

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144928.8 + phase: 0 /pseudo
(359 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
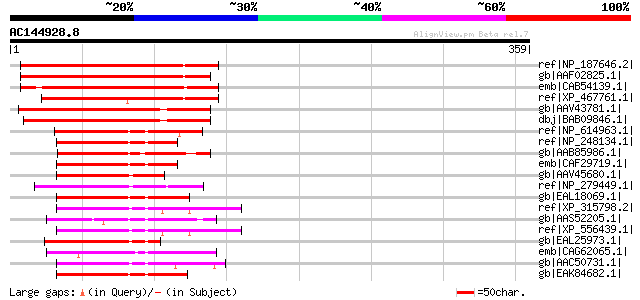
Score E
Sequences producing significant alignments: (bits) Value
ref|NP_187646.2| anion-transporting ATPase family protein [Arabi... 225 2e-57
gb|AAF02825.1| putative ATPase [Arabidopsis thaliana] 224 4e-57
emb|CAB54139.1| ATPase [Solanum tuberosum] 199 8e-50
ref|XP_467761.1| putative ATPase [Oryza sativa (japonica cultiva... 190 6e-47
gb|AAV43781.1| At5g60730 [Arabidopsis thaliana] gi|52627093|gb|A... 156 8e-37
dbj|BAB09846.1| arsenite translocating ATPase-like protein [Arab... 154 4e-36
ref|NP_614963.1| Arsenite transporting ATPase [Methanopyrus kand... 78 4e-13
ref|NP_248134.1| arsenical pump-driving ATPase (arsA) [Methanoca... 74 5e-12
gb|AAB85986.1| arsenical pump-driving ATPase [Methanothermobacte... 74 7e-12
emb|CAF29719.1| Putative arsenical pump-driving ATPase [Methanoc... 74 9e-12
gb|AAV45680.1| arsenical pump-driving ATPase [Haloarcula marismo... 73 2e-11
ref|NP_279449.1| ArsA1 [Halobacterium sp. NRC-1] gi|10579983|gb|... 72 2e-11
gb|EAL18069.1| hypothetical protein CNBK0900 [Cryptococcus neofo... 72 3e-11
ref|XP_315798.2| ENSANGP00000018739 [Anopheles gambiae str. PEST... 71 4e-11
gb|AAS52205.1| ADR285Wp [Ashbya gossypii ATCC 10895] gi|45188158... 71 4e-11
ref|XP_556439.1| ENSANGP00000029267 [Anopheles gambiae str. PEST... 71 4e-11
gb|EAL25973.1| GA14038-PA [Drosophila pseudoobscura] 70 7e-11
emb|CAG62065.1| unnamed protein product [Candida glabrata CBS138... 70 1e-10
gb|AAC50731.1| hASNA-I [Homo sapiens] 70 1e-10
gb|EAK84682.1| hypothetical protein UM03838.1 [Ustilago maydis 5... 69 2e-10
>ref|NP_187646.2| anion-transporting ATPase family protein [Arabidopsis thaliana]
Length = 411
Score = 225 bits (573), Expect = 2e-57
Identities = 110/137 (80%), Positives = 126/137 (91%), Gaps = 1/137 (0%)
Query: 8 LSVKSVAAPTESISVFDDMVAGTERKYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVS 67
L VKSVA+PTE+IS FD+MV+GT+RKYYMLGGKGGVGKTSCAASLAV+FANNGHPTLVVS
Sbjct: 63 LQVKSVASPTETISEFDEMVSGTKRKYYMLGGKGGVGKTSCAASLAVRFANNGHPTLVVS 122
Query: 68 TDPAHSLSDSFAQDLTGGALVQVDGPDYPLFALEINPEKAREDFRDVAKQNGGSTGVKDF 127
TDPAHSLSDSFAQDLTGG LV V+GP+ PLFALEINPEKARE+FR ++ NGG TGVKDF
Sbjct: 123 TDPAHSLSDSFAQDLTGGMLVPVEGPEAPLFALEINPEKAREEFRSASQMNGG-TGVKDF 181
Query: 128 MDGMGLGMIVDQTAKVR 144
MDGMGLGM+V+Q +++
Sbjct: 182 MDGMGLGMLVEQLGELK 198
>gb|AAF02825.1| putative ATPase [Arabidopsis thaliana]
Length = 386
Score = 224 bits (570), Expect = 4e-57
