

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126008.1 - phase: 0 /pseudo
(790 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
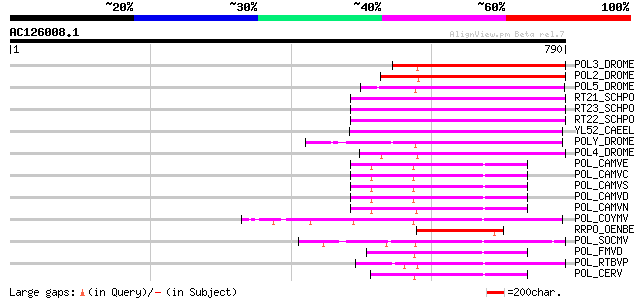
Sequences producing significant alignments: (bits) Value
POL3_DROME (P04323) Retrovirus-related Pol polyprotein from tran... 218 5e-56
POL2_DROME (P20825) Retrovirus-related Pol polyprotein from tran... 218 6e-56
POL5_DROME (Q8I7P9) Retrovirus-related Pol polyprotein from tran... 211 8e-54
RT21_SCHPO (Q05654) Retrotransposable element Tf2 155 kDa protei... 207 9e-53
RT23_SCHPO (Q9UR07) Retrotransposable element Tf2 155 kDa protei... 204 6e-52
RT22_SCHPO (Q9C0R2) Retrotransposable element Tf2 155 kDa protei... 204 6e-52
YL52_CAEEL (P34431) Hypothetical protein F44E2.2 in chromosome III 193 1e-48
POLY_DROME (P10401) Retrovirus-related Pol polyprotein from tran... 182 3e-45
POL4_DROME (P10394) Retrovirus-related Pol polyprotein from tran... 171 5e-42
POL_CAMVE (Q02964) Enzymatic polyprotein [Contains: Aspartic pro... 152 3e-36
POL_CAMVC (P03555) Enzymatic polyprotein [Contains: Aspartic pro... 152 3e-36
POL_CAMVS (P03554) Enzymatic polyprotein [Contains: Aspartic pro... 152 4e-36
POL_CAMVD (P03556) Enzymatic polyprotein [Contains: Aspartic pro... 149 3e-35
POL_CAMVN (Q00962) Enzymatic polyprotein [Contains: Aspartic pro... 147 8e-35
POL_COYMV (P19199) Putative polyprotein [Contains: Coat protein;... 146 2e-34
RRPO_OENBE (P31843) RNA-directed DNA polymerase homolog (Reverse... 145 3e-34
POL_SOCMV (P15629) Enzymatic polyprotein [Contains: Aspartic pro... 137 8e-32
POL_FMVD (P09523) Enzymatic polyprotein [Contains: Aspartic prot... 131 6e-30
POL_RTBVP (P27502) Polyprotein (P194 protein) [Contains: Coat pr... 131 8e-30
POL_CERV (P05400) Enzymatic polyprotein [Contains: Aspartic prot... 119 3e-26
>POL3_DROME (P04323) Retrovirus-related Pol polyprotein from
transposon 17.6 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1058
Score = 218 bits (555), Expect = 5e-56
Identities = 106/251 (42%), Positives = 166/251 (65%), Gaps = 5/251 (1%)
Query: 545 ELKKQLEELLEKQFIRPSVSPRGAPVLLV-KKKDGS----MRLCVDYRQLNKLTIKNRYP 599
E++ Q++++L + IR S SP +P+ +V KK+D S R+ +DYR+LN++T+ +R+P
Sbjct: 222 EVESQIQDMLNQGIIRTSNSPYNSPIWVVPKKQDASGKQKFRIVIDYRKLNEITVGDRHP 281
Query: 600 LPRIDD*MDQLVGAEVFSKIDLRSGYHQIRVKAEDISKTAFRTRYGHYEYSVMPFGVSNA 659
+P +D+ + +L F+ IDL G+HQI + E +SKTAF T++GHYEY MPFG+ NA
Sbjct: 282 IPNMDEILGKLGRCNYFTTIDLAKGFHQIEMDPESVSKTAFSTKHGHYEYLRMPFGLKNA 341
Query: 660 PGVFMEYMNRIFHPYLDRFVVVFIDDILVYSKSEEEHAEHLRIVLQVLKENQLCAKLSKC 719
P F MN I P L++ +V++DDI+V+S S +EH + L +V + L + L +L KC
Sbjct: 342 PATFQRCMNDILRPLLNKHCLVYLDDIIVFSTSLDEHLQSLGLVFEKLAKANLKLQLDKC 401
Query: 720 EFWLKEVSFLGHVISKGGISVDPSKVDAVLQWESPKSVFEIKAIWGLDGYYRRFIEGFSK 779
EF +E +FLGHV++ GI +P K++A+ ++ P EIKA GL GYYR+FI F+
Sbjct: 402 EFLKQETTFLGHVLTPDGIKPNPEKIEAIQKYPIPTKPKEIKAFLGLTGYYRKFIPNFAD 461
Query: 780 LALPLTQLTQK 790
+A P+T+ +K
Sbjct: 462 IAKPMTKCLKK 472
>POL2_DROME (P20825) Retrovirus-related Pol polyprotein from
transposon 297 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1059
Score = 218 bits (554), Expect = 6e-56
Identities = 109/267 (40%), Positives = 166/267 (61%), Gaps = 5/267 (1%)
Query: 529 SPISIAPYRMSASELGELKKQLEELLEKQFIRPSVSPRGAPVLLVKKKDGSM-----RLC 583
SPI Y ++ + E++ Q++E+L + IR S SP +P +V KK + R+
Sbjct: 205 SPIYSKQYPLAQTHEIEVENQVQEMLNQGLIRESNSPYNSPTWVVPKKPDASGANKYRVV 264
Query: 584 VDYRQLNKLTIKNRYPLPRIDD*MDQLVGAEVFSKIDLRSGYHQIRVKAEDISKTAFRTR 643
