

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147179.2 + phase: 0
(185 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
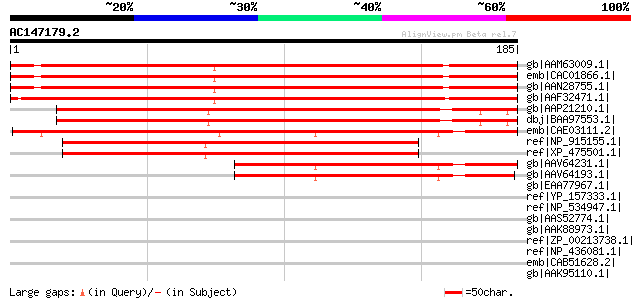
Score E
Sequences producing significant alignments: (bits) Value
gb|AAM63009.1| unknown [Arabidopsis thaliana] 257 1e-67
emb|CAC01866.1| putative protein [Arabidopsis thaliana] gi|15237... 255 4e-67
gb|AAN28755.1| At5g16250/T21H19_170 [Arabidopsis thaliana] gi|21... 254 8e-67
gb|AAF32471.1| unknown protein [Arabidopsis thaliana] gi|2159383... 243 1e-63
gb|AAP21210.1| At5g36710 [Arabidopsis thaliana] gi|62318562|dbj|... 200 1e-50
dbj|BAA97553.1| unnamed protein product [Arabidopsis thaliana] g... 200 1e-50
emb|CAE03111.2| OSJNBa0067K08.8 [Oryza sativa (japonica cultivar... 198 7e-50
ref|NP_915155.1| P0696G06.26 [Oryza sativa (japonica cultivar-gr... 159 5e-38
ref|XP_475501.1| putative mutator-like transposase [Oryza sativa... 154 1e-36
gb|AAV64231.1| unknown [Zea mays] 113 2e-24
gb|AAV64193.1| unknown [Zea mays] 111 1e-23
gb|EAA77967.1| hypothetical protein FG07773.1 [Gibberella zeae P... 38 0.12
ref|YP_157333.1| conserved hypothetical protein, predicted sulfa... 35 1.0
ref|NP_534947.1| ABC transporter, membrane spanning protein [iro... 35 1.0
gb|AAS52774.1| AER090Wp [Ashbya gossypii ATCC 10895] gi|45190696... 35 1.0
gb|AAK88973.1| AGR_L_801p [Agrobacterium tumefaciens str. C58] g... 35 1.0
ref|ZP_00213738.1| COG3302: DMSO reductase anchor subunit [Burkh... 35 1.0
ref|NP_436081.1| NuoN2 NADH I CHAIN N [Sinorhizobium meliloti 1... 35 1.3
emb|CAB51628.2| putative NADH-ubiquinone oxidoreductase subunit ... 35 1.3
gb|AAK95110.1| olfactory receptor [Homo sapiens] 35 1.3
>gb|AAM63009.1| unknown [Arabidopsis thaliana]
Length = 183
Score = 257 bits (656), Expect = 1e-67
Identities = 129/187 (68%), Positives = 150/187 (79%), Gaps = 6/187 (3%)
Query: 1 MGLSMNNPPATITPRHYNIHKLFLFCNYVLLGAASSCIFLTLSLRLVPSVCGFFLILLHI 60
MGL ++ P + HY+ HK+FLF NY+LLGAASSCIFLTLSLRL+PS+CGF LILLH
Sbjct: 1 MGLISSSSP--VEESHYHTHKIFLFSNYILLGAASSCIFLTLSLRLIPSICGFLLILLHA 58
Query: 61 FTIAGAVSGCAAV--GANRWYSAHMVATVLTAIFQGSVSVLVFTRTSDFLGELQSYVREE 118
TIA AVSGCAA G NRWY+AHMVATVLTAIFQGSVSVL+FT TS FLG L+SYVREE
Sbjct: 59 TTIAAAVSGCAAASCGRNRWYAAHMVATVLTAIFQGSVSVLIFTNTSKFLGSLKSYVREE 118
