQNYGAF 554
++ + +V+ + GS + V+IP G++T+D+L G NYG
Sbjct: 424 YLGDVRDVARVYVDQRPVGSVERRLQQVSLEVEIP----AGQHTLDVLVENSGRINYGTR 479
Query: 555 YDTVGAGITGPVILKGLKNGSSVDLTS-QQWTYQVGLQGEFVGLSSGNV-GQWNSQSNLP 612
AG+ PV+L S LT Q + + G + V G + L
Sbjct: 480 MADGRAGLVDPVLL------DSQQLTGWQAFPLPMRTPDSIRGWTGKAVQGPAFHRGTLR 533
Query: 613 ANQPLTWYKTNFVAPSGSNPVAIDFTGMGKGEAWVNGQSIGRYW------PTYISPNS 664
P Y +D GKG AW NG ++GR+W Y+ P+S
Sbjct: 534 IGTPTDTY--------------LDMRAFGKGFAWANGVNLGRHWNIGPQTALYLRPSS 577
>BGAL_CANFA (Q9TRY9) Beta-galactosidase precursor (EC 3.2.1.23)
(Lactase) (Acid beta-galactosidase) (Fragment)
Length = 662
Score = 158 bits (400), Expect = 5e-38
Identities = 111/340 (32%), Positives = 161/340 (46%), Gaps = 18/340 (5%)
Query: 26 VTYDHRALVIDGKRRVLMSGSIHYPRSTPQMWPDLIQKSKDGGIDVIETYVFWNLHEPVR 85
+ Y H + DG+ +SGSIHY R W D + K K G++ I+TYV WN HEP
Sbjct: 29 IDYSHNRFLKDGQPFRYISGSIHYSRVPRFYWKDRLLKMKMAGLNAIQTYVPWNFHEPQP 88
Query: 86 GQYNFEGRGDLVGFVKAVAAAGLYVHLRIGPYVCAEWNYGGFPLWLHFIAGIKFRTNNEP 145
GQY F G D+ F+K GL V LR GPY+CAEW+ GG P WL I R+++
Sbjct: 89 GQYQFSGEQDVEYFIKLAHELGLLVILRPGPYICAEWDMGGLPAWLLLKESIILRSSDPD 148
Query: 146 FKAEMKRFTAKIVDMMKQENLYASQGGPIILSQIENEYGNIDTHD-------ARAAKSYI 198
+ A + ++ ++ MK L GGPII Q+ENEYG+ T D + ++
Sbjct: 149 YLAAVDKWLGVLLPKMKP--LLYQNGGPIITMQVENEYGSYFTCDYDYLRFLQKLFHHHL 206
Query: 199 DWAASMATSLDTGVPWIMC---QQANAPDPIINTCNSFYCDQFTPNSDNK-PKMWTENWS 254
+ T+ ++ C Q A N Q S+ K P + +E ++
Sbjct: 207 GNDVLLFTTDGANEKFLQCGALQGLYATVDFGPGANITAAFQIQRKSEPKGPLVNSEFYT 266
Query: 255 GWFLAFGGAVPYRPVEDLAFAVARFFQRGGTFQNYYMYHGGTNFGRTTGG--PFIS--TS 310
GW +G E +A ++ G N YM+ GGTNF G P+ + TS
Sbjct: 267 GWLDHWGQPHSTVRTEVVASSLHDILAHGANV-NLYMFIGGTNFAYWNGANMPYQAQPTS 325
Query: 311 YDYDAPIDEYGDIRQPKWGHLKDLHKAIKLCEEALIASDP 350
YDYDAP+ E GD+ + + + + K K+ E + S P
Sbjct: 326 YDYDAPLSEAGDLTEKYFALREVIRKFEKVPEGFIPPSTP 365
Score = 52.0 bits (123), Expect = 6e-06
Identities = 65/236 (27%), Positives = 94/236 (39%), Gaps = 65/236 (27%)
Query: 530 ITL-VTGKN--TIDLLSLTVGLQNYGAFYD---------TVGAGITGPVILKGLKNGSSV 577
ITL +TGK T+DLL +G NYG + + T+G+ I ++ L +V
Sbjct: 456 ITLNITGKAGATLDLLVENMGRVNYGRYINDFKGLISNLTLGSSILTNWMIFPLNTEDAV 515
Query: 578 DLTSQQWTYQVGLQGEFVGLSSGNVGQWNSQSNLPANQPLTWYKTNFVAPSG----SNPV 633
++ G G G +S LPA +Y NF PSG
Sbjct: 516 R------SHLGGWHGPNNGRHDKTFAHRSSNYTLPA-----FYMGNFSIPSGIPDLPQDT 564
