

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140023.16 - phase: 0
(1060 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
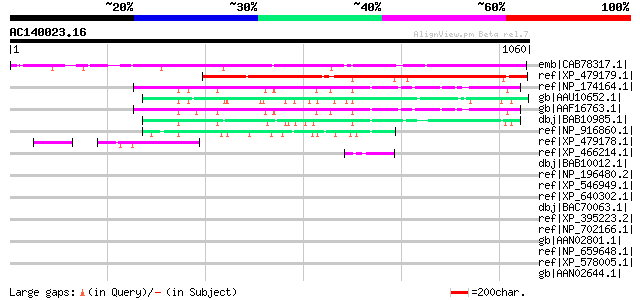
Score E
Sequences producing significant alignments: (bits) Value
emb|CAB78317.1| putative protein [Arabidopsis thaliana] gi|45862... 862 0.0
ref|XP_479179.1| homeobox transcription factor Hox7-like protein... 630 e-179
ref|NP_174164.1| homeobox transcription factor, putative [Arabid... 212 6e-53
gb|AAU10652.1| unknown protein [Oryza sativa (japonica cultivar-... 209 3e-52
gb|AAF16763.1| F3M18.14 [Arabidopsis thaliana] gi|25403342|pir||... 209 3e-52
dbj|BAB10985.1| unnamed protein product [Arabidopsis thaliana] g... 162 7e-38
ref|NP_916860.1| P0007F06.8 [Oryza sativa (japonica cultivar-gro... 146 4e-33
ref|XP_479178.1| unknown protein [Oryza sativa (japonica cultiva... 144 2e-32
ref|XP_466214.1| putative DDT domain-containing protein [Oryza s... 51 2e-04
dbj|BAB10012.1| unnamed protein product [Arabidopsis thaliana] 48 0.002
ref|NP_196480.2| DDT domain-containing protein [Arabidopsis thal... 48 0.002
ref|XP_546949.1| PREDICTED: similar to zonadhesin [Canis familia... 46 0.006
ref|XP_640302.1| hypothetical protein DDB0216395 [Dictyostelium ... 45 0.010
dbj|BAC70063.1| hypothetical protein [Streptomyces avermitilis M... 44 0.029
ref|XP_395223.2| PREDICTED: similar to bromodomain adjacent to z... 44 0.029
ref|NP_702166.1| coatamer protein, beta subunit, putative [Plasm... 44 0.037
gb|AAN02801.1| putative late transcription factor [lumpy skin di... 41 0.24
ref|NP_659648.1| Late transcription factor VLTF-4 [Sheeppox virus] 41 0.24
ref|XP_578005.1| PREDICTED: similar to putative DNA-directed RNA... 40 0.32
gb|AAN02644.1| putative late transcription factor [lumpy skin di... 40 0.54
>emb|CAB78317.1| putative protein [Arabidopsis thaliana] gi|4586251|emb|CAB40992.1|
putative protein [Arabidopsis thaliana]
gi|15235464|ref|NP_193011.1| expressed protein
[Arabidopsis thaliana] gi|7452049|pir||T06633
hypothetical protein T20K18.100 - Arabidopsis thaliana
Length = 1108
Score = 862 bits (2226), Expect = 0.0
Identities = 518/1128 (45%), Positives = 671/1128 (58%), Gaps = 145/1128 (12%)
Query: 3 GKGLIVPPCYSRGGKVLQREKVDATSSSSYEMVVNGNWKKNRKRKSVQELYTTGDIVNTV 62
G G P Y R L R +TSS + V ++ S Q L + I+ V
Sbjct: 36 GLGAKNPQLYDRS---LMRS---STSSRCVGVAVEERCIVGTRKASCQNLLPSSHILAKV 89
Query: 63 LLNDAPTLGSEFDSLPSGPKNYN-----SACQQDQEPVKRRKASKSAIQSHPNCNMKAPV 117
D P+LGSEFD LPSG + + S QQ Q+ ++RK S+ + +C +
Sbjct: 90 FRKDGPSLGSEFDHLPSGARKASWLGTSSVGQQKQKVARKRKISELMDHTSQDCIQE--- 146
Query: 118 ERHGMGKGLATNPNCKMKAPVKRHGMGKGLAT-----NPNCNMKAPVKRHGMGKGLAANP 172
A V +HG+GKGL T NPN +P + A P
Sbjct: 147 -----------------NATVMKHGIGKGLMTVWRVMNPNRRDVSPCV--DLLDERATLP 187
Query: 173 NSNMKAPVKRHGMGKGLMTIWRATNHDARDLPISFGSVDKDVHLTSNTKTPISVNRSQKA 232
S+ + P + + L +I + R S + +++
Sbjct: 188 QSSARNPPHQKKKQRQLASILKQKLLQKR-----------------------STEKKRRS 224
Query: 233 VTTNGESNQYATQNQLPIEKCELALDSSISDAGVDQISMLIDDEELELREIQEGSNLLIC 292
+ E N+ TQ + E CELA D + IS L+DDEELE+RE E N L C
Sbjct: 225 IHREAELNKDETQREFK-ENCELAADGEVFKETCQTISTLVDDEELEMRERHERGNPLTC 283
Query: 293 SDQLAANGMLGGSLC--------------PDVLVKFPPGDVKMKKPIHLQPWDSSPELVK 338
S ++G G LC PD+L KFPP V+M+ P L PW+SSPE VK
Sbjct: 284 SCHHPSSGSHGCFLCKGIAMRSSDSSLLFPDLLPKFPPNSVQMRMPFGLHPWNSSPESVK 343
Query: 339 KLFKDSMLLGQIHVALLTLLLSDIEVELSNGFCPHLNKSCNFLALLHSVENQEYSLDAWR 398
KLFKDS+LLG+IH++LL LLL D+E EL G +L+ SC FLALL SVE+Q LD WR