Query: 119 DGSVILKLSGGLAILIFCLEWVVLTLAFFLKYYACVEGGNTCRTVVLGSAKVQQDEDLKD 178
D +VILKL GGL I+IFCL+W+VL AFFLKYYA V+GG+ + + KVQ +E+ KD
Sbjct: 119 DAAVILKLGGGLCIVIFCLDWIVLVCAFFLKYYAYVDGGD--GVAMKRTGKVQSEENPKD 176
Query: 179 WPWPFQV 185
WPWPFQV
Sbjct: 177 WPWPFQV 183
>emb|CAC01866.1| putative protein [Arabidopsis thaliana]
gi|15237310|ref|NP_197129.1| expressed protein
[Arabidopsis thaliana] gi|11346160|pir||T51495
hypothetical protein T21H19_170 - Arabidopsis thaliana
Length = 183
Score = 255 bits (652), Expect = 4e-67
Identities = 128/187 (68%), Positives = 149/187 (79%), Gaps = 6/187 (3%)
Query: 1 MGLSMNNPPATITPRHYNIHKLFLFCNYVLLGAASSCIFLTLSLRLVPSVCGFFLILLHI 60
MG ++ P + HY+ HK+FLF NY+LLGAASSCIFLTLSLRL+PS+CGF LILLH
Sbjct: 1 MGFISSSSP--VEESHYHTHKIFLFSNYILLGAASSCIFLTLSLRLIPSICGFLLILLHA 58
Query: 61 FTIAGAVSGCAAV--GANRWYSAHMVATVLTAIFQGSVSVLVFTRTSDFLGELQSYVREE 118
TIA AVSGCAA G NRWY+AHMVATVLTAIFQGSVSVL+FT TS FLG L+SYVREE
Sbjct: 59 TTIAAAVSGCAAASCGRNRWYAAHMVATVLTAIFQGSVSVLIFTNTSKFLGSLKSYVREE 118
Query: 119 DGSVILKLSGGLAILIFCLEWVVLTLAFFLKYYACVEGGNTCRTVVLGSAKVQQDEDLKD 178
D +VILKL GGL I+IFCL+W+VL AFFLKYYA V+GG+ + + KVQ +E+ KD
Sbjct: 119 DAAVILKLGGGLCIVIFCLDWIVLVCAFFLKYYAYVDGGD--GVAMKRTGKVQSEENPKD 176
Query: 179 WPWPFQV 185
WPWPFQV
Sbjct: 177 WPWPFQV 183
>gb|AAN28755.1| At5g16250/T21H19_170 [Arabidopsis thaliana]
gi|21703131|gb|AAM74506.1| AT5g16250/T21H19_170
[Arabidopsis thaliana]
Length = 183
Score = 254 bits (649), Expect = 8e-67
Identities = 127/187 (67%), Positives = 149/187 (78%), Gaps = 6/187 (3%)
Query: 1 MGLSMNNPPATITPRHYNIHKLFLFCNYVLLGAASSCIFLTLSLRLVPSVCGFFLILLHI 60
MG ++ P + HY+ HK+FLF NY+LLGAASSCIFLTLSLRL+PS+CGF LILLH
Sbjct: 1 MGFISSSSP--VEESHYHTHKIFLFSNYILLGAASSCIFLTLSLRLIPSICGFLLILLHA 58
Query: 61 FTIAGAVSGCAAV--GANRWYSAHMVATVLTAIFQGSVSVLVFTRTSDFLGELQSYVREE 118
TIA AVSGCAA G NRWY+AHMVATVLTAIFQGSVSVL+FT TS FLG L+SYVR+E
Sbjct: 59 TTIAAAVSGCAAASCGRNRWYAAHMVATVLTAIFQGSVSVLIFTNTSKFLGSLKSYVRDE 118
Query: 119 DGSVILKLSGGLAILIFCLEWVVLTLAFFLKYYACVEGGNTCRTVVLGSAKVQQDEDLKD 178
D +VILKL GGL I+IFCL+W+VL AFFLKYYA V+GG+ + + KVQ +E+ KD
Sbjct: 119 DAAVILKLGGGLCIVIFCLDWIVLVCAFFLKYYAYVDGGD--GVAMKRTGKVQSEENPKD 176
Query: 179 WPWPFQV 185
WPWPFQV
Sbjct: 177 WPWPFQV 183
>gb|AAF32471.1| unknown protein [Arabidopsis thaliana] gi|21593832|gb|AAM65799.1|
