

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144405.9 - phase: 0
(166 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
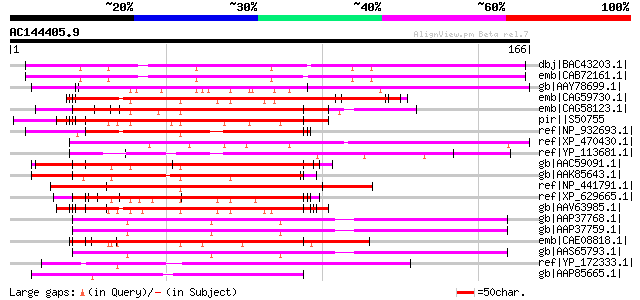
Score E
Sequences producing significant alignments: (bits) Value
dbj|BAC43203.1| putative serine/proline-rich protein [Arabidopsi... 115 4e-25
emb|CAB72161.1| serine/proline-rich [Arabidopsis thaliana] gi|15... 115 6e-25
gb|AAY78699.1| proline-rich family protein [Arabidopsis thaliana... 99 4e-20
emb|CAG59730.1| unnamed protein product [Candida glabrata CBS138... 94 1e-18
emb|CAG58123.1| unnamed protein product [Candida glabrata CBS138... 91 1e-17
pir||S50755 hypothetical protein VSP-3 - Chlamydomonas reinhardt... 91 2e-17
ref|NP_932693.1| unknown [Choristoneura fumiferana defective nuc... 89 3e-17
ref|XP_470430.1| unknown protein [Oryza sativa (japonica cultiva... 89 3e-17
ref|YP_113681.1| metalloprotease, putative [Methylococcus capsul... 88 1e-16
gb|AAC59091.1| unknown [Orgyia pseudotsugata multicapsid nucleop... 88 1e-16
gb|AAK85643.1| unknown [Epiphyas postvittana nucleopolyhedroviru... 86 3e-16
ref|NP_441791.1| hypothetical protein slr0442 [Synechocystis sp.... 85 8e-16
ref|XP_629665.1| hypothetical protein DDB0184442 [Dictyostelium ... 84 1e-15
gb|AAV63985.1| hydroxyproline-rich glycoprotein VSP-3 [Chlamydom... 84 2e-15
gb|AAP37768.1| At3g24600 [Arabidopsis thaliana] gi|13877617|gb|A... 83 3e-15
gb|AAP37759.1| At3g24550 [Arabidopsis thaliana] gi|22136228|gb|A... 83 3e-15
emb|CAE08818.1| conserved hypothetical protein [Synechococcus sp... 83 3e-15
gb|AAS65793.1| putative protein kinase [Arabidopsis thaliana] 83 3e-15
ref|YP_172333.1| hypothetical protein syc1623_d [Synechococcus e... 81 9e-15
gb|AAP85665.1| odv-e66 [Adoxophyes orana granulovirus] gi|326985... 80 2e-14
>dbj|BAC43203.1| putative serine/proline-rich protein [Arabidopsis thaliana]
gi|28372906|gb|AAO39935.1| At3g45230 [Arabidopsis
thaliana]
Length = 175
Score = 115 bits (288), Expect = 4e-25
Identities = 80/175 (45%), Positives = 105/175 (59%), Gaps = 19/175 (10%)
Query: 6 FSLVFLLLSFLVNIAS--SADSPAPTP-ATNSSLNSPSPTPIPTPSPANSPPAPTP---T 59
F +V ++LS ++ + SPAP+P +S L SP P+ +++ PA +P +
Sbjct: 5 FIIVAMMLSLVLVSGEILTKSSPAPSPDLADSPLIHASP---PSKLGSHNSPAESPIEYS 61
Query: 60 PTPSPHSDSPPAPSPDNSPSSSP-------SPSPSSSPAPSPDEAADNNAISHTGI-GED 111
P P ++ P+PSP NSPS SP SPS S+SP+PSPD A+D N TGI GE
Sbjct: 62 SPPEPETEHSPSPSPANSPSVSPPLPNDSQSPSSSASPSPSPD-ASDVNHSDITGIEGEK 120
Query: 112 GKS-SGGGMSSGKKAGIAVGVIAAVGVVALGAMVVKKRRQNIQRSEYGYTARREL 165
S SGGGMS GKK G+A G IAAV VV + V KKR++NI+RS YGY AR L
Sbjct: 121 LPSGSGGGMSGGKKVGVAFGAIAAVCVVGVAGFVYKKRQENIRRSRYGYAAREIL 175
>emb|CAB72161.1| serine/proline-rich [Arabidopsis thaliana]
gi|15230640|ref|NP_190109.1| hydroxyproline-rich
glycoprotein family protein [Arabidopsis thaliana]
gi|11358836|pir||T47463 serine/proline-rich protein -
Arabidopsis thaliana
Length = 175
Score = 115 bits (287), Expect = 6e-25
Identities = 80/175 (45%), Positives = 105/175 (59%), Gaps = 19/175 (10%)
Query: 6 FSLVFLLLSFLVNIAS--SADSPAPTP-ATNSSLNSPSPTPIPTPSPANSPPAPTP---T 59
F +V ++LS ++ + SPAP+P +S L SP P+ +++ PA +P +
Sbjct: 5 FIIVAMMLSLVLVSGEILTKSSPAPSPDLADSPLIHASP---PSKLGSHNSPAESPIEYS 61
Query: 60 PTPSPHSDSPPAPSPDNSPSSSP-------SPSPSSSPAPSPDEAADNNAISHTGI-GED 111
P P ++ P+PSP NSPS SP SPS S+SP+PSP EA+D N TGI GE
Sbjct: 62 SPPEPETEHSPSPSPANSPSVSPPLPNDSQSPSSSASPSPSP-EASDVNHSDITGIEGEK 120
Query: 112 GKS-SGGGMSSGKKAGIAVGVIAAVGVVALGAMVVKKRRQNIQRSEYGYTARREL 165
S SGGGMS GKK G+A G IAAV VV + V KKR++NI+RS YGY AR L
Sbjct: 121 LPSGSGGGMSGGKKVGVAFGAIAAVCVVGVAGFVYKKRQENIRRSRYGYAAREIL 175
>gb|AAY78699.1| proline-rich family protein [Arabidopsis thaliana]
gi|4432833|gb|AAD20682.1| En/Spm-like transposon protein
[Arabidopsis thaliana] gi|25407987|pir||H84684
En/Spm-like transposon protein [imported] - Arabidopsis
thaliana gi|15226398|ref|NP_180411.1| proline-rich
family protein [Arabidopsis thaliana]
Length = 268
Score = 99.0 bits (245), Expect = 4e-20
Identities = 67/157 (42%), Positives = 85/157 (53%), Gaps = 14/157 (8%)
Query: 23 ADSPAPTPATNSSLNSP-SPTPIPTP-SPANSPPAPTPTPTP-SPHSDSPPAPSPDNSPS 79
ADSP P P+++ NSP SP P P S A+SP P P P P SP S S P P+P +PS
Sbjct: 113 ADSPLP-PSSSPEANSPQSPASSPKPESLADSPSPPPPPPQPESPSSPSYPEPAPVPAPS 171
Query: 80 SSPS---PSPSS----SPAPSPDEAADNNAISHTGIGEDGKSSGG---GMSSGKKAGIAV 129
S P P + SPAPSP+ + + GE+ G GMS +KAGIA+
Sbjct: 172 DDDSDDDPEPETEYFPSPAPSPELGMAQDIKASDAAGEELNDERGEDYGMSGLEKAGIAI 231
Query: 130 GVIAAVGVVALGAMVVKKRRQNIQRSEYGYTARRELL 166
G I VG + +GA+V KKRR N+ R+ Y Y E L
Sbjct: 232 GTILGVGAIVIGALVYKKRRDNMTRARYTYFTEGEFL 268
Score = 47.0 bits (110), Expect = 2e-04
Identities = 34/95 (35%), Positives = 51/95 (52%), Gaps = 9/95 (9%)
Query: 8 LVFLLLSFLVNIASSADSPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTP-TPTPSPHS 66
+V L + L+ + A +P+PA S PS T P SP++SP +P +P+ SP
Sbjct: 8 IVMLSICLLIFDFAGAQEESPSPAAVSPGREPS-TDSPL-SPSSSPEEDSPLSPSSSPEE 65
Query: 67 DSP--PAPSPDN----SPSSSPSPSPSSSPAPSPD 95
DSP P+ SP+ +PSSSP +P+ SP+
Sbjct: 66 DSPLPPSSSPEEDSPLAPSSSPEVDSPLAPSSSPE 100
Score = 45.4 bits (106), Expect = 6e-04
Identities = 32/82 (39%), Positives = 45/82 (54%), Gaps = 8/82 (9%)
Query: 22 SADSPAPTPATNSSLN-SPSPTPIPTPSPANSPPAPTPTP-TPSPHSDSP--PAPSPD-- 75
+A SP P+T+S L+ S SP SP++SP +P P + SP DSP P+ SP+
Sbjct: 31 AAVSPGREPSTDSPLSPSSSPEEDSPLSPSSSPEEDSPLPPSSSPEEDSPLAPSSSPEVD 90
Query: 76 --NSPSSSPSPSPSSSPAPSPD 95
+PSSSP P+ SP+
Sbjct: 91 SPLAPSSSPEVDSPQPPSSSPE 112
Score = 38.1 bits (87), Expect = 0.091
Identities = 24/60 (40%), Positives = 30/60 (50%), Gaps = 10/60 (16%)
Query: 38 SPSPTPIPTPSPANSPPAPTP-TPTPSPHSDSPPAPSPDNSPSSSPSPSPSSSPAPSPDE 96
SPSP + SP P +P +P+ SP DSP SPSSSP P+ SP+E
Sbjct: 27 SPSPAAV---SPGREPSTDSPLSPSSSPEEDSPL------SPSSSPEEDSPLPPSSSPEE 77
>emb|CAG59730.1| unnamed protein product [Candida glabrata CBS138]
gi|50288747|ref|XP_446803.1| unnamed protein product
[Candida glabrata]
Length = 577
Score = 94.0 bits (232), Expect = 1e-18
Identities = 46/101 (45%), Positives = 65/101 (63%), Gaps = 1/101 (0%)
Query: 21 SSADSPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSS 80
S + SP+P+P+ + S SPSP+P P+PSP++ P+P+P+P+PSP P+PSP SPS
Sbjct: 121 SPSPSPSPSPSPSPS-PSPSPSPSPSPSPSSPSPSPSPSPSPSPSPSPSPSPSPSPSPSP 179
Query: 81 SPSPSPSSSPAPSPDEAADNNAISHTGIGEDGKSSGGGMSS 121
SPSPSPS SP+PSP + + + S + SS SS
Sbjct: 180 SPSPSPSPSPSPSPSPKSPSPSPSSSSSSSSMPSSSSSSSS 220
Score = 93.2 bits (230), Expect = 2e-18
Identities = 47/107 (43%), Positives = 66/107 (60%), Gaps = 1/107 (0%)
Query: 19 IASSADSPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSP 78
I ++ SP+P+P+ + S SPSP+P P+PSP+ SP +P+P+P+PSP P+PSP SP