+DYR+LN++TI +RYP+P +D+ + +L + F+ IDL G+HQI + E ISKTAF T+
Sbjct: 265 IDYRKLNEITIPDRYPIPNMDEILGKLGKCQYFTTIDLAKGFHQIEMDEESISKTAFSTK 324
Query: 644 YGHYEYSVMPFGVSNAPGVFMEYMNRIFHPYLDRFVVVFIDDILVYSKSEEEHAEHLRIV 703
GHYEY MPFG+ NAP F MN I P L++ +V++DDI+++S S EH +++V
Sbjct: 325 SGHYEYLRMPFGLRNAPATFQRCMNNILRPLLNKHCLVYLDDIIIFSTSLTEHLNSIQLV 384
Query: 704 LQVLKENQLCAKLSKCEFWLKEVSFLGHVISKGGISVDPSKVDAVLQWESPKSVFEIKAI 763
L + L +L KCEF KE +FLGH+++ GI +P KV A++ + P EI+A
Sbjct: 385 FTKLADANLKLQLDKCEFLKKEANFLGHIVTPDGIKPNPIKVKAIVSYPIPTKDKEIRAF 444
Query: 764 WGLDGYYRRFIEGFSKLALPLTQLTQK 790
GL GYYR+FI ++ +A P+T +K
Sbjct: 445 LGLTGYYRKFIPNYADIAKPMTSCLKK 471
>POL5_DROME (Q8I7P9) Retrovirus-related Pol polyprotein from
transposon opus [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1003
Score = 211 bits (536), Expect = 8e-54
Identities = 112/295 (37%), Positives = 175/295 (58%), Gaps = 6/295 (2%)
Query: 500 EFPEVFPEDITELPSEREVEFAIDLVPGTSPISIAPYRMSASELGELKKQLEELLEKQFI 559
EFP +F ++ + E V+ I PI Y + GE+++Q++ELL+ I
Sbjct: 94 EFPRIFEPPLSGMSVETAVKAEIR-TNTQDPIYAKSYPYPVNMRGEVERQIDELLQDGII 152
Query: 560 RPSVSPRGAPVLLVKKK-----DGSMRLCVDYRQLNKLTIKNRYPLPRIDD*MDQLVGAE 614
RPS SP +P+ +V KK + R+ VD+++LN +TI + YP+P I+ + L A+
Sbjct: 153 RPSNSPYNSPIWIVPKKPKPNGEKQYRMVVDFKRLNTVTIPDTYPIPDINATLASLGNAK 212
Query: 615 VFSKIDLRSGYHQIRVKAEDISKTAFRTRYGHYEYSVMPFGVSNAPGVFMEYMNRIFHPY 674
F+ +DL SG+HQI +K DI KTAF T G YE+ +PFG+ NAP +F ++ I +
Sbjct: 213 YFTTLDLTSGFHQIHMKESDIPKTAFSTLNGKYEFLRLPFGLKNAPAIFQRMIDDILREH 272
Query: 675 LDRFVVVFIDDILVYSKSEEEHAEHLRIVLQVLKENQLCAKLSKCEFWLKEVSFLGHVIS 734
+ + V+IDDI+V+S+ + H ++LR+VL L + L L K F +V FLG++++
Sbjct: 273 IGKVCYVYIDDIIVFSEDYDTHWKNLRLVLASLSKANLQVNLEKSHFLDTQVEFLGYIVT 332
Query: 735 KGGISVDPSKVDAVLQWESPKSVFEIKAIWGLDGYYRRFIEGFSKLALPLTQLTQ 789
GI DP KV A+ + P SV E+K G+ YYR+FI+ ++K+A PLT LT+
Sbjct: 333 ADGIKADPKKVRAISEMPPPTSVKELKRFLGMTSYYRKFIQDYAKVAKPLTNLTR 387
>RT21_SCHPO (Q05654) Retrotransposable element Tf2 155 kDa protein
type 1
Length = 1333
Score = 207 bits (527), Expect = 9e-53
Identities = 117/309 (37%), Positives = 178/309 (56%), Gaps = 3/309 (0%)
Query: 485 KVSGEKGASVLP-VVQEFPEVFPEDITE-LPSE-REVEFAIDLVPGTSPISIAPYRMSAS 541
KVS LP + +EF ++ E TE LP + +EF ++L + I Y +
Sbjct: 364 KVSNIVKEPELPDIYKEFKDITAETNTEKLPKPIKGLEFEVELTQENYRLPIRNYPLPPG 423
Query: 542 ELGELKKQLEELLEKQFIRPSVSPRGAPVLLVKKKDGSMRLCVDYRQLNKLTIKNRYPLP 601
++ + ++ + L+ IR S + PV+ V KK+G++R+ VDY+ LNK N YPLP
Sbjct: 424 KMQAMNDEINQGLKSGIIRESKAINACPVMFVPKKEGTLRMVVDYKPLNKYVKPNIYPLP 483
Query: 602 RIDD*MDQLVGAEVFSKIDLRSGYHQIRVKAEDISKTAFRTRYGHYEYSVMPFGVSNAPG 661
I+ + ++ G+ +F+K+DL+S YH IRV+ D K AFR G +EY VMP+G+S AP
Sbjct: 484 LIEQLLAKIQGSTIFTKLDLKSAYHLIRVRKGDEHKLAFRCPRGVFEYLVMPYGISTAPA 543
Query: 662 VFMEYMNRIFHPYLDRFVVVFIDDILVYSKSEEEHAEHLRIVLQVLKENQLCAKLSKCEF 721
F ++N I + VV ++DDIL++SKSE EH +H++ VLQ LK L +KCEF
Sbjct: 544 HFQYFINTILGEAKESHVVCYMDDILIHSKSESEHVKHVKDVLQKLKNANLIINQAKCEF 603
Query: 722 WLKEVSFLGHVISKGGISVDPSKVDAVLQWESPKSVFEIKAIWGLDGYYRRFIEGFSKLA 781
+V F+G+ IS+ G + +D VLQW+ PK+ E++ G Y R+FI S+L
Sbjct: 604 HQSQVKFIGYHISEKGFTPCQENIDKVLQWKQPKNRKELRQFLGSVNYLRKFIPKTSQLT 663
Query: 782 LPLTQLTQK 790
PL L +K
Sbjct: 664 HPLNNLLKK 672
>RT23_SCHPO (Q9UR07) Retrotransposable element Tf2 155 kDa protein
type 3
Length = 1333
Score = 204 bits (520), Expect = 6e-52
Identities = 116/309 (37%), Positives = 178/309 (57%), Gaps = 3/309 (0%)
Query: 485 KVSGEKGASVLP-VVQEFPEVFPEDITE-LPSE-REVEFAIDLVPGTSPISIAPYRMSAS 541