unknown [Arabidopsis thaliana]
gi|22136266|gb|AAM91211.1| unknown protein [Arabidopsis
thaliana] gi|17065096|gb|AAL32702.1| Unknown protein
[Arabidopsis thaliana] gi|18396262|ref|NP_566178.1|
expressed protein [Arabidopsis thaliana]
Length = 185
Score = 243 bits (621), Expect = 1e-63
Identities = 126/187 (67%), Positives = 147/187 (78%), Gaps = 4/187 (2%)
Query: 1 MGLSMNNPPATITPRHYNIHKLFLFCNYVLLGAASSCIFLTLSLRLVPSVCGFFLILLHI 60
MGL + P +I HY HKLFL NYVLLGA+SSCIFLTLSLRL+PS+CGFFLILLH
Sbjct: 1 MGL-IPQPQESIQESHYYTHKLFLTANYVLLGASSSCIFLTLSLRLIPSLCGFFLILLHA 59
Query: 61 FTIAGAVSGCAAV--GANRWYSAHMVATVLTAIFQGSVSVLVFTRTSDFLGELQSYVREE 118
TIA AVSGCAA G NRWY+AHM+ATVLTAIFQGSVSVL+FT TS+FL L SYVRE+
Sbjct: 60 TTIAAAVSGCAAASYGKNRWYAAHMIATVLTAIFQGSVSVLIFTNTSNFLESLNSYVREK 119
Query: 119 DGSVILKLSGGLAILIFCLEWVVLTLAFFLKYYACVEGGNTCRTVVLGSAKVQQDEDLKD 178
+ S+ILKL+GGL ++IFCLEW+VL LAFFLKYYA V+G N + + KVQ +E LK+
Sbjct: 120 EASMILKLAGGLCVVIFCLEWIVLVLAFFLKYYAYVDGDNN-GVAMKRTGKVQSEETLKN 178
Query: 179 WPWPFQV 185
PW FQV
Sbjct: 179 SPWAFQV 185
>gb|AAP21210.1| At5g36710 [Arabidopsis thaliana] gi|62318562|dbj|BAD94940.1|
putative protein [Arabidopsis thaliana]
gi|15239407|ref|NP_198496.1| expressed protein
[Arabidopsis thaliana] gi|42568153|ref|NP_198486.2|
expressed protein [Arabidopsis thaliana]
Length = 183
Score = 200 bits (509), Expect = 1e-50
Identities = 109/177 (61%), Positives = 131/177 (73%), Gaps = 13/177 (7%)
Query: 18 NIHKLFLFCNYVLLGAASSCIFLTLSLRLVPSVCGFFLILLHIFTIAGAVSGCA------ 71
N H +FL CNY+LLG+ASSCIFLT+SLRL PS+ G LI L+ TIA AVSGC+
Sbjct: 9 NTHNIFLLCNYILLGSASSCIFLTISLRLFPSLSGLSLIFLYTLTIATAVSGCSIFASST 68
Query: 72 -AVGANRWYSAHMVATVLTAIFQGSVSVLVFTRTSDFLGELQSYVREEDGSVILKLSGGL 130
A ++R Y +HMVATVLTAIFQG+VSVL+FTRT DFL L+SYVREEDG VILKLSGGL
Sbjct: 69 SATASDRLYGSHMVATVLTAIFQGAVSVLIFTRTGDFLRFLKSYVREEDGEVILKLSGGL 128
Query: 131 AILIFCLEWVVLTLAFFLKYYACVEGGNTCRTVVLGSAKV-QQDEDLKDWP-WPFQV 185
+L+FCLEW+VL LAF LKY ++ V KV +Q+EDLKDWP +PFQ+
Sbjct: 129 CVLMFCLEWIVLVLAFLLKYSDYLDES----VVDDDDFKVRRQEEDLKDWPSYPFQL 181
>dbj|BAA97553.1| unnamed protein product [Arabidopsis thaliana]
gi|8953700|dbj|BAA98058.1| unnamed protein product
[Arabidopsis thaliana]
Length = 218
Score = 200 bits (509), Expect = 1e-50