Sbjct: 344 KLFKDSLLLGKIHLSLLKLLLLDVETELERGSFSNLSISCKFLALLQSVESQILILDMWR 403
Query: 399 RSLNPLTWIEILRQVLVAAGFGSKQGAFQREGLGKEL----------------------- 435
SLN LTW E+LRQ+LVAAG+GS + A Q E L K+L
Sbjct: 404 NSLNSLTWTELLRQILVAAGYGSLKCAVQSEELSKQLASTCFVLGDRSVICGELKALARL 463
Query: 436 ---------DILVNYGLCPGTLKCELFKILSERGNNGCKVSELAKSMQIAELNLSSTTEE 486
++ YGL GTLK ELF++L+ +GNNG K+SELA + ++A LNL++ EE
Sbjct: 464 YFVIDDIHMKLMKKYGLRLGTLKGELFRMLNGQGNNGLKISELADAPEVAVLNLATVPEE 523
Query: 487 LESLIYSTLSSDITLFEKISSSAYRLRMSTVAKDDDDSQSDTEDSGSVDDELNDSDTCSS 546
E+ I STL+SDITLFEKIS S YR+R++ ++D D SQSD++DSGSVDDE +D + SS
Sbjct: 524 RENSICSTLASDITLFEKISESTYRVRVNCFSEDPDKSQSDSDDSGSVDDE-SDDCSISS 582
Query: 547 GDDFGSGSIHSNIRKLRRHNSRKAKHNKLKVYTEIDESHAGEVWLLGLMDSEYSDLKIEE 606
GD+ S + +RK++ RK K +V +EIDESH GE WLLGLM+ EYSDL +EE
Sbjct: 583 GDEIEHVSENPALRKVKCRKRRKHKSKMREVCSEIDESHPGEPWLLGLMEGEYSDLSVEE 642
Query: 607 KLNALAALTGLLSSGSSIRMKDPVKVTADCSSSIQLRGSGAKIKRSVNPI---------- 656
KL+ AL LLSSGS+IRM+D + ADC+ SI GSG KIKRS +
Sbjct: 643 KLDVFVALIDLLSSGSTIRMEDLPRAVADCAPSIYSHGSGGKIKRSSSNQYSYPRGSWVH 702
Query: 657 -EQMQCTKEVHMNSHACPVDSSLLVSKFHIQEASLEKRKVSAYSHPIQSVFLGSDRRYNR 715
++ K + +S + PVDSS +V F A L + + HP+QSV+LGSDRR+NR
Sbjct: 703 GGELYGIKALSKSSDSHPVDSSSIVGAF----AKLAGNRANNV-HPMQSVYLGSDRRFNR 757
Query: 716 YWLFLGPCNIDDPGHRRVYFESSEDGHWEVIDTEEALCALLSVLDDRGKREALLIESLER 775
YWLFLG CN +DPGHR V+FESSEDGHWEVI+ +EAL ALLSVLDDRG+REA LIESLE+
Sbjct: 758 YWLFLGTCNANDPGHRCVFFESSEDGHWEVINNKEALRALLSVLDDRGRREARLIESLEK 817
Query: 776 RQTSLCRSMSRIKVSNIGMGCMSHSDQSE----LDRVAEDSCSPVSDVD-NLNLTEITD- 829
R++ LC++M +V+ QSE D V EDS SPVSD+D NL L EI +
Sbjct: 818 RESFLCQAMLSRQVT-----------QSETAHFTDIVREDSSSPVSDIDNNLCLNEIAND 866
Query: 830 -YLPSPGAVVIEAGKKEEEQLHKWIRVQEYDSWIWNSFYLDLNVVKYGRRSYLDSLARCR 888
+ A+V E G K E+ L W +QE+D WIW +F +LN VK+ RRSYLDSL RC+
Sbjct: 867 QFSSQHAAIVFEIGSKREKSL-LWSLIQEFDDWIWANFNFNLNSVKHRRRSYLDSLTRCK 925
Query: 889 SCHDLYWRDERHCKICHMTFELDFDLEEKYAIHIAMCREKEDSNTFPNHKVLPSQIQSLK 948
SCHDLYWRDE+HCKICH TFE+D DLEE+YAIH A C KE+ +TFP+HKVL SQ+QSLK
Sbjct: 926 SCHDLYWRDEKHCKICHATFEVDIDLEERYAIHAATCMRKEECDTFPDHKVLSSQLQSLK 985
Query: 949 AAIYAIESVMPEDALVGAWRKSAHNLWIKRLRRTSTLVELLQVLADFVGAFNDSWLFQCK 1008
AA+YAIES MPEDAL+GAWRKSAH LW KRLRR+S++ E+ QV+ DFVGA N+ WL+ C
Sbjct: 986 AAVYAIESAMPEDALIGAWRKSAHRLWAKRLRRSSSVSEITQVIGDFVGAINEEWLWHCS 1045
Query: 1009 FP-DGVVEETIASFASMPHTSSALALWLVKLDAIIAPYLDRVQTQKSQ 1055
++ E I F SMP T+SA+ALWLVKLD +IAPY+++ ++ Q
Sbjct: 1046 DQGQTLMGEIINCFPSMPQTTSAIALWLVKLDTLIAPYVEKAPPERDQ 1093
>ref|XP_479179.1| homeobox transcription factor Hox7-like protein [Oryza sativa
(japonica cultivar-group)] gi|33146626|dbj|BAC79914.1|
homeobox transcription factor Hox7-like protein [Oryza
sativa (japonica cultivar-group)]
gi|33146880|dbj|BAC79878.1| homeobox transcription factor
Hox7-like protein [Oryza sativa (japonica
cultivar-group)]
Length = 706
Score = 630 bits (1625), Expect = e-179
Identities = 347/696 (49%), Positives = 448/696 (63%), Gaps = 50/696 (7%)
Query: 394 LDAWRRSLNPLTWIEILRQVLVAAGFGSKQGAFQREGLGKELDILVNYGLCPGTLKCELF 453
++ W +SLN LTW+EILRQVLVA+GFGSK R+ KE + +V YGL P TLK ELF
Sbjct: 1 MNFWIKSLNSLTWVEILRQVLVASGFGSKHHMLNRDFFNKEKNQMVKYGLRPRTLKGELF 60
Query: 454 KILSERGNNGCKVSELAKSMQIAELNLSSTTEELESLIYSTLSSDITLFEKISSSAYRLR 513
+LS++G+ G KVSELAKS +I +L++SST E+E LIYSTLSSDITLFEKI+ SAYRLR
Sbjct: 61 ALLSKKGSGGLKVSELAKSPEIVDLSISST--EIEQLIYSTLSSDITLFEKIAPSAYRLR 118
Query: 514 MSTVAKDDDDSQSDTEDSGSVDDELNDSDTCSSGDDFGSGSIHSNIRKLRRHNSRKAKHN 573
+ K +DS SDTEDSGSVDD + S D S + ++ R + H