Query: 634 AIDFTGMGKGEAWVNGQSIGRYWPTYISPNSGCTDSCNYRGTYSASKCLKNCGKPSQTLY 693
I F G KG+ W+NG ++GRYWP RG P TL+
Sbjct: 565 FIQFPGWTKGQVWINGFNLGRYWPA--------------RG-------------PQMTLF 597
Query: 694 HVPRAWLKPDS-NTFVLFE-------ESGGDPTKISFGTKQIESVCSHVTESHPPP 741
VPR L + NT ++ E +SG + + F + + + + T HPPP
Sbjct: 598 -VPRHILVTSTPNTIMVLELEHAPCGDSGPEVCTVEFVDRPV--IGAPPTPGHPPP 650
>BGAL_FELCA (O19015) Beta-galactosidase precursor (EC 3.2.1.23)
(Lactase) (Acid beta-galactosidase)
Length = 669
Score = 156 bits (395), Expect = 2e-37
Identities = 111/336 (33%), Positives = 160/336 (47%), Gaps = 19/336 (5%)
Query: 26 VTYDHRALVIDGKRRVLMSGSIHYPRSTPQMWPDLIQKSKDGGIDVIETYVFWNLHEPVR 85
+ Y H + DG+ +SGSIHY R W D + K K G++ I+TYV WN HEP
Sbjct: 35 IDYGHNRFLKDGQPFRYISGSIHYFRVPRFYWKDRLLKMKMAGLNAIQTYVPWNFHEPQP 94
Query: 86 GQYNFEGRGDLVGFVKAVAAAGLYVHLRIGPYVCAEWNYGGFPLWLHFIAGIKFRTNNEP 145
GQY F G D+ F+K GL V LR GPY+CAEW+ GG P WL I R+++
Sbjct: 95 GQYQFSGEHDVEYFLKLAHELGLLVILRPGPYICAEWDMGGLPAWLLLKESIILRSSDPD 154
Query: 146 FKAEMKRFTAKIVDMMKQENLYASQGGPIILSQIENEYGNIDTHD-------ARAAKSYI 198
+ A + ++ ++ MK L GGPII Q+ENEYG+ T D R + ++
Sbjct: 155 YLAAVDKWLGVLLPKMKP--LLYQNGGPIITVQVENEYGSYFTCDYDYLRFLQRRFRDHL 212
Query: 199 DWAASMATSLDTGVPWIMC---QQANAPDPIINTCNSFYCDQFTPNSDNK-PKMWTENWS 254
+ T+ ++ C Q A N Q S+ + P + +E ++
Sbjct: 213 GGDVLLFTTDGAHEKFLQCGALQGIYATVDFGPDANITAAFQIQRKSEPRGPLVNSEFYT 272
Query: 255 GWFLAFGGAVPYRPVEDLAFAVARFFQRGGTFQNYYMYHGGTNFGRTTGG--PF--ISTS 310
GW +G E +A ++ G N YM+ GGTNF G P+ TS
Sbjct: 273 GWLDHWGQPHSRVRTEVVASSLHDVLAHGANV-NLYMFIGGTNFAYWNGANIPYQPQPTS 331
Query: 311 YDYDAPIDEYGDIRQPKWGHLKDLHKAIKLCEEALI 346
YDYDAP+ E GD+ K+ L+D+ + + E I
Sbjct: 332 YDYDAPLSEAGDLTD-KYFALRDVIRKFEKVPEGFI 366
Score = 48.9 bits (115), Expect = 5e-05
Identities = 45/145 (31%), Positives = 62/145 (42%), Gaps = 28/145 (19%)
Query: 530 ITL-VTGKN--TIDLLSLTVGLQNYGAFYDTVGAGITGPVILKGLKNGSSV--------- 577
ITL +TG+ T+DLL +G NYG + + ++ L GSSV
Sbjct: 462 ITLNITGQAGATLDLLVENMGRVNYGRYINDFKG------LISNLTLGSSVLTDWMIFPL 515
Query: 578 DLTSQQWTYQVGLQGEFVGLSSGNV-GQWNSQSNLPANQPLTWYKTNFVAPSG----SNP 632
D ++ G G G +S LPA +Y NF PSG
Sbjct: 516 DTEDAVRSHLGGWHGRNHGRQDNKAFAHHSSNYTLPA-----FYAGNFSIPSGIPDLPQD 570
Query: 633 VAIDFTGMGKGEAWVNGQSIGRYWP 657
I F+G KG+ W+NG ++GRYWP
Sbjct: 571 TFIQFSGWTKGQVWINGFNLGRYWP 595