Identities = 109/177 (61%), Positives = 131/177 (73%), Gaps = 13/177 (7%)
Query: 18 NIHKLFLFCNYVLLGAASSCIFLTLSLRLVPSVCGFFLILLHIFTIAGAVSGCA------ 71
N H +FL CNY+LLG+ASSCIFLT+SLRL PS+ G LI L+ TIA AVSGC+
Sbjct: 44 NTHNIFLLCNYILLGSASSCIFLTISLRLFPSLSGLSLIFLYTLTIATAVSGCSIFASST 103
Query: 72 -AVGANRWYSAHMVATVLTAIFQGSVSVLVFTRTSDFLGELQSYVREEDGSVILKLSGGL 130
A ++R Y +HMVATVLTAIFQG+VSVL+FTRT DFL L+SYVREEDG VILKLSGGL
Sbjct: 104 SATASDRLYGSHMVATVLTAIFQGAVSVLIFTRTGDFLRFLKSYVREEDGEVILKLSGGL 163
Query: 131 AILIFCLEWVVLTLAFFLKYYACVEGGNTCRTVVLGSAKV-QQDEDLKDWP-WPFQV 185
+L+FCLEW+VL LAF LKY ++ V KV +Q+EDLKDWP +PFQ+
Sbjct: 164 CVLMFCLEWIVLVLAFLLKYSDYLDES----VVDDDDFKVRRQEEDLKDWPSYPFQL 216
>emb|CAE03111.2| OSJNBa0067K08.8 [Oryza sativa (japonica cultivar-group)]
gi|50925967|ref|XP_473031.1| OSJNBa0067K08.8 [Oryza
sativa (japonica cultivar-group)]
Length = 193
Score = 198 bits (503), Expect = 7e-50
Identities = 108/195 (55%), Positives = 133/195 (67%), Gaps = 15/195 (7%)
Query: 2 GLSMNNPPA--TITPRHYNIHKLFLFCNYVLLGAASSCIFLTLSLRLVPSVCGFFLILLH 59
GLS+ PPA + +P H + FL NY++LGAAS C FLTLSLRLVPSV GF LILLH
Sbjct: 3 GLSVIAPPAGDSASPAHRRARRAFLVSNYMILGAASGCGFLTLSLRLVPSVDGFLLILLH 62
Query: 60 IFTIAGAVSGCAAVGA-----NRWYSAHMVATVLTAIFQGSVSVLVFTRTSDFLGE-LQS 113
T+A AV+GCA + A R Y+ HM TV +I QG+ +VL F+RTSDFL + L+S
Sbjct: 63 AITVAAAVAGCAVIAAPDPPRGRVYTTHMAGTVFVSILQGAAAVLTFSRTSDFLADGLKS 122
Query: 114 YVREEDGSVILKLSGGLAILIFCLEWVVLTLAFFLKYYACVE---GGNTCRTVVLGSAKV 170
YVREEDG+VIL++ GGL + IFCLEW+ L LAF L+YYA V+ GGN R SAKV
Sbjct: 123 YVREEDGAVILRMIGGLGVAIFCLEWIALALAFVLRYYAYVDRECGGNPLRR----SAKV 178
Query: 171 QQDEDLKDWPWPFQV 185
++ WPWPFQV
Sbjct: 179 GGEDGAGTWPWPFQV 193
>ref|NP_915155.1| P0696G06.26 [Oryza sativa (japonica cultivar-group)]
gi|21952854|dbj|BAC06269.1| P0696G06.26 [Oryza sativa
(japonica cultivar-group)]
Length = 166
Score = 159 bits (401), Expect = 5e-38
Identities = 79/135 (58%), Positives = 95/135 (69%), Gaps = 5/135 (3%)
Query: 20 HKLFLFCNYVLLGAASSCIFLTLSLRLVPSVCGFFLILLHIFTIAGAVSGC-----AAVG 74
H+ FL CNY LLGAASSCIFLTLSLRL+PS CG L+ LH T + +GC A
Sbjct: 10 HRAFLLCNYALLGAASSCIFLTLSLRLLPSPCGLLLLFLHALTAVFSAAGCSGSFTAPAT 69
Query: 75 ANRWYSAHMVATVLTAIFQGSVSVLVFTRTSDFLGELQSYVREEDGSVILKLSGGLAILI 134
+W++AH LTAIFQG+V++L FTRTSDFL ELQSYVR++D +VILK+ GGL I
