

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
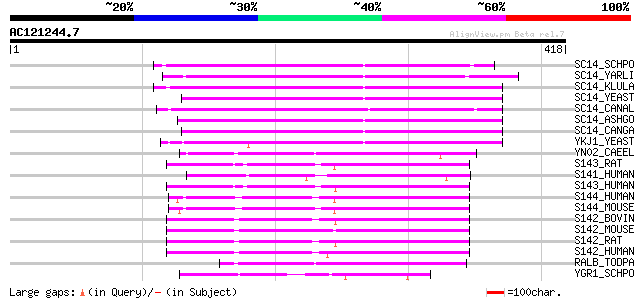
Query= AC121244.7 - phase: 0
(418 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
SC14_SCHPO (Q10137) Sec14 cytosolic factor (Phosphatidylinositol... 166 1e-40
SC14_YARLI (P45816) SEC14 cytosolic factor (Phosphatidylinositol... 165 2e-40
SC14_KLULA (P24859) SEC14 cytosolic factor (Phosphatidylinositol... 157 4e-38
SC14_YEAST (P24280) SEC14 cytosolic factor (Phosphatidylinositol... 157 5e-38
SC14_CANAL (P46250) SEC14 cytosolic factor (Phosphatidylinositol... 156 8e-38
SC14_ASHGO (Q75DK1) SEC14 cytosolic factor (Phosphatidylinositol... 154 4e-37
SC14_CANGA (P53989) SEC14 cytosolic factor (Phosphatidylinositol... 153 9e-37
YKJ1_YEAST (P33324) 36.1 kDa protein in BUD2-MIF2 intergenic region 142 2e-33
YN02_CAEEL (Q03606) Hypothetical protein T23G5.2 in chromosome III 104 4e-22
S143_RAT (Q9Z1J8) SEC14-like protein 3 (45 kDa secretory protein... 92 2e-18
S141_HUMAN (Q92503) SEC14-like protein 1 92 2e-18
S143_HUMAN (Q9UDX4) SEC14-like protein 3 (Tocopherol-associated ... 91 5e-18
S144_HUMAN (Q9UDX3) SEC14-like protein 4 (Tocopherol-associated ... 89 2e-17
S144_MOUSE (Q8R0F9) SEC14-like protein 4 89 2e-17
S142_BOVIN (P58875) SEC14-like protein 2 (Alpha-tocopherol assoc... 82 3e-15
S142_MOUSE (Q99J08) SEC14-like protein 2 (Alpha-tocopherol assoc... 78 4e-14
S142_RAT (Q99MS0) SEC14-like protein 2 (Alpha-tocopherol associa... 78 5e-14
S142_HUMAN (O76054) SEC14-like protein 2 (Alpha-tocopherol assoc... 77 6e-14
RALB_TODPA (P49193) Retinal-binding protein (RALBP) 71 5e-12
YGR1_SCHPO (Q9UUC2) Hypothetical protein C365.01 in chromosome II 60 1e-08
>SC14_SCHPO (Q10137) Sec14 cytosolic factor
(Phosphatidylinositol/phosphatidyl-choline transfer
protein) (PI/PC TP) (Sporulation-specific protein 20)
Length = 286
Score = 166 bits (419), Expect = 1e-40
Identities = 89/257 (34%), Positives = 146/257 (56%), Gaps = 5/257 (1%)
Query: 109 AVVDAFRKLLIMDELLPQKHDDYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIM 168
A +D+ R L + +L + D +LRFL+ARKF++ ++ M+ +WRKEFG D ++
Sbjct: 30 ATLDSMR--LELQKLGYTERLDDATLLRFLRARKFNLQQSLEMFIKCEKWRKEFGVDDLI 87
Query: 169 EDFEFNELNEVIKYNPHGYHGVDKEGRPVFIERFEKLDRNKLMQVTTIDRYVKYHAQRCE 228
++F ++E V KY P YH D +GRPV++E+ +D KL Q+TT +R ++ E
Sbjct: 88 KNFHYDEKEAVSKYYPQFYHKTDIDGRPVYVEQLGNIDLKKLYQITTPERMMQNLVYEYE 147
Query: 229 EMHAIKFPACTIASKRHIDSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQ 288
+ +FPAC+ + I++S TI+DL+G+G ++ +++ I D YP+ G+
Sbjct: 148 MLALKRFPACSRKAGGLIETSCTIMDLKGVGITSIHSV-YSYIRQASSISQDYYPERMGK 206
Query: 289 SFIINVGLELRSLRSICEYFMDPKVASKVHVIGDRYQRKLLKVIDASELPTFLGGTCTCA 348
++IN S ++ + F+D K+H++G Y+ LL+ I A LP LGG C C
Sbjct: 207 FYVINAPWGFSSAFNLIKGFLDEATVKKIHILGSNYKSALLEQIPADNLPAKLGGNCQC- 265
Query: 349 NQGGCLRSDKGPWNNPE 365
GGC SD GPW+ +
Sbjct: 266 -PGGCELSDAGPWHEEQ 281
>SC14_YARLI (P45816) SEC14 cytosolic factor