Sbjct: 119 VDPRIKGKEDSGSDTEDSGSVDDHSDASSGADESDGSHEMSFSEHEHRILRRKWKNG-HE 177
Query: 574 KLKVYTEIDESHAGEVWLLGLMDSEYSDLKIEEKLNALAALTGLLSSGSSI-RMKDPVKV 632
+ +EIDES++GE WLLGLM+ EYSDL I+EKL+ L AL ++S S R+++P +V
Sbjct: 178 NVNRCSEIDESYSGERWLLGLMEGEYSDLSIDEKLDCLVALMDVVSGADSAPRLEEPSRV 237
Query: 633 TADCSSSIQLRGSGAKIKRSVNPIEQMQCTKEVHMNSHACPVDSSLLV---SKFHIQEAS 689
+ Q SG KIK+S T+ + +S C S + S H Q S
Sbjct: 238 VPSIPRA-QPHVSGGKIKKS---------TRNICQSSDECFNASGSMYGLDSSMHEQSRS 287
Query: 690 LEKRKVSAYS----------HPIQSVFLGSDRRYNRYWLFLGPCNIDDPGHRRVYFESSE 739
L R AYS H Q V LGSDRRYN YWLFLGPC DDPGHRRVYFESSE
Sbjct: 288 LRSRDYVAYSGRNDTSTGVAHQPQVVLLGSDRRYNNYWLFLGPCRADDPGHRRVYFESSE 347
Query: 740 DGHWEVIDTEEALCALLSVLDDRGKREALLIESLERRQTSLCRSM-----SRIKVSNIGM 794
DGHWEVID+ + L +LL+ LD RG REA L+ S+++RQT L +M +R V
Sbjct: 348 DGHWEVIDSPQELLSLLASLDSRGTREAYLLASMKKRQTCLFEAMKKHYENRDAVQPAMP 407
Query: 795 GCMSHSDQSELDRVAE-----DSCSPVSDVDNLNL--TEITDYLPSPGAVVIEAGKKEEE 847
SHS+ S D + D SP SD+DN ++ + + + A+ IE G++ +E
Sbjct: 408 SDTSHSETSSGDGASPKLSSGDGASPTSDIDNASVPTNPAENMINASSAIAIEVGRRGDE 467
Query: 848 QLHKWIRVQEYDSWIWNSFYLDLNVVKYGRRSYLDSLARCRSCHDLYWRDERHCKICHMT 907
++ KW R Q +D WIW SFY L VK G++S+ +SL RC SCHDLYWRDE+HC+ICH T
Sbjct: 468 KILKWERSQTFDKWIWTSFYSCLTAVKCGKKSFKESLVRCESCHDLYWRDEKHCRICHST 527
Query: 908 FELDFDLEEKYAIHIAMCREKEDSNTFPNHKVLPSQIQSLKAAIYAIESVMPEDALVGAW 967
FE+ FDLEE+YAIH+A CR+ ED PNHKVLPSQ+Q+LKAAI+AIE+ MPE A G W
Sbjct: 528 FEVSFDLEERYAIHVATCRDPEDVYDVPNHKVLPSQLQALKAAIHAIEAHMPEAAFAGLW 587
Query: 968 RKSAHNLWIKRLRRTSTLVELLQVLADFVGAFNDSWLFQ-------CKFPDGVVEETIAS 1020
KS+H LW+KRLRRTS+L ELLQVL DFVGA ++ WL++ C + D +V
Sbjct: 588 MKSSHKLWVKRLRRTSSLAELLQVLVDFVGAMDEDWLYKSSSSVSFCSYLDDIV----IY 643
Query: 1021 FASMPHTSSALALWLVKLDAIIAPYLDRVQTQKSQG 1056
F +MP T+SA+ALW+VKLDA+I PYL+R + ++ G
Sbjct: 644 FQTMPQTTSAVALWVVKLDALITPYLERADSDRALG 679
>ref|NP_174164.1| homeobox transcription factor, putative [Arabidopsis thaliana]
Length = 1703
Score = 212 bits (539), Expect = 6e-53
Identities = 247/985 (25%), Positives = 399/985 (40%), Gaps = 220/985 (22%)
Query: 254 ELALDSSISDAGVDQISMLIDDEELELREIQEGSNLLICSDQLAANGMLGGSLCPDVLVK 313
+LA++ + + + LI+DE+LEL E+ S L QL + + + D L
Sbjct: 467 KLAIEKATARRIAKESMDLIEDEQLELMELAAISKGLPSVLQLDHDTLQNLEVYRDSLST 526
Query: 314 FPPGDVKMKKPIHLQPWDSSPELVKKLFK-----------------------------DS 344
FPP +++K P + PW S E V L DS
Sbjct: 527 FPPKSLQLKMPFAISPWKDSDETVGNLLMVWRFLISFSDVLDLWPFTLDEFIQAFHDYDS 586
Query: 345 MLLGQIHVALLTLLLSDIE-------VELSNGFCPHLNKSCNFLALLHSVENQEYSLDAW 397
LLG+IHV LL ++ D+E + N N ++ + + +W
Sbjct: 587 RLLGEIHVTLLRSIIRDVEDVARTPFSGIGNNQYTTANPEGGHPQIVEGAYAWGFDIRSW 646
Query: 398 RRSLNPLTWIEILRQVLVAAGFGSK----------------------------QGAFQRE 429
++ LNPLTW EILRQ+ ++AGFG K G
Sbjct: 647 KKHLNPLTWPEILRQLALSAGFGPKLKKKHSRLTNTGDKDEAKGCEDVISTIRNGTAAES 706
Query: 430 GLG--KELDILV----NYGLCPGTLKCELFKILSERGNNGCKVSELAKSMQIAELNLSST 483
+E +L + L PGT+K F +LS G+ G V ELA +Q + L +T
Sbjct: 707 AFASMREKGLLAPRKSRHRLTPGTVKFAAFHVLSLEGSKGLTVLELADKIQKSGLRDLTT 766
Query: 484 TEELESLIYSTLSSDITLFEKISSSAYRLRMSTVA--KD--------------------- 520
++ E+ I L+ D+ LFE+I+ S Y +R V KD
Sbjct: 767 SKTPEASISVALTRDVKLFERIAPSTYCVRAPYVKDPKDGEAILADARKKIRAFENGFTG 826
Query: 521 ---------DDDSQSDTEDSGSVDD--------------ELN--------------DSDT 543
D+D + D ++ VDD E N +D
Sbjct: 827 PEDVNDLERDEDFEIDIDEDPEVDDLATLASASKSAVLGEANVLSGKGVDTMFCDVKADV 886
Query: 544 CSSGDDFGSGSIHSNIRKLRRHNSRKAKHNKLK-VYTEIDESHAGEVWLLGLMDSEYSDL 602
S + S S ++ + +S + K+ + V IDES+ G+ W+ GL + +Y L