>BGAL_HUMAN (P16278) Beta-galactosidase precursor (EC 3.2.1.23)
(Lactase) (Acid beta-galactosidase)
Length = 677
Score = 149 bits (377), Expect = 2e-35
Identities = 105/340 (30%), Positives = 160/340 (46%), Gaps = 18/340 (5%)
Query: 26 VTYDHRALVIDGKRRVLMSGSIHYPRSTPQMWPDLIQKSKDGGIDVIETYVFWNLHEPVR 85
+ Y + + DG+ +SGSIHY R W D + K K G++ I+TYV WN HEP
Sbjct: 34 IDYSRDSFLKDGQPFRYISGSIHYSRVPRFYWKDRLLKMKMAGLNAIQTYVPWNFHEPWP 93
Query: 86 GQYNFEGRGDLVGFVKAVAAAGLYVHLRIGPYVCAEWNYGGFPLWLHFIAGIKFRTNNEP 145
GQY F D+ F++ GL V LR GPY+CAEW GG P WL I R+++
Sbjct: 94 GQYQFSEDHDVEYFLRLAHELGLLVILRPGPYICAEWEMGGLPAWLLEKESILLRSSDPD 153
Query: 146 FKAEMKRFTAKIVDMMKQENLYASQGGPIILSQIENEYGNIDTHD-------ARAAKSYI 198
+ A + ++ ++ MK L GGP+I Q+ENEYG+ D + + ++
Sbjct: 154 YLAAVDKWLGVLLPKMKP--LLYQNGGPVITVQVENEYGSYFACDFDYLRFLQKRFRHHL 211
Query: 199 DWAASMATSLDTGVPWIMCQQANAPDPIIN-TCNSFYCDQFTPNSDNKPK---MWTENWS 254
+ T+ ++ C ++ S D F +PK + +E ++
Sbjct: 212 GDDVVLFTTDGAHKTFLKCGALQGLYTTVDFGTGSNITDAFLSQRKCEPKGPLINSEFYT 271
Query: 255 GWFLAFGGAVPYRPVEDLAFAVARFFQRGGTFQNYYMYHGGTNFGRTTG--GPFIS--TS 310
GW +G E +A ++ RG + N YM+ GGTNF G P+ + TS
Sbjct: 272 GWLDHWGQPHSTIKTEAVASSLYDILARGASV-NLYMFIGGTNFAYWNGANSPYAAQPTS 330
Query: 311 YDYDAPIDEYGDIRQPKWGHLKDLHKAIKLCEEALIASDP 350
YDYDAP+ E GD+ + + + K K+ E + S P
Sbjct: 331 YDYDAPLSEAGDLTEKYFALRNIIQKFEKVPEGPIPPSTP 370
Score = 49.7 bits (117), Expect = 3e-05
Identities = 42/138 (30%), Positives = 62/138 (44%), Gaps = 15/138 (10%)
Query: 530 ITL-VTGKN--TIDLLSLTVGLQNYGAFYDTVGAGITGPVILKGLKNGSSVDLTSQQWTY 586
ITL +TGK T+DLL +G NYGA+ + ++ + + ++ +
Sbjct: 461 ITLNITGKAGATLDLLVENMGRVNYGAYINDFKGLVSNLTLSSNILTDWTIFPLDTEDAV 520
Query: 587 QVGLQGEFVGLSSGNVGQW---NSQSNLPANQPLTWYKTNFVAPSG----SNPVAIDFTG 639
+ L G S + W +S LPA +Y NF PSG I F G
Sbjct: 521 RSHLGGWGHRDSGHHDEAWAHNSSNYTLPA-----FYMGNFSIPSGIPDLPQDTFIQFPG 575
Query: 640 MGKGEAWVNGQSIGRYWP 657
KG+ W+NG ++GRYWP
Sbjct: 576 WTKGQVWINGFNLGRYWP 593
>BGAL_MOUSE (P23780) Beta-galactosidase precursor (EC 3.2.1.23)
(Lactase) (Acid beta-galactosidase)
Length = 647
Score = 145 bits (367), Expect = 3e-34
Identities = 108/350 (30%), Positives = 160/350 (44%), Gaps = 25/350 (7%)
Query: 15 GVYVPASFCSNVTYDHRALVIDGKRRVLMSGSIHYPRSTPQMWPDLIQKSKDGGIDVIET 74
G+Y + Y + DG+ +SGSIHY R W D + K K G++ I+
Sbjct: 24 GIYNVTQRTFKLDYSRDRFLKDGQPFRYISGSIHYFRIPRFYWEDRLLKMKMAGLNAIQM 83