KVS LP + +EF ++ E TE LP + +EF ++L + I Y +
Sbjct: 364 KVSNIVKEPELPDIYKEFKDITAETNTEKLPKPIKGLEFEVELTQENYRLPIRNYPLPPG 423
Query: 542 ELGELKKQLEELLEKQFIRPSVSPRGAPVLLVKKKDGSMRLCVDYRQLNKLTIKNRYPLP 601
++ + ++ + L+ IR S + PV+ V KK+G++R+ VDY+ LNK N YPLP
Sbjct: 424 KMQAMNDEINQGLKSGIIRESKAINACPVMFVPKKEGTLRMVVDYKPLNKYVKPNIYPLP 483
Query: 602 RIDD*MDQLVGAEVFSKIDLRSGYHQIRVKAEDISKTAFRTRYGHYEYSVMPFGVSNAPG 661
I+ + ++ G+ +F+K+DL+S YH IRV+ D K AFR G +EY VMP+G+S AP
Sbjct: 484 LIEQLLAKIQGSTIFTKLDLKSAYHLIRVRKGDEHKLAFRCPRGVFEYLVMPYGISIAPA 543
Query: 662 VFMEYMNRIFHPYLDRFVVVFIDDILVYSKSEEEHAEHLRIVLQVLKENQLCAKLSKCEF 721
F ++N I + VV ++D+IL++SKSE EH +H++ VLQ LK L +KCEF
Sbjct: 544 HFQYFINTILGEVKESHVVCYMDNILIHSKSESEHVKHVKDVLQKLKNANLIINQAKCEF 603
Query: 722 WLKEVSFLGHVISKGGISVDPSKVDAVLQWESPKSVFEIKAIWGLDGYYRRFIEGFSKLA 781
+V F+G+ IS+ G + +D VLQW+ PK+ E++ G Y R+FI S+L
Sbjct: 604 HQSQVKFIGYHISEKGFTPCQENIDKVLQWKQPKNRKELRQFLGSVNYLRKFIPKTSQLT 663
Query: 782 LPLTQLTQK 790
PL L +K
Sbjct: 664 HPLNNLLKK 672
>RT22_SCHPO (Q9C0R2) Retrotransposable element Tf2 155 kDa protein
type 2
Length = 1333
Score = 204 bits (520), Expect = 6e-52
Identities = 116/309 (37%), Positives = 178/309 (57%), Gaps = 3/309 (0%)
Query: 485 KVSGEKGASVLP-VVQEFPEVFPEDITE-LPSE-REVEFAIDLVPGTSPISIAPYRMSAS 541
KVS LP + +EF ++ E TE LP + +EF ++L + I Y +
Sbjct: 364 KVSNIVKEPELPDIYKEFKDITAETNTEKLPKPIKGLEFEVELTQENYRLPIRNYPLPPG 423
Query: 542 ELGELKKQLEELLEKQFIRPSVSPRGAPVLLVKKKDGSMRLCVDYRQLNKLTIKNRYPLP 601
++ + ++ + L+ IR S + PV+ V KK+G++R+ VDY+ LNK N YPLP
Sbjct: 424 KMQAMNDEINQGLKSGIIRESKAINACPVMFVPKKEGTLRMVVDYKPLNKYVKPNIYPLP 483
Query: 602 RIDD*MDQLVGAEVFSKIDLRSGYHQIRVKAEDISKTAFRTRYGHYEYSVMPFGVSNAPG 661
I+ + ++ G+ +F+K+DL+S YH IRV+ D K AFR G +EY VMP+G+S AP
Sbjct: 484 LIEQLLAKIQGSTIFTKLDLKSAYHLIRVRKGDEHKLAFRCPRGVFEYLVMPYGISIAPA 543
Query: 662 VFMEYMNRIFHPYLDRFVVVFIDDILVYSKSEEEHAEHLRIVLQVLKENQLCAKLSKCEF 721
F ++N I + VV ++D+IL++SKSE EH +H++ VLQ LK L +KCEF
Sbjct: 544 HFQYFINTILGEVKESHVVCYMDNILIHSKSESEHVKHVKDVLQKLKNANLIINQAKCEF 603
Query: 722 WLKEVSFLGHVISKGGISVDPSKVDAVLQWESPKSVFEIKAIWGLDGYYRRFIEGFSKLA 781
+V F+G+ IS+ G + +D VLQW+ PK+ E++ G Y R+FI S+L
Sbjct: 604 HQSQVKFIGYHISEKGFTPCQENIDKVLQWKQPKNRKELRQFLGSVNYLRKFIPKTSQLT 663
Query: 782 LPLTQLTQK 790
PL L +K
Sbjct: 664 HPLNNLLKK 672
>YL52_CAEEL (P34431) Hypothetical protein F44E2.2 in chromosome III
Length = 2186
Score = 193 bits (491), Expect = 1e-48
Identities = 102/304 (33%), Positives = 173/304 (56%)
Query: 484 LKVSGEKGASVLPVVQEFPEVFPEDITELPSEREVEFAIDLVPGTSPISIAPYRMSASEL 543
LK + V+++F +VF EL E I+L G PI P + +
Sbjct: 896 LKRENGDDRKIWDVIEQFQDVFAISDDELGRNSGTECVIELKEGAEPIRQKPRPIPLALK 955
Query: 544 GELKKQLEELLEKQFIRPSVSPRGAPVLLVKKKDGSMRLCVDYRQLNKLTIKNRYPLPRI 603
E++K ++++L ++ IR S SP +PV+LVKKKDGS+R+C+DYR++NK+ N +PLP I
Sbjct: 956 PEIRKMIQKMLNQKVIRESKSPWSSPVVLVKKKDGSIRMCIDYRKVNKVVKNNAHPLPNI 1015
Query: 604 DD*MDQLVGAEVFSKIDLRSGYHQIRVKAEDISKTAFRTRYGHYEYSVMPFGVSNAPGVF 663
+ + L G ++++ D+ +G+ QI + + TAF +E++V+PFG+ +P +F
Sbjct: 1016 EATLQSLAGKKLYTVFDMIAGFWQIPLDEKSKEITAFAIGSELFEWNVLPFGLVISPALF 1075
Query: 664 MEYMNRIFHPYLDRFVVVFIDDILVYSKSEEEHAEHLRIVLQVLKENQLCAKLSKCEFWL 723
M I L V++DD+L+ SK E+H + ++ L ++++ + + SKC
Sbjct: 1076 QGTMEEIIGDLLGVCAFVYVDDLLIASKDMEQHLQDVKEALTRIRKSGMKLRASKCHIAK 1135
Query: 724 KEVSFLGHVISKGGISVDPSKVDAVLQWESPKSVFEIKAIWGLDGYYRRFIEGFSKLALP 783
KEV +LGH ++ G+ K D + Q+ P +V E+++ GL GYYR+FI F+++A
Sbjct: 1136 KEVEYLGHKVTLDGVETQEVKTDKMKQFSRPTNVKELQSFLGLVGYYRKFILNFAQIASS 1195
Query: 784 LTQL 787