Sbjct: 887 KSELEKEFSSPPPSTMKSIVPQHSERHKNTVVGGVDAVIDESNQGQSWIQGLTEGDYCHL 946
Query: 603 KIEEKLNALAALTGLLSSGSSIR---------------------------MKDPVKVTAD 635
+EE+LNAL AL G+ + G+SIR M+D +K+
Sbjct: 947 SVEERLNALVALVGIANEGNSIRTGLEDRMEAANALKKQMWAEAQLDNSCMRDVLKLDLQ 1006
Query: 636 CSSSIQLRGS-GAKIKRSV---------NPIEQMQCTK---EVHMNSHACPVDSSLLVSK 682
+S + + G I +S +P + + TK ++ + H + +L+
Sbjct: 1007 NLASSKTESTIGLPIIQSSTRERDSFDRDPSQLLDETKPLEDLSNDLHKSSAERALINQD 1066
Query: 683 FHI-QEASLEKRKVSAYS----------HPIQSVFLGSDRRYNRYWLFLGPCNIDDPGHR 731
+I QE KR S +P +S+ LG DRR+NRYW F + DP R
Sbjct: 1067 ANISQENYASKRSRSQLKSYIGHKAEEVYPYRSLPLGQDRRHNRYWHFAVSVSKSDPCSR 1126
Query: 732 RVYFESSEDGHWEVIDTEEALCALLSVLDDRGKREALLIESLERRQTSLCRSMSR-IKVS 790
++ E DG W +ID+EEA L++ LD RG RE+ L L++ + S + + IK++
Sbjct: 1127 LLFVEL-HDGKWLLIDSEEAFDILVASLDMRGIRESHLRIMLQKIEGSFKENACKDIKLA 1185
Query: 791 NIGMGCMSHSDQSELDRVAEDSCSPVSD-VDNLNLTEITDYLPSPGAVVIEAGKKEEEQL 849
+++S ++ DS SP S + N +D + + ++ ++ G+ + E
Sbjct: 1186 RNPF----LTEKSVVNHSPTDSVSPSSSAISGSN----SDSMETSTSIRVDLGRNDTENK 1237
Query: 850 HKWIRVQEYDSWIWNSFYLDLNVV--KYGRRSYLDSLARCRSCHDLYWRDERHCKICHMT 907
+ R ++ W+W Y L KYG++ + LA C +C Y + C CH
Sbjct: 1238 NLSKRFHDFQRWMWTETYSSLPSCARKYGKKRS-ELLATCDACVASYLSEYTFCSSCHQR 1296
Query: 908 FELDFDLEEKYAIHIAMCREKEDSNTFPNHKVLPSQIQSLKAAIYAIESVMPEDALVGAW 967
++ D E +A+ LP ++ LK + +E+ +P++AL W
Sbjct: 1297 LDV-VDSSEILDSGLAV-------------SPLPFGVRLLKPLLVFLEASVPDEALESFW 1342
Query: 968 RKSAHNLWIKRLRRTSTLVELLQVLADFVGAFN----DSWLFQCKFPDGVV------EET 1017
+ W RL +S+ ELLQVL A S K G + +
Sbjct: 1343 TEDQRKKWGFRLNTSSSPGELLQVLTSLESAIKKESLSSNFMSAKELLGAANAEADDQGS 1402
Query: 1018 IASFASMPHTSSALALWLVKLDAII 1042
+ +P T SA+AL L +LDA I
Sbjct: 1403 VDVLPWIPKTVSAVALRLSELDASI 1427
>gb|AAU10652.1| unknown protein [Oryza sativa (japonica cultivar-group)]
Length = 1397
Score = 209 bits (533), Expect = 3e-52
Identities = 253/1030 (24%), Positives = 411/1030 (39%), Gaps = 255/1030 (24%)
Query: 272 LIDDEELELREIQEGSNLLICSDQLAANGMLGGSLCPDVLVKFPPGDVKMKKPIHLQPWD 331
L++DE LEL E+ S L L ++ + +L +FP V++K P ++PW
Sbjct: 98 LMEDERLELMELVSRSKGLPSMLSLDSDTLQQLDSFRGMLRQFPSEIVRLKVPFSIKPWT 157
Query: 332 SSPELVKKLFK-----------------------------DSMLLGQIHVALLTLLLSDI 362
SS + + L DS LLG++HVALL ++ DI
Sbjct: 158 SSEDNIGNLLMVWKFFITFADVLGIPSFTLDEFVQSLHDYDSRLLGELHVALLKSIIKDI 217
Query: 363 E-----VELSNGFCPHLNKSCNFLALLHSVENQEYSLDAWRRSLNPLTWIEILRQVLVAA 417
E +++G N ++ + +++ AW+R LN LTW EILRQ ++A
Sbjct: 218 EDVARTPSVASGMTA--NPGGGHPQIVEGAYDWGFNILAWQRHLNLLTWPEILRQFGLSA 275
Query: 418 GFGSKQGAFQREGLGKELD-----------------ILVN---------YG--------L 443
G G + E + D VN YG L
Sbjct: 276 GLGPQLRKRNAENVNNHDDNEGRNGEDVISILRSGSAAVNAAAKMKERGYGNRRRSRHRL 335
Query: 444 CPGTLKCELFKILSERGNNGCKVSELAKSMQIAELNLSSTTEELESLIYSTLSSDITLFE 503
PGT+K F +LS G+ G + E+A+ +Q + L +T++ E+ I + LS D LFE
Sbjct: 336 TPGTVKFAAFHVLSLEGSQGLTILEVAEKIQKSGLRDLTTSKTPEASISAALSRDSKLFE 395
Query: 504 KISSSAY------------------------RLRMSTV------------AKDDDDSQSD 527
+ + S Y R+ +T+ A+ D+DS+ D
Sbjct: 396 RTAPSTYCVKTPYRKDPADSEAVLAAAREKIRVFQNTISECEEVEKDVDDAERDEDSECD 455
Query: 528 TEDSGSVDDELN--DSDTCSS---GDDFGSGSIHSNIRK-----------------LRRH 565
D DE+N + D +S D G + +I+K
Sbjct: 456 DADDDPDGDEVNIEEKDVKTSLVKAQDGGMPTAVGDIKKETNSIVNSLTTPLIHTKSSES 515
Query: 566 NSRKAKHNKLKVYT-----------------------EIDESHAGEVWLLGLMDSEYSDL 602
+S + ++V T EIDES+ GE W+ GL + +Y DL
Sbjct: 516 SSLRTLDKSVQVRTTSDLPAEISSDNHEGASDSAQDAEIDESNQGESWVQGLAEGDYCDL 575
Query: 603 KIEEKLNAL--------------AALTGLLSSGSSIRMK-------DPVKVTADCSSSIQ 641