Sbjct: 115 IVTAWPSPSPSPSPSPS-PSPSPSPSPSPSPSPSPSSPSPSPSPSPSPSPSPSPSPSPSP 173
Query: 79 SSSPSPSPSSSPAPSPDEAADNNAISHTGIGEDGKSSGGGMSSGKKA 125
S SPSPSPS SP+PSP + + S + SS SS +
Sbjct: 174 SPSPSPSPSPSPSPSPSPSPSPKSPSPSPSSSSSSSSMPSSSSSSSS 220
Score = 89.4 bits (220), Expect = 3e-17
Identities = 43/101 (42%), Positives = 65/101 (63%), Gaps = 4/101 (3%)
Query: 21 SSADSPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSS 80
S + SP+P+P+ + S +SPSP+P P+PSP+ P+P+P+P+PSP P+PSP SPS
Sbjct: 133 SPSPSPSPSPSPSPSPSSPSPSPSPSPSPS---PSPSPSPSPSPSPSPSPSPSPSPSPSP 189
Query: 81 SPSPSPSSSPAPSPDEAADNNAISHTGIGEDGKSSGGGMSS 121
SPSPSP SP+PSP ++ ++++ + S SS
Sbjct: 190 SPSPSP-KSPSPSPSSSSSSSSMPSSSSSSSSMPSSSSSSS 229
Score = 85.1 bits (209), Expect = 6e-16
Identities = 43/103 (41%), Positives = 64/103 (61%), Gaps = 2/103 (1%)
Query: 21 SSADSPAPTPATNSSLNSPSPTPIPTPSPANSP-PAPTPTPTPSPHSDSPPAPSPDNSPS 79
S + SP+P+P+ +S SPSP+P P+PSP+ SP P+P+P+P+PSP P+PSP SP
Sbjct: 137 SPSPSPSPSPSPSSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPK 196
Query: 80 S-SPSPSPSSSPAPSPDEAADNNAISHTGIGEDGKSSGGGMSS 121
S SPSPS SSS + P ++ ++++ + S SS
Sbjct: 197 SPSPSPSSSSSSSSMPSSSSSSSSMPSSSSSSSSMPSSSSSSS 239
Score = 77.0 bits (188), Expect = 2e-13
Identities = 43/101 (42%), Positives = 59/101 (57%), Gaps = 2/101 (1%)
Query: 21 SSADSPAPTPATNSSLNSPSPTPIPTPSPANSP-PAPTPTPTPSPHSDSPPAPSPDNSPS 79
S + SP+P+ + S SPSP+P P+PSP+ SP P+P+P+P+PSP P+PSP SPS
Sbjct: 141 SPSPSPSPSSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSP-KSPS 199
Query: 80 SSPSPSPSSSPAPSPDEAADNNAISHTGIGEDGKSSGGGMS 120
SPS S SSS PS ++ + S + SS S
Sbjct: 200 PSPSSSSSSSSMPSSSSSSSSMPSSSSSSSSMPSSSSSSSS 240
Score = 75.1 bits (183), Expect = 7e-13
Identities = 43/106 (40%), Positives = 64/106 (59%), Gaps = 9/106 (8%)
Query: 22 SADSPAPTPATNSSLNSPSPTPIPTPSPANSP-PAPTPTPTPSPH-SDSP----PAPSPD 75
S SP+P+P+ + S PSP+P P+PSP+ SP P+P+P+P+PSP S SP P+PSP
Sbjct: 147 SPSSPSPSPSPSPS---PSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPKSPSPSPS 203
Query: 76 NSPSSSPSPSPSSSPAPSPDEAADNNAISHTGIGEDGKSSGGGMSS 121
+S SSS PS SSS + P ++ ++++ + S SS
Sbjct: 204 SSSSSSSMPSSSSSSSSMPSSSSSSSSMPSSSSSSSSMPSSSSSSS 249
Score = 73.9 bits (180), Expect = 1e-12
Identities = 43/107 (40%), Positives = 62/107 (57%), Gaps = 2/107 (1%)
Query: 21 SSADSPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSS 80
S + SP+P+P+ + S SPSP+P P+PSP+ SP +P+P+PS S S PS +S SS
Sbjct: 162 SPSPSPSPSPSPSPS-PSPSPSPSPSPSPSPSPSPKSPSPSPSSSSSSSSMPSSSSSSSS 220
Query: 81 SPSPSPSSSPAPSPDEAADNNAISHTGIGEDGKSSGGGMSSGKKAGI 127
PS S SSS PS ++ ++ S + SS M+ +KA I
Sbjct: 221 MPSSSSSSSSMPS-SSSSSSSMPSSSSSSSSMPSSSSSMTPSQKASI 266
Score = 53.9 bits (128), Expect = 2e-06
Identities = 41/134 (30%), Positives = 61/134 (44%), Gaps = 49/134 (36%)
Query: 22 SADSPAPTPATNSSLN-SPSPTPIPTPSPANSP-PAPTPTPTPSPHSDSP-PAPSPDNSP 78
S+ SP+P+P+ + S + SPSP+P P+PSP+ SP P+P+P+P+PSP SP P+PS +S
Sbjct: 149 SSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPKSPSPSPSSSSSS 208
Query: 79 SSSPS----------------------------------------------PSPSSSPAP 92
SS PS PS +S P
Sbjct: 209 SSMPSSSSSSSSMPSSSSSSSSMPSSSSSSSSMPSSSSSSSSMPSSSSSMTPSQKASIIP 268
Query: 93 SPDEAADNNAISHT 106
S + +++I T
Sbjct: 269 SSAAPSSSSSIVTT 282
Score = 49.3 bits (116), Expect = 4e-05
Identities = 38/138 (27%), Positives = 57/138 (40%), Gaps = 53/138 (38%)
Query: 20 ASSADSPAPTPATNSSLN-SPSPTPIPTPSPANSP-PAPTPTPTPSPHSDSP-------- 69
+S + SP+P+P+ + S + SPSP+P P+PSP+ SP P+P+P+P+PSP S SP
Sbjct: 149 SSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPKSPSPSPSSSSSS 208
Query: 70 -------------------------------------------PAPSPDNSPSSSPSPSP 86
P+ S +PS S P
Sbjct: 209 SSMPSSSSSSSSMPSSSSSSSSMPSSSSSSSSMPSSSSSSSSMPSSSSSMTPSQKASIIP 268
Query: 87 SSSPAPSPDEAADNNAIS 104
SS+ S ++IS
Sbjct: 269 SSAAPSSSSSIVTTSSIS 286
Score = 40.8 bits (94), Expect = 0.014
Identities = 36/131 (27%), Positives = 52/131 (39%), Gaps = 49/131 (37%)
Query: 21 SSADSPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPS----------------- 63
S + SP+P+P+ + S SPSP+P P+PSP+ SP +P+P+P+ S
Sbjct: 164 SPSPSPSPSPSPSPS-PSPSPSPSPSPSPSPSPKSPSPSPSSSSSSSSMPSSSSSSSSMP 222
Query: 64 -----------------------------PHSDSPPAPSPDNS--PSSSPSPSPSSSPAP 92
P S S PS S PSS+ S SS
Sbjct: 223 SSSSSSSSMPSSSSSSSSMPSSSSSSSSMPSSSSSMTPSQKASIIPSSAAPSSSSSIVTT 282
Query: 93 SPDEAADNNAI 103
S +AD + +
Sbjct: 283 SSISSADASPV 293
>emb|CAG58123.1| unnamed protein product [Candida glabrata CBS138]
gi|50285581|ref|XP_445219.1| unnamed protein product
[Candida glabrata]
Length = 790
Score = 90.9 bits (224), Expect = 1e-17
Identities = 44/76 (57%), Positives = 58/76 (75%), Gaps = 2/76 (2%)
Query: 21 SSADSPAPTPATNSSLN-SPSPTPIPTPSPANSP-PAPTPTPTPSPHSDSPPAPSPDNSP 78
SS+ SP+P+P + S + SPSP+P P+PSP+ SP P+P+P+P+PSP P+PSP SP
Sbjct: 331 SSSPSPSPSPKPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPKPSP 390
Query: 79 SSSPSPSPSSSPAPSP 94
S SPSPSPS SP+PSP
Sbjct: 391 SPSPSPSPSPSPSPSP 406
Score = 89.4 bits (220), Expect = 3e-17
Identities = 43/75 (57%), Positives = 56/75 (74%), Gaps = 2/75 (2%)
Query: 21 SSADSPAPTPATNSSLNSPSPTPIPTPSPANSP-PAPTPTPTPSPHSDSPPAPSPDNSPS 79
S SP+P+P+ + S SPSP+P P+PSP+ SP P+P+P+P+PSP P+PSP SPS
Sbjct: 339 SPKPSPSPSPSPSPS-PSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPKPSPSPSPSPS 397
Query: 80 SSPSPSPSSSPAPSP 94
SPSPSPS SP+PSP
Sbjct: 398 PSPSPSPSPSPSPSP 412
Score = 89.0 bits (219), Expect = 4e-17
Identities = 43/80 (53%), Positives = 57/80 (70%), Gaps = 6/80 (7%)
Query: 21 SSADSPAPTPATNSSLN-----SPSPTPIPTPSPANSP-PAPTPTPTPSPHSDSPPAPSP 74
S + SP+P+P+ + S + SPSP P P+PSP+ SP P+P+P+P+PSP P+PSP
Sbjct: 399 SPSPSPSPSPSPSPSFSPGPKPSPSPKPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSP 458
Query: 75 DNSPSSSPSPSPSSSPAPSP 94
SPS SPSPSPS SP+PSP
Sbjct: 459 SPSPSPSPSPSPSPSPSPSP 478
Score = 87.0 bits (214), Expect = 2e-16
Identities = 42/75 (56%), Positives = 54/75 (72%), Gaps = 2/75 (2%)
Query: 21 SSADSPAPTPATNSSLNSPSPTPIPTPSPANSP-PAPTPTPTPSPHSDSPPAPSPDNSPS 79
S + SP+P+P+ + S SPSP+P P+PSP+ SP P+P+P+P PSP P+PSP SPS
Sbjct: 347 SPSPSPSPSPSPSPS-PSPSPSPSPSPSPSPSPSPSPSPSPKPSPSPSPSPSPSPSPSPS 405
Query: 80 SSPSPSPSSSPAPSP 94
SPSPSPS SP P P
Sbjct: 406 PSPSPSPSFSPGPKP 420
Score = 87.0 bits (214), Expect = 2e-16
Identities = 42/75 (56%), Positives = 55/75 (73%), Gaps = 2/75 (2%)
Query: 21 SSADSPAPTPATNSSLNSPSPTPIPTPSPANSP-PAPTPTPTPSPHSDSPPAPSPDNSPS 79
S + SP+P+P+ + S SPSP+P P+PSP+ SP P+P+P+P+PSP P PSP PS
Sbjct: 437 SPSPSPSPSPSPSPS-PSPSPSPSPSPSPSPSPSPSPSPSPSPSPSFSPGPKPSPSPKPS 495
Query: 80 SSPSPSPSSSPAPSP 94
SPSPSPS SP+PSP
Sbjct: 496 PSPSPSPSPSPSPSP 510
Score = 86.3 bits (212), Expect = 3e-16
Identities = 43/85 (50%), Positives = 57/85 (66%), Gaps = 4/85 (4%)