LT L
Sbjct: 1196 LTSL 1199
>POLY_DROME (P10401) Retrovirus-related Pol polyprotein from
transposon gypsy [Contains: Reverse transcriptase (EC
2.7.7.49); Endonuclease]
Length = 1035
Score = 182 bits (462), Expect = 3e-45
Identities = 111/373 (29%), Positives = 185/373 (48%), Gaps = 20/373 (5%)
Query: 422 LKNMDVIFGMDWLLAFGVSINCLTKSVTFSKPVEELGREFLTAEQVKKSLDGEACVFMMF 481
L D I G+D L GV +N S+ + E+L + + V F
Sbjct: 85 LNAFDAIIGLDLLTQAGVKLNLAEDSLEYQGIAEKL--HYFSCPSVN---------FTDV 133
Query: 482 ASLKVSGEKGASVLPVVQEFPEVFPEDITELPSEREVEFAIDLVPGTSPISIA-PYRMSA 540
+ V + + F LP V I V S A P M
Sbjct: 134 NDIVVPDSVKKEFKDTIIRRKKAFSTTNEALPFNTAVTATIRTVDNEPVYSRAYPTLMGV 193
Query: 541 SELGELKKQLEELLEKQFIRPSVSPRGAPVLLVKKK------DGSMRLCVDYRQLNKLTI 594
S+ + ++++LL+ IRPS SP +P +V KK + + RL +D+R+LN+ TI
Sbjct: 194 SDF--VNNEVKQLLKDGIIRPSRSPYNSPTWVVDKKGTDAFGNPNKRLVIDFRKLNEKTI 251
Query: 595 KNRYPLPRIDD*MDQLVGAEVFSKIDLRSGYHQIRVKAEDISKTAFRTRYGHYEYSVMPF 654
+RYP+P I + L A+ F+ +DL+SGYHQI + D KT+F G YE+ +PF
Sbjct: 252 PDRYPMPSIPMILANLGKAKFFTTLDLKSGYHQIYLAEHDREKTSFSVNGGKYEFCRLPF 311
Query: 655 GVSNAPGVFMEYMNRIFHPYLDRFVVVFIDDILVYSKSEEEHAEHLRIVLQVLKENQLCA 714
G+ NA +F ++ + + + V++DD++++S++E +H H+ VL+ L + +
Sbjct: 312 GLRNASSIFQRALDDVLREQIGKICYVYVDDVIIFSENESDHVRHIDTVLKCLIDANMRV 371
Query: 715 KLSKCEFWLKEVSFLGHVISKGGISVDPSKVDAVLQWESPKSVFEIKAIWGLDGYYRRFI 774
K F+ + V +LG ++SK G DP KV A+ ++ P V+++++ GL YYR FI
Sbjct: 372 SQEKTRFFKESVEYLGFIVSKDGTKSDPEKVKAIQEYPEPDCVYKVRSFLGLASYYRVFI 431
Query: 775 EGFSKLALPLTQL 787
+ F+ +A P+T +
Sbjct: 432 KDFAAIARPITDI 444
>POL4_DROME (P10394) Retrovirus-related Pol polyprotein from
transposon 412 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1237
Score = 171 bits (434), Expect = 5e-42
Identities = 102/312 (32%), Positives = 160/312 (50%), Gaps = 20/312 (6%)
Query: 499 QEFPEVFPEDITELPSEREVEFAIDLVPGT--------------SPISIAPYRMSASELG 544
+ FPE+F + + SE FA++ P T P+ YR S++
Sbjct: 269 KNFPELFKSQLENICSEYIDIFALESEPITVNNLYKQQLRLKDDEPVYTKNYRSPHSQVE 328
Query: 545 ELKKQLEELLEKQFIRPSVSPRGAPVLLVKKKDG------SMRLCVDYRQLNKLTIKNRY 598
E++ Q+++L++ + + PSVS +P+LLV KK RL +DYRQ+NK + +++
Sbjct: 329 EIQAQVQKLIKDKIVEPSVSQYNSPLLLVPKKSSPNSDKKKWRLVIDYRQINKKLLADKF 388
Query: 599 PLPRIDD*MDQLVGAEVFSKIDLRSGYHQIRVKAEDISKTAFRTRYGHYEYSVMPFGVSN 658
PLPRIDD +DQL A+ FS +DL SG+HQI + T+F T G Y ++ +PFG+
Sbjct: 389 PLPRIDDILDQLGRAKYFSCLDLMSGFHQIELDEGSRDITSFSTSNGSYRFTRLPFGLKI 448
Query: 659 APGVFMEYMNRIFHPYLDRFVVVFIDDILVYSKSEEEHAEHLRIVLQVLKENQLCAKLSK 718
AP F M F +++DD++V SE+ ++L V +E L K
Sbjct: 449 APNSFQRMMTIAFSGIEPSQAFLYMDDLIVIGCSEKHMLKNLTEVFGKCREYNLKLHPEK 508
Query: 719 CEFWLKEVSFLGHVISKGGISVDPSKVDAVLQWESPKSVFEIKAIWGLDGYYRRFIEGFS 778
C F++ EV+FLGH + GI D K D + + P + YYRRFI+ F+
Sbjct: 509 CSFFMHEVTFLGHKCTDKGILPDDKKYDVIQNYPVPHDADSARRFVAFCNYYRRFIKNFA 568
Query: 779 KLALPLTQLTQK 790
+ +T+L +K
Sbjct: 569 DYSRHITRLCKK 580
>POL_CAMVE (Q02964) Enzymatic polyprotein [Contains: Aspartic
protease (EC 3.4.23.-); Endonuclease; Reverse
transcriptase (EC 2.7.7.49)]
Length = 679
Score = 152 bits (384), Expect = 3e-36
Identities = 85/261 (32%), Positives = 149/261 (56%), Gaps = 10/261 (3%)
Query: 485 KVSGEKGASVLPVVQEFPEVFPEDITELP-----SEREVEFAIDLVPGTSPISIAPYRMS 539
++S EK +Q+ E+ + +E P +++ ++ +I L + I + P + S
Sbjct: 196 RLSEEKLFITQQRMQKIEELLEKVCSENPLDPNKTKQWMKASIKLSDPSKAIKVKPMKYS 255
Query: 540 ASELGELKKQLEELLEKQFIRPSVSPRGAPVLLV----KKKDGSMRLCVDYRQLNKLTIK 595
+ E KQ++ELL+ + I+PS SP AP LV +K+ G R+ V+Y+ +NK TI