Sbjct: 70 PAQWHNAHTAGAALTAIFQGAVALLAFTRTSDFLAELQSYVRDDDAAVILKMVGGLGTAI 129
Query: 135 FCLEWVVLTLAFFLK 149
F LEW L LAF L+
Sbjct: 130 FVLEWAALALAFSLR 144
>ref|XP_475501.1| putative mutator-like transposase [Oryza sativa (japonica
cultivar-group)] gi|46981280|gb|AAT07598.1| putative
mutator-like transposase [Oryza sativa (japonica
cultivar-group)]
Length = 1725
Score = 154 bits (389), Expect = 1e-36
Identities = 79/135 (58%), Positives = 93/135 (68%), Gaps = 5/135 (3%)
Query: 20 HKLFLFCNYVLLGAASSCIFLTLSLRLVPSVCGFFLILLHIFTIAGAVSGC-----AAVG 74
H+ FL CNY+LLGAAS CIFLTLSLRL+PS CG L+ LH T A + C A G
Sbjct: 7 HRAFLLCNYLLLGAASGCIFLTLSLRLLPSPCGLLLVFLHALTAVLAAAACSGSFTAPGG 66
Query: 75 ANRWYSAHMVATVLTAIFQGSVSVLVFTRTSDFLGELQSYVREEDGSVILKLSGGLAILI 134
++AH + VLTAIFQG+ ++L FTRT DFL EL+SYVREEDG VIL+L GGL I
Sbjct: 67 GGGTHTAHTASAVLTAIFQGAAALLAFTRTGDFLAELRSYVREEDGEVILELVGGLGAAI 126
Query: 135 FCLEWVVLTLAFFLK 149
F LEW L LAF L+
Sbjct: 127 FVLEWAALALAFALR 141
>gb|AAV64231.1| unknown [Zea mays]
Length = 133
Score = 113 bits (283), Expect = 2e-24
Identities = 59/107 (55%), Positives = 76/107 (70%), Gaps = 8/107 (7%)
Query: 83 MVATVLTAIFQGSVSVLVFTRTSDFLGE-LQSYVREEDGSVILKLSGGLAILIFCLEWVV 141
M ATV+ +I QG+ +VL F+RTS+FL + L+SYVREEDG+VIL++ GGL + IFCLEWV
Sbjct: 1 MSATVVVSILQGAAAVLAFSRTSEFLSDGLKSYVREEDGAVILRMVGGLGVAIFCLEWVA 60
Query: 142 LTLAFFLKYYACVE---GGNTCRTVVLGSAKVQQDEDLKDWPWPFQV 185
L LAF L+YY V+ GGN R SAKV ++ WPWPFQ+
Sbjct: 61 LALAFVLRYYVYVDRECGGNPMRR----SAKVGGEDGAGTWPWPFQI 103
>gb|AAV64193.1| unknown [Zea mays]
Length = 151
Score = 111 bits (277), Expect = 1e-23
Identities = 59/106 (55%), Positives = 75/106 (70%), Gaps = 8/106 (7%)
Query: 83 MVATVLTAIFQGSVSVLVFTRTSDFLGE-LQSYVREEDGSVILKLSGGLAILIFCLEWVV 141
M ATV+ +I QG+ +VL F+RTS+FL + L+SYVREEDG+VIL++ GGL + IFCLEWV
Sbjct: 1 MSATVVVSILQGAAAVLAFSRTSEFLSDGLKSYVREEDGAVILRMVGGLGVGIFCLEWVA 60
Query: 142 LTLAFFLKYYACVE---GGNTCRTVVLGSAKVQQDEDLKDWPWPFQ 184
L LAF L+YY V+ GGN R SAKV ++ WPWPFQ
Sbjct: 61 LALAFVLRYYVYVDRECGGNPMRR----SAKVGGEDGAGTWPWPFQ 102
>gb|EAA77967.1| hypothetical protein FG07773.1 [Gibberella zeae PH-1]
gi|46126791|ref|XP_387949.1| hypothetical protein
FG07773.1 [Gibberella zeae PH-1]
Length = 887
Score = 38.1 bits (87), Expect = 0.12