Query: 21 SSADSPAPTPATNSSLNSPSPTPIPTPSPANSP---PAPTPTPTPSPHSDSPPAPSPDNS 77
S + SP+P+P+ + S SPSP+P P+PSP+ SP P P P+P+P P P+PSP S
Sbjct: 449 SPSPSPSPSPSPSPS-PSPSPSPSPSPSPSPSPSFSPGPKPSPSPKPSPSPSPSPSPSPS 507
Query: 78 PSSSPSPSPSSSPAPSPDEAADNNA 102
PS SPSPSPS SP PSP+ +N+
Sbjct: 508 PSPSPSPSPSPSPYPSPNPFPISNS 532
Score = 85.9 bits (211), Expect = 4e-16
Identities = 41/75 (54%), Positives = 54/75 (71%), Gaps = 2/75 (2%)
Query: 21 SSADSPAPTPATNSSLNSPSPTPIPTPSPANSP-PAPTPTPTPSPHSDSPPAPSPDNSPS 79
S + SP+P+P+ + S SPSP+P P+PSP+ SP P+P+P+P+PSP P+PSP SPS
Sbjct: 355 SPSPSPSPSPSPSPS-PSPSPSPSPSPSPSPSPKPSPSPSPSPSPSPSPSPSPSPSPSPS 413
Query: 80 SSPSPSPSSSPAPSP 94
SP P PS SP PSP
Sbjct: 414 FSPGPKPSPSPKPSP 428
Score = 85.1 bits (209), Expect = 6e-16
Identities = 41/75 (54%), Positives = 52/75 (68%), Gaps = 2/75 (2%)
Query: 21 SSADSPAPTPATNSSLNSPSPTPIPTPSPANSP-PAPTPTPTPSPHSDSPPAPSPDNSPS 79
S P+P+P + S SPSP+P P+PSP+ SP P+P+P+P+PSP P+PSP SPS
Sbjct: 415 SPGPKPSPSPKPSPS-PSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPS 473
Query: 80 SSPSPSPSSSPAPSP 94
SPSPSPS SP P P
Sbjct: 474 PSPSPSPSFSPGPKP 488
Score = 84.3 bits (207), Expect = 1e-15
Identities = 40/75 (53%), Positives = 53/75 (70%), Gaps = 2/75 (2%)
Query: 21 SSADSPAPTPATNSSLNSPSPTPIPTPSPANSP-PAPTPTPTPSPHSDSPPAPSPDNSPS 79
S + SP+P+P+ + S SPSP+P P+PSP SP P+P+P+P+PSP P+PSP SP
Sbjct: 359 SPSPSPSPSPSPSPS-PSPSPSPSPSPSPKPSPSPSPSPSPSPSPSPSPSPSPSPSFSPG 417
Query: 80 SSPSPSPSSSPAPSP 94
PSPSP SP+PSP
Sbjct: 418 PKPSPSPKPSPSPSP 432
Score = 82.0 bits (201), Expect = 5e-15
Identities = 40/75 (53%), Positives = 54/75 (71%), Gaps = 2/75 (2%)
Query: 21 SSADSPAPTPATNSSLNSPSPTPIPTPSPANSP-PAPTPTPTPSPHSDSPPAPSPDNSPS 79
S + SP+P+P+ + S SPSP+P P+PSP+ SP P+P+P+P+PSP +P P SPS
Sbjct: 365 SPSPSPSPSPSPSPS-PSPSPSPKPSPSPSPSPSPSPSPSPSPSPSPSPSFSPGPKPSPS 423
Query: 80 SSPSPSPSSSPAPSP 94
PSPSPS SP+PSP
Sbjct: 424 PKPSPSPSPSPSPSP 438
Score = 81.6 bits (200), Expect = 7e-15
Identities = 40/75 (53%), Positives = 53/75 (70%), Gaps = 2/75 (2%)
Query: 21 SSADSPAPTPATNSSLNSPSPTPIPTPSPANSP-PAPTPTPTPSPHSDSPPAPSPDNSPS 79
S + SP+P+P+ + S SPSP+P P PSP+ SP P+P+P+P+PSP P+ SP PS
Sbjct: 363 SPSPSPSPSPSPSPS-PSPSPSPSPKPSPSPSPSPSPSPSPSPSPSPSPSPSFSPGPKPS 421
Query: 80 SSPSPSPSSSPAPSP 94
SP PSPS SP+PSP
Sbjct: 422 PSPKPSPSPSPSPSP 436
Score = 79.7 bits (195), Expect = 3e-14
Identities = 40/67 (59%), Positives = 49/67 (72%), Gaps = 7/67 (10%)
Query: 28 PTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPSPS 87
PTP SS SPSP+P P+PSP+ P+P+P+P+PSP P+PSP SPS SPSPSPS
Sbjct: 325 PTPPGLSSSPSPSPSPKPSPSPS---PSPSPSPSPSPS----PSPSPSPSPSPSPSPSPS 377
Query: 88 SSPAPSP 94
SP+PSP
Sbjct: 378 PSPSPSP 384
Score = 69.3 bits (168), Expect = 4e-11
Identities = 36/64 (56%), Positives = 47/64 (73%), Gaps = 6/64 (9%)
Query: 33 NSSLNSPSPTPIPT-PSPANSP-PAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPSPSSSP 90
NSS +P+P + + PSP+ SP P+P+P+P+PSP P+PSP SPS SPSPSPS SP
Sbjct: 319 NSSCYTPTPPGLSSSPSPSPSPKPSPSPSPSPSPS----PSPSPSPSPSPSPSPSPSPSP 374
Query: 91 APSP 94
+PSP
Sbjct: 375 SPSP 378
Score = 68.9 bits (167), Expect = 5e-11
Identities = 31/59 (52%), Positives = 39/59 (65%)
Query: 36 LNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPSPSSSPAPSP 94
+NS TP P ++ P+P+P P+PSP P+PSP SPS SPSPSPS SP+PSP
Sbjct: 318 INSSCYTPTPPGLSSSPSPSPSPKPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSP 376
Score = 68.6 bits (166), Expect = 6e-11
Identities = 45/124 (36%), Positives = 67/124 (53%), Gaps = 18/124 (14%)
Query: 21 SSADSPAPTPATNSSLNSPSPTPIPTPSPANSP-------------PAPTPTPTPSPHSD 67
S + SP+P+P+ + S SPSP+P P+PSP+ SP P+P+P+P+PSP
Sbjct: 453 SPSPSPSPSPSPSPS-PSPSPSPSPSPSPSFSPGPKPSPSPKPSPSPSPSPSPSPSPSPS 511
Query: 68 SPPAPSPDNSPSSSPSPSPSSSPAPSPDEAADNNAIS-HTGIGEDGKSSGGGMSSGKKAG 126
P+PSP PS +P P +SS + SP + ++ S +TG D +S S
Sbjct: 512 PSPSPSPSPYPSPNPFPISNSSTSLSPSNISMHSYSSLYTG---DPESPTNTKSIPTSID 568
Query: 127 IAVG 130
+AVG
Sbjct: 569 LAVG 572
Score = 67.4 bits (163), Expect = 1e-10
Identities = 33/86 (38%), Positives = 46/86 (53%)
Query: 9 VFLLLSFLVNIASSADSPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDS 68
V + S + +A+ TP+ P + TP+P +P+P+P+P P
Sbjct: 287 VLPITSTIYYVATPCADVPETPSCQIGFWDPINSSCYTPTPPGLSSSPSPSPSPKPSPSP 346
Query: 69 PPAPSPDNSPSSSPSPSPSSSPAPSP 94
P+PSP SPS SPSPSPS SP+PSP
Sbjct: 347 SPSPSPSPSPSPSPSPSPSPSPSPSP 372
>pir||S50755 hypothetical protein VSP-3 - Chlamydomonas reinhardtii
gi|530876|gb|AAB53953.1| amino acid feature: Rod protein
domain, aa 266 .. 468; amino acid feature: globular
protein domain, aa 32 .. 265
Length = 473
Score = 90.5 bits (223), Expect = 2e-17
Identities = 41/74 (55%), Positives = 54/74 (72%), Gaps = 1/74 (1%)
Query: 22 SADSPAPTPATN-SSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSS 80
++ SP+P+P+ +S SPSP+P P+PSP SPP P+P+P+PSP P+PSP SPS
Sbjct: 331 ASPSPSPSPSVQPASKPSPSPSPSPSPSPRPSPPLPSPSPSPSPSPSPSPSPSPKPSPSP 390
Query: 81 SPSPSPSSSPAPSP 94
SPSPSPS P+PSP
Sbjct: 391 SPSPSPSPKPSPSP 404
Score = 88.2 bits (217), Expect = 8e-17
Identities = 42/83 (50%), Positives = 55/83 (65%), Gaps = 1/83 (1%)
Query: 21 SSADSPAPTPATNSSLNSPSPTPIPTPSPANSP-PAPTPTPTPSPHSDSPPAPSPDNSPS 79
S + SP+P+P + L SPSP+P P+PSP+ SP P P+P+P+PSP P+PSP SPS
Sbjct: 350 SPSPSPSPSPRPSPPLPSPSPSPSPSPSPSPSPSPKPSPSPSPSPSPSPKPSPSPSPSPS 409
Query: 80 SSPSPSPSSSPAPSPDEAADNNA 102
SPSP S SP+PSP + A
Sbjct: 410 PSPSPKVSPSPSPSPSPSPSPKA 432
Score = 87.4 bits (215), Expect = 1e-16
Identities = 41/75 (54%), Positives = 52/75 (68%), Gaps = 1/75 (1%)
Query: 21 SSADSPAPTPATNSSLNSPSPTPIPTPSPANSP-PAPTPTPTPSPHSDSPPAPSPDNSPS 79
S + SP+P+P+ S PSP+P P+PSP+ SP P+P P+P+PSP P PSP SPS
Sbjct: 348 SPSPSPSPSPSPRPSPPLPSPSPSPSPSPSPSPSPSPKPSPSPSPSPSPSPKPSPSPSPS 407
Query: 80 SSPSPSPSSSPAPSP 94
SPSPSP SP+PSP
Sbjct: 408 PSPSPSPKVSPSPSP 422
Score = 85.5 bits (210), Expect = 5e-16
Identities = 43/76 (56%), Positives = 55/76 (71%), Gaps = 3/76 (3%)
Query: 20 ASSADSPAPTPATNSSLNSPSPTPIPTPSPANSP-PAPTPTPTPSPHSDSPPAPSPDNSP 78
AS + SP+P+P+ + SPSP+P P+ PA+ P P+P+P+P+PSP SPP PSP SP
Sbjct: 315 ASPSPSPSPSPSPSPKA-SPSPSPSPSVQPASKPSPSPSPSPSPSPR-PSPPLPSPSPSP 372
Query: 79 SSSPSPSPSSSPAPSP 94
S SPSPSPS SP PSP
Sbjct: 373 SPSPSPSPSPSPKPSP 388
Score = 81.6 bits (200), Expect = 7e-15
Identities = 39/72 (54%), Positives = 52/72 (72%), Gaps = 2/72 (2%)
Query: 25 SPAPTP-ATNSSLNSPSPTPIPTPSPANSP-PAPTPTPTPSPHSDSPPAPSPDNSPSSSP 82
SP+P+P A+ S SP +P P+PSP SP P+P+P+P+PSP + P+PSP P+S P
Sbjct: 288 SPSPSPKASPSPSPSPKASPSPSPSPKASPSPSPSPSPSPSPKASPSPSPSPSVQPASKP 347
Query: 83 SPSPSSSPAPSP 94
SPSPS SP+PSP
Sbjct: 348 SPSPSPSPSPSP 359
Score = 79.7 bits (195), Expect = 3e-14
Identities = 38/76 (50%), Positives = 52/76 (68%), Gaps = 2/76 (2%)
