

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136472.7 + phase: 0
(123 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
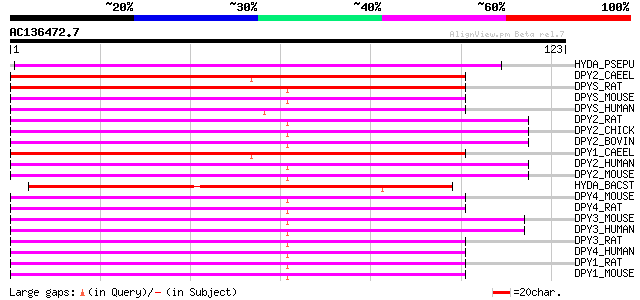
Score E
Sequences producing significant alignments: (bits) Value
HYDA_PSEPU (Q59699) D-hydantoinase (EC 3.5.2.2) (Dihydropyrimidi... 96 2e-20
DPY2_CAEEL (Q18677) Dihydropyrimidinase 2 (EC 3.5.2.2) 92 2e-19
DPYS_RAT (Q63150) Dihydropyrimidinase (EC 3.5.2.2) (DHPase) (Hyd... 88 3e-18
DPYS_MOUSE (Q9EQF5) Dihydropyrimidinase (EC 3.5.2.2) (DHPase) (H... 88 3e-18
DPYS_HUMAN (Q14117) Dihydropyrimidinase (EC 3.5.2.2) (DHPase) (H... 86 1e-17
DPY2_RAT (P47942) Dihydropyrimidinase related protein-2 (DRP-2) ... 84 6e-17
DPY2_CHICK (Q90635) Dihydropyrimidinase related protein-2 (DRP-2... 84 6e-17
DPY2_BOVIN (O02675) Dihydropyrimidinase related protein-2 (DRP-2... 84 6e-17
DPY1_CAEEL (Q21773) Dihydropyrimidinase 1 (EC 3.5.2.2) 84 6e-17
DPY2_HUMAN (Q16555) Dihydropyrimidinase related protein-2 (DRP-2... 83 1e-16
DPY2_MOUSE (O08553) Dihydropyrimidinase related protein-2 (DRP-2... 82 2e-16
HYDA_BACST (Q45515) D-hydantoinase (EC 3.5.2.2) (Dihydropyrimidi... 80 6e-16
DPY4_MOUSE (O35098) Dihydropyrimidinase related protein-4 (DRP-4... 79 1e-15
DPY4_RAT (Q62951) Dihydropyrimidinase related protein-4 (DRP-4) ... 79 2e-15
DPY3_MOUSE (Q62188) Dihydropyrimidinase related protein-3 (DRP-3... 79 2e-15
DPY3_HUMAN (Q14195) Dihydropyrimidinase related protein-3 (DRP-3... 79 2e-15
DPY3_RAT (Q62952) Dihydropyrimidinase related protein-3 (DRP-3) ... 79 2e-15
DPY4_HUMAN (O14531) Dihydropyrimidinase related protein-4 (DRP-4... 77 7e-15
DPY1_RAT (Q62950) Dihydropyrimidinase related protein-1 (DRP-1) ... 76 2e-14
DPY1_MOUSE (P97427) Dihydropyrimidinase related protein-1 (DRP-1... 76 2e-14
>HYDA_PSEPU (Q59699) D-hydantoinase (EC 3.5.2.2)
(Dihydropyrimidinase) (DHPase)
Length = 495
Score = 95.5 bits (236), Expect = 2e-20
Identities = 51/108 (47%), Positives = 63/108 (58%)
Query: 2 SIDAMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKAL 61
S +A++E+ AR GQ V GE + L LDDS PD+ TAA YVMSPP R + H +AL
Sbjct: 242 SREALDEITYARAKGQPVYGEVLPGHLLLDDSVYRDPDWATAAGYVMSPPFRPREHQEAL 301
Query: 62 QAALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNGDHSNLRYL 109
L +G L TDHC F + QKA G DDF +IPNG G + L
Sbjct: 302 WRGLQSGNLHTTATDHCCFCAEQKAMGRDDFSRIPNGTAGIEDRMAVL 349
>DPY2_CAEEL (Q18677) Dihydropyrimidinase 2 (EC 3.5.2.2)
Length = 520
Score = 92.0 bits (227), Expect = 2e-19
Identities = 50/102 (49%), Positives = 62/102 (60%), Gaps = 1/102 (0%)
Query: 1 MSIDAMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPI-RKQGHDK 59
MS A +A RK G V GEP+ +GLA D S ++ D+ AA+YVMSPP+ R
Sbjct: 243 MSKGAAAAIAHHRKKGAVVFGEPIAAGLATDGSHYYNEDWLHAARYVMSPPLSRDPSTPS 302
Query: 60 ALQAALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNG 101
AL L+ G L L TD+C F+ QK+ G DDF KIPNGVNG
Sbjct: 303 ALMKLLAAGELHLTATDNCTFDCQQKSLGKDDFTKIPNGVNG 344
>DPYS_RAT (Q63150) Dihydropyrimidinase (EC 3.5.2.2) (DHPase)
(Hydantoinase) (DHP)
Length = 519