Identities = 27/123 (21%), Positives = 56/123 (44%), Gaps = 15/123 (12%)
Query: 54 FLILLHIFTIAGAVSGCAAVGANRWY---SAHMVATVLTAIFQGSVSVLVFTRTSDFLGE 110
++ L ++ ++ GA S + W+ S H+V V +AI G +S+ + F+G
Sbjct: 23 YVALSYLISLVGAASTLELINRRSWFNGISNHLVL-VSSAITMGGISIW----SMHFIGN 77
Query: 111 LQSYVREEDGSVILKLSGGLAILIFCLEWVVLTLAFF-------LKYYACVEGGNTCRTV 163
+ + + + + S G+ +L FC+ V L AF+ + ++ + G C +
Sbjct: 78 QALTLGQGEAEMQVDYSVGITVLSFCMPVVFLLAAFYAIGISNGIAWWRVITAGILCGSA 137
Query: 164 VLG 166
+ G
Sbjct: 138 ICG 140
>ref|YP_157333.1| conserved hypothetical protein, predicted sulfatase family
[Azoarcus sp. EbN1] gi|56311787|emb|CAI06432.1|
conserved hypothetical protein,predicted sulfatase
family [Azoarcus sp. EbN1]
Length = 690
Score = 35.0 bits (79), Expect = 1.0
Identities = 29/109 (26%), Positives = 48/109 (43%), Gaps = 10/109 (9%)
Query: 40 LTLSLRLVPSVCGFFLILLHIFTIAGAVSGCAAVGANRWYSAHMVATVLTAIFQGSVSVL 99
L LRLV +C ++ + GAV AA A+R A + A A+F G + +
Sbjct: 70 LRFDLRLVVYLCAPLILAM------GAVRAMAARSAHR---AWLTAAASIALFLGLIELD 120
Query: 100 VFTRTSDFLGELQ-SYVREEDGSVILKLSGGLAILIFCLEWVVLTLAFF 147
+ L L Y+RE+ +V+ L G +L + W++ + F
Sbjct: 121 FYREFHQRLNSLVFQYLREDPATVLSMLWHGFPVLRYLAAWLLASWGLF 169
>ref|NP_534947.1| ABC transporter, membrane spanning protein [iron] [Agrobacterium
tumefaciens str. C58] gi|17742948|gb|AAL45263.1| ABC
transporter, membrane spanning protein [iron]
[Agrobacterium tumefaciens str. C58]
gi|25316173|pir||AI3105 hypothetical protein sitC
[imported] - Agrobacterium tumefaciens (strain C58,
Dupont)
Length = 286
Score = 35.0 bits (79), Expect = 1.0
Identities = 35/144 (24%), Positives = 66/144 (45%), Gaps = 15/144 (10%)
Query: 25 FCNYVLLGAASSCIFLTLSLRLVPSVCGFFLILLHIFTIAGAVSGCAAVGA-------NR 77
F + L+ S I LS +VP V G +++ L F++ +SG A G+ R
Sbjct: 30 FLSAYLMLKGWSLIGDALSHAIVPGVAGAYMLGLP-FSLGAFLSGGLAAGSMLFLNQKTR 88
Query: 78 WYSAHMVATVLTAIFQGSVSVLVFTRTSD-----FLGELQSYVREEDGSVILKLSGGLAI 132
++ + T+ F + ++ + TS LG + + E+ ++ L L GG+++
Sbjct: 89 LKEDAIIGLIFTSFFGLGLFMVSLSPTSVNIQTIVLGNILAITAED--TIQLALIGGISL 146
Query: 133 LIFCLEWVVLTLAFFLKYYACVEG 156
L+ L+W + FF + +A G
Sbjct: 147 LVLALKWRDFMVVFFDENHARTVG 170
>gb|AAS52774.1| AER090Wp [Ashbya gossypii ATCC 10895] gi|45190696|ref|NP_984950.1|
AER090Wp [Eremothecium gossypii]
Length = 676
Score = 35.0 bits (79), Expect = 1.0