Query: 21 SSADSPAPTPATNSSLN-SPSPTPIPTPSPANSP-PAPTPTPTPSPHSDSPPAPSPDNSP 78
S + SP+P+P+ + S SPSP+P P+PSP SP P+P+P+P+PSP P+PSP SP
Sbjct: 369 SPSPSPSPSPSPSPSPKPSPSPSPSPSPSPKPSPSPSPSPSPSPSPKVSPSPSPSPSPSP 428
Query: 79 SSSPSPSPSSSPAPSP 94
S SPSP+ P+P P
Sbjct: 429 SPKASPSPAKKPSPPP 444
Score = 79.0 bits (193), Expect = 5e-14
Identities = 39/87 (44%), Positives = 55/87 (62%), Gaps = 4/87 (4%)
Query: 19 IASSADSPAPTPATNSSLNSPSPTPIPTPSPANSP---PAPTPTPTPSPHSDSPPAPSPD 75
+ S + SP+P+P+ + S SP P+P P+PSP+ SP P+P+P+P+PSP P+PSP
Sbjct: 365 LPSPSPSPSPSPSPSPS-PSPKPSPSPSPSPSPSPKPSPSPSPSPSPSPSPKVSPSPSPS 423
Query: 76 NSPSSSPSPSPSSSPAPSPDEAADNNA 102
SPS SP SPS + PSP + A
Sbjct: 424 PSPSPSPKASPSPAKKPSPPPPVEEGA 450
Score = 77.0 bits (188), Expect = 2e-13
Identities = 42/80 (52%), Positives = 51/80 (63%), Gaps = 11/80 (13%)
Query: 25 SPAPTPATNSSLNSPSPTPIPTPSPANSP---------PAPTPTPTPSPHSDSPPAPSPD 75
SP+P+P+ +S SPSP+P P+PSP SP PA P+P+PSP P PSP
Sbjct: 306 SPSPSPSPKAS-PSPSPSPSPSPSPKASPSPSPSPSVQPASKPSPSPSPSPSPSPRPSPP 364
Query: 76 -NSPSSSPSPSPSSSPAPSP 94
SPS SPSPSPS SP+PSP
Sbjct: 365 LPSPSPSPSPSPSPSPSPSP 384
Score = 75.5 bits (184), Expect = 5e-13
Identities = 39/81 (48%), Positives = 52/81 (64%), Gaps = 2/81 (2%)
Query: 16 LVNIASSADSPAPTP-ATNSSLNSPSPTPIPTPSPA-NSPPAPTPTPTPSPHSDSPPAPS 73
L N + SP+P+P A+ S SPSP+P +PSP+ + P+P+P+P SP P S
Sbjct: 257 LANFNRTGASPSPSPKASPSPKVSPSPSPKASPSPSPKASPSPSPSPKASPSPSPSPKAS 316
Query: 74 PDNSPSSSPSPSPSSSPAPSP 94
P SPS SPSPSP +SP+PSP
Sbjct: 317 PSPSPSPSPSPSPKASPSPSP 337
Score = 73.2 bits (178), Expect = 3e-12
Identities = 38/73 (52%), Positives = 49/73 (67%), Gaps = 4/73 (5%)
Query: 25 SPAPTPATNSSLNSPSPTPIPTPSPANSP-PAPTPTPTPSPHSDSPPAPSPDNSPSSSPS 83
SP+P+P + S SP +P P+PSP SP P+P+P +PSP P+PSP SPS SPS
Sbjct: 280 SPSPSPKASPS-PSPKASPSPSPSPKASPSPSPSPKASPSPSPSPSPSPSPKASPSPSPS 338
Query: 84 PS--PSSSPAPSP 94
PS P+S P+PSP
Sbjct: 339 PSVQPASKPSPSP 351
Score = 70.5 bits (171), Expect = 2e-11
Identities = 39/106 (36%), Positives = 59/106 (54%), Gaps = 5/106 (4%)
Query: 2 AIPRFSLVFLLLSFLVNIASSADSPAPTPATNSSLN----SPSPTPIPTPSPANSP-PAP 56
++ + ++ F+ SF + + + T ++ N SPSP+P +PSP SP P+P
Sbjct: 226 SVGKGAITFIGSSFAMPHLKGYEDMSGVAVTLANFNRTGASPSPSPKASPSPKVSPSPSP 285
Query: 57 TPTPTPSPHSDSPPAPSPDNSPSSSPSPSPSSSPAPSPDEAADNNA 102
+P+PSP + P+PSP SPS SPSP S SP+PSP + A
Sbjct: 286 KASPSPSPKASPSPSPSPKASPSPSPSPKASPSPSPSPSPSPSPKA 331
Score = 63.9 bits (154), Expect = 2e-09
Identities = 34/82 (41%), Positives = 47/82 (56%), Gaps = 9/82 (10%)
Query: 21 SSADSPAPTPATNSSLNSPSPTPIPTPSPANSP-PAPTPTPTPSPHSD-------SPPAP 72
S + SP+P+P+ S SPSP+P P+PSP SP P+P+P+P+PSP + SPP P
Sbjct: 387 SPSPSPSPSPSPKPS-PSPSPSPSPSPSPKVSPSPSPSPSPSPSPKASPSPAKKPSPPPP 445
Query: 73 SPDNSPSSSPSPSPSSSPAPSP 94
+ +P P P AP P
Sbjct: 446 VEEGAPPPIEGPPPMEEAAPPP 467
>ref|NP_932693.1| unknown [Choristoneura fumiferana defective nucleopolyhedrovirus]
gi|37499308|gb|AAQ91707.1| unknown [Choristoneura
fumiferana defective nucleopolyhedrovirus]
Length = 218
Score = 89.4 bits (220), Expect = 3e-17
Identities = 44/98 (44%), Positives = 59/98 (59%), Gaps = 9/98 (9%)
Query: 6 FSLVFLLLSFLVNIA------SSADSPAPTPATNSSLNSPSPTPIPTPSPANSP---PAP 56
F ++FL L L + + + +P+PTP + + +PSPTP PTPSP SP P P
Sbjct: 11 FIIIFLTLYLLATLKRVPEPPTPSPTPSPTPPSPTPPPTPSPTPSPTPSPTPSPTPPPTP 70
Query: 57 TPTPTPSPHSDSPPAPSPDNSPSSSPSPSPSSSPAPSP 94
PTP+P+P PP P P SP+ SP+PSP+ SP PSP
Sbjct: 71 LPTPSPTPSPTPPPTPLPTPSPTPSPTPSPTPSPTPSP 108
Score = 80.1 bits (196), Expect = 2e-14
Identities = 37/78 (47%), Positives = 50/78 (63%), Gaps = 8/78 (10%)
Query: 25 SPAPTPATNSSLNSPSPTPIPTPSPANSP-------PAPTPTPTPSPHSDSPPAPSPDNS 77
+P+PTP+ S +P PTP+PTPSP SP P P+PTP+P+P P PSP
Sbjct: 53 TPSPTPSPTPS-PTPPPTPLPTPSPTPSPTPPPTPLPTPSPTPSPTPSPTPSPTPSPTPP 111
Query: 78 PSSSPSPSPSSSPAPSPD 95
P+ SP+P+PS +P PSP+
Sbjct: 112 PTPSPTPTPSPTPPPSPE 129
Score = 76.3 bits (186), Expect = 3e-13
Identities = 37/73 (50%), Positives = 46/73 (62%), Gaps = 2/73 (2%)
Query: 25 SPAPTPATNSSLNSPSPTPIPTPSPANSP-PAPTPTPTPSPHSDSPPAPSPDNSPSSSPS 83
+P PTP+ S +P PTP+PTPSP SP P+PTP+PTPSP P+P+P SP+ PS
Sbjct: 69 TPLPTPSPTPS-PTPPPTPLPTPSPTPSPTPSPTPSPTPSPTPPPTPSPTPTPSPTPPPS 127
Query: 84 PSPSSSPAPSPDE 96
P P P P E
Sbjct: 128 PEPLGEPMYFPAE 140
Score = 75.1 bits (183), Expect = 7e-13
Identities = 37/71 (52%), Positives = 48/71 (67%), Gaps = 6/71 (8%)
Query: 25 SPAPTPATNSSLNSPSPTPIPTPSPANSP-PAPTPTPTPSPHSDSPPAPSPDNSPSSSPS 83
+P PTP S +PSPTP PTP P SP P+PTP+PTPSP P+P+P +PS +P+
Sbjct: 65 TPPPTPLPTPS-PTPSPTPPPTPLPTPSPTPSPTPSPTPSP----TPSPTPPPTPSPTPT 119
Query: 84 PSPSSSPAPSP 94
PSP+ P+P P
Sbjct: 120 PSPTPPPSPEP 130
Score = 44.7 bits (104), Expect = 0.001
Identities = 21/45 (46%), Positives = 30/45 (66%), Gaps = 2/45 (4%)
Query: 19 IASSADSPAPTPATNSSLNSPSPTPIPTPSPANSP-PAPTPTPTP 62
+ + + +P+PTP+ S +PSPTP PTPSP +P P P P+P P
Sbjct: 87 LPTPSPTPSPTPSPTPS-PTPSPTPPPTPSPTPTPSPTPPPSPEP 130
>ref|XP_470430.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|27573359|gb|AAO20077.1| unknown protein [Oryza sativa
(japonica cultivar-group)]
Length = 219
Score = 89.4 bits (220), Expect = 3e-17
Identities = 64/168 (38%), Positives = 88/168 (52%), Gaps = 22/168 (13%)
Query: 20 ASSADSPAPTPATNSSLNSPSPTP---IPTPSPANSPPAP---TPTPTPSPHSD-SPPAP 72
A+ A S P+ A++S+ P P P P+P++SPP P P P+P HS S PAP
Sbjct: 53 AALAPSQPPSEASSSAAALSPPAPPETSPLPAPSHSPPVPHSAAPEPSPMEHSAASAPAP 112
Query: 73 SP------------DNSPSSSPSPSPSS-SPAPSPDEAADNNAISHTGIGEDGKSSGGGM 119
S D+ PS+ SPAP+ +E A G EDG++ +
Sbjct: 113 SAAKAKQGGDDEEDDDDKEKDKEEKPSTPSPAPAAEEIKAATAGDKAG-EEDGETERHEL 171
Query: 120 SSGKKAGIAVGVIAAVGVVALGAMVVKKRRQNIQRSEYG-YTARRELL 166
+ GKKAG+ VG +A VV L A+V KKR+ NI+RS Y Y+AR EL+
Sbjct: 172 NGGKKAGVVVGAFSAAAVVGLAAVVWKKRQANIRRSRYADYSARLELV 219
>ref|YP_113681.1| metalloprotease, putative [Methylococcus capsulatus str. Bath]
gi|53758245|gb|AAU92536.1| putative metalloprotease
[Methylococcus capsulatus str. Bath]
Length = 839
Score = 87.8 bits (216), Expect = 1e-16
Identities = 53/124 (42%), Positives = 71/124 (56%), Gaps = 15/124 (12%)
Query: 20 ASSADSPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPS 79
AS + SP+P+P SPSP+P P+PSP+ P+P+P+P+PSP P+PSP SPS
Sbjct: 693 ASCSGSPSPSP-------SPSPSPSPSPSPS---PSPSPSPSPSP----SPSPSPSPSPS 738
Query: 80 SSPSPSPSSSPAPSPDEAADNNAISHTGIGEDG-KSSGGGMSSGKKAGIAVGVIAAVGVV 138
SPSPSPS SP+PSP A +++ T SS G+S G A A+ +V
Sbjct: 739 PSPSPSPSPSPSPSPAPTAYTLSVTKTNSSRGAVNSSPSGISCGTSCSSASASFASGSMV 798
Query: 139 ALGA 142
L A
Sbjct: 799 TLTA 802
Score = 68.6 bits (166), Expect = 6e-11
Identities = 47/128 (36%), Positives = 69/128 (53%), Gaps = 14/128 (10%)
Query: 38 SPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPSPSSSPAPSPDEA 97
S SP+P P+PSP+ P+P+P+P+PSP +PSP SPS SPSPSPS SP+PSP +
Sbjct: 696 SGSPSPSPSPSPS---PSPSPSPSPSP------SPSPSPSPSPSPSPSPSPSPSPSPSPS 746
Query: 98 ---ADNNAISHTGIGEDGKSSGGGMSSGKKAGIAVGV--IAAVGVVALGAMVVKKRRQNI 152
+ + A + + +S G + +GI+ G +A A G+MV I
Sbjct: 747 PSPSPSPAPTAYTLSVTKTNSSRGAVNSSPSGISCGTSCSSASASFASGSMVTLTASPFI 806
Query: 153 QRSEYGYT 160
R G++
Sbjct: 807 FRKFTGWS 814
>gb|AAC59091.1| unknown [Orgyia pseudotsugata multicapsid nucleopolyhedrovirus]
gi|2493240|sp|O10341|Y091_NPVOP Hypothetical 29.3 kDa
protein (ORF92) gi|7515481|pir||T10361 hypothetical
protein 92 - Orgyia pseudotsugata nuclear polyhedrosis
virus gi|9630030|ref|NP_046248.1| unknown [Orgyia
pseudotsugata multicapsid nucleopolyhedrovirus]
Length = 279
Score = 87.8 bits (216), Expect = 1e-16
Identities = 42/76 (55%), Positives = 53/76 (69%), Gaps = 2/76 (2%)
Query: 21 SSADSPAPTPA-TNSSLNSPSPTPIPTPSPANSP-PAPTPTPTPSPHSDSPPAPSPDNSP 78
S SP PTP+ T S +PSPTP PTP+P+ +P P+PTP+PTPSP P PSP +P
Sbjct: 102 SPTPSPTPTPSPTPSPTPTPSPTPSPTPTPSPTPTPSPTPSPTPSPTPTPSPTPSPTPTP 161
Query: 79 SSSPSPSPSSSPAPSP 94
S +PSP+P+ SP PSP
Sbjct: 162 SPTPSPTPTPSPTPSP 177
Score = 85.9 bits (211), Expect = 4e-16
Identities = 41/78 (52%), Positives = 53/78 (67%), Gaps = 4/78 (5%)
Query: 21 SSADSPAPTPA---TNSSLNSPSPTPIPTPSPANSP-PAPTPTPTPSPHSDSPPAPSPDN 76
S +P+PTP+ T S SP+PTP PTP+P+ +P P P+PTPTPSP P PSP
Sbjct: 106 SPTPTPSPTPSPTPTPSPTPSPTPTPSPTPTPSPTPSPTPSPTPTPSPTPSPTPTPSPTP 165
Query: 77 SPSSSPSPSPSSSPAPSP 94
SP+ +PSP+PS +P PSP
Sbjct: 166 SPTPTPSPTPSPTPPPSP 183
Score = 85.9 bits (211), Expect = 4e-16
Identities = 41/88 (46%), Positives = 58/88 (65%), Gaps = 2/88 (2%)
Query: 9 VFLLLSFLVNIASSADSPAPTPA-TNSSLNSPSPTPIPTPSPANSP-PAPTPTPTPSPHS 66
++LL S NI + P+PTP T S +P+P+P PTP+P +P P PTP+PTP+P
Sbjct: 18 LWLLTSLKQNIETKPPLPSPTPTPTPSPTPTPTPSPTPTPTPTPTPTPTPTPSPTPTPAL 77
Query: 67 DSPPAPSPDNSPSSSPSPSPSSSPAPSP 94
P PSP SP+ SP+P+PS +P+P+P
Sbjct: 78 SPTPTPSPTLSPTPSPTPTPSPTPSPTP 105
Score = 84.3 bits (207), Expect = 1e-15
Identities = 41/83 (49%), Positives = 51/83 (61%), Gaps = 4/83 (4%)
Query: 21 SSADSPAPTPATNSSLN---SPSPTPIPTPSPANSP-PAPTPTPTPSPHSDSPPAPSPDN 76
S SP PTP+ S SP+PTP PTPSP SP P P+PTP+P+P P+P+P
Sbjct: 112 SPTPSPTPTPSPTPSPTPTPSPTPTPSPTPSPTPSPTPTPSPTPSPTPTPSPTPSPTPTP 171
Query: 77 SPSSSPSPSPSSSPAPSPDEAAD 99
SP+ SP+P PS +P PSP D
Sbjct: 172 SPTPSPTPPPSPTPPPSPSPLGD 194
Score = 84.3 bits (207), Expect = 1e-15
Identities = 42/75 (56%), Positives = 49/75 (65%), Gaps = 6/75 (8%)
Query: 25 SPAPTPA-----TNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPS 79
SP PTPA T S SP+P+P PTPSP SP P+PTPTPSP P PSP SP+
Sbjct: 70 SPTPTPALSPTPTPSPTLSPTPSPTPTPSPTPSP-TPSPTPTPSPTPSPTPTPSPTPSPT 128
Query: 80 SSPSPSPSSSPAPSP 94
+PSP+P+ SP PSP
Sbjct: 129 PTPSPTPTPSPTPSP 143
Score = 76.3 bits (186), Expect = 3e-13
Identities = 38/87 (43%), Positives = 49/87 (55%), Gaps = 4/87 (4%)
Query: 21 SSADSPAPTPA---TNSSLNSPSPTPIPTPSPANSP-PAPTPTPTPSPHSDSPPAPSPDN 76
S SP PTP+ T S SP+P+P PTPSP SP P P+PTP+P+P P+P+P
Sbjct: 122 SPTPSPTPTPSPTPTPSPTPSPTPSPTPTPSPTPSPTPTPSPTPSPTPTPSPTPSPTPPP 181
Query: 77 SPSSSPSPSPSSSPAPSPDEAADNNAI 103
SP+ PSPSP P P + +
Sbjct: 182 SPTPPPSPSPLGDPMYFPSSVGTTDQL 208
Score = 71.6 bits (174), Expect = 7e-12
Identities = 37/79 (46%), Positives = 47/79 (58%), Gaps = 4/79 (5%)
Query: 25 SPAPTPA-TNSSLNSPSPTPIPTPSPANSP---PAPTPTPTPSPHSDSPPAPSPDNSPSS 80
SP PTP+ T S SP+PTP PTPSP +P P+PTPTP+P+P PP+P+P SPS
Sbjct: 132 SPTPTPSPTPSPTPSPTPTPSPTPSPTPTPSPTPSPTPTPSPTPSPTPPPSPTPPPSPSP 191
Query: 81 SPSPSPSSSPAPSPDEAAD 99
P S + D+ D
Sbjct: 192 LGDPMYFPSSVGTTDQLRD 210
Score = 67.4 bits (163), Expect = 1e-10
Identities = 34/87 (39%), Positives = 48/87 (55%), Gaps = 16/87 (18%)
Query: 8 LVFLLLSFLVNIASSADSPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSD 67
++ L L L ++ + ++ P P SPTP PTPSP TPTPTPSP
Sbjct: 13 IILLTLWLLTSLKQNIETKPPLP---------SPTPTPTPSP-------TPTPTPSPTPT 56
Query: 68 SPPAPSPDNSPSSSPSPSPSSSPAPSP 94
P P+P +P+ SP+P+P+ SP P+P
Sbjct: 57 PTPTPTPTPTPTPSPTPTPALSPTPTP 83
Score = 56.6 bits (135), Expect = 2e-07
Identities = 21/45 (46%), Positives = 32/45 (70%)
Query: 53 PPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPSPSSSPAPSPDEA 97
PP P+PTPTP+P P PSP +P+ +P+P+P+ +P+P+P A
Sbjct: 32 PPLPSPTPTPTPSPTPTPTPSPTPTPTPTPTPTPTPTPSPTPTPA 76
>gb|AAK85643.1| unknown [Epiphyas postvittana nucleopolyhedrovirus]
gi|15320736|ref|NP_203248.1| unknown [Epiphyas
postvittana nucleopolyhedrovirus]
Length = 234
Score = 86.3 bits (212), Expect = 3e-16
Identities = 44/100 (44%), Positives = 61/100 (61%), Gaps = 13/100 (13%)
Query: 8 LVFLLLSFLVNIASS---------ADSPAPTPA-TNSSLNSPSPTPIPTPSPANSP---P 54
++ L+L L++I +P+PTP T S SPSPTP PTP P+ +P P
Sbjct: 13 IIILILKLLLSIKKQNKCNSEKCLPPTPSPTPLPTPSPTPSPSPTPSPTPPPSPTPSPTP 72
Query: 55 APTPTPTPSPHSDSPPAPSPDNSPSSSPSPSPSSSPAPSP 94
P+PTP+P+P PP+P+P +PS +PSP+PS SP PSP
Sbjct: 73 PPSPTPSPTPSPTPPPSPTPSPTPSPTPSPTPSPSPLPSP 112
Score = 83.6 bits (205), Expect = 2e-15
Identities = 42/77 (54%), Positives = 51/77 (65%), Gaps = 4/77 (5%)
Query: 21 SSADSPAPTPA---TNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNS 77
S + +P+PTP T S PSPTP PTPSP PP+PTP+PTPSP P+PSP S
Sbjct: 53 SPSPTPSPTPPPSPTPSPTPPPSPTPSPTPSPT-PPPSPTPSPTPSPTPSPTPSPSPLPS 111
Query: 78 PSSSPSPSPSSSPAPSP 94
P+ SP+PSP+ P PSP
Sbjct: 112 PTPSPTPSPTLWPTPSP 128
Score = 77.4 bits (189), Expect = 1e-13
Identities = 43/81 (53%), Positives = 52/81 (64%), Gaps = 8/81 (9%)
Query: 21 SSADSPAPTPA-TNSSLNSPSPTPIPTPSPANSP---PAPTPTPTPSPHSDSP---PAPS 73
S P+PTP+ T S PSPTP PTPSP SP P+P P+PTPSP + SP P PS
Sbjct: 69 SPTPPPSPTPSPTPSPTPPPSPTPSPTPSPTPSPTPSPSPLPSPTPSP-TPSPTLWPTPS 127
Query: 74 PDNSPSSSPSPSPSSSPAPSP 94
P SP+ P+PSPS P+P+P
Sbjct: 128 PTPSPTLWPTPSPSPPPSPNP 148
Score = 77.0 bits (188), Expect = 2e-13
Identities = 37/74 (50%), Positives = 48/74 (64%), Gaps = 2/74 (2%)
Query: 21 SSADSPAPTPATNSSLNSPSPTPIPTPSPANSP-PAPTPTPTPSPHSDSPPAPSPDNSPS 79
S SP+PTP+ SP+P+P P PSP SP P+PTP P+P+P P PSP SPS
Sbjct: 49 SPTPSPSPTPSPTPP-PSPTPSPTPPPSPTPSPTPSPTPPPSPTPSPTPSPTPSPTPSPS 107
Query: 80 SSPSPSPSSSPAPS 93
PSP+PS +P+P+
Sbjct: 108 PLPSPTPSPTPSPT 121
Score = 63.9 bits (154), Expect = 2e-09
Identities = 36/82 (43%), Positives = 43/82 (51%), Gaps = 4/82 (4%)
Query: 21 SSADSPAPTPA-TNSSLNSPSPTPIPTPSPANSP---PAPTPTPTPSPHSDSPPAPSPDN 76
S SP P P+ T S SP+P+P P+PSP SP P P+PT P+P P P
Sbjct: 79 SPTPSPTPPPSPTPSPTPSPTPSPTPSPSPLPSPTPSPTPSPTLWPTPSPTPSPTLWPTP 138
Query: 77 SPSSSPSPSPSSSPAPSPDEAA 98
SPS PSP+P P P E A
Sbjct: 139 SPSPPPSPNPLGDPMYFPSEIA 160
>ref|NP_441791.1| hypothetical protein slr0442 [Synechocystis sp. PCC 6803]
gi|1653557|dbj|BAA18470.1| slr0442 [Synechocystis sp.