Sbjct: 256 PMDREEFDKQIKELLDLKVIKPSKSPHMAPAFLVNNEAEKRRGKKRMVVNYKAMNKATIG 315
Query: 596 NRYPLPRIDD*MDQLVGAEVFSKIDLRSGYHQIRVKAEDISKTAFRTRYGHYEYSVMPFG 655
+ Y LP D+ + + G ++FS D +SG+ Q+ + E TAF GHYE++V+PFG
Sbjct: 316 DAYNLPNKDELLTLIRGKKIFSSFDCKSGFWQVLLDQESRPLTAFTCPQGHYEWNVVPFG 375
Query: 656 VSNAPGVFMEYMNRIFHPYLDRFVVVFIDDILVYSKSEEEHAEHLRIVLQVLKENQLCAK 715
+ AP +F +M+ F + +F V++DDILV+S +EE+H H+ ++LQ ++ +
Sbjct: 376 LKQAPSIFQRHMDEAFRVF-RKFCCVYVDDILVFSNNEEDHLLHVAMILQKCNQHGIILS 434
Query: 716 LSKCEFWLKEVSFLGHVISKG 736
K + + K+++FLG I +G
Sbjct: 435 KKKAQLFKKKINFLGLEIDEG 455
>POL_CAMVC (P03555) Enzymatic polyprotein [Contains: Aspartic
protease (EC 3.4.23.-); Endonuclease; Reverse
transcriptase (EC 2.7.7.49)]
Length = 679
Score = 152 bits (384), Expect = 3e-36
Identities = 85/261 (32%), Positives = 149/261 (56%), Gaps = 10/261 (3%)
Query: 485 KVSGEKGASVLPVVQEFPEVFPEDITELP-----SEREVEFAIDLVPGTSPISIAPYRMS 539
++S EK +Q+ E+ + +E P +++ ++ +I L + I + P + S
Sbjct: 196 RLSEEKLFITQQRMQKIEELLEKVCSENPLDPNKTKQWMKASIKLSDPSKAIKVKPMKYS 255
Query: 540 ASELGELKKQLEELLEKQFIRPSVSPRGAPVLLV----KKKDGSMRLCVDYRQLNKLTIK 595
+ E KQ++ELL+ + I+PS SP AP LV +K+ G R+ V+Y+ +NK TI
Sbjct: 256 PMDREEFDKQIKELLDLKVIKPSKSPHMAPAFLVNNEAEKRRGKKRMVVNYKAMNKATIG 315
Query: 596 NRYPLPRIDD*MDQLVGAEVFSKIDLRSGYHQIRVKAEDISKTAFRTRYGHYEYSVMPFG 655
+ Y LP D+ + + G ++FS D +SG+ Q+ + E TAF GHYE++V+PFG
Sbjct: 316 DAYNLPNKDELLTLIRGKKIFSSFDCKSGFWQVLLDQESRPLTAFTCPQGHYEWNVVPFG 375
Query: 656 VSNAPGVFMEYMNRIFHPYLDRFVVVFIDDILVYSKSEEEHAEHLRIVLQVLKENQLCAK 715
+ AP +F +M+ F + +F V++DDILV+S +EE+H H+ ++LQ ++ +
Sbjct: 376 LKQAPSIFQRHMDEAFRVF-RKFCCVYVDDILVFSNNEEDHLLHVAMILQKCNQHGIILS 434
Query: 716 LSKCEFWLKEVSFLGHVISKG 736
K + + K+++FLG I +G
Sbjct: 435 KKKAQLFKKKINFLGLEIDEG 455
>POL_CAMVS (P03554) Enzymatic polyprotein [Contains: Aspartic
protease (EC 3.4.23.-); Endonuclease; Reverse
transcriptase (EC 2.7.7.49)]
Length = 679
Score = 152 bits (383), Expect = 4e-36
Identities = 84/261 (32%), Positives = 149/261 (56%), Gaps = 10/261 (3%)
Query: 485 KVSGEKGASVLPVVQEFPEVFPEDITELP-----SEREVEFAIDLVPGTSPISIAPYRMS 539
++S EK +Q+ E+ + +E P +++ ++ +I L + I + P + S
Sbjct: 196 RLSEEKLFITQQRMQKIEELLEKVCSENPLDPNKTKQWMKASIKLSDPSKAIKVKPMKYS 255
Query: 540 ASELGELKKQLEELLEKQFIRPSVSPRGAPVLLV----KKKDGSMRLCVDYRQLNKLTIK 595
+ E KQ++ELL+ + I+PS SP AP LV +K+ G R+ V+Y+ +NK T+
Sbjct: 256 PMDREEFDKQIKELLDLKVIKPSKSPHMAPAFLVNNEAEKRRGKKRMVVNYKAMNKATVG 315
Query: 596 NRYPLPRIDD*MDQLVGAEVFSKIDLRSGYHQIRVKAEDISKTAFRTRYGHYEYSVMPFG 655
+ Y LP D+ + + G ++FS D +SG+ Q+ + E TAF GHYE++V+PFG
Sbjct: 316 DAYNLPNKDELLTLIRGKKIFSSFDCKSGFWQVLLDQESRPLTAFTCPQGHYEWNVVPFG 375
Query: 656 VSNAPGVFMEYMNRIFHPYLDRFVVVFIDDILVYSKSEEEHAEHLRIVLQVLKENQLCAK 715
+ AP +F +M+ F + +F V++DDILV+S +EE+H H+ ++LQ ++ +
Sbjct: 376 LKQAPSIFQRHMDEAFRVF-RKFCCVYVDDILVFSNNEEDHLLHVAMILQKCNQHGIILS 434
Query: 716 LSKCEFWLKEVSFLGHVISKG 736
K + + K+++FLG I +G
Sbjct: 435 KKKAQLFKKKINFLGLEIDEG 455
>POL_CAMVD (P03556) Enzymatic polyprotein [Contains: Aspartic
protease (EC 3.4.23.-); Endonuclease; Reverse
transcriptase (EC 2.7.7.49)]
Length = 674
Score = 149 bits (376), Expect = 3e-35
Identities = 83/261 (31%), Positives = 148/261 (55%), Gaps = 10/261 (3%)
Query: 485 KVSGEKGASVLPVVQEFPEVFPEDITELP-----SEREVEFAIDLVPGTSPISIAPYRMS 539
++S EK +Q+ E+ + +E P +++ ++ +I L + I + P + S
Sbjct: 191 RLSEEKLFITQQRMQKIEELLEKVCSENPLDPNKTKQWMKASIKLSDPSKAIKVKPMKYS 250
Query: 540 ASELGELKKQLEELLEKQFIRPSVSPRGAPVLLV----KKKDGSMRLCVDYRQLNKLTIK 595
+ E KQ++ELL+ + I+PS SP AP LV +K+ G R+ V+Y+ +NK T+