Identities = 110/132 (83%), Positives = 123/132 (92%), Gaps = 1/132 (0%)
Query: 8 LSVKSVAAPTESISVFDDMVAGTERKYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVS 67
L VKSVA+PTE+IS FD+MV+GT+RKYYMLGGKGGVGKTSCAASLAV+FANNGHPTLVVS
Sbjct: 63 LQVKSVASPTETISEFDEMVSGTKRKYYMLGGKGGVGKTSCAASLAVRFANNGHPTLVVS 122
Query: 68 TDPAHSLSDSFAQDLTGGALVQVDGPDYPLFALEINPEKAREDFRDVAKQNGGSTGVKDF 127
TDPAHSLSDSFAQDLTGG LV V+GP+ PLFALEINPEKARE+FR ++ NGG TGVKDF
Sbjct: 123 TDPAHSLSDSFAQDLTGGMLVPVEGPEAPLFALEINPEKAREEFRSASQMNGG-TGVKDF 181
Query: 128 MDGMGLGMIVDQ 139
MDGMGLGM+V+Q
Sbjct: 182 MDGMGLGMLVEQ 193
>emb|CAB54139.1| ATPase [Solanum tuberosum]
Length = 369
Score = 199 bits (507), Expect = 8e-50
Identities = 99/137 (72%), Positives = 117/137 (85%), Gaps = 5/137 (3%)
Query: 8 LSVKSVAAPTESISVFDDMVAGTERKYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVS 67
L V++VA+P FD+MV+GTERKYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVS
Sbjct: 50 LQVRAVASPAG----FDEMVSGTERKYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVS 105
Query: 68 TDPAHSLSDSFAQDLTGGALVQVDGPDYPLFALEINPEKAREDFRDVAKQNGGSTGVKDF 127
TDPAHSLSDSFAQDL GG LV V+GP PLFALE+NPEKA+E+FR + +GGS G+KDF
Sbjct: 106 TDPAHSLSDSFAQDLAGGTLVPVEGPYSPLFALELNPEKAKEEFRSATQISGGS-GIKDF 164
Query: 128 MDGMGLGMIVDQTAKVR 144
MDGMGLG++ +Q +++
Sbjct: 165 MDGMGLGVLAEQLGELK 181
>ref|XP_467761.1| putative ATPase [Oryza sativa (japonica cultivar-group)]
gi|46390644|dbj|BAD16127.1| putative ATPase [Oryza
sativa (japonica cultivar-group)]
gi|46390107|dbj|BAD15543.1| putative ATPase [Oryza
sativa (japonica cultivar-group)]
Length = 406
Score = 190 bits (482), Expect = 6e-47
Identities = 95/135 (70%), Positives = 112/135 (82%), Gaps = 14/135 (10%)
Query: 23 FDDMVAGTERKYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSDSFAQ-- 80
F++M AGT R+YYMLGGKGGVGKTSCAASLAV+FANNGHPTLVVSTDPAHSLSDSFAQ
Sbjct: 60 FEEMAAGTRRRYYMLGGKGGVGKTSCAASLAVRFANNGHPTLVVSTDPAHSLSDSFAQVA 119
Query: 81 -----------DLTGGALVQVDGPDYPLFALEINPEKAREDFRDVAKQNGGSTGVKDFMD 129
DL+GGALV V+GP+ PLFALEINPEKARE+FR +++NGG TGVKDFMD
Sbjct: 120 SPVEHLLSRFEDLSGGALVPVEGPEAPLFALEINPEKAREEFRAASQKNGG-TGVKDFMD 178
Query: 130 GMGLGMIVDQTAKVR 144
GMGLG++ +Q +++
Sbjct: 179 GMGLGVLAEQLGELK 193
>gb|AAV43781.1| At5g60730 [Arabidopsis thaliana] gi|52627093|gb|AAU84673.1|
At5g60730 [Arabidopsis thaliana]
gi|30697424|ref|NP_200881.2| anion-transporting ATPase
family protein [Arabidopsis thaliana]
Length = 391
Score = 156 bits (395), Expect = 8e-37
Identities = 81/133 (60%), Positives = 97/133 (72%), Gaps = 4/133 (3%)