PCC 6803] gi|7470231|pir||S76211 hypothetical protein
slr0442 - Synechocystis sp. (strain PCC 6803)
Length = 611
Score = 84.7 bits (208), Expect = 8e-16
Identities = 42/88 (47%), Positives = 58/88 (65%), Gaps = 1/88 (1%)
Query: 14 SFLVNIASSADSPAPTPATNSSLNSPSPTPIPTPSPANSP-PAPTPTPTPSPHSDSPPAP 72
S L + + + P T + ++ +SPSP+P P+PSP+ SP P+P+P+P+PSP P+P
Sbjct: 496 SSLADDDAITEPPIYTGSFLTASSSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSP 555
Query: 73 SPDNSPSSSPSPSPSSSPAPSPDEAADN 100
SP SPS SPSPSPS SP+PSP N
Sbjct: 556 SPSPSPSPSPSPSPSPSPSPSPTPVTVN 583
Score = 84.3 bits (207), Expect = 1e-15
Identities = 43/86 (50%), Positives = 55/86 (63%), Gaps = 1/86 (1%)
Query: 32 TNSSLNSPSPTPIPTPSPANSP-PAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPSPSSSP 90
T SS SPSP+P P+PSP+ SP P+P+P+P+PSP P+PSP SPS SPSPSPS SP
Sbjct: 516 TASSSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSPSP 575
Query: 91 APSPDEAADNNAISHTGIGEDGKSSG 116
+P+P N + +G SG
Sbjct: 576 SPTPVTVNVQNKKACDDLGGTYSQSG 601
>ref|XP_629665.1| hypothetical protein DDB0184442 [Dictyostelium discoideum]
gi|60463036|gb|EAL61232.1| hypothetical protein
DDB0184442 [Dictyostelium discoideum]
Length = 644
Score = 84.0 bits (206), Expect = 1e-15
Identities = 39/76 (51%), Positives = 51/76 (66%), Gaps = 2/76 (2%)
Query: 21 SSADSPAPTPATNSSLN-SPSPTPIPTPSPANSP-PAPTPTPTPSPHSDSPPAPSPDNSP 78
S SP P+P +L+ +PSPTP PTPSP +P P+PT TPTPSP P+P+P +P
Sbjct: 196 SPTPSPTPSPTQTPTLSPTPSPTPSPTPSPTQTPTPSPTQTPTPSPTPSPTPSPTPSPTP 255
Query: 79 SSSPSPSPSSSPAPSP 94
S +PSP+P+ P PSP
Sbjct: 256 SPTPSPTPTPFPTPSP 271
Score = 79.3 bits (194), Expect = 4e-14
Identities = 42/81 (51%), Positives = 53/81 (64%), Gaps = 16/81 (19%)
Query: 25 SPAPTPATNSSLNSPSPTPIPTPSPANSP-----PAPTPTPTPSPH---SDSP---PAPS 73
+P+PTP+ +PSPTP PTPSP +P P+PTP+PTPSP + SP P PS
Sbjct: 186 TPSPTPSP-----TPSPTPSPTPSPTQTPTLSPTPSPTPSPTPSPTQTPTPSPTQTPTPS 240
Query: 74 PDNSPSSSPSPSPSSSPAPSP 94
P SP+ SP+PSP+ SP PSP
Sbjct: 241 PTPSPTPSPTPSPTPSPTPSP 261
Score = 78.2 bits (191), Expect = 8e-14
Identities = 39/74 (52%), Positives = 52/74 (69%), Gaps = 4/74 (5%)
Query: 25 SPAPTPA-TNSSLNSPSPTPI--PTPSPANSP-PAPTPTPTPSPHSDSPPAPSPDNSPSS 80
+P+PTP+ T S SP+ TP PTPSP SP P+PT TPTPSP P+P+P +PS
Sbjct: 190 TPSPTPSPTPSPTPSPTQTPTLSPTPSPTPSPTPSPTQTPTPSPTQTPTPSPTPSPTPSP 249
Query: 81 SPSPSPSSSPAPSP 94
+PSP+PS +P+P+P
Sbjct: 250 TPSPTPSPTPSPTP 263
Score = 75.9 bits (185), Expect = 4e-13
Identities = 34/78 (43%), Positives = 49/78 (62%), Gaps = 10/78 (12%)
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPANSP---------PAPTPTPTPSPHSDSPPAPSPDN 76
P P+P T + SP+P+P P+P+P+ +P P P+PTP+P+P P PSP
Sbjct: 177 PTPSP-TQTPTPSPTPSPTPSPTPSPTPSPTQTPTLSPTPSPTPSPTPSPTQTPTPSPTQ 235
Query: 77 SPSSSPSPSPSSSPAPSP 94
+P+ SP+PSP+ SP PSP
Sbjct: 236 TPTPSPTPSPTPSPTPSP 253
Score = 70.5 bits (171), Expect = 2e-11
Identities = 32/61 (52%), Positives = 41/61 (66%), Gaps = 1/61 (1%)
Query: 36 LNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPSPSSSPAPSPD 95
L +PSPT PTPSP SP P+PTP+P+P P SP SP+ SP+PSP+ +P PSP
Sbjct: 176 LPTPSPTQTPTPSPTPSP-TPSPTPSPTPSPTQTPTLSPTPSPTPSPTPSPTQTPTPSPT 234
Query: 96 E 96
+
Sbjct: 235 Q 235
Score = 67.0 bits (162), Expect = 2e-10
Identities = 36/82 (43%), Positives = 47/82 (56%), Gaps = 4/82 (4%)
Query: 15 FLVNIASSADSPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSP 74
FLV S S +SSL P +PTPSP +P P+PTP+P+P P PSP
Sbjct: 150 FLVTSLSENPSCTYNFLIHSSLACPY---LPTPSPTQTP-TPSPTPSPTPSPTPSPTPSP 205
Query: 75 DNSPSSSPSPSPSSSPAPSPDE 96
+P+ SP+PSP+ SP PSP +
Sbjct: 206 TQTPTLSPTPSPTPSPTPSPTQ 227
Score = 57.8 bits (138), Expect = 1e-07
Identities = 30/66 (45%), Positives = 41/66 (61%), Gaps = 9/66 (13%)
Query: 29 TPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPSPSS 88
T T S+ +PTP PTPSP P+PT TPTP+ PSP +PS +PSP+P+
Sbjct: 407 TIETVSACVGYTPTPSPTPSPT---PSPTRTPTPT------RTPSPTRTPSPTPSPTPTP 457
Query: 89 SPAPSP 94
+P+P+P
Sbjct: 458 NPSPNP 463
Score = 57.0 bits (136), Expect = 2e-07
Identities = 36/82 (43%), Positives = 48/82 (57%), Gaps = 7/82 (8%)
Query: 25 SPAPTPA-TNSSLNSPS--PTPIPTPSPANSP---PAPTPTPTPSPHSDSPPAPSPDNS- 77
+P+PTP+ T + SP+ PTP PTPSP SP P P+PTP+P+P P+P+P+N
Sbjct: 218 TPSPTPSPTQTPTPSPTQTPTPSPTPSPTPSPTPSPTPSPTPSPTPTPFPTPSPTPNNPI 277
Query: 78 PSSSPSPSPSSSPAPSPDEAAD 99
PSS S SP A+D
Sbjct: 278 PSSCQIKFGSQEIDLSPLIASD 299
Score = 48.1 bits (113), Expect = 9e-05
Identities = 19/40 (47%), Positives = 27/40 (67%)
Query: 56 PTPTPTPSPHSDSPPAPSPDNSPSSSPSPSPSSSPAPSPD 95
PTP+PTPSP P+P +PS + +PSP+ SP P+P+
Sbjct: 419 PTPSPTPSPTPSPTRTPTPTRTPSPTRTPSPTPSPTPTPN 458
Score = 35.4 bits (80), Expect = 0.59
Identities = 20/48 (41%), Positives = 24/48 (49%), Gaps = 10/48 (20%)
Query: 21 SSADSPAPTPA-------TNSSLNSPSPTPIPTPSPANSPPAPTPTPT 61
S SP P+P T S +PSPTP PTP+P P+P P T
Sbjct: 422 SPTPSPTPSPTRTPTPTRTPSPTRTPSPTPSPTPTP---NPSPNPMDT 466
Score = 33.5 bits (75), Expect = 2.2
Identities = 21/56 (37%), Positives = 29/56 (51%), Gaps = 3/56 (5%)
Query: 25 SPAPTPA-TNSSLNSPSPTPI--PTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNS 77
+P+PTP+ T S SP+PTP P+P+P N P+ S D P + D S
Sbjct: 246 TPSPTPSPTPSPTPSPTPTPFPTPSPTPNNPIPSSCQIKFGSQEIDLSPLIASDYS 301
>gb|AAV63985.1| hydroxyproline-rich glycoprotein VSP-3 [Chlamydomonas incerta]
Length = 465
Score = 83.6 bits (205), Expect = 2e-15
Identities = 41/77 (53%), Positives = 54/77 (69%), Gaps = 8/77 (10%)
Query: 25 SPAPTPATNSSLNSPSPTPIPTPSPANSP-------PAPTPTPTPSPHSDSPPAPSPDNS 77
SP+PTP+ +S SPSP+P +PSP SP P+P+P+P+PSP + P+PSP S
Sbjct: 282 SPSPTPSPKAS-PSPSPSPKASPSPFPSPKASPSPSPSPSPSPSPSPSPKASPSPSPSPS 340
Query: 78 PSSSPSPSPSSSPAPSP 94
PS SPSPSP +SP+PSP
Sbjct: 341 PSPSPSPSPKASPSPSP 357
Score = 82.0 bits (201), Expect = 5e-15
Identities = 44/89 (49%), Positives = 58/89 (64%), Gaps = 8/89 (8%)
Query: 22 SADSPAPTP-ATNSSLNSPSPTPIPTPSPANSP-----PAPTPTPTPSPHSDSPPAPSPD 75
++ SP P+P A+ S SPSP+P P+PSP SP P+P+P+P+PSP + P+PSP
Sbjct: 301 ASPSPFPSPKASPSPSPSPSPSPSPSPSPKASPSPSPSPSPSPSPSPSPKASPSPSPSPS 360
Query: 76 NSPSS--SPSPSPSSSPAPSPDEAADNNA 102
PSS SPSPSPS SP+PSP + A
Sbjct: 361 VQPSSKPSPSPSPSPSPSPSPSPSPSPKA 389
Score = 81.6 bits (200), Expect = 7e-15
Identities = 40/77 (51%), Positives = 54/77 (69%), Gaps = 4/77 (5%)
Query: 22 SADSPAPTPATN-SSLNSPSPTPIPTPSPANSP---PAPTPTPTPSPHSDSPPAPSPDNS 77
++ SP+P+P+ SS SPSP+P P+PSP+ SP P +P+P+PSP +PSP S
Sbjct: 351 ASPSPSPSPSVQPSSKPSPSPSPSPSPSPSPSPSPSPKASPSPSPSPSPSPKVSPSPSPS 410
Query: 78 PSSSPSPSPSSSPAPSP 94
PS SPSPSP +SP+PSP
Sbjct: 411 PSPSPSPSPKASPSPSP 427
Score = 79.3 bits (194), Expect = 4e-14
Identities = 44/91 (48%), Positives = 60/91 (65%), Gaps = 12/91 (13%)
Query: 20 ASSADSPAPTPATNSSLN---SPSPTPIPTPSPANSP---PAPTPTPTPSPH-SDSP--- 69
AS + SP+P+P+ + S + SPSP+P P+ P++ P P+P+P+P+PSP S SP
Sbjct: 331 ASPSPSPSPSPSPSPSPSPKASPSPSPSPSVQPSSKPSPSPSPSPSPSPSPSPSPSPKAS 390