Score = 88.2 bits (217), Expect = 3e-18
Identities = 45/102 (44%), Positives = 62/102 (60%), Gaps = 1/102 (0%)
Query: 1 MSIDAMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKA 60
MS A + +A A++ G+ V GEP+ +GL D + W+ ++ AA +VM PP+R
Sbjct: 250 MSKSAAKVIADAKREGKVVYGEPIAAGLGTDGTQYWNKEWRHAAHHVMGPPLRPDPSTPG 309
Query: 61 -LQAALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNG 101
L L+ G L G+D+C FN+ QKA G DDF KIPNGVNG
Sbjct: 310 FLMNLLANGDLTTTGSDNCTFNTCQKALGKDDFTKIPNGVNG 351
>DPYS_MOUSE (Q9EQF5) Dihydropyrimidinase (EC 3.5.2.2) (DHPase)
(Hydantoinase) (DHP)
Length = 519
Score = 88.2 bits (217), Expect = 3e-18
Identities = 47/102 (46%), Positives = 60/102 (58%), Gaps = 1/102 (0%)
Query: 1 MSIDAMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKA 60
MS A + VA AR++G V GEP+ +GL D W ++ AA +VM PP+R
Sbjct: 250 MSKSAAKVVADARRAGNVVYGEPIAAGLGTDGRQYWSEEWSHAAHHVMGPPLRPDPLTPG 309
Query: 61 -LQAALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNG 101
L L+ G L G+D+C FN+ QKA G DDF KIPNGVNG
Sbjct: 310 FLMDLLANGDLTTTGSDNCTFNTCQKALGKDDFTKIPNGVNG 351
>DPYS_HUMAN (Q14117) Dihydropyrimidinase (EC 3.5.2.2) (DHPase)
(Hydantoinase) (DHP)
Length = 519
Score = 85.9 bits (211), Expect = 1e-17
Identities = 45/102 (44%), Positives = 60/102 (58%), Gaps = 1/102 (0%)
Query: 1 MSIDAMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQ-GHDK 59
MS A + +A AR+ G+ V GEP+ + L D + W+ ++ AA +VM PP+R
Sbjct: 250 MSKSAAKVIADARRDGKVVYGEPIAASLGTDGTHYWNKEWHHAAHHVMGPPLRPDPSTPD 309
Query: 60 ALQAALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNG 101
L L+ L GTD+C FN+ QKA G DDF KIPNGVNG
Sbjct: 310 FLMNLLANDDLTTTGTDNCTFNTCQKALGKDDFTKIPNGVNG 351
>DPY2_RAT (P47942) Dihydropyrimidinase related protein-2 (DRP-2)
(Turned on after division, 64 kDa protein) (TOAD-64)
(Collapsin response mediator protein 2) (CRMP-2)
Length = 572
Score = 84.0 bits (206), Expect = 6e-17
Identities = 46/116 (39%), Positives = 63/116 (53%), Gaps = 1/116 (0%)
Query: 1 MSIDAMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKA 60
MS A E +A+ARK G V GEP+ + L D S W ++ AA +V SPP+
Sbjct: 256 MSKSAAEVIAQARKKGTVVYGEPITASLGTDGSHYWSKNWAKAAAFVTSPPLSPDPTTPD 315
Query: 61 -LQAALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNGDHSNLRYLRPKYIL 115
L + LS G LQ+ G+ HC FN+ QKA G D+F IP G NG + + K ++
Sbjct: 316 FLNSLLSCGDLQVTGSAHCTFNTAQKAVGKDNFTLIPEGTNGTEERMSVIWDKAVV 371
>DPY2_CHICK (Q90635) Dihydropyrimidinase related protein-2 (DRP-2)
(Collapsin response mediator protein CRMP-62)
Length = 572
Score = 84.0 bits (206), Expect = 6e-17
Identities = 46/116 (39%), Positives = 63/116 (53%), Gaps = 1/116 (0%)
Query: 1 MSIDAMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKA 60
MS A E +A+ARK G V GEP+ + L D S W ++ AA +V SPP+
Sbjct: 256 MSKSAAEVIAQARKKGTVVYGEPITASLGTDGSHYWSKNWAKAAAFVTSPPLSPDPTTPD 315
Query: 61 -LQAALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNGDHSNLRYLRPKYIL 115
L + LS G LQ+ G+ HC FN+ QKA G D+F IP G NG + + K ++
Sbjct: 316 FLNSLLSCGDLQVTGSAHCTFNTAQKAVGKDNFTLIPEGTNGTEERMSIIWDKAVV 371