Query: 7 LLSVKSVAAPTESISVFDDMVAGTERKYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVV 66
L V+S+A E S F++MV+ +RKYY+LGGKGGVGKTSCAASLAVKFA++GHPT+VV
Sbjct: 44 LFRVRSLATLAEGASHFNEMVSVNQRKYYLLGGKGGVGKTSCAASLAVKFASHGHPTIVV 103
Query: 67 STDPAHSLSDSFAQDLTGGALVQVDGPDYPLFALEINPEKAREDFRDVAKQNGGSTGVKD 126
STDPAHSLSDSF+QDL+GG L V G D PL ALEI P E +D K+ G VK+
Sbjct: 104 STDPAHSLSDSFSQDLSGGVLKPVQGVDSPLLALEITP----EIMKDEIKRQTGDKSVKN 159
Query: 127 FMDGMGLGMIVDQ 139
MD MGLGM +
Sbjct: 160 MMDSMGLGMFAGE 172
>dbj|BAB09846.1| arsenite translocating ATPase-like protein [Arabidopsis thaliana]
Length = 417
Score = 154 bits (389), Expect = 4e-36
Identities = 79/130 (60%), Positives = 96/130 (73%), Gaps = 4/130 (3%)
Query: 10 VKSVAAPTESISVFDDMVAGTERKYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTD 69
++S+A E S F++MV+ +RKYY+LGGKGGVGKTSCAASLAVKFA++GHPT+VVSTD
Sbjct: 73 LRSLATLAEGASHFNEMVSVNQRKYYLLGGKGGVGKTSCAASLAVKFASHGHPTIVVSTD 132
Query: 70 PAHSLSDSFAQDLTGGALVQVDGPDYPLFALEINPEKAREDFRDVAKQNGGSTGVKDFMD 129
PAHSLSDSF+QDL+GG L V G D PL ALEI P E +D K+ G VK+ MD
Sbjct: 133 PAHSLSDSFSQDLSGGVLKPVQGVDSPLLALEITP----EIMKDEIKRQTGDKSVKNMMD 188
Query: 130 GMGLGMIVDQ 139
MGLGM +
Sbjct: 189 SMGLGMFAGE 198
>ref|NP_614963.1| Arsenite transporting ATPase [Methanopyrus kandleri AV19]
gi|19888411|gb|AAM02893.1| Arsenite transporting ATPase
[Methanopyrus kandleri AV19]
Length = 333
Score = 78.2 bits (191), Expect = 4e-13
Identities = 48/106 (45%), Positives = 66/106 (61%), Gaps = 6/106 (5%)
Query: 32 RKYYMLGGKGGVGKTSCAASLAVKFA-NNGHPTLVVSTDPAHSLSDSFAQDLTGGALVQV 90
++Y GGKGGVGKT+CAA+ AV + G LVVSTDPAHSLSD F Q++ G +
Sbjct: 13 QRYVFFGGKGGVGKTTCAAATAVWLSEEEGKEVLVVSTDPAHSLSDIFDQNI-GSEPTPI 71
Query: 91 DGPDYPLFALEINPEKAREDFRDVAK---QNGGSTGVKDFMDGMGL 133
+G + L A+EI+PEKA E++ +V K + G++D G L
Sbjct: 72 EGVE-GLKAIEIDPEKAAEEYVEVMKRVYEMSKDKGMEDLFGGEDL 116
>ref|NP_248134.1| arsenical pump-driving ATPase (arsA) [Methanocaldococcus jannaschii
DSM 2661] gi|1591774|gb|AAB99142.1| arsenical
pump-driving ATPase (arsA) [Methanocaldococcus
jannaschii DSM 2661] gi|2127768|pir||E64442 probable
arsenical pump-driving ATPase (EC 3.6.1.-) -
Methanococcus jannaschii gi|6647440|sp|Q58542|ARSA_METJA
Putative arsenical pump-driving ATPase
(Arsenite-translocating ATPase) (Arsenical resistance
ATPase) (Arsenite-transporting ATPase)
Length = 349
Score = 74.3 bits (181), Expect = 5e-12
Identities = 40/84 (47%), Positives = 55/84 (64%), Gaps = 2/84 (2%)
Query: 33 KYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSDSFAQDLTGGALVQVDG 92
KY M GGKGGVGKT+ +A+ V A G ++VSTDPAHSL D F Q+ G +V G
