

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147875.9 + phase: 0
(122 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
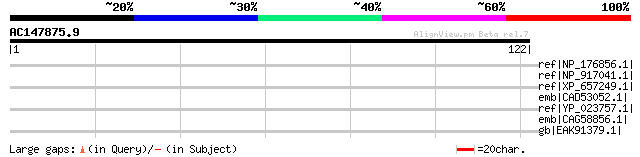
Score E
Sequences producing significant alignments: (bits) Value
ref|NP_176856.1| expressed protein [Arabidopsis thaliana] gi|125... 37 0.14
ref|NP_917041.1| myosin heavy chain-like protein [Oryza sativa (... 34 0.93
ref|XP_657249.1| protein kinase, putative [Entamoeba histolytica... 33 1.6
emb|CAD53052.1| hypothetical protein [Trypanosoma brucei] 32 4.6
ref|YP_023757.1| iron-dependent repressor [Picrophilus torridus ... 32 4.6
emb|CAG58856.1| unnamed protein product [Candida glabrata CBS138... 31 7.9
gb|EAK91379.1| potential ER biogenesis protein fragment [Candida... 31 7.9
>ref|NP_176856.1| expressed protein [Arabidopsis thaliana] gi|12597773|gb|AAG60086.1|
hypothetical protein [Arabidopsis thaliana]
Length = 607
Score = 36.6 bits (83), Expect = 0.14
Identities = 23/63 (36%), Positives = 36/63 (56%), Gaps = 9/63 (14%)
Query: 1 MKNLSSMIEESSYKAKAKMKDIETLEKKGKGQHGVMVVRRNENYEYAQVMRELEYLKKEL 60
+K L+ IEES+ + K++ DIE + + R + N Y ++MRELE +K+EL
Sbjct: 71 VKELTLRIEESNRRLKSRRIDIEAVMNES---------RIDGNGGYVRIMRELEDMKQEL 121
Query: 61 FKL 63
KL
Sbjct: 122 SKL 124
>ref|NP_917041.1| myosin heavy chain-like protein [Oryza sativa (japonica
cultivar-group)]
Length = 1051
Score = 33.9 bits (76), Expect = 0.93
Identities = 17/62 (27%), Positives = 33/62 (52%)
Query: 2 KNLSSMIEESSYKAKAKMKDIETLEKKGKGQHGVMVVRRNENYEYAQVMRELEYLKKELF 61
K L IE+++ KA ++ +++ + + G + YA+V++EL+ KKEL
Sbjct: 506 KELERQIEQTTAKATSQRSELQAMWAARTRRKGTDAPGAERDARYAEVVQELDQAKKELL 565
Query: 62 KL 63
+L
Sbjct: 566 RL 567
>ref|XP_657249.1| protein kinase, putative [Entamoeba histolytica HM-1:IMSS]
gi|56474496|gb|EAL51863.1| protein kinase, putative
[Entamoeba histolytica HM-1:IMSS]
Length = 420
Score = 33.1 bits (74), Expect = 1.6
Identities = 17/48 (35%), Positives = 25/48 (51%)
Query: 6 SMIEESSYKAKAKMKDIETLEKKGKGQHGVMVVRRNENYEYAQVMREL 53
S ++SS K K++D ETL+ GKG +G V E+ RE+
Sbjct: 102 SKSDQSSNALKPKLEDFETLKLIGKGTYGKKAVVETNEVEHTMAEREV 149
>emb|CAD53052.1| hypothetical protein [Trypanosoma brucei]
Length = 761
Score = 31.6 bits (70), Expect = 4.6
Identities = 17/62 (27%), Positives = 29/62 (46%)
Query: 1 MKNLSSMIEESSYKAKAKMKDIETLEKKGKGQHGVMVVRRNENYEYAQVMRELEYLKKEL 60
+++L ++ SY KA + KK + + R NE +EY Q + L L++E
Sbjct: 543 LRHLQRQHQQDSYDLKALRLQVGEERKKRQDAEARLAERENETHEYQQRLERLTVLQEEC 602
Query: 61 FK 62
K
Sbjct: 603 EK 604
>ref|YP_023757.1| iron-dependent repressor [Picrophilus torridus DSM 9790]
gi|48430699|gb|AAT43564.1| iron-dependent repressor
[Picrophilus torridus DSM 9790]
Length = 131
Score = 31.6 bits (70), Expect = 4.6
Identities = 16/49 (32%), Positives = 30/49 (60%), Gaps = 2/49 (4%)
Query: 4 LSSMIEESSYKAKAKMKDIETLEKKG--KGQHGVMVVRRNENYEYAQVM 50
LS + + K ++ +E LE+KG K +HG++++ N N EY+++M
Sbjct: 25 LSDLAVGLNIKPPTAIELLERLERKGLIKKEHGMIILTDNGNNEYSKIM 73
>emb|CAG58856.1| unnamed protein product [Candida glabrata CBS138]
gi|50287015|ref|XP_445937.1| unnamed protein product
[Candida glabrata]
Length = 358
Score = 30.8 bits (68), Expect = 7.9
Identities = 14/47 (29%), Positives = 26/47 (54%)
Query: 8 IEESSYKAKAKMKDIETLEKKGKGQHGVMVVRRNENYEYAQVMRELE 54
+E+S+Y K +KD E K GKG+ G + R++ + ++ +E
Sbjct: 86 LEKSTYNNKLSLKDFEVGRKLGKGKFGKVYCVRHKKSGFICALKAIE 132
>gb|EAK91379.1| potential ER biogenesis protein fragment [Candida albicans SC5314]
Length = 805
Score = 30.8 bits (68), Expect = 7.9
Identities = 20/58 (34%), Positives = 33/58 (56%), Gaps = 1/58 (1%)
Query: 2 KNLSSMIEESSYKAKAKMKDIETLEKKGKGQHGVMVVRRNENYEYAQVMRELEYLKKE 59
K SS EE + +A+ + KD+E+ EKK + V +NEN E + + ++E +KE
Sbjct: 141 KETSSKNEEKAEEAEGQSKDLESTEKKELNPKDNVNVNKNEN-EKSGLKDKVETNEKE 197
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.313 0.129 0.347
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 132,168,376
Number of Sequences: 2540612
Number of extensions: 3192762
Number of successful extensions: 8778
Number of sequences better than 10.0: 7
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 8773
Number of HSP's gapped (non-prelim): 7
length of query: 122
length of database: 863,360,394
effective HSP length: 98
effective length of query: 24
effective length of database: 614,380,418
effective search space: 14745130032
effective search space used: 14745130032
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 68 (30.8 bits)
Medicago: description of AC147875.9