(Phosphatidylinositol/phosphatidylcholine transfer
protein) (PI/PC TP)
Length = 492
Score = 165 bits (417), Expect = 2e-40
Identities = 91/268 (33%), Positives = 150/268 (55%), Gaps = 4/268 (1%)
Query: 116 KLLIMDELLPQKHDDYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNE 175
K++++ + + DD +LRFL+ARKFD+ A+ MW + +WRKEFG +TI+EDF + E
Sbjct: 40 KMILLTKGYEDRTDDA-TLLRFLRARKFDVPLAQEMWENCEKWRKEFGTNTILEDFWYKE 98
Query: 176 LNEVIKYNPHGYHGVDKEGRPVFIERFEKLDRNKLMQVTTIDRYVKYHAQRCEEMHAIKF 235
EV K P YH DK+GRPV++E K++ +++ ++TT +R ++ E +
Sbjct: 99 KKEVAKLYPQYYHKTDKDGRPVYVENVGKVNIHEMYKITTQERMLRNLVWEYESFVRHRL 158
Query: 236 PACTIASKRHIDSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVG 295
PAC+ I++S TILDL+G+ + + +K I + YP+ G+ ++IN
Sbjct: 159 PACSRVVGHLIETSCTILDLKGVSLSSASQV-YGFLKDASNIGQNYYPERMGKFYLINAP 217
Query: 296 LELRSLRSICEYFMDPKVASKVHVIGDRYQRKLLKVIDASELPTFLGGTCTCANQGGCLR 355
++ S+ + F+DP SK+HV G Y+ KLL + A LP GG +++ G
Sbjct: 218 FGFSTVFSVIKRFLDPVTVSKIHVYGSNYKEKLLAQVPAYNLPIKFGG--QSSSKIGVEL 275
Query: 356 SDKGPWNNPEILKVKGSDTLTAESSSEA 383
SD GPW +P+ + +G + E + A
Sbjct: 276 SDDGPWRDPQFVGPEGLAPVAGERPTGA 303
>SC14_KLULA (P24859) SEC14 cytosolic factor
(Phosphatidylinositol/phosphatidylcholine transfer
protein) (PI/PC TP)
Length = 301
Score = 157 bits (398), Expect = 4e-38
Identities = 92/264 (34%), Positives = 143/264 (53%), Gaps = 4/264 (1%)
Query: 109 AVVDAFRKLLIMDELLPQKHDDYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIM 168
A + FR+LL + L ++ D +LRFL+ARKFD+ +K M+ + +WRKEFG DTI
Sbjct: 33 AKLKEFRELL--ESLGYKERLDDSTLLRFLRARKFDLEASKIMYENCEKWRKEFGVDTIF 90
Query: 169 EDFEFNELNEVIKYNPHGYHGVDKEGRPVFIERFEKLDRNKLMQVTTIDRYVKYHAQRCE 228
EDF + E V KY P YH D +GRPV+IE ++ ++ ++TT +R +K E
Sbjct: 91 EDFHYEEKPLVAKYYPQYYHKTDNDGRPVYIEELGSVNLTQMYKITTQERMLKNLVWEYE 150
Query: 229 EMHAIKFPACTIASKRHIDSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQ 288
+ PAC+ + +++S TILDL+GI + + ++ I + YP+ G+
Sbjct: 151 AFVRYRLPACSRKAGYLVETSCTILDLKGISISSAAQV-LSYVREASNIGQNYYPERMGK 209
Query: 289 SFIINVGLELRSLRSICEYFMDPKVASKVHVIGDRYQRKLLKVIDASELPTFLGGTCTCA 348
++IN + + + F+DP SK+ ++G YQ+ LLK I A LP GG +
Sbjct: 210 FYLINAPFGFSTAFRLFKPFLDPVTVSKIFILGSSYQKDLLKQIPAENLPKKFGGQSEVS 269
Query: 349 N-QGGCLRSDKGPWNNPEILKVKG 371
+GG SD GPW E + +G
Sbjct: 270 EAEGGLYLSDIGPWREEEYIGPEG 293
>SC14_YEAST (P24280) SEC14 cytosolic factor
(Phosphatidylinositol/phosphatidylcholine transfer
protein) (PI/PC TP)
Length = 303
Score = 157 bits (397), Expect = 5e-38
Identities = 83/243 (34%), Positives = 139/243 (57%), Gaps = 2/243 (0%)
Query: 130 DYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNELNEVIKYNPHGYHG 189
D +LRFL+ARKFD+ AK M+ + +WRK++G DTI++DF ++E + K+ P YH
Sbjct: 53 DDSTLLRFLRARKFDVQLAKEMFENCEKWRKDYGTDTILQDFHYDEKPLIAKFYPQYYHK 112
Query: 190 VDKEGRPVFIERFEKLDRNKLMQVTTIDRYVKYHAQRCEEMHAIKFPACTIASKRHIDSS 249
DK+GRPV+ E ++ +++ +VT+ +R +K E + + PAC+ A+ +++S