Sbjct: 27 KYIMFGGKGGVGKTTMSAATGVYLAEKGLKVVIVSTDPAHSLRDIFEQEF-GHEPTKVKG 85
Query: 93 PDYPLFALEINPEKAREDFRDVAK 116
D L+ +EI+P+KA E++++ K
Sbjct: 86 YD-NLYVVEIDPQKAMEEYKEKLK 108
>gb|AAB85986.1| arsenical pump-driving ATPase [Methanothermobacter
thermautotrophicus str. Delta H]
gi|15679508|ref|NP_276625.1| arsenical pump-driving
ATPase [Methanothermobacter thermautotrophicus str.
Delta H] gi|7428325|pir||F69068 probable arsenical
pump-driving ATPase (EC 3.6.1.-) - Methanobacterium
thermoautotrophicum (strain Delta H)
gi|6647437|sp|O27555|ARSA_METTH Putative arsenical
pump-driving ATPase (Arsenite-translocating ATPase)
(Arsenical resistance ATPase) (Arsenite-transporting
ATPase)
Length = 324
Score = 73.9 bits (180), Expect = 7e-12
Identities = 43/106 (40%), Positives = 65/106 (60%), Gaps = 10/106 (9%)
Query: 34 YYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSDSFAQDLTGGALVQVDGP 93
+ +GGKGGVGKT+ +A+ A+ A +G TLV+STDPAHSLSDS +++ G ++
Sbjct: 16 FVFIGGKGGVGKTTISAATALWMARSGKKTLVISTDPAHSLSDSLEREI-GHTPTKI--- 71
Query: 94 DYPLFALEINPEKAREDFRDVAKQNGGSTGVKDFMDGMGLGMIVDQ 139
L+A+EI+PE A E+++ ++ GMGL M+ DQ
Sbjct: 72 TENLYAVEIDPEVAMEEYQAKLQEQAAMN------PGMGLDMLQDQ 111
>emb|CAF29719.1| Putative arsenical pump-driving ATPase [Methanococcus maripaludis
S2] gi|45357726|ref|NP_987283.1| Putative arsenical
pump-driving ATPase [Methanococcus maripaludis S2]
Length = 345
Score = 73.6 bits (179), Expect = 9e-12
Identities = 40/84 (47%), Positives = 55/84 (64%), Gaps = 2/84 (2%)
Query: 33 KYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSDSFAQDLTGGALVQVDG 92
KY M GGKGGVGKT+ +A+ + A G T+VVSTDPAHSL DSF Q G +V+G
Sbjct: 25 KYIMFGGKGGVGKTTMSAATGIYCAEQGLKTVVVSTDPAHSLKDSFEQSF-GHEPTKVNG 83
Query: 93 PDYPLFALEINPEKAREDFRDVAK 116
+ L+ +EI+PE A ++++ K
Sbjct: 84 ME-NLYVVEIDPEVAMSEYKEKLK 106
>gb|AAV45680.1| arsenical pump-driving ATPase [Haloarcula marismortui ATCC 43049]
gi|55377536|ref|YP_135386.1| arsenical pump-driving
ATPase [Haloarcula marismortui ATCC 43049]
Length = 362
Score = 72.8 bits (177), Expect = 2e-11
Identities = 40/75 (53%), Positives = 50/75 (66%), Gaps = 2/75 (2%)
Query: 33 KYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSDSFAQDLTGGALVQVDG 92
+Y + GGKGGVGKT+CAA+ A+ A + TLVVSTDPAHSLSD+ D+ A
Sbjct: 22 EYVLYGGKGGVGKTTCAAATALASARDDTATLVVSTDPAHSLSDTLDADIP--ATPTRIR 79
Query: 93 PDYPLFALEINPEKA 107
D PL+A EI+PE A
Sbjct: 80 EDIPLYAAEIDPEAA 94
>ref|NP_279449.1| ArsA1 [Halobacterium sp. NRC-1] gi|10579983|gb|AAG18929.1|
arsenical pump-driving ATPase; ArsA1 [Halobacterium sp.
