

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
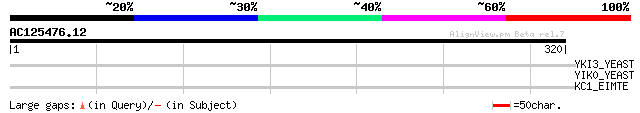
Query= AC125476.12 - phase: 0 /pseudo
(320 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
YKI3_YEAST (P36079) Hypothetical 23.7 kDa protein in MDH1-VMA5 i... 35 0.33
YIK0_YEAST (P40490) Hypothetical 13.5 kDa protein in MOB1-SGA1 i... 33 0.95
KC1_EIMTE (Q6QNL9) Casein kinase I (EC 2.7.1.-) 31 3.6
>YKI3_YEAST (P36079) Hypothetical 23.7 kDa protein in MDH1-VMA5
intergenic region
Length = 204
Score = 34.7 bits (78), Expect = 0.33
Identities = 12/21 (57%), Positives = 18/21 (85%)
Query: 75 LLKMIYIYIYIYIYIYIYIYI 95
+ + IYIY+Y+YIY++ YIYI
Sbjct: 5 IYRYIYIYMYLYIYVHTYIYI 25
Score = 33.5 bits (75), Expect = 0.73
Identities = 12/16 (75%), Positives = 15/16 (93%)
Query: 80 YIYIYIYIYIYIYIYI 95
YIY YIYIY+Y+YIY+
Sbjct: 4 YIYRYIYIYMYLYIYV 19
Score = 32.3 bits (72), Expect = 1.6
Identities = 11/16 (68%), Positives = 15/16 (93%)
Query: 79 IYIYIYIYIYIYIYIY 94
IY+Y+YIY++ YIYIY
Sbjct: 11 IYMYLYIYVHTYIYIY 26
>YIK0_YEAST (P40490) Hypothetical 13.5 kDa protein in MOB1-SGA1
intergenic region
Length = 117
Score = 33.1 bits (74), Expect = 0.95
Identities = 16/38 (42%), Positives = 23/38 (60%), Gaps = 4/38 (10%)
Query: 79 IYIYIYIYIYIYIYIYIERERERER----GREGESKGR 112
IYIYIY+Y+ +YI + I R E+ ++ E KGR
Sbjct: 33 IYIYIYVYVNVYIQLVISRAEWAEKHSIVEKKREEKGR 70
Score = 32.7 bits (73), Expect = 1.2
Identities = 14/31 (45%), Positives = 19/31 (61%)
Query: 79 IYIYIYIYIYIYIYIYIERERERERGREGES 109
IY +IYIYIY+Y+ +YI+ R E S
Sbjct: 29 IYAHIYIYIYVYVNVYIQLVISRAEWAEKHS 59
Score = 30.0 bits (66), Expect = 8.0
Identities = 10/16 (62%), Positives = 14/16 (87%)
Query: 80 YIYIYIYIYIYIYIYI 95
YIY +IYIYIY+Y+ +
Sbjct: 28 YIYAHIYIYIYVYVNV 43
Score = 30.0 bits (66), Expect = 8.0
Identities = 11/17 (64%), Positives = 14/17 (81%)
Query: 79 IYIYIYIYIYIYIYIYI 95
I YIY +IYIYIY+Y+
Sbjct: 25 ICTYIYAHIYIYIYVYV 41
>KC1_EIMTE (Q6QNL9) Casein kinase I (EC 2.7.1.-)
Length = 335
Score = 31.2 bits (69), Expect = 3.6
Identities = 14/37 (37%), Positives = 23/37 (61%), Gaps = 1/37 (2%)
Query: 82 YIYIYIYIYIYIYIERERERERGREGESKGRMQGQMW 118
Y Y +I+ + +++ ERERER+R R ++G G W
Sbjct: 282 YQYDFIFDWTFLHAERERERQR-RSMVNQGAESGNQW 317
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.361 0.162 0.622
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 30,519,958
Number of Sequences: 164201
Number of extensions: 1050051
Number of successful extensions: 4328
Number of sequences better than 10.0: 3
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 1
Number of HSP's that attempted gapping in prelim test: 4300
Number of HSP's gapped (non-prelim): 13
length of query: 320
length of database: 59,974,054
effective HSP length: 110
effective length of query: 210
effective length of database: 41,911,944
effective search space: 8801508240
effective search space used: 8801508240
T: 11
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.9 bits)
S2: 66 (30.0 bits)
Medicago: description of AC125476.12