Sbjct: 113 TDKDGRPVYFEELGAVNLHEMNKVTSEERMLKNLVWEYESVVQYRLPACSRAAGHLVETS 172
Query: 250 ITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVGLELRSLRSICEYFM 309
TI+DL+GI + ++ I + YP+ G+ +IIN + + + F+
Sbjct: 173 CTIMDLKGISISSAYSV-MSYVREASYISQNYYPERMGKFYIINAPFGFSTAFRLFKPFL 231
Query: 310 DPKVASKVHVIGDRYQRKLLKVIDASELPTFLGGTCTC-ANQGGCLRSDKGPWNNPEILK 368
DP SK+ ++G YQ++LLK I A LP GG ++GG SD GPW +P+ +
Sbjct: 232 DPVTVSKIFILGSSYQKELLKQIPAENLPVKFGGKSEVDESKGGLYLSDIGPWRDPKYIG 291
Query: 369 VKG 371
+G
Sbjct: 292 PEG 294
>SC14_CANAL (P46250) SEC14 cytosolic factor
(Phosphatidylinositol/phosphatidylcholine transfer
protein) (PI/PC TP)
Length = 301
Score = 156 bits (395), Expect = 8e-38
Identities = 92/261 (35%), Positives = 143/261 (54%), Gaps = 4/261 (1%)
Query: 111 VDAFRKLLIMDELLPQKHDDYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMED 170
+D FR+ L EL + D +LRFL+ARKFDI KA M+ +WR++FG +TI++D
Sbjct: 37 LDIFRQQLT--ELGYKDRLDDASLLRFLRARKFDIQKAIDMFVACEKWREDFGVNTILKD 94
Query: 171 FEFNELNEVIKYNPHGYHGVDKEGRPVFIERFEKLDRNKLMQVTTIDRYVKYHAQRCEEM 230
F + E V K P YH DK+GRPV+ E K+D K++++TT +R +K E M
Sbjct: 95 FHYEEKPIVAKMYPTYYHKTDKDGRPVYFEELGKVDLVKMLKITTQERMLKNLVWEYEAM 154
Query: 231 HAIKFPACTIASKRHIDSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSF 290
+ PAC+ + +++S T+LDL GI + + + KI D YP+ G+ +
Sbjct: 155 CQYRLPACSRKAGYLVETSCTVLDLSGISVTSAYNVIGYV-REASKIGQDYYPERMGKFY 213
Query: 291 IINVGLELRSLRSICEYFMDPKVASKVHVIGDRYQRKLLKVIDASELPTFLGGTCTCANQ 350
+IN + + + F+DP SK+H++G Y+++LLK I LP GG ++
Sbjct: 214 LINAPFGFSTAFKLFKPFLDPVTVSKIHILGYSYKKELLKQIPPQNLPVKFGGMSDVSDD 273
Query: 351 GGCLRSDKGPWNNPEILKVKG 371
L D GPW +PE + +G
Sbjct: 274 -DLLLKDVGPWRDPEFIGPEG 293
>SC14_ASHGO (Q75DK1) SEC14 cytosolic factor
(Phosphatidylinositol/phosphatidylcholine transfer
protein) (PI/PC TP)
Length = 311
Score = 154 bits (389), Expect = 4e-37
Identities = 86/246 (34%), Positives = 134/246 (53%), Gaps = 2/246 (0%)
Query: 127 KHDDYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNELNEVIKYNPHG 186
K D +LRFL+ARKFD+ A+ M+ + +WRKE G DTI EDF + E V K+ P
Sbjct: 52 KRLDDSTLLRFLRARKFDVAAARAMFENCEKWRKENGVDTIFEDFHYEEKPLVAKFYPQY 111
Query: 187 YHGVDKEGRPVFIERFEKLDRNKLMQVTTIDRYVKYHAQRCEEMHAIKFPACTIASKRHI 246
YH DK+GRPV+IE ++ ++ ++TT +R +K E + PA + + +
Sbjct: 112 YHKTDKDGRPVYIEELGAVNLTEMYKITTQERMLKNLIWEYESFSRYRLPASSRQADCLV 171
Query: 247 DSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVGLELRSLRSICE 306
++S TILDL+GI + ++ I + YP+ G+ ++IN + + +
Sbjct: 172 ETSCTILDLKGISISAAAQV-LSYVREASNIGQNYYPERMGKFYMINAPFGFSAAFRLFK 230
Query: 307 YFMDPKVASKVHVIGDRYQRKLLKVIDASELPTFLGGTCTCAN-QGGCLRSDKGPWNNPE 365
F+DP SK+ ++G YQ++LLK I A LP GG + +GG SD GPW NP+
Sbjct: 231 PFLDPVTVSKIFILGSSYQKELLKQIPAENLPVKFGGQSDVSEAEGGLYLSDIGPWRNPK 290
Query: 366 ILKVKG 371
+ +G
Sbjct: 291 YIGPEG 296
>SC14_CANGA (P53989) SEC14 cytosolic factor
(Phosphatidylinositol/phosphatidylcholine transfer
protein) (PI/PC TP)
Length = 302