+EE+LNAL A L L + S+++ + D + + SS +Q
Sbjct: 576 SVEERLNALVALIGVATEGNSIRAVLEERLEAASALKKQMWAEAQLDKRRSREEFSSKMQ 635
Query: 642 LRGSGAKIKRSV---NPIEQMQCT------KEVHMN----SHACPVDSSLLVSKFHI--- 685
SG +K V N + + T K+ + N ++ PVD + +
Sbjct: 636 Y-DSGMGLKTDVDQQNTLAESNLTPVHNLVKDSNGNGSLVNNELPVDQQSQPNACSVVHE 694
Query: 686 -----QEASLEKRKVSAYS----------------------HPIQSVFLGSDRRYNRYWL 718
QE S +S H +S+ LG DRR NRYW
Sbjct: 695 RNGVRQEFSANPENLSGQQYVTSEKTRSQLKSYIGHKAEQLHVYRSLPLGQDRRRNRYWQ 754
Query: 719 FLGPCNIDDPGHRRVYFESSEDGHWEVIDTEEALCALLSVLDDRGKREALLIESLERRQT 778
F + DDPG R++FE S DG+W +ID+ E AL+S LD RG RE+ L L+ +
Sbjct: 755 FSTSASPDDPGSGRIFFE-SRDGYWRLIDSIETFDALVSSLDTRGIRESHLHSMLQSIEP 813
Query: 779 SLCRSMSRIKVSNI--GMGCMSHSDQSEL--DRVAEDSCSPVSDVDNLNLTEITDYLPSP 834
+ ++ R + ++I G + + SE+ + + SP S + + Y S
Sbjct: 814 TFKEAIGRKRCASIEPSAGRVLKNGTSEIISPNHSNEFGSPCSTLSGVATDSAMAYSDS- 872
Query: 835 GAVVIEAGKKEEEQLHKWIRVQEYDSWIWN--SFYLDLNVVKYGRRSYLDSLARCRSCHD 892
IE G+ + E+ R + W+W + + +K+G++ + + C C+
Sbjct: 873 --FRIELGRNDVEKTAISERADLFIKWMWKECNNHQPTCAMKHGKKRCSELIQCCDFCYQ 930
Query: 893 LYWRDERHCKICHMTFELDFDLEEKYAIHIAMCREKEDSNTFPNHKV------LPSQIQS 946
+Y +E HC CH TF+ ++ E H + C EK T PN K+ +P ++
Sbjct: 931 IYLAEETHCASCHKTFKSIHNISE----HSSQCEEKR--RTDPNWKMQISDYSVPVGLRL 984
Query: 947 LKAAIYAIESVMPEDALVGAWRKSAHNLWIKRLRRTSTLVELLQVLADFVGAFNDSW--- 1003
LK + +E+ +P +AL W W +L TS+ E+ ++L GA +
Sbjct: 985 LKLLLATVEASVPAEALEPFWTDVYRKSWGVKLYSTSSTKEVFEMLTILEGAIRRDFLSS 1044
Query: 1004 -------LFQCKFPDGVVEETIASFAS------MPHTSSALALWLVKLDAIIAPYL-DRV 1049
L D T+ S +P T +A+ L L+ LD+ I+ L +V
Sbjct: 1045 DFETTTELLNLSTQDSASRNTVPRSGSADVLPWVPDTVAAVVLRLLDLDSAISYTLRQKV 1104
Query: 1050 QTQKSQGIGK 1059
+ K +G G+
Sbjct: 1105 GSNKERGAGE 1114
>gb|AAF16763.1| F3M18.14 [Arabidopsis thaliana] gi|25403342|pir||E86410 protein
F3M18.14 [imported] - Arabidopsis thaliana
Length = 1819
Score = 209 bits (533), Expect = 3e-52
Identities = 249/991 (25%), Positives = 399/991 (40%), Gaps = 226/991 (22%)
Query: 254 ELALDSSISDAGVDQISMLIDDEELELREIQEGSNLLICSDQLAANGMLGGSLCPDVLVK 313
+LA++ + + + LI+DE+LEL E+ S L QL + + + D L
Sbjct: 577 KLAIEKATARRIAKESMDLIEDEQLELMELAAISKGLPSVLQLDHDTLQNLEVYRDSLST 636
Query: 314 FPPGDVKMKKPIHLQPWDSSPELVKKLFK-----------------------------DS 344
FPP +++K P + PW S E V L DS
Sbjct: 637 FPPKSLQLKMPFAISPWKDSDETVGNLLMVWRFLISFSDVLDLWPFTLDEFIQAFHDYDS 696
Query: 345 MLLGQIHVALLTLLLSDIEVELSNGFCPHLNKSCNF-------------LALLHSVENQE 391
LLG+IHV LL ++ D+E F N +A S
Sbjct: 697 RLLGEIHVTLLRSIIRDVEDVARTPFSGIGNNQYTTANPEGGHPQIVEGVAFFVSAYAWG 756
Query: 392 YSLDAWRRSLNPLTWIEILRQVLVAAGFGSK----------------------------Q 423
+ + +W++ LNPLTW EILRQ+ ++AGFG K
Sbjct: 757 FDIRSWKKHLNPLTWPEILRQLALSAGFGPKLKKKHSRLTNTGDKDEAKGCEDVISTIRN 816
Query: 424 GAFQREGLG--KELDILV----NYGLCPGTLKCELFKILSERGNNGCKVSELAKSMQIAE 477
G +E +L + L PGT+K F +LS G+ G V ELA +Q +
Sbjct: 817 GTAAESAFASMREKGLLAPRKSRHRLTPGTVKFAAFHVLSLEGSKGLTVLELADKIQKSG 876
Query: 478 LNLSSTTEELESLIYSTLSSDITLFEKISSSAYRLRMSTVA--KD--------------- 520
L +T++ E+ I L+ D+ LFE+I+ S Y +R V KD
Sbjct: 877 LRDLTTSKTPEASISVALTRDVKLFERIAPSTYCVRAPYVKDPKDGEAILADARKKIRAF 936
Query: 521 ---------------DDDSQSDTEDSGSVDD--------------ELN------------ 539
D+D + D ++ VDD E N
Sbjct: 937 ENGFTGPEDVNDLERDEDFEIDIDEDPEVDDLATLASASKSAVLGEANVLSGKGVDTMFC 996
Query: 540 --DSDTCSSGDDFGSGSIHSNIRKLRRHNSRKAKHNKLK-VYTEIDESHAGEVWLLGLMD 596
+D S + S S ++ + +S + K+ + V IDES+ G+ W+ GL +
Sbjct: 997 DVKADVKSELEKEFSSPPPSTMKSIVPQHSERHKNTVVGGVDAVIDESNQGQSWIQGLTE 1056
