

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC142094.8 - phase: 0 /pseudo
(82 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
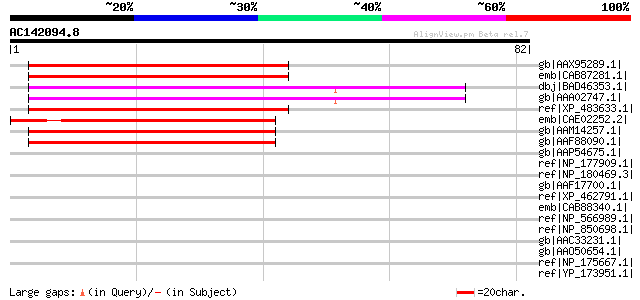
Score E
Sequences producing significant alignments: (bits) Value
gb|AAX95289.1| Rhomboid family, putative [Oryza sativa (japonica... 47 9e-05
emb|CAB87281.1| membrane protein [Arabidopsis thaliana] gi|20466... 47 1e-04
dbj|BAD46353.1| putative membrane protein [Oryza sativa (japonic... 47 1e-04
gb|AAA02747.1| membrane protein [Saccharum hybrid cultivar H65-7... 45 5e-04
ref|XP_483633.1| putative membrane protein [Oryza sativa (japoni... 45 6e-04
emb|CAE02252.2| OSJNBb0032E06.11 [Oryza sativa (japonica cultiva... 45 6e-04
gb|AAM14257.1| putative membrane protein [Arabidopsis thaliana] ... 45 6e-04
gb|AAF88090.1| T12C24.28 [Arabidopsis thaliana] gi|25402744|pir|... 45 6e-04
gb|AAP54675.1| putative membrane protein [Oryza sativa (japonica... 44 0.001
ref|NP_177909.1| rhomboid family protein [Arabidopsis thaliana] 44 0.001
ref|NP_180469.3| rhomboid family protein [Arabidopsis thaliana] 44 0.001
gb|AAF17700.1| F28K19.7 [Arabidopsis thaliana] 44 0.001
ref|XP_462791.1| putative membrane protein [Oryza sativa (japoni... 44 0.001
emb|CAB88340.1| putative protein [Arabidopsis thaliana] gi|11357... 43 0.002
ref|NP_566989.1| rhomboid family protein [Arabidopsis thaliana] 43 0.002
ref|NP_850698.1| rhomboid family protein [Arabidopsis thaliana] 43 0.002
gb|AAC33231.1| hypothetical protein [Arabidopsis thaliana] gi|74... 42 0.003
gb|AAO50654.1| putative membrane protein [Arabidopsis thaliana] ... 42 0.005
ref|NP_175667.1| rhomboid family protein [Arabidopsis thaliana] ... 40 0.015
ref|YP_173951.1| PTS system, fructose specific enzyme II, C comp... 39 0.026
>gb|AAX95289.1| Rhomboid family, putative [Oryza sativa (japonica cultivar-group)]
Length = 374
Score = 47.4 bits (111), Expect = 9e-05
Identities = 23/41 (56%), Positives = 30/41 (73%)
Query: 4 LSIGMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCK 44
L++ V V NL +GI+P V+NF IGGLI GFLLGFV+ +
Sbjct: 210 LTLVFVIVVNLALGILPRVDNFAHIGGLISGFLLGFVMFIR 250
>emb|CAB87281.1| membrane protein [Arabidopsis thaliana] gi|20466143|gb|AAM19993.1|
AT5g07250/T28J14_190 [Arabidopsis thaliana]
gi|15240744|ref|NP_196342.1| rhomboid family protein
[Arabidopsis thaliana] gi|16648762|gb|AAL25572.1|
AT5g07250/T28J14_190 [Arabidopsis thaliana]
gi|11358506|pir||T48496 membrane protein - Arabidopsis
thaliana
Length = 346
Score = 47.0 bits (110), Expect = 1e-04
Identities = 21/41 (51%), Positives = 30/41 (72%)
Query: 4 LSIGMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCK 44
L++ V + NL IGI+P V+NF +GG + GFLLGF+LL +
Sbjct: 230 LTLLFVILINLAIGILPHVDNFAHVGGFVTGFLLGFILLAR 270
>dbj|BAD46353.1| putative membrane protein [Oryza sativa (japonica cultivar-group)]
gi|52076044|dbj|BAD46497.1| putative membrane protein
[Oryza sativa (japonica cultivar-group)]
Length = 323