Query: 75 YVFWNLHEPVRGQYNFEGRGDLVGFVKAVAAAGLYVHLRIGPYVCAEWNYGGFPLWLHFI 134
YV WN HEP GQY F G D+ F++ GL V LR GPY+CAEW+ GG P WL
Sbjct: 84 YVPWNFHEPQPGQYEFSGDRDVEHFIQLAHELGLLVILRPGPYICAEWDMGGLPAWLLEK 143
Query: 135 AGIKFRTNNEPFKAEMKRFTAKIVDMMKQENLYASQGGPIILSQIENEYGNIDTHD---- 190
I R+++ + + ++ A ++ MK L GGPII Q+ENEYG+ D
Sbjct: 144 QSIVLRSSDPDYLVAVDKWLAVLLPKMKP--LLYQNGGPIITVQVENEYGSYFACDYDYL 201
Query: 191 ---ARAAKSYIDWAASMATSLDTGVPWIMCQQANAPDPII------NTCNSFYCD-QFTP 240
+ ++ + T+ + C + N +F +F P
Sbjct: 202 RFLVHRFRYHLGNDVILFTTDGASEKMLKCGTLQDLYATVDFGTGNNITQAFLVQRKFEP 261
Query: 241 NSDNKPKMWTENWSGWFLAFGGAVPYRPVEDLAFAVARFFQRGGTFQNYYMYHGGTNFGR 300
P + +E ++GW +G + LA ++ RG N YM+ GGTNF
Sbjct: 262 KG---PLINSEFYTGWLDHWGKPHSTVKTKTLATSLYNLLARGANV-NLYMFIGGTNFAY 317
Query: 301 TTGG--PF--ISTSYDYDAPIDEYGDIRQPKWGHLKDLHKAIKLCEEALI 346
G P+ TSYDYDAP+ E GD+ + K+ L+++ + K E I
Sbjct: 318 WNGANTPYEPQPTSYDYDAPLSEAGDLTK-KYFALREVIQMFKEVPEGPI 366
Score = 55.1 bits (131), Expect = 8e-07
Identities = 42/132 (31%), Positives = 59/132 (43%), Gaps = 20/132 (15%)
Query: 538 TIDLLSLTVGLQNYGAFYDTVGAGITGPVILKG-LKNGSSVDLTSQQ------WTYQVGL 590
T+D+L +G NYG F + I+ I L N + L ++ W +
Sbjct: 474 TLDILVENMGRVNYGRFINDFKGLISNMTINSTVLTNWTVFPLNTEAMVRNHLWGREASD 533
Query: 591 QGEFVGLSSGNVGQWNSQSNLPANQPLTWYKTNFVAPSG----SNPVAIDFTGMGKGEAW 646
+G G S+ N +S LP T+Y NF PSG I F G KG+ W
Sbjct: 534 EGHLDGRSTSN----SSDLILP-----TFYVGNFSIPSGIPDLPQDTFIQFPGWSKGQVW 584
Query: 647 VNGQSIGRYWPT 658
+NG ++GRYWPT
Sbjct: 585 INGFNLGRYWPT 596
>BGAL_ASPNG (P29853) Beta-galactosidase precursor (EC 3.2.1.23)
(Lactase)
Length = 1006
Score = 116 bits (291), Expect = 2e-25
Identities = 122/456 (26%), Positives = 188/456 (40%), Gaps = 73/456 (16%)
Query: 26 VTYDHRALVIDGKRRVLMSGSIH-YPRSTPQMWPDLIQKSKDGGIDVIETYVFWNLHEPV 84
VT+D ++L I+G+R ++ SG H + ++ D+ QK K G + + YV W L E
Sbjct: 46 VTWDDKSLFINGERIMIFSGEFHPFRLPVKELQLDIFQKVKALGFNCVSFYVDWALVEGK 105
Query: 85 RGQYNFEGRGDLVGFVKAVAAAGLYVHLRIGPYVCAEWNYGGFPLWLHFIAGIKFRTNNE 144
G+Y +G DL F A + AG+Y+ R GPY+ AE + GGFP WL + G R++++
Sbjct: 106 PGEYRADGIFDLEPFFDAASEAGIYLLARPGPYINAESSGGGFPGWLQRVNG-TLRSSDK 164
Query: 145 PFKAEMKRFTAKIVDMMKQENLYASQGGPIILSQIENEYGNIDTHDARAAKSYIDWAASM 204
+ + + + + + + + GGPIIL Q ENEY + + Y+ +
Sbjct: 165 AYLDATDNYVSHVAATIAKYQI--TNGGPIILYQPENEYTSGCSGVEFPDPVYMQYVEDQ 222