NRC-1] gi|25290844|pir||E84195 arsenical pump-driving
ATPase [imported] - Halobacterium sp. NRC-1
Length = 347
Score = 72.4 bits (176), Expect = 2e-11
Identities = 46/117 (39%), Positives = 68/117 (57%), Gaps = 3/117 (2%)
Query: 18 ESISVFDDMVAGTERKYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSDS 77
E++ D + +Y + GGKGGVGKT+ AA+ AV A++G TLVVSTDPAHSLSD+
Sbjct: 7 EAVDSLDPGITVDTAEYVLYGGKGGVGKTTMAAATAVASASDGTNTLVVSTDPAHSLSDT 66
Query: 78 FAQDLTGGALVQVDGPDYPLFALEINPEKAREDFRDVAKQNGGSTGVKDFMDGMGLG 134
++ + + PL+A+EI+P+ A D + Q+G +G + M G G
Sbjct: 67 LEAEIP--SRPHRIRENVPLWAVEIDPDDAL-DRTGMFGQDGALSGTLETMLGGDAG 120
>gb|EAL18069.1| hypothetical protein CNBK0900 [Cryptococcus neoformans var.
neoformans B-3501A] gi|57229944|gb|AAW46346.1| conserved
hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21] gi|58260906|ref|XP_567863.1| conserved
hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21]
Length = 325
Score = 72.0 bits (175), Expect = 3e-11
Identities = 40/92 (43%), Positives = 60/92 (64%), Gaps = 2/92 (2%)
Query: 33 KYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSDSFAQDLTGGALVQVDG 92
K+ GGKGGVGKT+ + SLAV+ A L++STDPAH+LSD+F+Q G +V+G
Sbjct: 21 KWIFCGGKGGVGKTTTSCSLAVQLAACRESVLLISTDPAHNLSDAFSQKF-GKDATKVNG 79
Query: 93 PDYPLFALEINPEKAREDFRDVAKQNGGSTGV 124
D L+A+EI+P + ++ + + Q GG G+
Sbjct: 80 FD-NLYAMEIDPNGSLQEMIESSDQTGGMGGM 110
>ref|XP_315798.2| ENSANGP00000018739 [Anopheles gambiae str. PEST]
gi|55238584|gb|EAA11891.2| ENSANGP00000018739 [Anopheles
gambiae str. PEST]
Length = 336
Score = 71.2 bits (173), Expect = 4e-11
Identities = 50/137 (36%), Positives = 73/137 (52%), Gaps = 11/137 (8%)
Query: 33 KYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSDSFAQDLTGGALVQVDG 92
K+ +GGKGGVGKT+C+ SLA++ A L++STDPAH++SD+F Q T +V+G
Sbjct: 22 KWVFVGGKGGVGKTTCSCSLAIQLAQKRESVLIISTDPAHNISDAFDQKFT-KVPTKVNG 80
Query: 93 PDYPLFALEINP-----EKAREDFRDVAKQNGGSTG----VKDFMDGMGLGMIVDQTAKV 143
D LFA+EI+P E E F D A G V + G+ M + K+
Sbjct: 81 FD-NLFAMEIDPNVGISELPDEYFEDEASPLNVGKGMLQEVIGTLPGIDEAMSYAEVMKL 139
Query: 144 RRAKTGRVTGYTSSWTG 160
+A V + ++ TG
Sbjct: 140 VKAMNFSVVVFDTAPTG 156
>gb|AAS52205.1| ADR285Wp [Ashbya gossypii ATCC 10895] gi|45188158|ref|NP_984381.1|
ADR285Wp [Eremothecium gossypii]
Length = 349