>DPY2_BOVIN (O02675) Dihydropyrimidinase related protein-2 (DRP-2)
(Neural specific protein NSP60)
Length = 572
Score = 84.0 bits (206), Expect = 6e-17
Identities = 46/116 (39%), Positives = 63/116 (53%), Gaps = 1/116 (0%)
Query: 1 MSIDAMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKA 60
MS A E +A+ARK G V GEP+ + L D S W ++ AA +V SPP+
Sbjct: 256 MSKSAAEVIAQARKKGTVVYGEPITASLGTDGSHYWSKNWAKAAAFVTSPPLSPDPTTPD 315
Query: 61 -LQAALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNGDHSNLRYLRPKYIL 115
L + LS G LQ+ G+ HC FN+ QKA G D+F IP G NG + + K ++
Sbjct: 316 FLNSLLSCGDLQVTGSAHCTFNTAQKAVGKDNFTLIPEGTNGTEERMSVIWDKAVV 371
>DPY1_CAEEL (Q21773) Dihydropyrimidinase 1 (EC 3.5.2.2)
Length = 489
Score = 84.0 bits (206), Expect = 6e-17
Identities = 44/102 (43%), Positives = 62/102 (60%), Gaps = 1/102 (0%)
Query: 1 MSIDAMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPI-RKQGHDK 59
M+ A ++ R G V GEP+ +GLALD S ++ D+ AA+YVMSPP+ R +
Sbjct: 247 MTKGAASAISHHRAQGSIVFGEPIAAGLALDGSHYYNEDWLHAARYVMSPPLSRDPTTPE 306
Query: 60 ALQAALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNG 101
L L+ G L L GTD+C ++ QK+ G +F KIPNG+NG
Sbjct: 307 LLMKLLAAGELHLTGTDNCTYDCRQKSLGKGNFTKIPNGING 348
>DPY2_HUMAN (Q16555) Dihydropyrimidinase related protein-2 (DRP-2)
(Collapsin response mediator protein 2) (CRMP-2) (N2A3)
Length = 572
Score = 82.8 bits (203), Expect = 1e-16
Identities = 45/116 (38%), Positives = 63/116 (53%), Gaps = 1/116 (0%)
Query: 1 MSIDAMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKA 60
MS + E +A+ARK G V GEP+ + L D S W ++ AA +V SPP+
Sbjct: 256 MSKSSAEVIAQARKKGTVVYGEPITASLGTDGSHYWSKNWAKAAAFVTSPPLSPDPTTPD 315
Query: 61 -LQAALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNGDHSNLRYLRPKYIL 115
L + LS G LQ+ G+ HC FN+ QKA G D+F IP G NG + + K ++
Sbjct: 316 FLNSLLSCGDLQVTGSAHCTFNTAQKAVGKDNFTLIPEGTNGTEERMSVIWDKAVV 371
>DPY2_MOUSE (O08553) Dihydropyrimidinase related protein-2 (DRP-2)
(ULIP 2 protein)
Length = 572
Score = 82.0 bits (201), Expect = 2e-16
Identities = 45/116 (38%), Positives = 62/116 (52%), Gaps = 1/116 (0%)
Query: 1 MSIDAMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKA 60
M A E +A+ARK G V GEP+ + L D S W ++ AA +V SPP+
Sbjct: 256 MPKSAAEVIAQARKKGTVVYGEPITASLGTDGSHYWSKNWAKAAAFVTSPPLSPDPTTPD 315
Query: 61 -LQAALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNGDHSNLRYLRPKYIL 115
L + LS G LQ+ G+ HC FN+ QKA G D+F IP G NG + + K ++
Sbjct: 316 FLNSLLSCGDLQVTGSAHCTFNTAQKAVGKDNFTLIPEGTNGTEERMSVIWDKAVV 371
>HYDA_BACST (Q45515) D-hydantoinase (EC 3.5.2.2)
(Dihydropyrimidinase) (DHPase)
Length = 471
Score = 80.5 bits (197), Expect = 6e-16
Identities = 45/95 (47%), Positives = 59/95 (61%), Gaps = 2/95 (2%)
Query: 5 AMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKALQAA 64
A+E++A+AR G V GE L LD S+L P+F+ AKYV SPP+R++ H + L A
Sbjct: 245 AVEKIAEARNKGLNVWGETCPQYLVLDQSYLEKPNFE-GAKYVWSPPLREKWHQEVLWNA 303
Query: 65 LSTGVLQLVGTDHCVFN-STQKAFGIDDFRKIPNG 98
L G LQ +G+D C F+ QK G DF KIPNG