Query: 205 ATSLDTGVPWI----MCQQANAPDPIINTCNSFYCDQFTPNSD-NKPKMWTE-----NWS 254
A + +P I NAP + + D + D P +W N+
Sbjct: 223 ARNAGVVIPLINNDASASGNNAPGTGKGAVDIYGHDSYPLGFDCANPTVWPSGDLPTNFR 282
Query: 255 GWFLAFGGAVPYRPVE----------DLAFAVAR----------FFQRGGTFQ----NYY 290
L PY VE FA F++ +FQ N Y
Sbjct: 283 TLHLEQSPTTPYAIVEFQGGSYDPWGGPGFAACSELLNNEFERVFYKNDFSFQIAIMNLY 342
Query: 291 MYHGGTNFGRTTGGPFISTSYDYDAPIDEYGDIRQPKWGHLKDLHKAIKLCEEALIASDP 350
M GGTN+G G P TSYDY + + E +I + K+ LK L K+ L AS
Sbjct: 343 MIFGGTNWG-NLGYPNGYTSYDYGSAVTESRNITREKYSELKLLGNFAKVSPGYLTASPG 401
Query: 351 TITSPG----------------------------PNLETAVYKTGAVCSAFLANIGMSDA 382
+T+ G + E+ YK SA I
Sbjct: 402 NLTTSGYADTTDLTVTPLLGNSTGSFFVVRHSDYSSEESTSYKLRLPTSAGSVTIPQLGG 461
Query: 383 TVTFNG--NSYHLPGWSVSILPDCKNVVLNTAKVNT 416
T+T NG + H+ +VS N++ +TA+V T
Sbjct: 462 TLTLNGRDSKIHVTDHNVS----GTNIIYSTAEVFT 493
>BGAM_HUMAN (P16279) Beta-galactosidase-related protein precursor
(Beta-galactosidase-like protein) (S-Gal)
(Elastin-binding protein) (EBP)
Length = 546
Score = 51.6 bits (122), Expect = 8e-06
Identities = 40/130 (30%), Positives = 61/130 (46%), Gaps = 8/130 (6%)
Query: 228 NTCNSFYCDQFTPNSDNKPK---MWTENWSGWFLAFGGAVPYRPVEDLAFAVARFFQRGG 284
N+ S D F +PK + +E ++GW +G E +A ++ RG
Sbjct: 111 NSAGSNITDAFLSQRKCEPKGPLINSEFYTGWLDHWGQPHSTIKTEAVASSLYDILARGA 170
Query: 285 TFQNYYMYHGGTNFGRTTGG--PFIS--TSYDYDAPIDEYGDIRQPKWGHLKDLHKAIKL 340
+ N YM+ GGTNF G P+ + TSYDYDAP+ E GD+ + + + K K+
Sbjct: 171 SV-NLYMFIGGTNFAYWNGANSPYAAQPTSYDYDAPLSEAGDLTEKYFALRNIIQKFEKV 229
Query: 341 CEEALIASDP 350
E + S P
Sbjct: 230 PEGPIPPSTP 239
Score = 49.7 bits (117), Expect = 3e-05
Identities = 42/138 (30%), Positives = 62/138 (44%), Gaps = 15/138 (10%)
Query: 530 ITL-VTGKN--TIDLLSLTVGLQNYGAFYDTVGAGITGPVILKGLKNGSSVDLTSQQWTY 586
ITL +TGK T+DLL +G NYGA+ + ++ + + ++ +
Sbjct: 330 ITLNITGKAGATLDLLVENMGRVNYGAYINDFKGLVSNLTLSSNILTDWTIFPLDTEDAV 389
Query: 587 QVGLQGEFVGLSSGNVGQW---NSQSNLPANQPLTWYKTNFVAPSG----SNPVAIDFTG 639
+ L G S + W +S LPA +Y NF PSG I F G
Sbjct: 390 RSHLGGWGHRDSGHHDEAWAHNSSNYTLPA-----FYMGNFSIPSGIPDLPQDTFIQFPG 444
Query: 640 MGKGEAWVNGQSIGRYWP 657
KG+ W+NG ++GRYWP
Sbjct: 445 WTKGQVWINGFNLGRYWP 462
Score = 35.0 bits (79), Expect = 0.81
Identities = 17/49 (34%), Positives = 26/49 (52%)
Query: 26 VTYDHRALVIDGKRRVLMSGSIHYPRSTPQMWPDLIQKSKDGGIDVIET 74
