

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
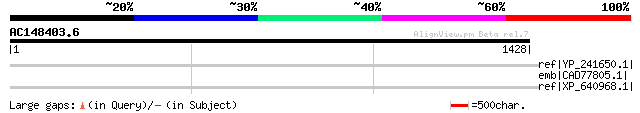
Query= AC148403.6 - phase: 0 /pseudo
(1428 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
ref|YP_241650.1| hypothetical protein XC_0549 [Xanthomonas campe... 39 1.7
emb|CAD77805.1| hypothetical protein-signal peptide and transmem... 38 2.2
ref|XP_640968.1| hypothetical protein DDB0215238 [Dictyostelium ... 37 4.8
>ref|YP_241650.1| hypothetical protein XC_0549 [Xanthomonas campestris pv. campestris
str. 8004] gi|66572220|gb|AAY47630.1| conserved
hypothetical protein [Xanthomonas campestris pv.
campestris str. 8004]
Length = 103
Score = 38.5 bits (88), Expect = 1.7
Identities = 28/80 (35%), Positives = 51/80 (63%)
Query: 292 LRQRTRIV*SEVLAILIILL*LLLLILVLHIVLLLSIVFLLWVLIFLI*MERWLSKLQLR 351
+ RT ++ +L +L++LL LLLL+L+L ++LLL ++ LL +L+ L+ + L L L
Sbjct: 1 MAMRTAMLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLL 60
Query: 352 VR*LLLSYV*NVLCLCLVVI 371
+ LLL + +L L L+++
Sbjct: 61 LLLLLLLLLLLLLLLLLLLL 80
Score = 38.1 bits (87), Expect = 2.2
Identities = 32/90 (35%), Positives = 56/90 (61%)
Query: 282 LLVESLL*LVLRQRTRIV*SEVLAILIILL*LLLLILVLHIVLLLSIVFLLWVLIFLI*M 341
LL+ LL L+L ++ +L +L++LL LLLL+L+L ++LLL ++ LL +L+ L+ +
Sbjct: 13 LLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLL 72
Query: 342 ERWLSKLQLRVR*LLLSYV*NVLCLCLVVI 371
L L L + LLL + +L L L+++
Sbjct: 73 LLLLLLLLLLLLLLLLLLLLLLLLLLLLLL 102
Score = 37.7 bits (86), Expect = 2.8
Identities = 33/97 (34%), Positives = 59/97 (60%), Gaps = 4/97 (4%)
Query: 275 LSLRGRRLLVESLL*LVLRQRTRIV*SEVLAILIILL*LLLLILVLHIVLLLSIVFLLWV 334
+++R LL+ LL L+L ++ L +L++LL LLLL+L+L ++LLL ++ LL +
Sbjct: 1 MAMRTAMLLLLLLLLLLLLLLLLLL----LLLLLLLLLLLLLLLLLLLLLLLLLLLLLLL 56
Query: 335 LIFLI*MERWLSKLQLRVR*LLLSYV*NVLCLCLVVI 371
L+ L+ + L L L + LLL + +L L L+++
Sbjct: 57 LLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLL 93
Score = 36.2 bits (82), Expect = 8.3
Identities = 33/98 (33%), Positives = 57/98 (57%)
Query: 254 ASVRTLCVLTAMERVTLVHSVLSLRGRRLLVESLL*LVLRQRTRIV*SEVLAILIILL*L 313
A + L +L + + L+ +L L LL+ LL L+L ++ +L +L++LL L
Sbjct: 6 AMLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLL 65
Query: 314 LLLILVLHIVLLLSIVFLLWVLIFLI*MERWLSKLQLR 351
LLL+L+L ++LLL ++ LL +L+ L+ + L L LR
Sbjct: 66 LLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLLR 103
>emb|CAD77805.1| hypothetical protein-signal peptide and transmembrane prediction
[Rhodopirellula baltica SH 1]
gi|32477734|ref|NP_870728.1| hypothetical protein-signal
peptide and transmembrane prediction [Rhodopirellula
baltica SH 1]
Length = 122
Score = 38.1 bits (87), Expect = 2.2
Identities = 18/45 (40%), Positives = 34/45 (75%)
Query: 303 VLAILIILL*LLLLILVLHIVLLLSIVFLLWVLIFLI*MERWLSK 347
VL ++++L+ +L+L+LVL +VL+L +V +L +++ L+ RW SK
Sbjct: 7 VLVLVLVLVLVLVLVLVLVLVLVLVLVLVLVLVLVLVISVRWASK 51
>ref|XP_640968.1| hypothetical protein DDB0215238 [Dictyostelium discoideum]
gi|60468990|gb|EAL66989.1| hypothetical protein
DDB0215238 [Dictyostelium discoideum]
Length = 230
Score = 37.0 bits (84), Expect = 4.8
Identities = 18/38 (47%), Positives = 31/38 (81%)
Query: 302 EVLAILIILL*LLLLILVLHIVLLLSIVFLLWVLIFLI 339
E+L +L++LL LLLL+L+L ++LLL ++FL +L+ L+
Sbjct: 170 ELLLLLLLLLLLLLLLLLLLLLLLLLLLFLFLLLLLLL 207
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.368 0.165 0.613
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,766,312,550
Number of Sequences: 2540612
Number of extensions: 57893925
Number of successful extensions: 256397
Number of sequences better than 10.0: 3
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 256255
Number of HSP's gapped (non-prelim): 28
length of query: 1428
length of database: 863,360,394
effective HSP length: 141
effective length of query: 1287
effective length of database: 505,134,102
effective search space: 650107589274
effective search space used: 650107589274
T: 11
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.7 bits)
S2: 82 (36.2 bits)
Medicago: description of AC148403.6