Query: 597 SEYSDLKIEEKLNALAALTGLLSSGSSIR---------------------------MKDP 629
+Y L +EE+LNAL AL G+ + G+SIR M+D
Sbjct: 1057 GDYCHLSVEERLNALVALVGIANEGNSIRTGLEDRMEAANALKKQMWAEAQLDNSCMRDV 1116
Query: 630 VKVTADCSSSIQLRGS-GAKIKRSV---------NPIEQMQCTK---EVHMNSHACPVDS 676
+K+ +S + + G I +S +P + + TK ++ + H +
Sbjct: 1117 LKLDLQNLASSKTESTIGLPIIQSSTRERDSFDRDPSQLLDETKPLEDLSNDLHKSSAER 1176
Query: 677 SLLVSKFHI-QEASLEKRKVSAYS----------HPIQSVFLGSDRRYNRYWLFLGPCNI 725
+L+ +I QE KR S +P +S+ LG DRR+NRYW F +
Sbjct: 1177 ALINQDANISQENYASKRSRSQLKSYIGHKAEEVYPYRSLPLGQDRRHNRYWHFAVSVSK 1236
Query: 726 DDPGHRRVYFESSEDGHWEVIDTEEALCALLSVLDDRGKREALLIESLERRQTSLCRSMS 785
DP R ++ E DG W +ID+EEA L++ LD RG RE+ L L++ + S +
Sbjct: 1237 SDPCSRLLFVEL-HDGKWLLIDSEEAFDILVASLDMRGIRESHLRIMLQKIEGSFKENAC 1295
Query: 786 R-IKVSNIGMGCMSHSDQSELDRVAEDSCSPVSD-VDNLNLTEITDYLPSPGAVVIEAGK 843
+ IK++ +++S ++ DS SP S + N +D + + ++ ++ G+
Sbjct: 1296 KDIKLARNPF----LTEKSVVNHSPTDSVSPSSSAISGSN----SDSMETSTSIRVDLGR 1347
Query: 844 KEEEQLHKWIRVQEYDSWIWNSFYLDLNVV--KYGRRSYLDSLARCRSCHDLYWRDERHC 901
+ E + R ++ W+W Y L KYG++ + LA C +C Y + C
Sbjct: 1348 NDTENKNLSKRFHDFQRWMWTETYSSLPSCARKYGKKRS-ELLATCDACVASYLSEYTFC 1406
Query: 902 KICHMTFELDFDLEEKYAIHIAMCREKEDSNTFPNHKVLPSQIQSLKAAIYAIESVMPED 961
CH ++ D E +A+ LP ++ LK + +E+ +P++
Sbjct: 1407 SSCHQRLDV-VDSSEILDSGLAV-------------SPLPFGVRLLKPLLVFLEASVPDE 1452
Query: 962 ALVGAWRKSAHNLWIKRLRRTSTLVELLQVLADFVGAFN----DSWLFQCKFPDGVV--- 1014
AL W + W RL +S+ ELLQVL A S K G
Sbjct: 1453 ALESFWTEDQRKKWGFRLNTSSSPGELLQVLTSLESAIKKESLSSNFMSAKELLGAANAE 1512
Query: 1015 ---EETIASFASMPHTSSALALWLVKLDAII 1042
+ ++ +P T SA+AL L +LDA I
Sbjct: 1513 ADDQGSVDVLPWIPKTVSAVALRLSELDASI 1543
>dbj|BAB10985.1| unnamed protein product [Arabidopsis thaliana]
gi|15241428|ref|NP_199231.1| homeobox transcription
factor, putative [Arabidopsis thaliana]
Length = 1694
Score = 162 bits (409), Expect = 7e-38
Identities = 229/984 (23%), Positives = 366/984 (36%), Gaps = 244/984 (24%)
Query: 272 LIDDEELELREIQEGSNLLICSDQLAANGMLGGSLCPDVLVKFPPGDVKMKKPIHLQPWD 331
LI+DE LEL E+ + L L + D FPP VK+KKP ++PW+
Sbjct: 452 LIEDERLELMEVAALTKGLPSMLALDFETLQNLDEYRDKQAIFPPTSVKLKKPFAVKPWN 511
Query: 332 SSPELVKKLFK-----------------------------DSMLLGQIHVALLTLLLSDI 362
S E V L D L+G+IH+ LL ++ DI
Sbjct: 512 GSDENVANLLMVWRFLITFADVLGLWPFTLDEFAQAFHDYDPRLMGEIHIVLLKTIIKDI 571
Query: 363 EV---ELSNGFCPHLNKSCN----FLALLHSVENQEYSLDAWRRSLNPLTWIEILRQVLV 415
E LS G + N + N ++ + + +WR++LN TW EILRQ+ +
Sbjct: 572 EGVVRTLSTGVGANQNVAANPGGGHPHVVEGAYAWGFDIRSWRKNLNVFTWPEILRQLAL 631
Query: 416 AAGFGSK---------------------------------QGAF---QREGLGKELDILV 439
+AG G + + AF Q GL
Sbjct: 632 SAGLGPQLKKMNIRTVSVHDDNEANNSENVIFNLRKGVAAENAFAKMQERGLSNPRRS-- 689
Query: 440 NYGLCPGTLKCELFKILSERGNNGCKVSELAKSMQIAELNLSSTTEELESLIYSTLSSDI 499
+ L PGT+K F +LS G G + E+A+ +Q + L +T+ E+ + + LS D
Sbjct: 690 RHRLTPGTVKFAAFHVLSLEGEKGLNILEVAEKIQKSGLRDLTTSRTPEASVAAALSRDT 749
Query: 500 TLFEKISSSAYRLRMSTVAKDDDDSQ--------------SDTEDSGSVDDELNDSDTCS 545
LFE+++ S Y +R S KD D++ S D VDD D D S
Sbjct: 750 KLFERVAPSTYCVRAS-YRKDAGDAETIFAEARERIRAFKSGITDVEDVDDAERDED--S 806
Query: 546 SGDDFGSGSIHSNIRK-----LRRHNS-------RKAKHNKLKVYTEI------------ 581
D + N++K L+ N K + + + TE+
Sbjct: 807 ESDVGEDPEVDVNLKKEDPNPLKVENLIGVEPLLENGKLDTVPMKTELGLPLTPSLPEEM 866
Query: 582 ---------------------------DESHAGEVWLLGLMDSEYSDLK----------- 603
DES GE W+ GL++ +YS+L
Sbjct: 867 KDEKRDDTLADQSLEDAVANGEDSACFDESKLGEQWVQGLVEGDYSNLSSEERLNALVAL 926