Sbjct: 251 PMDREEFDKQIKELLDLKVIKPSKSPHMAPAFLVNNEAEKRRGKKRMVVNYKAMNKATVG 310
Query: 596 NRYPLPRIDD*MDQLVGAEVFSKIDLRSGYHQIRVKAEDISKTAFRTRYGHYEYSVMPFG 655
+ Y P D+ + + G ++FS D +SG+ Q+ + E TAF GHYE++V+PFG
Sbjct: 311 DAYNPPNKDELLTLIRGKKIFSSFDCKSGFWQVLLDQESRPLTAFTCPQGHYEWNVVPFG 370
Query: 656 VSNAPGVFMEYMNRIFHPYLDRFVVVFIDDILVYSKSEEEHAEHLRIVLQVLKENQLCAK 715
+ AP +F +M+ F + +F V++DDILV+S +EE+H H+ ++LQ ++ +
Sbjct: 371 LKQAPSIFQRHMDEAFRVF-RKFCCVYVDDILVFSNNEEDHLLHVAMILQKCNQHGIILS 429
Query: 716 LSKCEFWLKEVSFLGHVISKG 736
K + + K+++FLG I +G
Sbjct: 430 KKKAQLFKKKINFLGLEIDEG 450
>POL_CAMVN (Q00962) Enzymatic polyprotein [Contains: Aspartic
protease (EC 3.4.23.-); Endonuclease; Reverse
transcriptase (EC 2.7.7.49)]
Length = 680
Score = 147 bits (372), Expect = 8e-35
Identities = 82/261 (31%), Positives = 148/261 (56%), Gaps = 10/261 (3%)
Query: 485 KVSGEKGASVLPVVQEFPEVFPEDITELP-----SEREVEFAIDLVPGTSPISIAPYRMS 539
++S EK +Q+ E+ + +E P +++ ++ +I L + I + P + S
Sbjct: 197 RLSEEKLFITQQRMQKTEELLEKVCSENPLDPNKTKQWMKASIKLSDPSKAIKVKPMKYS 256
Query: 540 ASELGELKKQLEELLEKQFIRPSVSPRGAPVLLVKKKD----GSMRLCVDYRQLNKLTIK 595
+ E KQ++ELL+ + I+PS SP AP LV + G+ R+ V+Y+ +NK T+
Sbjct: 257 PMDREEFDKQIKELLDLKVIKPSKSPHMAPAFLVNNEAENGRGNKRMVVNYKAMNKATVG 316
Query: 596 NRYPLPRIDD*MDQLVGAEVFSKIDLRSGYHQIRVKAEDISKTAFRTRYGHYEYSVMPFG 655
+ Y LP D+ + + G ++FS D +SG+ Q+ + E TAF GHYE++V+PFG
Sbjct: 317 DAYNLPNKDELLTLIRGKKIFSSFDCKSGFWQVLLDQESRPLTAFTCPQGHYEWNVVPFG 376
Query: 656 VSNAPGVFMEYMNRIFHPYLDRFVVVFIDDILVYSKSEEEHAEHLRIVLQVLKENQLCAK 715
+ AP +F +M+ F + +F V++DDI+V+S +EE+H H+ ++LQ ++ +
Sbjct: 377 LKQAPSIFQRHMDEAFRVF-RKFCCVYVDDIVVFSNNEEDHLLHVAMILQKCNQHGIILS 435
Query: 716 LSKCEFWLKEVSFLGHVISKG 736
K + + K+++FLG I +G
Sbjct: 436 KKKAQLFKKKINFLGLEIDEG 456
>POL_COYMV (P19199) Putative polyprotein [Contains: Coat protein;
Protease (EC 3.4.23.-); Reverse transcriptase (EC
2.7.7.49); Ribonuclease H (EC 3.1.26.4)]
Length = 1886
Score = 146 bits (369), Expect = 2e-34
Identities = 125/486 (25%), Positives = 228/486 (46%), Gaps = 45/486 (9%)
Query: 331 NAKEVEQPDNLIRGMCFINSTPLIAIIDTGATHSFISASCVERL---GLVVTPLLRGMVI 387
N K +PDN+ +IN AI+DTGAT I S + VT R ++
Sbjct: 1200 NVKVGIKPDNM--EPYYIN-----AIVDTGATACLIQISAIPENYYEDAKVTVNFRSVL- 1251
Query: 388 DTPASGSGTTSFMCAKCPVNFGDVDFKLDLVCLPLKNMD----VIFGMDWLLAFGVSINC 443
G GT++ M + G+ F++ + + + +I G ++ + +
Sbjct: 1252 -----GIGTSTQMIKAGRILIGEQYFRMPVTYVMNMGLSPGIQMIIGCSFIRSLEGGLRI 1306
Query: 444 LTKSVTFSKPVE--ELGREFLTAEQVKKSLDGEACVFMMFASLKVSG-------EKGASV 494
+TF K V E R A +++ E + AS++ K +
Sbjct: 1307 EKDIITFYKLVTSIETSRTTQVANSIEELELSEDEYLNIAASVETPSFLDQEFARKNKDL 1366
Query: 495 LPVVQEFPEVFPEDITELPSEREVEFAIDLVPGTSPISIAPYR-MSASELGELKKQLEEL 553
L ++E + E+ E +++ ++++ I P + ++ + + +Q+ L
Sbjct: 1367 LKEMKEMKYI-GENPMEFWKNNKIKCKLNIINPDIKIMGRPIKHVTPGDEEAMTRQINLL 1425
Query: 554 LEKQFIRPSVSPRGAPVLLV-----------KKKDGSMRLCVDYRQLNKLTIKNRYPLPR 602
L+ + IRPS S + +V K+K G R+ +Y+ LN+ T ++Y LP
Sbjct: 1426 LQMKVIRPSESKHRSTAFIVRSGTEIDPITGKEKKGKERMVFNYKLLNENTESDQYSLPG 1485
Query: 603 IDD*MDQLVGAEVFSKIDLRSGYHQIRVKAEDISKTAFRTRYGHYEYSVMPFGVSNAPGV 662
I+ + ++ ++++SK DL+SG+ Q+ ++ E + TAF YE+ VMPFG+ NAP +
Sbjct: 1486 INTIISKVGRSKIYSKFDLKSGFWQVAMEEESVPWTAFLAGNKLYEWLVMPFGLKNAPAI 1545
Query: 663 FMEYMNRIFHPYLDRFVVVFIDDILVYSKSEEEHAEHLRIVLQVLKENQLCAKLSKCEFW 722
F M+ +F ++F+ V+IDDILV+S++ E+H++HL +LQ+ KEN L +K +
Sbjct: 1546 FQRKMDNVFKG-TEKFIAVYIDDILVFSETAEQHSQHLYTMLQLCKENGLILSPTKMKIG 1604