+ Y + + DG+ +SGSIHY R W D + K K G++ I+T
Sbjct: 34 IDYSRDSFLKDGQPFRYISGSIHYSRVPRFYWKDRLLKMKMAGLNAIQT 82
>LEG_ANTCR (P22031) D-galactoside-specific lectin (Sea urchin egg
lectin) (SUEL)
Length = 105
Score = 36.6 bits (83), Expect = 0.28
Identities = 22/73 (30%), Positives = 35/73 (47%), Gaps = 8/73 (10%)
Query: 759 LSLECPYPNQAISSIKFASFGTPRG-TCGNY-----NHGSCSSNRALSIVQKACIGSSSC 812
L++ CP + I A +G RG C + C S+ + +V+ +C G SSC
Sbjct: 19 LTISCPEGEGIV--IYDAIYGRKRGEVCPGLFGAFTKNRKCRSSNSQQVVENSCEGKSSC 76
Query: 813 NIGVSINTFGNPC 825
+ S + FG+PC
Sbjct: 77 TVLASNSVFGDPC 89
>YP69_CAEEL (Q09218) Hypothetical protein B0495.9 in chromosome II
Length = 267
Score = 33.5 bits (75), Expect = 2.4
Identities = 16/44 (36%), Positives = 23/44 (51%)
Query: 30 HRALVIDGKRRVLMSGSIHYPRSTPQMWPDLIQKSKDGGIDVIE 73
HR VIDGK S + YP++ Q W D+ + + GI + E
Sbjct: 174 HRGAVIDGKAASEASVIVEYPQNAQQFWADVRRAEGEIGIGLTE 217
>CU63_PANTR (Q68US5) Protein C21orf63 homolog precursor
Length = 441
Score = 32.7 bits (73), Expect = 4.0
Identities = 13/33 (39%), Positives = 19/33 (57%)
Query: 793 CSSNRALSIVQKACIGSSSCNIGVSINTFGNPC 825
C S AL ++ + C G C I V+ + FG+PC
Sbjct: 213 CLSYSALQVLSRRCYGKQRCKIIVNNHHFGSPC 245
>CU63_HUMAN (P58658) Protein C21orf63 precursor (Protein PRED34)
(SUE21) (UNQ2504/PRO5993)
Length = 441
Score = 32.7 bits (73), Expect = 4.0
Identities = 13/33 (39%), Positives = 19/33 (57%)
Query: 793 CSSNRALSIVQKACIGSSSCNIGVSINTFGNPC 825
C S AL ++ + C G C I V+ + FG+PC
Sbjct: 213 CLSYSALQVLSRRCYGKQRCKIIVNNHHFGSPC 245
>AMPR_YEREN (P45461) HTH-type transcriptional activator ampR
Length = 294
Score = 32.7 bits (73), Expect = 4.0
Identities = 17/46 (36%), Positives = 22/46 (46%), Gaps = 5/46 (10%)
Query: 225 PIINTCNSFYCDQFTPNSDNKPKMW-----TENWSGWFLAFGGAVP 265
P+ C +FTP +D + M NWS WF A GG+VP
Sbjct: 164 PLAQLCAPSISKRFTPPTDLQRFMLLGSYRAMNWSAWFAAAGGSVP 209
>AG43_ECOLI (P39180) Antigen 43 precursor (AG43) (Fluffing protein)
Length = 1039
Score = 32.7 bits (73), Expect = 4.0
Identities = 52/223 (23%), Positives = 93/223 (41%), Gaps = 37/223 (16%)
Query: 408 VLNTAKVNTASMISSFATESLKEKVDSLDSSSSGWSWISEPVGISTPDAFTKSGLLEQIN 467
+++T+ A +++ S+K D+ D + GW W+ + P + T + +LE
Sbjct: 831 LVHTSSGLWADIVAQGTRHSMKASSDNNDFRARGWGWLGS-LETGLPFSITDNLMLE--- 886
Query: 468 TTADRSDYLWYSLSI-VYEDNAGDQPVLHIESLGHALHA-----------FVNGKLAGSK 515