Score = 153 bits (386), Expect = 9e-37
Identities = 83/243 (34%), Positives = 135/243 (55%), Gaps = 2/243 (0%)
Query: 130 DYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNELNEVIKYNPHGYHG 189
D +LRFL+ARKFD+ AK M+ + +WRKE+G +TIM+DF ++E V KY P YH
Sbjct: 52 DDSTLLRFLRARKFDVALAKEMFENCEKWRKEYGTNTIMQDFHYDEKPLVAKYYPQYYHK 111
Query: 190 VDKEGRPVFIERFEKLDRNKLMQVTTIDRYVKYHAQRCEEMHAIKFPACTIASKRHIDSS 249
DK+GRPV+ E ++ ++ ++TT +R +K E + + PAC+ A+ +++S
Sbjct: 112 TDKDGRPVYFEELGAVNLTEMEKITTQERMLKNLVWEYESVVNYRLPACSRAAGYLVETS 171
Query: 250 ITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVGLELRSLRSICEYFM 309
T++DL+GI + ++ I + YP+ G+ ++IN + + + F+
Sbjct: 172 CTVMDLKGISISSAYSV-LSYVREASYISQNYYPERMGKFYLINAPFGFSTAFRLFKPFL 230
Query: 310 DPKVASKVHVIGDRYQRKLLKVIDASELPTFLGGTCTCAN-QGGCLRSDKGPWNNPEILK 368
DP SK+ ++G YQ +LLK I A LP+ GG GG SD GPW + + +
Sbjct: 231 DPVTVSKIFILGSSYQSELLKQIPAENLPSKFGGKSEVDEAAGGLYLSDIGPWRDAKYIG 290
Query: 369 VKG 371
+G
Sbjct: 291 PEG 293
>YKJ1_YEAST (P33324) 36.1 kDa protein in BUD2-MIF2 intergenic region
Length = 310
Score = 142 bits (358), Expect = 2e-33
Identities = 86/265 (32%), Positives = 143/265 (53%), Gaps = 10/265 (3%)
Query: 114 FRKLLIMDELLPQKHDDYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEF 173
FR +L+ ++ ++ DD +LRFL+ARKFDI + M+ + WR+E+GA+TI+ED+E
Sbjct: 36 FRSILL-EKNYKERLDD-STLLRFLRARKFDINASVEMFVETERWREEYGANTIIEDYEN 93
Query: 174 NELNE------VIKYNPHGYHGVDKEGRPVFIERFEKLDRNKLMQVTTIDRYVKYHAQRC 227
N+ E + K P YH VDK+GRP++ E ++ K+ ++TT + ++ +
Sbjct: 94 NKEAEDKERIKLAKMYPQYYHHVDKDGRPLYFEELGGINLKKMYKITTEKQMLRNLVKEY 153
Query: 228 EEMHAIKFPACTIASKRHIDSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGG 287
E + PAC+ + I++S T+LDL+GI N +K I + YP+ G
Sbjct: 154 ELFATYRVPACSRRAGYLIETSCTVLDLKGISLSNAYHV-LSYIKDVADISQNYYPERMG 212
Query: 288 QSFIINVGLELRSLRSICEYFMDPKVASKVHVIGDRYQRKLLKVIDASELPTFLGGTCTC 347
+ +II+ ++ + + F+DP SK+ ++G Y+++LLK I LP GGT
Sbjct: 213 KFYIIHSPFGFSTMFKMVKPFLDPVTVSKIFILGSSYKKELLKQIPIENLPVKYGGTSVL 272
Query: 348 ANQGG-CLRSDKGPWNNPEILKVKG 371
N SD GPW +P + +G
Sbjct: 273 HNPNDKFYYSDIGPWRDPRYIGPEG 297
>YN02_CAEEL (Q03606) Hypothetical protein T23G5.2 in chromosome III
Length = 743
Score = 104 bits (260), Expect = 4e-22
Identities = 70/226 (30%), Positives = 112/226 (48%), Gaps = 7/226 (3%)
Query: 129 DDYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNELNEVIKYNPHGYH 188
+D H+ LRFL+AR FD+ KAK M + WRK+ D I+E E+ + +Y P +H
Sbjct: 300 NDAHL-LRFLRARDFDVAKAKDMVHASIIWRKQHNVDKILE--EWTRPTVIKQYFPGCWH 356
Query: 189 GVDKEGRPVFIERFEKLDRNKLMQVTTIDRYVKYHAQRCEEMHAIKFPACTIASKRHIDS 248
DK GRP++I RF +LD +++ ++ VK CE+ + T I S
Sbjct: 357 NSDKAGRPMYILRFGQLDTKGMLRSCGVENLVKLTLSICED-GLQRAAEATRKLGTPISS 415
Query: 249 SITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVGLELRSLRSICEYF 308
++DL G+ +L + + + ++I+ NYP+T GQ ++ L ++ F
Sbjct: 416 WSLVVDLDGLSMRHLWRPGVQCLLKIIEIVEANYPETMGQVLVVRAPRVFPVLWTLISPF 475