Score = 71.2 bits (173), Expect = 4e-11
Identities = 47/122 (38%), Positives = 70/122 (56%), Gaps = 10/122 (8%)
Query: 26 MVAGTERKYYMLGGKGGVGKTSCAASLAVKFANNGHPT---LVVSTDPAHSLSDSFAQDL 82
++ T K+ +GGKGGVGKT+ + S+A++ A PT L++STDPAH+LSD+F +
Sbjct: 13 LINSTTHKWIFVGGKGGVGKTTSSCSIAIQMA-LAQPTKQFLLISTDPAHNLSDAFNEKF 71
Query: 83 TGGALVQVDGPDYPLFALEINPEKAREDFRDVAKQNGG-STGVKDFMDGMGLGMIVDQTA 141
G +V G D L +EI+P A +D D+A NGG G+ + G G + D T
Sbjct: 72 -GKDARKVTGMD-NLSCMEIDPSAALKDVNDMAIANGGDDDGLSGLLQG---GALADLTG 126
Query: 142 KV 143
+
Sbjct: 127 SI 128
>ref|XP_556439.1| ENSANGP00000029267 [Anopheles gambiae str. PEST]
gi|55238585|gb|EAL39917.1| ENSANGP00000029267 [Anopheles
gambiae str. PEST]
Length = 297
Score = 71.2 bits (173), Expect = 4e-11
Identities = 50/137 (36%), Positives = 73/137 (52%), Gaps = 11/137 (8%)
Query: 33 KYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSDSFAQDLTGGALVQVDG 92
K+ +GGKGGVGKT+C+ SLA++ A L++STDPAH++SD+F Q T +V+G
Sbjct: 22 KWVFVGGKGGVGKTTCSCSLAIQLAQKRESVLIISTDPAHNISDAFDQKFT-KVPTKVNG 80
Query: 93 PDYPLFALEINP-----EKAREDFRDVAKQNGGSTG----VKDFMDGMGLGMIVDQTAKV 143
D LFA+EI+P E E F D A G V + G+ M + K+
Sbjct: 81 FD-NLFAMEIDPNVGISELPDEYFEDEASPLNVGKGMLQEVIGTLPGIDEAMSYAEVMKL 139
Query: 144 RRAKTGRVTGYTSSWTG 160
+A V + ++ TG
Sbjct: 140 VKAMNFSVVVFDTAPTG 156
>gb|EAL25973.1| GA14038-PA [Drosophila pseudoobscura]
Length = 336
Score = 70.5 bits (171), Expect = 7e-11
Identities = 38/80 (47%), Positives = 55/80 (68%), Gaps = 2/80 (2%)
Query: 25 DMVAGTERKYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSDSFAQDLTG 84
++V K+ +GGKGGVGKT+C++SLAV+ A L++STDPAH++SD+F Q T
Sbjct: 14 NLVEQDSLKWIFVGGKGGVGKTTCSSSLAVQLAKVRESVLIISTDPAHNISDAFDQKFT- 72
Query: 85 GALVQVDGPDYPLFALEINP 104
+V+G D LFA+EI+P
Sbjct: 73 KVPTKVNGFD-NLFAMEIDP 91
>emb|CAG62065.1| unnamed protein product [Candida glabrata CBS138]
gi|50293367|ref|XP_449095.1| unnamed protein product
[Candida glabrata]
Length = 350
Score = 70.1 bits (170), Expect = 1e-10
Identities = 42/120 (35%), Positives = 65/120 (54%), Gaps = 4/120 (3%)
Query: 26 MVAGTERKYYMLGGKGGVGKT--SCAASLAVKFANNGHPTLVVSTDPAHSLSDSFAQDLT 83
++ + K+ +GGKGGVGKT SC+ ++ + A L++STDPAH+LSD+F +
Sbjct: 12 LITSSTHKWIFVGGKGGVGKTTSSCSIAIQMALAQPHKKFLLISTDPAHNLSDAFGEKFD 71