Query: 70 --PAPSPDNSPSSSPSPSPSSSPAPSPDEAA 98
P+PSP SP SPSPSPS SP+PSP A
Sbjct: 391 PSPSPSPSPSPKVSPSPSPSPSPSPSPSPKA 421
Score = 78.2 bits (191), Expect = 8e-14
Identities = 39/72 (54%), Positives = 49/72 (67%), Gaps = 2/72 (2%)
Query: 25 SPAPTP-ATNSSLNSPSPTPIPTPSPANSP-PAPTPTPTPSPHSDSPPAPSPDNSPSSSP 82
SP+P+P A+ S SP +P P+PSP SP P P+P +PSP P+PSP SP +SP
Sbjct: 274 SPSPSPKASPSPTPSPKASPSPSPSPKASPSPFPSPKASPSPSPSPSPSPSPSPSPKASP 333
Query: 83 SPSPSSSPAPSP 94
SPSPS SP+PSP
Sbjct: 334 SPSPSPSPSPSP 345
Score = 77.4 bits (189), Expect = 1e-13
Identities = 41/85 (48%), Positives = 53/85 (62%), Gaps = 6/85 (7%)
Query: 16 LVNIASSADSPAPTPATNSSLN-SPSP--TPIPTPSPANSP---PAPTPTPTPSPHSDSP 69
L N + P P+P+ S + SPSP +P PTPSP SP P+P +P+P P +
Sbjct: 253 LANFNRTGPVPTPSPSPKKSPSPSPSPKASPSPTPSPKASPSPSPSPKASPSPFPSPKAS 312
Query: 70 PAPSPDNSPSSSPSPSPSSSPAPSP 94
P+PSP SPS SPSPSP +SP+PSP
Sbjct: 313 PSPSPSPSPSPSPSPSPKASPSPSP 337
Score = 73.2 bits (178), Expect = 3e-12
Identities = 39/78 (50%), Positives = 52/78 (66%), Gaps = 4/78 (5%)
Query: 21 SSADSPAPTPATNSSLN-SPSPTPIPTPSPANSP-PAPTPTPTPSPHSDSPPAPSPDNSP 78
SS SP+P+P+ + S + SPSP+P +PSP+ SP P+P +P+PSP P+PSP SP
Sbjct: 364 SSKPSPSPSPSPSPSPSPSPSPSPKASPSPSPSPSPSPKVSPSPSPSPSPSPSPSPKASP 423
Query: 79 SSSPSPSPSSSPAPSPDE 96
SPSP P SP+P P E
Sbjct: 424 --SPSPVPKKSPSPPPPE 439
Score = 70.5 bits (171), Expect = 2e-11
Identities = 36/77 (46%), Positives = 48/77 (61%), Gaps = 2/77 (2%)
Query: 21 SSADSPAPTPATNSSLN-SPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPS 79
S + SP+P+P+ + S SPSP+P P+PSP SP +P+P+P+PSP +PSP P
Sbjct: 372 SPSPSPSPSPSPSPSPKASPSPSPSPSPSPKVSP-SPSPSPSPSPSPSPKASPSPSPVPK 430
Query: 80 SSPSPSPSSSPAPSPDE 96
SPSP P AP P E
Sbjct: 431 KSPSPPPPEELAPPPVE 447
Score = 63.2 bits (152), Expect = 3e-09
Identities = 34/82 (41%), Positives = 49/82 (59%), Gaps = 8/82 (9%)
Query: 21 SSADSPAPTP-ATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPS 79
S + SP+P+P A+ S SPSP+P +PSP+ P+P+P+P+PSP + P+P P SPS
Sbjct: 378 SPSPSPSPSPKASPSPSPSPSPSPKVSPSPS---PSPSPSPSPSPKASPSPSPVPKKSPS 434
Query: 80 SSP----SPSPSSSPAPSPDEA 97
P +P P P P + A
Sbjct: 435 PPPPEELAPPPVEGPPPMEESA 456
Score = 60.8 bits (146), Expect = 1e-08
Identities = 31/67 (46%), Positives = 39/67 (57%), Gaps = 7/67 (10%)
Query: 32 TNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPSPSSSPA 91
T ++ N P P P+PSP SP +P+PSP + P PSP SPS PSPSP +SP+
Sbjct: 252 TLANFNRTGPVPTPSPSPKKSP-----SPSPSPKASPSPTPSPKASPS--PSPSPKASPS 304
Query: 92 PSPDEAA 98
P P A
Sbjct: 305 PFPSPKA 311
Score = 58.5 bits (140), Expect = 6e-08
Identities = 30/75 (40%), Positives = 42/75 (56%), Gaps = 4/75 (5%)
Query: 20 ASSADSPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPS 79
AS + SP+P+P+ S PSP+P P+PSP+ SP A +P+P+P P P P + +P
Sbjct: 389 ASPSPSPSPSPSPKVS---PSPSPSPSPSPSPSPKA-SPSPSPVPKKSPSPPPPEELAPP 444
Query: 80 SSPSPSPSSSPAPSP 94
P P AP P
Sbjct: 445 PVEGPPPMEESAPPP 459
Score = 41.6 bits (96), Expect = 0.008
Identities = 22/45 (48%), Positives = 26/45 (56%), Gaps = 4/45 (8%)
Query: 58 PTPTPSPHSDSPPAPSPDNSPSSSPSPSPSSSPAPSPDEAADNNA 102
P PTPSP P+PSP SP +SPSP+PS P SP + A
Sbjct: 261 PVPTPSPSPKKSPSPSP--SPKASPSPTPS--PKASPSPSPSPKA 301
Score = 31.6 bits (70), Expect = 8.5
Identities = 18/57 (31%), Positives = 26/57 (45%), Gaps = 13/57 (22%)
Query: 21 SSADSPAPTPATNSSLN-SPSPTPIP------------TPSPANSPPAPTPTPTPSP 64
S + SP+P+P+ + S SPSP+P+P P P PP + P P
Sbjct: 404 SPSPSPSPSPSPSPSPKASPSPSPVPKKSPSPPPPEELAPPPVEGPPPMEESAPPPP 460
>gb|AAP37768.1| At3g24600 [Arabidopsis thaliana] gi|13877617|gb|AAK43886.1| protein
kinase-like protein [Arabidopsis thaliana]
Length = 652
Score = 82.8 bits (203), Expect = 3e-15
Identities = 50/145 (34%), Positives = 78/145 (53%), Gaps = 12/145 (8%)
Query: 21 SSADSPAPTPATNSSL---NSPSPTPIPTPSPANSPPAPTPTPTPS-PHSDSPPAPSPDN 76
++ SP P+P+TNS+ +SP P +P PSP S P P P+PS P + SPP+P+ +
Sbjct: 38 TTPSSPPPSPSTNSTSPPPSSPLPPSLPPPSPPGSLTPPIPQPSPSAPITPSPPSPTTPS 97
Query: 77 SPSSSPSPS--PSSSPAPSPDEAADNNAISHTGIGEDGKSSGGGMSSGKKAGIAVGVIAA 134
+P S PSP+ P ++P+ S N S S G+S+G GIA+G +A
Sbjct: 98 NPRSPPSPNQGPPNTPSGSTPRTPSNTKPS------PPSDSSDGLSTGVVVGIAIGGVAI 151
Query: 135 VGVVALGAMVVKKRRQNIQRSEYGY 159
+ ++ L ++ KK+R+ E Y
Sbjct: 152 LVILTLICLLCKKKRRRRHDDEAAY 176
Score = 34.7 bits (78), Expect = 1.0
Identities = 33/110 (30%), Positives = 45/110 (40%), Gaps = 15/110 (13%)
Query: 27 APTPATNSSLNS-PSPTPIPTPSPANSPPAPTPTPT------PSPHSDSP--PAPSPDNS 77
A P+ N + S P P P PSP PP P P P S +SD P P PSP
Sbjct: 204 ASRPSDNHVVTSLPPPKP---PSPPRKPPPPPPPPAFMSSSGGSDYSDLPVLPPPSPGLV 260
Query: 78 PSSSPSPSPSSSPAPSPDEAADNNAISHTGIGEDGKSSGGGMSSGKKAGI 127
S S + + + ++ N + G G K G + SGK+ +
Sbjct: 261 LGFSKSTFTYEELSRATNGFSEANLLGQGGFGYVHK---GILPSGKEVAV 307
>gb|AAP37759.1| At3g24550 [Arabidopsis thaliana] gi|22136228|gb|AAM91192.1| protein
kinase-like protein [Arabidopsis thaliana]
gi|9294050|dbj|BAB02007.1| protein kinase-like protein
[Arabidopsis thaliana] gi|20260332|gb|AAM13064.1|
unknown protein [Arabidopsis thaliana]
gi|16649063|gb|AAL24383.1| protein kinase-like protein
[Arabidopsis thaliana] gi|15983765|gb|AAL10479.1|
AT3g24550/MOB24_8 [Arabidopsis thaliana]
gi|15230130|ref|NP_189098.1| protein kinase family
protein [Arabidopsis thaliana]
Length = 652
Score = 82.8 bits (203), Expect = 3e-15
Identities = 50/145 (34%), Positives = 78/145 (53%), Gaps = 12/145 (8%)
Query: 21 SSADSPAPTPATNSSL---NSPSPTPIPTPSPANSPPAPTPTPTPS-PHSDSPPAPSPDN 76
++ SP P+P+TNS+ +SP P +P PSP S P P P+PS P + SPP+P+ +
Sbjct: 38 TTPSSPPPSPSTNSTSPPPSSPLPPSLPPPSPPGSLTPPLPQPSPSAPITPSPPSPTTPS 97
Query: 77 SPSSSPSPS--PSSSPAPSPDEAADNNAISHTGIGEDGKSSGGGMSSGKKAGIAVGVIAA 134
+P S PSP+ P ++P+ S N S S G+S+G GIA+G +A
Sbjct: 98 NPRSPPSPNQGPPNTPSGSTPRTPSNTKPS------PPSDSSDGLSTGVVVGIAIGGVAI 151
Query: 135 VGVVALGAMVVKKRRQNIQRSEYGY 159
+ ++ L ++ KK+R+ E Y
Sbjct: 152 LVILTLICLLCKKKRRRRHDDEAAY 176
Score = 34.7 bits (78), Expect = 1.0
Identities = 33/110 (30%), Positives = 45/110 (40%), Gaps = 15/110 (13%)
Query: 27 APTPATNSSLNS-PSPTPIPTPSPANSPPAPTPTPT------PSPHSDSP--PAPSPDNS 77
A P+ N + S P P P PSP PP P P P S +SD P P PSP
Sbjct: 204 ASRPSDNHVVTSLPPPKP---PSPPRKPPPPPPPPAFMSSSGGSDYSDLPVLPPPSPGLV 260
Query: 78 PSSSPSPSPSSSPAPSPDEAADNNAISHTGIGEDGKSSGGGMSSGKKAGI 127
S S + + + ++ N + G G K G + SGK+ +
Sbjct: 261 LGFSKSTFTYEELSRATNGFSEANLLGQGGFGYVHK---GILPSGKEVAV 307
>emb|CAE08818.1| conserved hypothetical protein [Synechococcus sp. WH 8102]
gi|33866833|ref|NP_898392.1| hypothetical protein
SYNW2303 [Synechococcus sp. WH 8102]
Length = 2014
Score = 82.8 bits (203), Expect = 3e-15
Identities = 46/107 (42%), Positives = 66/107 (60%), Gaps = 11/107 (10%)
Query: 20 ASSADSPAPTPA-TNSSLNSPSPTPIPTPSPANSP-PAPTPTPTPSPHSDSPPAPS--PD 75
A+ +P PTP T S+ +PSP+ PTPSP+ +P P+P+ TPTPSP + P+PS P