Query: 309 MDPKVASKVHVIGDR---YQRKLLKVIDASELPTFLGGTCTCANQG 351
+D K K V G + +L K I+ +P FLGG+C N G
Sbjct: 476 IDEKTRKKFMVSGGSGGDLKEELRKHIEEKFIPDFLGGSCLTTNCG 521
>S143_RAT (Q9Z1J8) SEC14-like protein 3 (45 kDa secretory protein)
(rsec45)
Length = 400
Score = 92.4 bits (228), Expect = 2e-18
Identities = 65/231 (28%), Positives = 115/231 (49%), Gaps = 10/231 (4%)
Query: 119 IMDELLPQKHDDYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNELNE 178
+ D L + D + +LR+L+AR FD+ K++ M +E+RK D I+ D++ E+
Sbjct: 23 VQDVLPALPNPDDYFLLRWLRARNFDLQKSEAMLRKYMEFRKTMDIDHIL-DWQPPEV-- 79
Query: 179 VIKYNPHGYHGVDKEGRPVFIERFEKLDRNKLMQVTTIDRYVKYHAQRCEEMHAIKFPAC 238
+ KY P G G D++G PV+ + LD L+ T +K + CE + C
Sbjct: 80 IQKYMPGGLCGYDRDGCPVWYDIIGPLDPKGLLFSVTKQDLLKTKMRDCERI----LHEC 135
Query: 239 TIASK---RHIDSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVG 295
+ ++ R I++ + I D +G+G + + E+ + F +L +NYP+T I+
Sbjct: 136 DLQTERLGRKIETIVMIFDCEGLGLKHFWKPLVEVYQEFFGLLEENYPETLKFMLIVKAT 195
Query: 296 LELRSLRSICEYFMDPKVASKVHVIGDRYQRKLLKVIDASELPTFLGGTCT 346
++ + F+ K+ V+G+ ++ LLK+I ELP GGT T
Sbjct: 196 KLFPVGYNLMKPFLSEDTRRKIVVLGNSWKEGLLKLISPEELPAHFGGTLT 246
>S141_HUMAN (Q92503) SEC14-like protein 1
Length = 715
Score = 92.0 bits (227), Expect = 2e-18
Identities = 65/224 (29%), Positives = 109/224 (48%), Gaps = 20/224 (8%)
Query: 134 MLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNELNEVIKYNPHGYHGVDKE 193
+LRFL+AR F+I KA+ + L WRK+ D I+E + ++ + Y G+H DK+
Sbjct: 280 ILRFLRARDFNIDKAREIMCQSLTWRKQHQVDYILETWTPPQVLQ--DYYAGGWHHHDKD 337
Query: 194 GRPVFIERFEKLDRNKLMQVTTIDRYVKY-------HAQRCEEMHAIKFPACTIASKRHI 246
GRP+++ R ++D L++ + ++Y +RCEE T R I
Sbjct: 338 GRPLYVLRLGQMDTKGLVRALGEEALLRYVLSVNEERLRRCEEN--------TKVFGRPI 389
Query: 247 DSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVGLELRSLRSICE 306
S ++DL+G+ +L + + R ++++ NYP+T G+ I+ L ++
Sbjct: 390 SSWTCLVDLEGLNMRHLWRPGVKALLRIIEVVEANYPETLGRLLILRAPRVFPVLWTLVS 449
Query: 307 YFMDPKVASKVHV-IGDRYQRK--LLKVIDASELPTFLGGTCTC 347
F+D K + G+ YQ LL ID +P FL G C C
Sbjct: 450 PFIDDNTRRKFLIYAGNDYQGPGGLLDYIDKEIIPDFLSGECMC 493
>S143_HUMAN (Q9UDX4) SEC14-like protein 3 (Tocopherol-associated
protein 2)
Length = 400
Score = 90.9 bits (224), Expect = 5e-18
Identities = 64/231 (27%), Positives = 115/231 (49%), Gaps = 10/231 (4%)
Query: 119 IMDELLPQKHDDYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNELNE 178
+ D L + D + +LR+L+AR FD+ K++ + +E+RK D I+ D++ E+
Sbjct: 23 VQDVLPALPNPDDYFLLRWLRARNFDLQKSEALLRKYMEFRKTMDIDHIL-DWQPPEV-- 79
Query: 179 VIKYNPHGYHGVDKEGRPVFIERFEKLDRNKLMQVTTIDRYVKYHAQRCEEMHAIKFPAC 238
+ KY P G G D++G PV+ + LD L+ T +K + CE + C
Sbjct: 80 IQKYMPGGLCGYDRDGCPVWYDIIGPLDPKGLLFSVTKQDLLKTKMRDCERI----LHEC 135
Query: 239 TIASKR---HIDSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVG 295
+ ++R I++ + I D +G+G + + E+ + F +L +NYP+T I+
Sbjct: 136 DLQTERLGKKIETIVMIFDCEGLGLKHFWKPLVEVYQEFFGLLEENYPETLKFMLIVKAT 195