Query: 723 LKEVSFLGHVISKGGISVDPSKVDAVLQWESPKSVFE--IKAIWGLDGYYRRFIEGFSKL 780
E+ FLG + I + P + + + K +++ G+ Y R +I+ KL
Sbjct: 1605 TPEIDFLGASLGCTKIKLQPHIISKICDFSDEKLATPEGMRSWLGILSYARNYIQDIGKL 1664
Query: 781 ALPLTQ 786
PL Q
Sbjct: 1665 VQPLRQ 1670
Score = 33.9 bits (76), Expect = 1.7
Identities = 12/17 (70%), Positives = 13/17 (75%)
Query: 265 KCYRCGTLGHYANECMN 281
KCY CG GHYAN+C N
Sbjct: 880 KCYICGQEGHYANQCRN 896
>RRPO_OENBE (P31843) RNA-directed DNA polymerase homolog (Reverse
transcriptase homolog)
Length = 142
Score = 145 bits (367), Expect = 3e-34
Identities = 68/128 (53%), Positives = 92/128 (71%), Gaps = 3/128 (2%)
Query: 579 SMRLCVDYRQLNKLTIKNRYPLPRIDD*MDQLVGAEVFSKIDLRSGYHQIRVKAEDISKT 638
S+R+C+DYR L K+TIKN+YP+PR+DD D+L A F+K+DLRSGY Q+R+ D KT
Sbjct: 5 SLRMCIDYRALTKVTIKNKYPIPRVDDLFDRLAQATWFTKLDLRSGYWQVRIAKGDEPKT 64
Query: 639 AFRTRYGHYEYSVMPFGVSNAPGVFMEYMNRIFHPYLDRFVVVFIDDILV---YSKSEEE 695
TRYG +E+ VMPFG++NA F MN + + YLD FVVV++DD++V YS S E
Sbjct: 65 TCVTRYGSFEFRVMPFGLTNALATFCNLMNNVLYEYLDHFVVVYLDDLVVYTIYSNSLHE 124
Query: 696 HAEHLRIV 703
H +HLR+V
Sbjct: 125 HIKHLRVV 132
>POL_SOCMV (P15629) Enzymatic polyprotein [Contains: Aspartic
protease (EC 3.4.23.-); Endonuclease; Reverse
transcriptase (EC 2.7.7.49)]
Length = 692
Score = 137 bits (346), Expect = 8e-32
Identities = 111/398 (27%), Positives = 198/398 (48%), Gaps = 33/398 (8%)
Query: 412 DFKLDLVCLPLKNMDVIFGMDWLLAFGVSINCLT------KSVTFSKPVEELGREFLTAE 465
+F + ++ L +D+I G ++L + I L K++ K + + + LT
Sbjct: 86 EFLIPIIYLHDSGLDLIIGNNFLKLYQPFIQRLETIELRWKNLNNPKESQMISTKILTKN 145
Query: 466 QVKKSLDGEACVFMMFASLKVSGEKGASVLPVVQEFPEVFPED-ITELPSEREVEFAIDL 524
+V K + F + + EK + ++ EV E + E ++ + I L
Sbjct: 146 EVLK---------LSFEKIHICLEKYLFFKTIEEQLEEVCSEHPLDETKNKNGLLIEIRL 196
Query: 525 VPGTSPISIA---PYRMSASELGELKKQLEELLEKQFIRPSVSPRGAPVLLVKK----KD 577
I++ PY + ++ E K++ E+LL+K IR S SP AP V+ K
Sbjct: 197 KDPLQEINVTNRIPYTIR--DVQEFKEECEDLLKKGLIRESQSPHSAPAFYVENHNEIKR 254
Query: 578 GSMRLCVDYRQLNKLTIKNRYPLPRIDD*MDQLVGAEVFSKIDLRSGYHQIRVKAEDISK 637
G R+ ++Y+++N+ TI + Y LPR D ++++ G+ FS +D +SGY+Q+R+
Sbjct: 255 GKRRMVINYKKMNEATIGDSYKLPRKDFILEKIKGSLWFSSLDAKSGYYQLRLHENTKPL 314
Query: 638 TAFR-TRYGHYEYSVMPFGVSNAPGVFMEYMNRIFHPYLDRFVVVFIDDILVYSK-SEEE 695
TAF HYE++V+ FG+ AP ++ +M++ L+ + +IDDIL+++K S+E+
Sbjct: 315 TAFSCPPQKHYEWNVLSFGLKQAPSIYQRFMDQSLKG-LEHICLAYIDDILIFTKGSKEQ 373
Query: 696 HAEHLRIVLQVLKENQLCAKLSKCEFWLKEVSFLG-HVISKGGISVDPSKVDAVLQW-ES 753
H +RIVLQ +KE + K + +E+ +LG + G I + P + +LQ+ +
Sbjct: 374 HVNDVRIVLQRIKEKGIIISKKKSKLIQQEIEYLGLKIQGNGEIDLSPHTQEKILQFPDE 433
Query: 754 PKSVFEIKAIWGLDGYYRRFIEGFSK-LALPLTQLTQK 790
+ +I+ G Y EGF K LAL L +K
Sbjct: 434 LEDRKQIQRFLGCINYIAN--EGFFKNLALERKHLQKK 469
>POL_FMVD (P09523) Enzymatic polyprotein [Contains: Aspartic
protease (EC 3.4.23.-); Endonuclease; Reverse
transcriptase (EC 2.7.7.49)]
Length = 666
Score = 131 bits (330), Expect = 6e-30
Identities = 71/232 (30%), Positives = 129/232 (55%), Gaps = 5/232 (2%)
Query: 509 ITELPSEREVEFAIDLVPGTSPISIAPYRMSASELGELKKQLEELLEKQFIRPSVSPRGA 568
I + S++ ++ +I L+ I + P S + KQ++ELL+ I PS S +
Sbjct: 218 IDPIKSKQWMKASIKLIDPLKVIRVKPMSYSPQDREGFAKQIKELLDLGLIIPSKSQHMS 277
Query: 569 PVLLVK----KKDGSMRLCVDYRQLNKLTIKNRYPLPRIDD*MDQLVGAEVFSKIDLRSG 624
P LV+ ++ G R+ V+Y+ +N+ TI + + LP + + + L G +FS D +SG
Sbjct: 278 PAFLVENEAERRRGKKRMVVNYKAINQATIGDSHNLPNMQELLTLLRGKSIFSSFDCKSG 337
Query: 625 YHQIRVKAEDISKTAFRTRYGHYEYSVMPFGVSNAPGVFMEYMNRIFHPYLDRFVVVFID 684
+ Q+ + E TAF GH+++ V+PFG+ AP +F +M + D+F +V++D