Identities = 33/138 (23%), Positives = 63/138 (44%), Gaps = 10/138 (7%)
Query: 41 TLSLRLVPSVCGFFLILLHIFTIAGAVSGCAAVGANRWYSAHMVATVLTAIFQGSVSVLV 100
++ L + G + +FT+AGA++G ++G + W + +VA V + + +
Sbjct: 532 SIRAHLAFDILGLLSAFILLFTLAGAMTGALSLGMHAWVNRLLVAAVAGFALLSATVIQI 591
Query: 101 FTRTSDFLGELQSY-VREEDGSVILKLSGGLAILIFCLEWVVLTLAFFLKYYACVEGGNT 159
T + ++ +Y V + G+ LS + F LE +V+ L + YYA GG +
Sbjct: 592 VTAVVKWGAKIDNYGVVYKQGTAQATLSW----VAFVLEAIVV-LIVWGAYYA---GGKS 643
Query: 160 CRTVVLGSAKVQ-QDEDL 176
V + + DED+
Sbjct: 644 QGVVNVEEGRASTSDEDI 661
>gb|AAK88973.1| AGR_L_801p [Agrobacterium tumefaciens str. C58]
gi|25316162|pir||C98181 sitC protein (AF128999)
[imported] - Agrobacterium tumefaciens (strain C58,
Cereon) gi|15890516|ref|NP_356188.1| hypothetical
protein AGR_L_801 [Agrobacterium tumefaciens str. C58]
Length = 301
Score = 35.0 bits (79), Expect = 1.0
Identities = 35/144 (24%), Positives = 66/144 (45%), Gaps = 15/144 (10%)
Query: 25 FCNYVLLGAASSCIFLTLSLRLVPSVCGFFLILLHIFTIAGAVSGCAAVGA-------NR 77
F + L+ S I LS +VP V G +++ L F++ +SG A G+ R
Sbjct: 45 FLSAYLMLKGWSLIGDALSHAIVPGVAGAYMLGLP-FSLGAFLSGGLAAGSMLFLNQKTR 103
Query: 78 WYSAHMVATVLTAIFQGSVSVLVFTRTSD-----FLGELQSYVREEDGSVILKLSGGLAI 132
++ + T+ F + ++ + TS LG + + E+ ++ L L GG+++
Sbjct: 104 LKEDAIIGLIFTSFFGLGLFMVSLSPTSVNIQTIVLGNILAITAED--TIQLALIGGISL 161
Query: 133 LIFCLEWVVLTLAFFLKYYACVEG 156
L+ L+W + FF + +A G
Sbjct: 162 LVLALKWRDFMVVFFDENHARTVG 185
>ref|ZP_00213738.1| COG3302: DMSO reductase anchor subunit [Burkholderia cepacia
R18194]
Length = 317
Score = 35.0 bits (79), Expect = 1.0
Identities = 19/51 (37%), Positives = 27/51 (52%), Gaps = 1/51 (1%)
Query: 27 NYVLLGAASSCIFLTLSLR-LVPSVCGFFLILLHIFTIAGAVSGCAAVGAN 76
N+VLLG AS C T L PS+ G + + T+AG V+ A++ N
Sbjct: 150 NFVLLGCASGCTLATACAAWLAPSLVGRLAVAACVLTLAGGVARLASLARN 200
>ref|NP_436081.1| NuoN2 NADH I CHAIN N [Sinorhizobium meliloti 1021]
gi|14523965|gb|AAK65493.1| NuoN2 NADH I CHAIN N
[Sinorhizobium meliloti 1021]
gi|17380560|sp|P56911|NUN2_RHIME NADH-quinone
oxidoreductase chain N 2 (NADH dehydrogenase I, chain N
2) (NDH-1, chain N 2) gi|25285083|pir||C95366 NADH2
dehydrogenase (ubiquinone) (EC 1.6.5.3) chain N NuoN2
[imported] - Sinorhizobium meliloti (strain 1021)
magaplasmid pSymA
Length = 479