Score = 47.0 bits (110), Expect = 1e-04
Identities = 27/80 (33%), Positives = 42/80 (51%), Gaps = 11/80 (13%)
Query: 4 LSIGMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCKKDPFVL-----------PD 52
+++ + NL IGI+P +NF IGG + GFLLGFVLL + + P
Sbjct: 209 ITLLFIIAINLAIGILPHADNFAHIGGFVTGFLLGFVLLARPQFGWMERHELPQTNQPPK 268
Query: 53 QKLHKRCLPIICFILLSTGW 72
K ++ L ++ F+LL G+
Sbjct: 269 YKAYQYVLWVVAFVLLLVGF 288
>gb|AAA02747.1| membrane protein [Saccharum hybrid cultivar H65-7052]
Length = 325
Score = 45.1 bits (105), Expect = 5e-04
Identities = 27/80 (33%), Positives = 41/80 (50%), Gaps = 11/80 (13%)
Query: 4 LSIGMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCKKDPFVL-----------PD 52
+++ + NL IGI+P V+NF IGG GFLLGFVLL + + P
Sbjct: 211 ITLLFIIALNLAIGILPHVDNFAHIGGFATGFLLGFVLLARPQFSWMERHELPQTNQPPK 270
Query: 53 QKLHKRCLPIICFILLSTGW 72
K ++ L ++ +LL G+
Sbjct: 271 YKAYQYILWVVALVLLLVGF 290
>ref|XP_483633.1| putative membrane protein [Oryza sativa (japonica cultivar-group)]
gi|42408095|dbj|BAD09236.1| putative membrane protein
[Oryza sativa (japonica cultivar-group)]
Length = 323
Score = 44.7 bits (104), Expect = 6e-04
Identities = 20/41 (48%), Positives = 28/41 (67%)
Query: 4 LSIGMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCK 44
+++ + NL IGI+P +NF IGG + GFLLGFVLL +
Sbjct: 209 ITLLFIIAINLAIGILPHADNFAHIGGFVTGFLLGFVLLAR 249
>emb|CAE02252.2| OSJNBb0032E06.11 [Oryza sativa (japonica cultivar-group)]
gi|50928039|ref|XP_473547.1| OSJNBb0032E06.11 [Oryza
sativa (japonica cultivar-group)]
Length = 342
Score = 44.7 bits (104), Expect = 6e-04
Identities = 24/42 (57%), Positives = 28/42 (66%), Gaps = 2/42 (4%)
Query: 1 MFNLSIGMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLL 42
M NL I + NL +GI+P V+NF IGG GFLLGFVLL
Sbjct: 231 MVNLII--IAAINLALGILPRVDNFAHIGGFATGFLLGFVLL 270
>gb|AAM14257.1| putative membrane protein [Arabidopsis thaliana]
gi|17529034|gb|AAL38727.1| putative membrane protein
[Arabidopsis thaliana] gi|15222058|ref|NP_172735.1|
rhomboid family protein [Arabidopsis thaliana]
Length = 307
Score = 44.7 bits (104), Expect = 6e-04
Identities = 21/39 (53%), Positives = 27/39 (68%)
Query: 4 LSIGMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLL 42
LS + NL IG++P V+NF IGGL+ GF LGF+LL
Sbjct: 193 LSFLFIIAINLAIGLLPWVDNFAHIGGLLTGFCLGFILL 231
>gb|AAF88090.1| T12C24.28 [Arabidopsis thaliana] gi|25402744|pir||G86260 protein
T12C24.28 [imported] - Arabidopsis thaliana
Length = 302
Score = 44.7 bits (104), Expect = 6e-04
Identities = 21/39 (53%), Positives = 27/39 (68%)
Query: 4 LSIGMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLL 42
LS + NL IG++P V+NF IGGL+ GF LGF+LL
Sbjct: 188 LSFLFIIAINLAIGLLPWVDNFAHIGGLLTGFCLGFILL 226
>gb|AAP54675.1| putative membrane protein [Oryza sativa (japonica cultivar-group)]
gi|37536172|ref|NP_922388.1| putative membrane protein
[Oryza sativa (japonica cultivar-group)]
gi|22122915|gb|AAM92298.1| putative membrane protein