Sbjct: 338 FWQVVLDEESQKLTAFTCPQGHFQWKVVPFGLKQAPSIFQRHMQTALNG-ADKFCMVYVD 396
Query: 685 DILVYSKSEEEHAEHLRIVLQVLKENQLCAKLSKCEFWLKEVSFLGHVISKG 736
DI+V+S SE +H H+ VL+++++ + K + ++++FLG I KG
Sbjct: 397 DIIVFSNSELDHYNHVYAVLKIVEKYGIILSKKKANLFKEKINFLGLEIDKG 448
>POL_RTBVP (P27502) Polyprotein (P194 protein) [Contains: Coat
protein; Protease (EC 3.4.23.-); Reverse transcriptase
(EC 2.7.7.49); Ribonuclease H (EC 3.1.26.4)]
Length = 1675
Score = 131 bits (329), Expect = 8e-30
Identities = 93/308 (30%), Positives = 162/308 (52%), Gaps = 14/308 (4%)
Query: 493 SVLPVVQEFPEV--FPEDITELPSEREVEFAIDLVPGTSPISIAPYRMSASELGELKKQL 550
S++ +V+E + +DIT+ + +F I + PY + E+ E KQ+
Sbjct: 1146 SLITMVKELEALGFIGDDITKNRTTWVCDFKIINPDINITCATIPYTPADKEVFE--KQI 1203
Query: 551 EELLEKQFIR---PSVSPRGAPVLLVKKKDG---SMRLCVDYRQLNKLTIKNRYPLPRID 604
+ELL+ + I+ P+ R A ++ + R+ +Y++LN + + +P
Sbjct: 1204 KELLDNKLIKKADPTCRHRTAAFIVRNHSEEVAQKPRIVYNYKRLNDNMHTDPFNIPHKI 1263
Query: 605 D*MDQLVGAEVFSKIDLRSGYHQIRVKAEDISKTAFRTRYGHYEYSVMPFGVSNAPGVFM 664
++ + A +FSK DL++G+H +++K + T F G Y ++V PFG++NAP F
Sbjct: 1264 SMINLIQKANIFSKFDLKAGFHHMKLKDDFKDWTTFTCSEGLYTWNVCPFGIANAPCAFQ 1323
Query: 665 EYMNRIFHPYLDRFVVVFIDDILVYSKSEEEHAEHLRIVLQVLKENQLCAKLSKCEFWLK 724
+M F +F +++IDDIL+ S +E+EH EHL+I +KE K + +LK
Sbjct: 1324 RFMQESFGDL--KFALLYIDDILIASNNEKEHIEHLKIFFNRVKEVGCVLSKKKSKMFLK 1381
Query: 725 EVSFLGHVISKGGISVDPSKVDAVLQWESPK--SVFEIKAIWGLDGYYRRFIEGFSKLAL 782
EV +LG I +G IS+ P VD + +++ K ++ ++A GL Y R +I+ SKL
Sbjct: 1382 EVEYLGVEIKEGKISLQPHIVDKIKKFDKNKLNTLKGLQAYLGLLNYARGYIKDLSKLVG 1441
Query: 783 PLTQLTQK 790
PL + T K
Sbjct: 1442 PLYKKTGK 1449
>POL_CERV (P05400) Enzymatic polyprotein [Contains: Aspartic
protease (EC 3.4.23.-); Endonuclease; Reverse
transcriptase (EC 2.7.7.49)]
Length = 659
Score = 119 bits (298), Expect = 3e-26
Identities = 69/229 (30%), Positives = 130/229 (56%), Gaps = 7/229 (3%)
Query: 514 SEREVEFAIDLVPGTSPISIAPYRMSASELGELKKQLEELLEKQFIRPSVSPRGAPVLLV 573
S++ + I+L+ + + + P S S+ E +Q++ELLE + I+PS S +P LV
Sbjct: 212 SKQWMTATIELIDPKTVVKVKPMSYSPSDREEFDRQIKELLELKVIKPSKSTHMSPAFLV 271
Query: 574 K----KKDGSMRLCVDYRQLNKLTIKNRYPLPRIDD*MDQLVGAEVFSKIDLRSGYHQIR 629
+ ++ G R+ V+Y+ +NK T + + LP D+ + + G +++S D +SG Q+
Sbjct: 272 ENEAERRRGKKRMVVNYKAMNKATKGDAHNLPNKDELLTLVRGKKIYSSFDCKSGLWQVL 331
Query: 630 VKAEDISKTAFRTRYGHYEYSVMPFGVSNAPGVFME-YMNRIFHPYLDRFVVVFIDDILV 688
+ E TAF GHY+++V+PFG+ AP +F + Y N + Y ++ V++DDILV
Sbjct: 332 LDKESQLLTAFTCPQGHYQWNVVPFGLKQAPSIFPKTYANSHSNQY-SKYCCVYVDDILV 390
Query: 689 YSKS-EEEHAEHLRIVLQVLKENQLCAKLSKCEFWLKEVSFLGHVISKG 736
+S + +EH H+ +L+ ++ + K + + ++++FLG I +G
Sbjct: 391 FSNTGRKEHYIHVLNILRRCEKLGIILSKKKAQLFKEKINFLGLEIDQG 439
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.321 0.138 0.414
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 94,233,162
Number of Sequences: 164201
Number of extensions: 4159797
Number of successful extensions: 10799
Number of sequences better than 10.0: 243
Number of HSP's better than 10.0 without gapping: 111
Number of HSP's successfully gapped in prelim test: 132
Number of HSP's that attempted gapping in prelim test: 9989
Number of HSP's gapped (non-prelim): 532
length of query: 790
length of database: 59,974,054
effective HSP length: 118
effective length of query: 672
effective length of database: 40,598,336
effective search space: 27282081792
effective search space used: 27282081792
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 70 (31.6 bits)
Medicago: description of AC126008.1