Query: 84 GGALVQVDGPDYPLFALEINPEKAREDFRDVAKQNGGSTGVKDFMDGMGLGMIVDQTAKV 143
A +V G D L +EI+P A +D D+A + G G D + G + D T +
Sbjct: 72 KDAR-KVTGMD-NLSCMEIDPSAALKDMNDMAVSSAGENGNDDLGGLLQGGALADLTGSI 129
>gb|AAC50731.1| hASNA-I [Homo sapiens]
Length = 332
Score = 70.1 bits (170), Expect = 1e-10
Identities = 47/122 (38%), Positives = 67/122 (54%), Gaps = 7/122 (5%)
Query: 33 KYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSDSFAQDLTGGALVQVDG 92
K+ +GGKGGVGKT+C+ SLAV+ + L++STDPAH++SD+F Q + +V G
Sbjct: 22 KWIFVGGKGGVGKTTCSCSLAVQLSKGRESVLIISTDPAHNISDAFDQKFS-KVPTKVKG 80
Query: 93 PDYPLFALEINPEKAREDFRD--VAKQNGGSTGVKDFMDGMGLGMIVDQT---AKVRRAK 147
D LFA+EI+P D D + N S G K + M +D+ A+V R
Sbjct: 81 YD-NLFAMEIDPSLGVADVPDEFFEEDNMLSMGKKMMQEAMSAFPGIDEAMSYAEVMRLV 139
Query: 148 TG 149
G
Sbjct: 140 KG 141
>gb|EAK84682.1| hypothetical protein UM03838.1 [Ustilago maydis 521]
gi|49074670|ref|XP_401453.1| hypothetical protein
UM03838.1 [Ustilago maydis 521]
Length = 332
Score = 69.3 bits (168), Expect = 2e-10
Identities = 37/91 (40%), Positives = 58/91 (63%), Gaps = 2/91 (2%)
Query: 33 KYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSDSFAQDLTGGALVQVDG 92
K+ +GGKGGVGKT+ + SLA++ + L++STDPAH+LSD+F Q G +V+G
Sbjct: 25 KWLFVGGKGGVGKTTTSCSLAIQLSKVRESVLLISTDPAHNLSDAFGQKF-GKEATKVNG 83
Query: 93 PDYPLFALEINPEKAREDFRDVAKQNGGSTG 123
D L A+EI+P + ++ + + GG+ G
Sbjct: 84 FD-NLSAMEIDPNSSIQEMIEQSDSQGGAMG 113
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.347 0.152 0.531
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 545,323,484
Number of Sequences: 2540612
Number of extensions: 20911490
Number of successful extensions: 87637
Number of sequences better than 10.0: 643
Number of HSP's better than 10.0 without gapping: 563
Number of HSP's successfully gapped in prelim test: 80
Number of HSP's that attempted gapping in prelim test: 86968
Number of HSP's gapped (non-prelim): 682
length of query: 359
length of database: 863,360,394
effective HSP length: 129
effective length of query: 230
effective length of database: 535,621,446
effective search space: 123192932580
effective search space used: 123192932580
T: 11
A: 40
X1: 14 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.7 bits)
S2: 76 (33.9 bits)
Medicago: description of AC144928.8