Query: 604 -------------IEEKLNALAALTGLL----------SSGSSIRMKDPVKVTADCSSSI 640
+EE+L +AL + S IR TA +I
Sbjct: 927 IGIATEGNTIRIALEERLEVASALKKQMWGEVQLDKRWKEESLIRANYLSYPTAKPGLNI 986
Query: 641 QLRGSGAKIKRS--VNPIEQMQCTKEVHMNSHACPVDSSLLVSKFHIQEASLEKRKVSAY 698
SG + S V PI ++ + SL + + +L+ ++ Y
Sbjct: 987 ATPASGNQESSSADVTPISSQDPVSLPQIDVNNVIAGPSLQLQENVPGVENLQYQQQQGY 1046
Query: 699 S---------------------HPIQSVFLGSDRRYNRYWLFLGPCNIDDPGHRRVYFES 737
+ + +S+ LG DRR NRYW F + +DPG R++ E
Sbjct: 1047 TADRERLRAQLKAYVGYKAEELYVYRSLPLGQDRRRNRYWRFSASASRNDPGCGRIFVEL 1106
Query: 738 SEDGHWEVIDTEEALCALLSVLDDRGKREALLIESLERRQTSLCRSMSRIKVSNIGMGCM 797
+DG W +ID+EEA L+ LD RG RE+ L L + + S ++ R +N G+ +
Sbjct: 1107 -QDGRWRLIDSEEAFDYLVKSLDVRGVRESHLHFMLLKIEASFKEALRRNVAANPGVCSI 1165
Query: 798 SHSDQSELDRVAEDSCSPVSDVDNLNLTEITDYLPSPGAVVIEAGKKEEEQLHKWIRVQE 857
S S S+ AE S + ++ + N E L R
Sbjct: 1166 SSSLDSD---TAEISTTFKIELGDSNAVERCSVLQ---------------------RFHS 1201
Query: 858 YDSWIWNSFY--LDLNVVKYGRRSYLDSLARCRSCHDLYWRDERHCKICHMTFELDFDLE 915
++ W+W++ L+ KYG + CR C +L++ + C C E
Sbjct: 1202 FEKWMWDNMLHPSALSAFKYGAKQSSPLFRICRICAELHFVGDICCPSCGQMHAGPDVGE 1261
Query: 916 EKYAIHIAMCRE---KEDSNTFPNHKVL-PSQIQSLKAAIYAIESVMPEDALVGAWRKSA 971
+A +A + + D+ +L P +I+ LK + +E+ +P + L W ++
Sbjct: 1262 LCFAEQVAQLGDNLRRGDTGFILRSSILSPLRIRLLKVQLALVEASLPPEGLEAFWTENL 1321
Query: 972 HNLWIKRLRRTSTLVELLQVLADFVGAFNDSWLFQCKFPD-----GVVEETIASFAS--- 1023
W +L +S+ +L QVL A +L F G+ E +AS +
Sbjct: 1322 RKSWGMKLLSSSSHEDLYQVLTTLEAALKRDFL-SSNFETTSELLGLQEGALASDLTCGV 1380
Query: 1024 -----MPHTSSALALWLVKLDAII 1042
+P T+ +AL L D+ I
Sbjct: 1381 NVLPWIPKTAGGVALRLFDFDSSI 1404
>ref|NP_916860.1| P0007F06.8 [Oryza sativa (japonica cultivar-group)]
Length = 1790
Score = 146 bits (368), Expect = 4e-33
Identities = 174/734 (23%), Positives = 279/734 (37%), Gaps = 227/734 (30%)
Query: 272 LIDDEELELREIQEGSN-----LLICSDQLAANGMLGGSLCPDVLVKFPPGDVKMKKPIH 326
L++DE LEL E+ S L + SD L G L P FPP V++K+P
Sbjct: 608 LVEDECLELMELAAQSKGLPSMLSLDSDTLQQLDSFRGMLTP-----FPPEPVRLKEPFS 662
Query: 327 LQPWDSSPELVKKLFK-----------------------------DSMLLGQIHVALLTL 357
++PW S + V L DS LLG++H+ALL
Sbjct: 663 IKPWTVSEDNVGNLLMVWKFSITFADVLGLSSVTFDEFVQSLHDYDSRLLGELHIALLKS 722
Query: 358 LLSDIE-VELSNGFCPHLNKSCNFLALLHSVENQEYSLDAWRRSLNPLTWIEILRQVLVA 416
++ DIE V + +N + ++ +++ +W+R LN LTW EILRQ ++
Sbjct: 723 IIKDIEDVSRTPSVALAVNPAGGHPQIVEGAYAWGFNIRSWQRHLNVLTWPEILRQFALS 782
Query: 417 AGFGSKQGAFQREGL----------GKELDILVNYG------------------------ 442
AGFG + E + G+++ + G
Sbjct: 783 AGFGPQLKKRNAEDVYYRDDNEGHDGQDVISTLRNGSAAVHAAALMKERGYTHRRRSRHR 842
Query: 443 LCPGTLKCELFKILSERGNNGCKVSELAKSMQ---IAELNLSSTTEEL------------ 487
L PGT+K F +LS G+ G + E+A+ +Q + +L S T E
Sbjct: 843 LTPGTVKFAAFHVLSLEGSKGLTILEVAERIQKSGLRDLTTSKTPEASIAAALSRDTKLF 902
Query: 488 --------------------ESLIYSTLSSDITLFEKISSSAYRLRMSTVAKDDDDSQSD 527
++ S+ I F+ + S + + + A+ D+DS+ D
Sbjct: 903 ERTAPSTYCVKSPYRKDPADSEVVLSSAREKIRAFQNVISDSEAEKEANDAERDEDSECD 962
Query: 528 TEDSG----------------------------------------SVDDELNDSDTCSSG 547
D VD SD +SG
Sbjct: 963 DADDDPDGDDVNIDVGDGKDPLIGVKEQDGVPITTIVDSTKREKEKVDALTQSSDLTTSG 1022
Query: 548 DDFGSGSI----------HSNIRKLRRHNSRKAKHNKLKVYTEIDESHAGEVWLLGLMDS 597
+ S+ S +R ++ + K EIDES+ GE W+ GL +
Sbjct: 1023 KEAPKPSLGKPSSANTSSDSPVRASSEYHEVPPTDAEDK---EIDESNQGESWVHGLAEG 1079
Query: 598 EYSDLKIEEKLNALAALTGL--------------LSSGSSIRMK-------DPVKVTADC 636
+Y DL +EE+LNAL AL + L S ++++ + D + +
Sbjct: 1080 DYCDLSVEERLNALVALVSVANEGNFIRAVLEERLESANALKKQMLAEAQLDKRRSKEEF 1139