[Oryza sativa (japonica cultivar-group)]
Length = 329
Score = 43.9 bits (102), Expect = 0.001
Identities = 20/37 (54%), Positives = 26/37 (70%)
Query: 8 MVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCK 44
+V NL IGI+P V+NF IGG + GFLLGF+ L +
Sbjct: 217 IVIAINLAIGILPHVDNFAHIGGFLTGFLLGFIFLMR 253
>ref|NP_177909.1| rhomboid family protein [Arabidopsis thaliana]
Length = 319
Score = 43.9 bits (102), Expect = 0.001
Identities = 20/32 (62%), Positives = 23/32 (71%)
Query: 13 NLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCK 44
N IG +P ++NF IGG I GFLLGFVLL K
Sbjct: 227 NFLIGFLPFIDNFANIGGFISGFLLGFVLLFK 258
>ref|NP_180469.3| rhomboid family protein [Arabidopsis thaliana]
Length = 389
Score = 43.9 bits (102), Expect = 0.001
Identities = 30/96 (31%), Positives = 45/96 (46%), Gaps = 18/96 (18%)
Query: 4 LSIGMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCKKD---------------PF 48
L++ + NL +GI+P V+NF +GG GFLLGFV L + P
Sbjct: 225 LTLIFIIAINLAVGILPHVDNFAHLGGFTSGFLLGFVFLIRPQYGYFNQRNNPRGYAAPS 284
Query: 49 VLPDQKLHKRCLPIICFILLSTGWTG---VLTKGSE 81
K ++ L I +LL G+T VL +G++
Sbjct: 285 AKSKHKPYQYVLWITSLVLLIAGYTAGLVVLLRGTD 320
>gb|AAF17700.1| F28K19.7 [Arabidopsis thaliana]
Length = 735
Score = 43.9 bits (102), Expect = 0.001
Identities = 20/32 (62%), Positives = 23/32 (71%)
Query: 13 NLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCK 44
N IG +P ++NF IGG I GFLLGFVLL K
Sbjct: 241 NFLIGFLPFIDNFANIGGFISGFLLGFVLLFK 272
>ref|XP_462791.1| putative membrane protein [Oryza sativa (japonica cultivar-group)]
Length = 393
Score = 43.5 bits (101), Expect = 0.001
Identities = 19/41 (46%), Positives = 29/41 (70%)
Query: 4 LSIGMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCK 44
L++ ++ + NL +GI+P V+NF +GG GF LGFVLL +
Sbjct: 245 LTLVIIILINLAVGILPHVDNFAHLGGFTSGFFLGFVLLVR 285
>emb|CAB88340.1| putative protein [Arabidopsis thaliana] gi|11357824|pir||T45918
hypothetical protein F5K20.80 - Arabidopsis thaliana
Length = 361
Score = 42.7 bits (99), Expect = 0.002
Identities = 19/41 (46%), Positives = 29/41 (70%)
Query: 4 LSIGMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCK 44
+++ ++ NL +G++P V+NF IGG GFLLGFVLL +
Sbjct: 202 VTLVLIVAVNLGLGVLPGVDNFAHIGGFATGFLLGFVLLIR 242
>ref|NP_566989.1| rhomboid family protein [Arabidopsis thaliana]
Length = 270
Score = 42.7 bits (99), Expect = 0.002
Identities = 19/41 (46%), Positives = 29/41 (70%)
Query: 4 LSIGMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCK 44
+++ ++ NL +G++P V+NF IGG GFLLGFVLL +
Sbjct: 111 VTLVLIVAVNLGLGVLPGVDNFAHIGGFATGFLLGFVLLIR 151
>ref|NP_850698.1| rhomboid family protein [Arabidopsis thaliana]
Length = 394
Score = 42.7 bits (99), Expect = 0.002
Identities = 19/41 (46%), Positives = 29/41 (70%)
Query: 4 LSIGMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCK 44
+++ ++ NL +G++P V+NF IGG GFLLGFVLL +
Sbjct: 235 VTLVLIVAVNLGLGVLPGVDNFAHIGGFATGFLLGFVLLIR 275
>gb|AAC33231.1| hypothetical protein [Arabidopsis thaliana] gi|7487872|pir||T02735