Query: 296 LELRSLRSICEYFMDPKVASKVHVIGDRYQRKLLKVIDASELPTFLGGTCT 346
++ + F+ K+ V+G+ ++ LLK+I ELP GGT T
Sbjct: 196 KLFPVGYNLMKPFLSEDTRRKIIVLGNNWKEGLLKLISPEELPAQFGGTLT 246
>S144_HUMAN (Q9UDX3) SEC14-like protein 4 (Tocopherol-associated
protein 3)
Length = 406
Score = 89.4 bits (220), Expect = 2e-17
Identities = 65/233 (27%), Positives = 116/233 (48%), Gaps = 15/233 (6%)
Query: 120 MDELLP--QKHDDYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNELN 177
+ +LLP DDY +LR+L+AR FD+ K++ M +E+RK+ D I+ +
Sbjct: 23 LQDLLPILPNADDY-FLLRWLRARNFDLQKSEDMLRRHMEFRKQQDLDNIVTW----QPP 77
Query: 178 EVIK-YNPHGYHGVDKEGRPVFIERFEKLDRNKLMQVTTIDRYVKYHAQRCEEMHAIKFP 236
EVI+ Y+ G G D EG PV+ LD L+ + ++ + CE +
Sbjct: 78 EVIQLYDSGGLCGYDYEGCPVYFNIIGSLDPKGLLLSASKQDMIRKRIKVCE----LLLH 133
Query: 237 ACTIASK---RHIDSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIIN 293
C + ++ R I+ ++ + D++G+ +L + E+ ++F IL NYP+T +I
Sbjct: 134 ECELQTQKLGRKIEMALMVFDMEGLSLKHLWKPAVEVYQQFFSILEANYPETLKNLIVIR 193
Query: 294 VGLELRSLRSICEYFMDPKVASKVHVIGDRYQRKLLKVIDASELPTFLGGTCT 346
++ + FM + K+ ++GD ++++L K I +LP GGT T
Sbjct: 194 APKLFPVAFNLVKSFMSEETRRKIVILGDNWKQELTKFISPDQLPVEFGGTMT 246
>S144_MOUSE (Q8R0F9) SEC14-like protein 4
Length = 403
Score = 89.0 bits (219), Expect = 2e-17
Identities = 64/233 (27%), Positives = 115/233 (48%), Gaps = 15/233 (6%)
Query: 120 MDELLPQ--KHDDYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNELN 177
+ +LLP K DDY +LR+L+AR FD+ K++ M +E+R + D I+ +
Sbjct: 23 LQDLLPTLPKADDY-FLLRWLRARNFDLKKSEDMLRKHVEFRNQQNLDQILTW----QAP 77
Query: 178 EVIK-YNPHGYHGVDKEGRPVFIERFEKLDRNKLMQVTTIDRYVKYHAQRCEEMHAIKFP 236
EVI+ Y+ G G D EG PV+ + +D L + ++ + CE +
Sbjct: 78 EVIQLYDSGGLSGYDYEGCPVWFDIIGTMDPKGLFMSASKQDMIRKRIKVCEML----LH 133
Query: 237 ACTIASK---RHIDSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIIN 293
C + S+ R I+ + + D++G+ +L + E+ ++F IL NYP+T II
Sbjct: 134 ECELQSQKLGRKIERMVMVFDMEGLSLRHLWKPAVEVYQQFFAILEANYPETVKNLIIIR 193
Query: 294 VGLELRSLRSICEYFMDPKVASKVHVIGDRYQRKLLKVIDASELPTFLGGTCT 346
++ + FM + K+ ++G ++++L+K + +LP GGT T
Sbjct: 194 APKLFPVAFNLVKSFMGEETQKKIVILGGNWKQELVKFVSPDQLPVEFGGTMT 246
>S142_BOVIN (P58875) SEC14-like protein 2 (Alpha-tocopherol
associated protein) (TAP) (bTAP) (Fragment)
Length = 387
Score = 82.0 bits (201), Expect = 3e-15
Identities = 61/231 (26%), Positives = 107/231 (45%), Gaps = 10/231 (4%)
Query: 119 IMDELLPQKHDDYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNELNE 178
+ D L + D + +LR+L+AR F++ K++ M +E+RK+ D IM +
Sbjct: 23 VQDVLPALPNPDDYFLLRWLRARNFNLQKSEAMLRKHVEFRKQKDIDNIMS---WQPPEV 79
Query: 179 VIKYNPHGYHGVDKEGRPVFIERFEKLDRNKLMQVTTIDRYVKYHAQRCEEMHAIKFPAC 238
V +Y G G D EG P++ + LD L+ + K + CE + C
Sbjct: 80 VQQYLSGGMCGYDLEGSPIWYDIIGPLDAKGLLLSASKQDLFKTKMRDCE----LLLQEC 135
Query: 239 TIASKR---HIDSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVG 295
+++ I+++ I D +G+G +L + E FL + +NYP+T + FI+
Sbjct: 136 VRQTEKMGKKIEATTLIYDCEGLGLKHLWKPAVEAYGEFLCMFEENYPETLKRLFIVKAP 195