Score = 34.7 bits (78), Expect = 1.3
Identities = 31/106 (29%), Positives = 45/106 (42%), Gaps = 12/106 (11%)
Query: 66 AVSGCAAVGANRWYSAHMVATVLTAIFQGSVSVLVFTRTSDFLGELQSYVREEDGSVILK 125
A+ G A G + S + + T + G ++ R DF GE+ ED + K
Sbjct: 311 AIFGVVAGGQDGIASVMLYLLIYTFMNLGIFGAVIMMRNGDFSGEV-----IEDYAGFAK 365
Query: 126 LSGGLAIL----IFCLEWVVLTLAFFLKYY---ACVEGGNTCRTVV 164
GLA+L +F L + T FF K+Y A VE G V+
Sbjct: 366 FHPGLALLMLLYLFSLAGIPPTAGFFAKFYVLVALVERGFVALAVI 411
>emb|CAB51628.2| putative NADH-ubiquinone oxidoreductase subunit [Sinorhizobium
meliloti]
Length = 479
Score = 34.7 bits (78), Expect = 1.3
Identities = 31/106 (29%), Positives = 45/106 (42%), Gaps = 12/106 (11%)
Query: 66 AVSGCAAVGANRWYSAHMVATVLTAIFQGSVSVLVFTRTSDFLGELQSYVREEDGSVILK 125
A+ G A G + S + + T + G ++ R DF GE+ ED + K
Sbjct: 311 AIFGVVAGGQDGIASVMLYLLIYTFMNLGIFGAVIMMRNGDFSGEV-----IEDYAGFAK 365
Query: 126 LSGGLAIL----IFCLEWVVLTLAFFLKYY---ACVEGGNTCRTVV 164
GLA+L +F L + T FF K+Y A VE G V+
Sbjct: 366 FHPGLALLMLLYLFSLAGIPPTAGFFAKFYVLVALVERGFVALAVI 411
>gb|AAK95110.1| olfactory receptor [Homo sapiens]
Length = 216
Score = 34.7 bits (78), Expect = 1.3
Identities = 21/62 (33%), Positives = 30/62 (47%), Gaps = 5/62 (8%)
Query: 28 YVLLGAASSCIFLTLSLRLVPSVCGFFLILLHIFTIAGAVSGCAAVGANRWYSAHMVATV 87
+ GAA C+ T++ ++C LH I G +S CA + A W+S VATV
Sbjct: 37 FFFFGAAECCLLATMAYDRYVAICD----PLHYPVIMGHIS-CAQLAAASWFSGFSVATV 91
Query: 88 LT 89
T
Sbjct: 92 QT 93
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.328 0.142 0.449
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 308,761,203
Number of Sequences: 2540612
Number of extensions: 11280696
Number of successful extensions: 36095
Number of sequences better than 10.0: 44
Number of HSP's better than 10.0 without gapping: 12
Number of HSP's successfully gapped in prelim test: 32
Number of HSP's that attempted gapping in prelim test: 36055
Number of HSP's gapped (non-prelim): 50
length of query: 185
length of database: 863,360,394
effective HSP length: 120
effective length of query: 65
effective length of database: 558,486,954
effective search space: 36301652010
effective search space used: 36301652010
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 71 (32.0 bits)
Medicago: description of AC147179.2