Sbjct: 304 LKNGQLQTLGSDQCSFDFKGQKELGRGDFTKIPNG 338
>DPY4_MOUSE (O35098) Dihydropyrimidinase related protein-4 (DRP-4)
(Collapsin response mediator protein 3) (CRMP-3)
(UNC33-like phosphoprotein 4) (ULIP4 protein)
Length = 572
Score = 79.3 bits (194), Expect = 1e-15
Identities = 44/102 (43%), Positives = 59/102 (57%), Gaps = 1/102 (0%)
Query: 1 MSIDAMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKA 60
MS A + VA+A++ G V GEP+ + L D S W ++ AA +V SPPI
Sbjct: 256 MSKGAADMVAQAKRRGVVVFGEPITASLGTDGSHYWSKNWAKAAAFVTSPPINPDPTTAD 315
Query: 61 -LQAALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNG 101
L + LS+G LQ+ G+ HC F + QKA G D+F IP GVNG
Sbjct: 316 HLTSLLSSGDLQVTGSAHCTFTTAQKAVGKDNFTLIPEGVNG 357
>DPY4_RAT (Q62951) Dihydropyrimidinase related protein-4 (DRP-4)
(Collapsin response mediator protein 3) (CRMP-3)
(UNC33-like phosphoprotein 4) (ULIP4 protein) (Fragment)
Length = 564
Score = 79.0 bits (193), Expect = 2e-15
Identities = 43/102 (42%), Positives = 59/102 (57%), Gaps = 1/102 (0%)
Query: 1 MSIDAMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKA 60
MS A + VA+A++ G V GEP+ + L D S W ++ AA +V SPPI
Sbjct: 248 MSKGAADMVAQAKRRGVVVFGEPITASLGTDGSHYWSKNWAKAAAFVTSPPINPDPTTAD 307
Query: 61 -LQAALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNG 101
L + LS+G LQ+ G+ HC F + QKA G D+F IP G+NG
Sbjct: 308 HLTSLLSSGDLQVTGSAHCTFTTAQKAVGKDNFTLIPEGING 349
>DPY3_MOUSE (Q62188) Dihydropyrimidinase related protein-3 (DRP-3)
(Unc-33-like phosphoprotein) (ULIP protein)
Length = 570
Score = 79.0 bits (193), Expect = 2e-15
Identities = 41/115 (35%), Positives = 64/115 (55%), Gaps = 1/115 (0%)
Query: 1 MSIDAMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKA 60
MS A + +++ARK G V GEP+ + L +D + W ++ AA +V SPP+
Sbjct: 256 MSKSAADLISQARKKGNVVFGEPITASLGIDGTHYWSKNWAKAAAFVTSPPLSPDPTTPD 315
Query: 61 -LQAALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNGDHSNLRYLRPKYI 114
+ + L++G LQL G+ HC F++ QKA G D+F IP G NG + + K +
Sbjct: 316 YINSLLASGDLQLSGSAHCTFSTAQKAIGKDNFTAIPEGTNGVEERMSVIWDKAV 370
>DPY3_HUMAN (Q14195) Dihydropyrimidinase related protein-3 (DRP-3)
(Unc-33-like phosphoprotein) (ULIP protein) (Collapsin
response mediator protein 4) (CRMP-4)
Length = 570
Score = 79.0 bits (193), Expect = 2e-15
Identities = 41/115 (35%), Positives = 64/115 (55%), Gaps = 1/115 (0%)
Query: 1 MSIDAMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKA 60
MS A + +++ARK G V GEP+ + L +D + W ++ AA +V SPP+
Sbjct: 256 MSKSAADLISQARKKGNVVFGEPITASLGIDGTHYWSKNWAKAAAFVTSPPLSPDPTTPD 315
Query: 61 -LQAALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNGDHSNLRYLRPKYI 114
+ + L++G LQL G+ HC F++ QKA G D+F IP G NG + + K +
Sbjct: 316 YINSLLASGDLQLSGSAHCTFSTAQKAIGKDNFTAIPEGTNGVEERMSVIWDKAV 370
>DPY3_RAT (Q62952) Dihydropyrimidinase related protein-3 (DRP-3)
(Collapsin response mediator protein 4) (CRMP-4)
(Fragment)
Length = 358
Score = 78.6 bits (192), Expect = 2e-15
Identities = 40/102 (39%), Positives = 60/102 (58%), Gaps = 1/102 (0%)
Query: 1 MSIDAMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKA 60