Query: 296 LELRSLRSICEYFMDPKVASKVHVIGDRYQRKLLKVIDASELPTFLGGTCT 346
++ + F+ K+ V+G ++ LLK I +LP GGT T
Sbjct: 196 KLFPVAYNLVKPFLSEDTRKKIQVLGANWKEVLLKYISPDQLPVEYGGTMT 246
>S142_MOUSE (Q99J08) SEC14-like protein 2 (Alpha-tocopherol
associated protein) (TAP)
Length = 403
Score = 78.2 bits (191), Expect = 4e-14
Identities = 59/228 (25%), Positives = 106/228 (45%), Gaps = 4/228 (1%)
Query: 119 IMDELLPQKHDDYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNELNE 178
+ D L + D + +LR+L+AR FD+ K++ M +E+RK+ D I+ +
Sbjct: 23 VQDVLPTLPNPDDYFLLRWLRARSFDLQKSEAMLRKHVEFRKQKDIDKIIS---WQPPEV 79
Query: 179 VIKYNPHGYHGVDKEGRPVFIERFEKLDRNKLMQVTTIDRYVKYHAQRCEEMHAIKFPAC 238
+ +Y G G D +G PV+ + LD L+ + ++ + CE +
Sbjct: 80 IQQYLSGGRCGYDLDGCPVWYDIIGPLDAKGLLFSASKQDLLRTKMRDCELLLQECIQQT 139
Query: 239 TIASKRHIDSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVGLEL 298
T K+ I++ I D +G+G +L + E FL + +NYP+T + F++
Sbjct: 140 TKLGKK-IETITMIYDCEGLGLKHLWKPAVEAYGEFLTMFEENYPETLKRLFVVKAPKLF 198
Query: 299 RSLRSICEYFMDPKVASKVHVIGDRYQRKLLKVIDASELPTFLGGTCT 346
++ + F+ K+ V+G ++ LLK I +LP GGT T
Sbjct: 199 PVAYNLIKPFLSEDTRRKIMVLGANWKEVLLKHISPDQLPVEYGGTMT 246
>S142_RAT (Q99MS0) SEC14-like protein 2 (Alpha-tocopherol associated
protein) (TAP) (Supernatant protein factor) (SPF)
(Squalene transfer protein)
Length = 403
Score = 77.8 bits (190), Expect = 5e-14
Identities = 59/231 (25%), Positives = 107/231 (45%), Gaps = 10/231 (4%)
Query: 119 IMDELLPQKHDDYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNELNE 178
+ D L + D + +LR+L+AR FD+ K++ M +E+RK+ D I+ +
Sbjct: 23 VQDVLPALPNPDDYFLLRWLRARSFDLQKSEAMLRKHVEFRKQKDIDKIIS---WQPPEV 79
Query: 179 VIKYNPHGYHGVDKEGRPVFIERFEKLDRNKLMQVTTIDRYVKYHAQRCEEMHAIKFPAC 238
+ +Y G G D +G PV+ + LD L+ + ++ + CE + C
Sbjct: 80 IQQYLSGGRCGYDLDGCPVWYDIIGPLDAKGLLFSASKQDLLRTKMRDCE----LLLQEC 135
Query: 239 TIASKR---HIDSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVG 295
T + + I++ I D +G+G +L + E FL + +NYP+T + F++
Sbjct: 136 TQQTAKLGKKIETITMIYDCEGLGLKHLWKPAVEAYGEFLTMFEENYPETLKRLFVVKAP 195
Query: 296 LELRSLRSICEYFMDPKVASKVHVIGDRYQRKLLKVIDASELPTFLGGTCT 346
++ + F+ K+ V+G ++ LLK I +LP GGT T
Sbjct: 196 KLFPVAYNLIKPFLSEDTRKKIMVLGANWKEVLLKHISPDQLPVEYGGTMT 246
>S142_HUMAN (O76054) SEC14-like protein 2 (Alpha-tocopherol
associated protein) (TAP) (hTAP) (Supernatant protein
factor) (SPF) (Squalene transfer protein)
Length = 403
Score = 77.4 bits (189), Expect = 6e-14
Identities = 58/231 (25%), Positives = 106/231 (45%), Gaps = 10/231 (4%)
Query: 119 IMDELLPQKHDDYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNELNE 178
+ D L + D + +LR+L+AR FD+ K++ M +E+RK+ D I+ +
Sbjct: 23 VQDVLPALPNPDDYFLLRWLRARSFDLQKSEAMLRKHVEFRKQKDIDNIIS---WQPPEV 79
Query: 179 VIKYNPHGYHGVDKEGRPVFIERFEKLDRNKLMQVTTIDRYVKYHAQRCEEMHAIKFPAC 238
+ +Y G G D +G PV+ + LD L+ + ++ + CE + C
Sbjct: 80 IQQYLSGGMCGYDLDGCPVWYDIIGPLDAKGLLFSASKQDLLRTKMRECE----LLLQEC 135