hypothetical protein At2g29050 [imported] - Arabidopsis
thaliana
Length = 372
Score = 42.4 bits (98), Expect = 0.003
Identities = 19/41 (46%), Positives = 27/41 (65%)
Query: 4 LSIGMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCK 44
L++ + NL +GI+P V+NF +GG GFLLGFV L +
Sbjct: 279 LTLIFIIAINLAVGILPHVDNFAHLGGFTSGFLLGFVFLIR 319
>gb|AAO50654.1| putative membrane protein [Arabidopsis thaliana]
gi|28393358|gb|AAO42103.1| putative membrane protein
[Arabidopsis thaliana] gi|15221690|ref|NP_176500.1|
rhomboid family protein [Arabidopsis thaliana]
gi|12323258|gb|AAG51610.1| membrane protein, putative;
61952-60281 [Arabidopsis thaliana]
gi|25404407|pir||F96656 probable membrane protein
F16M19.4 [imported] - Arabidopsis thaliana
Length = 317
Score = 41.6 bits (96), Expect = 0.005
Identities = 18/41 (43%), Positives = 28/41 (67%)
Query: 4 LSIGMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCK 44
+++ + NL +G++P V+NF IGG + GF LGFVLL +
Sbjct: 206 ITLLFIIAINLALGMLPRVDNFAHIGGFLTGFCLGFVLLVR 246
>ref|NP_175667.1| rhomboid family protein [Arabidopsis thaliana]
gi|5903047|gb|AAD55606.1| F6D8.20 [Arabidopsis thaliana]
gi|25405571|pir||E96566 F6D8.20 [imported] - Arabidopsis
thaliana
Length = 309
Score = 40.0 bits (92), Expect = 0.015
Identities = 17/41 (41%), Positives = 28/41 (67%)
Query: 4 LSIGMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCK 44
+++ ++ V NL++G +P V+N GG + GF LGFVLL +
Sbjct: 204 MTLILIIVLNLSVGFLPRVDNSAHFGGFLAGFFLGFVLLLR 244
>ref|YP_173951.1| PTS system, fructose specific enzyme II, C component [Bacillus
clausii KSM-K16] gi|56908463|dbj|BAD62990.1| PTS system,
fructose specific enzyme II, C component [Bacillus
clausii KSM-K16]
Length = 361
Score = 39.3 bits (90), Expect = 0.026
Identities = 24/75 (32%), Positives = 41/75 (54%), Gaps = 4/75 (5%)
Query: 4 LSIGMVPVF--NLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCKKDPFVLPDQKLHKRCLP 61
+SI P F L IG + G IGG++ GFL+G+V+L K +P + +P
Sbjct: 73 MSIADRPAFAPGLVIGFIANEIQAGFIGGILGGFLVGYVVLAIKKYVKVPKSMI--GLMP 130
Query: 62 IICFILLSTGWTGVL 76
++ L++T ++G+L
Sbjct: 131 VMIIPLIATAFSGLL 145
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.331 0.154 0.501
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 151,601,005
Number of Sequences: 2540612
Number of extensions: 5988587
Number of successful extensions: 14946
Number of sequences better than 10.0: 64
Number of HSP's better than 10.0 without gapping: 51
Number of HSP's successfully gapped in prelim test: 13
Number of HSP's that attempted gapping in prelim test: 14892
Number of HSP's gapped (non-prelim): 66
length of query: 82
length of database: 863,360,394
effective HSP length: 58
effective length of query: 24
effective length of database: 716,004,898
effective search space: 17184117552
effective search space used: 17184117552
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 68 (30.8 bits)
Medicago: description of AC142094.8