MS A + +++ARK G V GEP+ + L +D + W ++ AA +V SPP+
Sbjct: 256 MSKSAADLISQARKKGNVVFGEPITASLGIDGTHYWSKNWAKAAAFVTSPPLSPDPTTPD 315
Query: 61 -LQAALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNG 101
+ + L++G LQL G+ HC F++ QKA G D+F IP G NG
Sbjct: 316 YINSLLASGDLQLSGSAHCTFSTAQKAIGKDNFTAIPEGTNG 357
>DPY4_HUMAN (O14531) Dihydropyrimidinase related protein-4 (DRP-4)
(Collapsin response mediator protein 3) (CRMP-3)
(UNC33-like phosphoprotein 4) (ULIP4 protein)
Length = 572
Score = 77.0 bits (188), Expect = 7e-15
Identities = 41/102 (40%), Positives = 57/102 (55%), Gaps = 1/102 (0%)
Query: 1 MSIDAMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKA 60
MS A + +A+A++ G V GEP+ + L D S W ++ AA +V SPP+
Sbjct: 256 MSKGAADAIAQAKRRGVVVFGEPITASLGTDGSHYWSKNWAKAAAFVTSPPVNPDPTTAD 315
Query: 61 -LQAALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNG 101
L LS+G LQ+ G+ HC F + QKA G D+F IP G NG
Sbjct: 316 HLTCLLSSGDLQVTGSAHCTFTTAQKAVGKDNFALIPEGTNG 357
>DPY1_RAT (Q62950) Dihydropyrimidinase related protein-1 (DRP-1)
(Collapsin response mediator protein 1) (CRMP-1)
Length = 572
Score = 75.9 bits (185), Expect = 2e-14
Identities = 41/102 (40%), Positives = 58/102 (56%), Gaps = 1/102 (0%)
Query: 1 MSIDAMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKA 60
MS A + +A ARK G V GEP+ + L D + W ++ AA +V SPP+
Sbjct: 256 MSKSAADIIALARKKGPLVFGEPIAASLGTDGTHYWSKNWAKAAAFVTSPPLSPDPTTPD 315
Query: 61 -LQAALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNG 101
L + L+ G LQ+ G+ HC +++ QKA G D+F IP GVNG
Sbjct: 316 YLTSLLACGDLQVTGSGHCPYSTAQKAVGKDNFTLIPEGVNG 357
>DPY1_MOUSE (P97427) Dihydropyrimidinase related protein-1 (DRP-1)
(Collapsin response mediator protein 1) (CRMP-1) (ULIP3
protein)
Length = 572
Score = 75.9 bits (185), Expect = 2e-14
Identities = 41/102 (40%), Positives = 58/102 (56%), Gaps = 1/102 (0%)
Query: 1 MSIDAMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKA 60
MS A + +A ARK G V GEP+ + L D + W ++ AA +V SPP+
Sbjct: 256 MSKSAADIIALARKKGPLVFGEPIAASLGTDGTHYWSKNWAKAAAFVTSPPLSPDPTTPD 315
Query: 61 -LQAALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNG 101
L + L+ G LQ+ G+ HC +++ QKA G D+F IP GVNG
Sbjct: 316 YLTSLLACGDLQVTGSGHCPYSTAQKAVGKDNFTLIPEGVNG 357
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.137 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,667,043
Number of Sequences: 164201
Number of extensions: 547204
Number of successful extensions: 1259
Number of sequences better than 10.0: 72
Number of HSP's better than 10.0 without gapping: 61
Number of HSP's successfully gapped in prelim test: 11
Number of HSP's that attempted gapping in prelim test: 1167
Number of HSP's gapped (non-prelim): 72
length of query: 123
length of database: 59,974,054
effective HSP length: 99
effective length of query: 24
effective length of database: 43,718,155
effective search space: 1049235720
effective search space used: 1049235720
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC136472.7