Query: 239 ---TIASKRHIDSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVG 295
T R +++ I D +G+G +L + E FL + +NYP+T + F++
Sbjct: 136 AHQTTKLGRKVETITIIYDCEGLGLKHLWKPAVEAYGEFLCMFEENYPETLKRLFVVKAP 195
Query: 296 LELRSLRSICEYFMDPKVASKVHVIGDRYQRKLLKVIDASELPTFLGGTCT 346
++ + F+ K+ V+G ++ LLK I ++P GGT T
Sbjct: 196 KLFPVAYNLIKPFLSEDTRKKIMVLGANWKEVLLKHISPDQVPVEYGGTMT 246
>RALB_TODPA (P49193) Retinal-binding protein (RALBP)
Length = 342
Score = 71.2 bits (173), Expect = 5e-12
Identities = 49/186 (26%), Positives = 90/186 (48%), Gaps = 3/186 (1%)
Query: 159 RKEFGADTIMEDFEFNELNEVIKYNPHGYHGVDKEGRPVFIERFEKLDRNKLMQVTTIDR 218
R++ GADT++ E+ + + K+ G G DK+G + IE + LD +M
Sbjct: 3 REQMGADTLIA--EYTPPDVIQKFMTGGDVGHDKDGSVLRIEPWGYLDMKGIMYSCKKSD 60
Query: 219 YVKYHAQRCEEMHAIKFPACTIASKRHIDSSITILDLQGIGFCNLEEADREIMKRFLKIL 278
K +CE+ H A + + + D++ +G ++ + ++ +++L
Sbjct: 61 LEKSKLLQCEK-HLKDLEAQSEKVGKPCTGLTVVFDMENVGSKHMWKPGLDMYLYLVQVL 119
Query: 279 IDNYPQTGGQSFIINVGLELRSLRSICEYFMDPKVASKVHVIGDRYQRKLLKVIDASELP 338
DNYP+ + F+IN L + + + + +K+ V+G Y+ LL+ IDA ELP
Sbjct: 120 EDNYPEMMKRLFVINAPTLFPVLYKLVKPLLSEDMKNKIFVLGGDYKDTLLEYIDAEELP 179
Query: 339 TFLGGT 344
+LGGT
Sbjct: 180 AYLGGT 185
>YGR1_SCHPO (Q9UUC2) Hypothetical protein C365.01 in chromosome II
Length = 355
Score = 60.1 bits (144), Expect = 1e-08
Identities = 55/198 (27%), Positives = 94/198 (46%), Gaps = 24/198 (12%)
Query: 129 DDYHM-MLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNELNE-VIKYNPHG 186
DD+ + +LRFLKARKF + + M A+ + WR++ +IM E N LN+ +K + +
Sbjct: 49 DDFDLTLLRFLKARKFVVTDSSDMLANAIVWRQQANLRSIMVRGE-NGLNQNFVKASMYF 107
Query: 187 YHGVDKEGRPVFIERFEKLDRNKLMQVTTIDRYVKYHAQRCEEMHAIKFPACTIASKRHI 246
G DK+GR + K + + EE+ A+ A A + +
Sbjct: 108 IWGQDKKGRAIVFLNLHNFIPPK-------------NTKDMEELKALILYAMENA-RLFL 153
Query: 247 DSSIT----ILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVGLEL---R 299
DS +L L + + + + D + + F + + YP+ GQ+ I+ G +
Sbjct: 154 DSEQNAAKGVLGLVDLTYFSRKNIDLDFARVFAETFQNYYPEILGQALIVGSGFRMALFE 213
Query: 300 SLRSICEYFMDPKVASKV 317
+ SI +YF+DP+V SKV
Sbjct: 214 GVWSIGKYFLDPEVRSKV 231
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.324 0.140 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 47,828,339
Number of Sequences: 164201
Number of extensions: 2008479
Number of successful extensions: 6395
Number of sequences better than 10.0: 50
Number of HSP's better than 10.0 without gapping: 29
Number of HSP's successfully gapped in prelim test: 21
Number of HSP's that attempted gapping in prelim test: 6314
Number of HSP's gapped (non-prelim): 55
length of query: 418
length of database: 59,974,054
effective HSP length: 113
effective length of query: 305
effective length of database: 41,419,341
effective search space: 12632899005
effective search space used: 12632899005
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 67 (30.4 bits)
Medicago: description of AC121244.7