+ Y W LS+ +DNAG H G A H G+ S+
Sbjct: 887 ---PQLQYTWQGLSLDDGKDNAGYVKFGH----GSAQHVRAGFRLGSHNDMTFGEGTSSR 939
Query: 516 AGSSGNAKVNV-DIPITLVTGKNTIDLLSLTVGLQNYGAFYDTVGAGITGPVILKGLKNG 574
A +AK +V ++P+ + I S + G G T G+G+T +NG
Sbjct: 940 APLRDSAKHSVSELPVNWWVQPSVIRTFS-SRGDMRVGT--STAGSGMT----FSPSQNG 992
Query: 575 SSVDLTS-----QQWTYQVGLQGEFVGLSSGNVGQ-WNSQSNL 611
+S+DL + + +G+Q + SG+ + +N Q+ L
Sbjct: 993 TSLDLQAGLEARVRENITLGVQAGYAHSVSGSSAEGYNGQATL 1035
>SEC8_NEUCR (Q9HE88) Probable exocyst complex component sec8
Length = 1111
Score = 32.3 bits (72), Expect = 5.2
Identities = 34/126 (26%), Positives = 53/126 (41%), Gaps = 28/126 (22%)
Query: 130 WLHF-IAGIKFRTNNE--------PFKAEMKRFTAKIVDMMKQENLYASQGGPIILSQIE 180
WL I G++ T NE P KAE KR+T ++ + P+ L E
Sbjct: 807 WLSVKIHGLRHITRNETDSSKSSFPTKAEKKRWT-----LLNDPSKATGGEAPVYLPMTE 861
Query: 181 NEYGNIDT----HDARAAKSYIDWAASMAT----SLDTGV------PWIMCQQANAPDPI 226
N D+ +D A+ + + + T SL T + P+++ Q+ N PDP
Sbjct: 862 ETVENFDSILVSYDELASTALLTLHLEIRTRILHSLQTALSPLTTAPYLLDQEVNEPDPE 921
Query: 227 INTCNS 232
I + NS
Sbjct: 922 ILSLNS 927
>YE62_MYCLE (Q49682) Hypothetical UPF0051 protein ML0594
Length = 392
Score = 32.0 bits (71), Expect = 6.9
Identities = 25/83 (30%), Positives = 40/83 (48%), Gaps = 7/83 (8%)
Query: 568 LKGLKNGSSV--DLTSQQWTYQVGLQGEFVGLSSGNVGQWNSQSNLPANQPLTWYKTNFV 625
L+GL +GS+V + + Q G+Q E VG G +GQ ++ A Q + +++ V
Sbjct: 44 LRGLHDGSAVANGKVHIRVSQQSGVQTEIVGRGDGRLGQGGIPTDRVAAQAFSSFQSATV 103
Query: 626 APSG-----SNPVAIDFTGMGKG 643
G + P+ I TG KG
Sbjct: 104 VTVGRDMQIATPINIAVTGPDKG 126
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.317 0.135 0.424
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 107,940,372
Number of Sequences: 164201
Number of extensions: 4999767
Number of successful extensions: 10402
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 16
Number of HSP's successfully gapped in prelim test: 12
Number of HSP's that attempted gapping in prelim test: 10315
Number of HSP's gapped (non-prelim): 43
length of query: 826
length of database: 59,974,054
effective HSP length: 119
effective length of query: 707
effective length of database: 40,434,135
effective search space: 28586933445
effective search space used: 28586933445
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 70 (31.6 bits)
Medicago: description of AC139344.4