Sbjct: 1628 ATPTPTPTPTPTPTPSATPTPSPSATPTPSPSATPTPSPSATPTPSPSATPTPSPSATPT 1687
Query: 76 NSPSSSPSPSPSSSPAPSPD-------EAADNNAISHTGIGEDGKSS 115
SPS++P+PSPS++P PSP A+D + + +G D SS
Sbjct: 1688 PSPSATPTPSPSATPTPSPSATPDPTPTASDELPVDVSDLGSDDISS 1734
Score = 78.6 bits (192), Expect = 6e-14
Identities = 37/73 (50%), Positives = 52/73 (70%), Gaps = 4/73 (5%)
Query: 25 SPAPTPATNSSLNSPSPTPIPTPSPANSP---PAPTPTPTPSPHSDSPPAPSPDNSPSSS 81
+P PTP T S+ +PSP+ PTPSP+ +P P+ TPTPTP+P P+ +P SPS++
Sbjct: 1595 TPTPTP-TPSATPTPSPSATPTPSPSATPTPSPSATPTPTPTPTPTPTPSATPTPSPSAT 1653
Query: 82 PSPSPSSSPAPSP 94
P+PSPS++P PSP
Sbjct: 1654 PTPSPSATPTPSP 1666
Score = 77.0 bits (188), Expect = 2e-13
Identities = 37/78 (47%), Positives = 51/78 (64%), Gaps = 8/78 (10%)
Query: 25 SPAPTPATN-----SSLNSPSPTPIPTPSPANSP---PAPTPTPTPSPHSDSPPAPSPDN 76
+P PTP+ S+ +PSP+ PTPSP+ +P P PTPTPTPS P+ +P
Sbjct: 1597 TPTPTPSATPTPSPSATPTPSPSATPTPSPSATPTPTPTPTPTPTPSATPTPSPSATPTP 1656
Query: 77 SPSSSPSPSPSSSPAPSP 94
SPS++P+PSPS++P PSP
Sbjct: 1657 SPSATPTPSPSATPTPSP 1674
Score = 77.0 bits (188), Expect = 2e-13
Identities = 36/74 (48%), Positives = 54/74 (72%), Gaps = 4/74 (5%)
Query: 25 SPAPTP-ATNSSLNSPSPTPIPTPSPANSP-PAPTPTPTPSPHSDSPPAPS--PDNSPSS 80
+P PTP AT + +P+PTP PTP+P+ +P P+P+ TPTPSP + P+PS P +P+
Sbjct: 1577 TPTPTPSATPTPTPTPTPTPTPTPTPSATPTPSPSATPTPSPSATPTPSPSATPTPTPTP 1636
Query: 81 SPSPSPSSSPAPSP 94
+P+P+PS++P PSP
Sbjct: 1637 TPTPTPSATPTPSP 1650
Score = 76.3 bits (186), Expect = 3e-13
Identities = 39/82 (47%), Positives = 55/82 (66%), Gaps = 8/82 (9%)
Query: 21 SSADSPAPTP-ATNSSLNSPSPTPIPTPSPANSP-----PAPTPTPTPSPHSDSPPAPS- 73
S + +P P+P AT + S +PTP PTP+P +P P+P+ TPTPSP + P+PS
Sbjct: 1609 SPSATPTPSPSATPTPSPSATPTPTPTPTPTPTPSATPTPSPSATPTPSPSATPTPSPSA 1668
Query: 74 -PDNSPSSSPSPSPSSSPAPSP 94
P SPS++P+PSPS++P PSP
Sbjct: 1669 TPTPSPSATPTPSPSATPTPSP 1690
Score = 75.9 bits (185), Expect = 4e-13
Identities = 35/75 (46%), Positives = 50/75 (66%), Gaps = 6/75 (8%)
Query: 21 SSADSPAPTPATNSSLNSPSPTPIPTPSPANSP-PAPTPTPTPSPHSDSPPAPSPDNSPS 79
S + +P PTP+ +P+PTP TP+P +P P PTPTPTPS P+ +P SPS
Sbjct: 1565 SDSTTPTPTPSV-----TPTPTPSATPTPTPTPTPTPTPTPTPSATPTPSPSATPTPSPS 1619
Query: 80 SSPSPSPSSSPAPSP 94
++P+PSPS++P P+P
Sbjct: 1620 ATPTPSPSATPTPTP 1634
Score = 70.1 bits (170), Expect = 2e-11
Identities = 35/70 (50%), Positives = 45/70 (64%), Gaps = 3/70 (4%)
Query: 27 APTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPS--PDNSPSSSPSP 84
A + T S +P+PTP TP+P S PTPTPTP+P P PS P SPS++P+P
Sbjct: 1558 ARSSGTTSDSTTPTPTPSVTPTPTPSA-TPTPTPTPTPTPTPTPTPSATPTPSPSATPTP 1616
Query: 85 SPSSSPAPSP 94
SPS++P PSP
Sbjct: 1617 SPSATPTPSP 1626
Score = 64.7 bits (156), Expect = 9e-10
Identities = 32/70 (45%), Positives = 43/70 (60%), Gaps = 3/70 (4%)
Query: 25 SPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSP 84
+P A +S S S TP PTPS P P+ TPTP+P P P+P +PS++P+P
Sbjct: 1552 TPGFVAARSSGTTSDSTTPTPTPS-VTPTPTPSATPTPTP--TPTPTPTPTPTPSATPTP 1608
Query: 85 SPSSSPAPSP 94
SPS++P PSP
Sbjct: 1609 SPSATPTPSP 1618
Score = 50.1 bits (118), Expect = 2e-05
Identities = 21/62 (33%), Positives = 36/62 (57%), Gaps = 2/62 (3%)
Query: 35 SLNSPSPTPIPTPSPANSPPAPTPTP--TPSPHSDSPPAPSPDNSPSSSPSPSPSSSPAP 92
++ +P + + PTPTP TP+P + P P+P +P+ +P+P+PS++P P
Sbjct: 1549 TIRTPGFVAARSSGTTSDSTTPTPTPSVTPTPTPSATPTPTPTPTPTPTPTPTPSATPTP 1608
Query: 93 SP 94
SP
Sbjct: 1609 SP 1610
Score = 42.4 bits (98), Expect = 0.005
Identities = 27/84 (32%), Positives = 41/84 (48%), Gaps = 5/84 (5%)
Query: 20 ASSADSPAPTPATNSSLN-SPSPTPIPTPSPANSPPAPTPTPTPSP---HSDSPPAPSPD 75
A+ SP+ TP + S +PSP+ PTPSP+ + P P+P+ TP P SD P D
Sbjct: 1668 ATPTPSPSATPTPSPSATPTPSPSATPTPSPS-ATPTPSPSATPDPTPTASDELPVDVSD 1726
Query: 76 NSPSSSPSPSPSSSPAPSPDEAAD 99
S +P + + ++ D
Sbjct: 1727 LGSDDISSLTPDAVSELTAEQVND 1750
>gb|AAS65793.1| putative protein kinase [Arabidopsis thaliana]
Length = 217
Score = 82.8 bits (203), Expect = 3e-15
Identities = 50/145 (34%), Positives = 78/145 (53%), Gaps = 12/145 (8%)
Query: 21 SSADSPAPTPATNSSL---NSPSPTPIPTPSPANSPPAPTPTPTPS-PHSDSPPAPSPDN 76
++ SP P+P+TNS+ +SP P +P PSP S P P P+PS P + SPP+P+ +
Sbjct: 31 TTPSSPPPSPSTNSTSPPPSSPLPPSLPPPSPPGSLTPPLPQPSPSAPITPSPPSPTTPS 90
Query: 77 SPSSSPSPS--PSSSPAPSPDEAADNNAISHTGIGEDGKSSGGGMSSGKKAGIAVGVIAA 134
+P S PSP+ P ++P+ S N S S G+S+G GIA+G +A
Sbjct: 91 NPRSPPSPNQGPPNTPSGSTPRTPSNTKPS------PPSDSSDGLSTGVVVGIAIGGVAI 144
Query: 135 VGVVALGAMVVKKRRQNIQRSEYGY 159
+ ++ L ++ KK+R+ E Y
Sbjct: 145 LVILTLICLLCKKKRRRRHDDEAAY 169
>ref|YP_172333.1| hypothetical protein syc1623_d [Synechococcus elongatus PCC 6301]
gi|56686591|dbj|BAD79813.1| unknown protein
[Synechococcus elongatus PCC 6301]
gi|25019697|gb|AAN71783.1| unknown [Synechococcus sp.
PCC 7942]
Length = 410
Score = 81.3 bits (199), Expect = 9e-15
Identities = 47/128 (36%), Positives = 69/128 (53%), Gaps = 16/128 (12%)
Query: 11 LLLSFLVNIASSADSPAPTPATN----------SSLNSPSPTPIPTPSPANSPPAPTPTP 60
+L++FL N+ + SP TN + +SPSP+P PTP+P P+P+P
Sbjct: 220 VLIAFLRNVGNGG-SPGTDCVTNIVAARAIDPANIASSPSPSPTPTPTPT-----PSPSP 273
Query: 61 TPSPHSDSPPAPSPDNSPSSSPSPSPSSSPAPSPDEAADNNAISHTGIGEDGKSSGGGMS 120
TPSP P P+P SP+ SPSP+P+ +P PSP A +IS I G +S G++
Sbjct: 274 TPSPSPTPTPTPTPSPSPTPSPSPTPTPTPTPSPSPAPTPVSISIASISGGGNNSTVGLN 333
Query: 121 SGKKAGIA 128
G + +A
Sbjct: 334 LGSLSLLA 341
>gb|AAP85665.1| odv-e66 [Adoxophyes orana granulovirus]
gi|32698567|ref|NP_872482.1| odv-e66 [Adoxophyes orana
granulovirus]
Length = 747
Score = 80.5 bits (197), Expect = 2e-14
Identities = 36/89 (40%), Positives = 53/89 (59%), Gaps = 5/89 (5%)
Query: 8 LVFLLLSFLVNIASSADS--PAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPH 65
++ +L F++ S D+ P P P N + P PTP PTP+P PP PTP PTP+P
Sbjct: 21 IIIILFFFIIQKISVFDNCEPTPIPPVNPPIVIPPPTPPPTPTP---PPTPTPPPTPTPP 77
Query: 66 SDSPPAPSPDNSPSSSPSPSPSSSPAPSP 94
PP P+P +P+ P+P+P+ +P P+P
Sbjct: 78 PTPPPTPTPPPTPTPPPTPTPTPTPPPTP 106
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.305 0.124 0.354
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 355,443,730
Number of Sequences: 2540612
Number of extensions: 23868500
Number of successful extensions: 1081079
Number of sequences better than 10.0: 34797
Number of HSP's better than 10.0 without gapping: 16760
Number of HSP's successfully gapped in prelim test: 19678
Number of HSP's that attempted gapping in prelim test: 385569
Number of HSP's gapped (non-prelim): 280412
length of query: 166
length of database: 863,360,394
effective HSP length: 118
effective length of query: 48
effective length of database: 563,568,178
effective search space: 27051272544
effective search space used: 27051272544
T: 11
A: 40
X1: 16 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 43 (21.9 bits)
S2: 70 (31.6 bits)
Medicago: description of AC144405.9