Query: 637 SSSIQLRGSGAKIKRSVNPIEQMQCTK------EVHMNSHACPVDSS---LLVSKFHIQE 687
+ +Q S +K VN + T + H + + VD++ ++ +
Sbjct: 1140 AGRVQYN-SNMNLKADVNQENATESTPTPFHNVDKHNDGNTGVVDNNNNEIIDHNSNAAN 1198
Query: 688 ASLEKR--------------------------KVSAY-SHPIQSVF------LGSDRRYN 714
AS E+ ++ AY H + +F LG DRR N
Sbjct: 1199 ASYERNGLGQDIAATPDTLSVQQYAYADKTRSQLRAYIGHRAEQLFVYRSLPLGQDRRRN 1258
Query: 715 RYWLFLGPCNIDDPGHRRVYFESSEDGHWEVIDTEEALCALLSVLDDRGKREALLIESLE 774
RYW F + +DPG R++FE DG+W V+DTEEA +L++ LD RG REA L L+
Sbjct: 1259 RYWQFSTSASPNDPGSGRIFFEC-RDGYWRVLDTEEAFDSLVASLDTRGSREAQLHSMLQ 1317
Query: 775 RRQTSLCRSMSRIK 788
R + + ++ R K
Sbjct: 1318 RIEPTFKEAIKRKK 1331
>ref|XP_479178.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|33146625|dbj|BAC79913.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
gi|33146879|dbj|BAC79877.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
Length = 330
Score = 144 bits (363), Expect = 2e-32
Identities = 89/234 (38%), Positives = 124/234 (52%), Gaps = 27/234 (11%)
Query: 179 PVKRHGMGKGLMTIWRATNHDARDLPISFGSVDKDVHLTSNTKTPIS------------- 225
PV++HG GKGLMT+W A + + +D+ L S P+
Sbjct: 89 PVRKHGAGKGLMTVWHAMYSHSSKIQDGSNFIDETGCLRS--LRPLDDCGRIEDCDDGKL 146
Query: 226 -----VNRSQKAVTTNGESNQYATQNQL-------PIEKCELALDSSISDAGVDQISMLI 273
+ R + T SN+ + P +C L++D S S L+
Sbjct: 147 IQKKVLARKKVVKRTRPPSNKRKVPSSRVTDPKKHPPMECHLSVDESQSPVLQANQVTLV 206
Query: 274 DDEELELREIQEGSNLLICSDQLAANGMLGGSLCPDVLVKFPPGDVKMKKPIHLQPWDSS 333
DDEELELRE+Q G N L CS L+++G G LC D+L +FPP VKMK+P +PW SS
Sbjct: 207 DDEELELRELQAGPNPLRCSAHLSSSGRHGCPLCKDLLSRFPPSSVKMKQPFSTRPWGSS 266
Query: 334 PELVKKLFKDSMLLGQIHVALLTLLLSDIEVELSNGFCPHLNKSCNFLALLHSV 387
PE+VKKLF+DSMLLG++HV LL LLL + E ++ F P +K C FL+ ++ V
Sbjct: 267 PEMVKKLFQDSMLLGEVHVNLLKLLLLNTERGSNDVFVPRSSKDCRFLSFVNFV 320
Score = 49.7 bits (117), Expect = 5e-04
Identities = 33/85 (38%), Positives = 41/85 (47%), Gaps = 6/85 (7%)
Query: 50 QELYTTGDIVNTVLLNDAPTLGSEFDSLP-SGPKNYNSACQ----QDQEPVKRRKASKSA 104
Q L+ I+ V D P LGSEFD LP S P + Q+Q +K+RK +
Sbjct: 17 QVLFPKDYILRKVFRKDGPPLGSEFDPLPHSAPGHLRDTTDDHFYQNQRVIKKRKIVEPT 76
Query: 105 IQ-SHPNCNMKAPVERHGMGKGLAT 128
Q S C PV +HG GKGL T
Sbjct: 77 TQRSSLPCGDNGPVRKHGAGKGLMT 101
>ref|XP_466214.1| putative DDT domain-containing protein [Oryza sativa (japonica
cultivar-group)] gi|46389867|dbj|BAD15468.1| putative
DDT domain-containing protein [Oryza sativa (japonica
cultivar-group)]
Length = 617
Score = 50.8 bits (120), Expect = 2e-04
Identities = 31/102 (30%), Positives = 48/102 (46%), Gaps = 13/102 (12%)
Query: 685 IQEASLEKRKVSAYSHPIQSVFLGSDRRYNRYWLFLGPCNIDDPGHRRVYFESSEDGHWE 744
+Q E K+S S P LG DR YNRYW F R++ ES++ W
Sbjct: 485 VQHLETEIEKLSIRSSP-----LGKDRHYNRYWFF--------RREGRLFVESADSKEWG 531
Query: 745 VIDTEEALCALLSVLDDRGKREALLIESLERRQTSLCRSMSR 786
T+E L L+S L+ +G RE L L++ + + ++ +
Sbjct: 532 YYSTKEELDVLMSSLNVKGLRERALKRQLDKLYSKISNALEK 573
>dbj|BAB10012.1| unnamed protein product [Arabidopsis thaliana]
Length = 723
Score = 48.1 bits (113), Expect = 0.002
Identities = 26/80 (32%), Positives = 40/80 (49%), Gaps = 8/80 (10%)
Query: 707 LGSDRRYNRYWLFLGPCNIDDPGHRRVYFESSEDGHWEVIDTEEALCALLSVLDDRGKRE 766
LG DR YNRYW F + R++ E+S+ W +E L AL+ L+ +G+RE
Sbjct: 608 LGKDRDYNRYWWFRS--------NGRIFVENSDSEEWGYYTAKEELDALMGSLNRKGERE 659
Query: 767 ALLIESLERRQTSLCRSMSR 786
L LE +C ++ +
Sbjct: 660 LSLYTQLEIFYDRICSTLQK 679
>ref|NP_196480.2| DDT domain-containing protein [Arabidopsis thaliana]
Length = 723
Score = 48.1 bits (113), Expect = 0.002
Identities = 26/80 (32%), Positives = 40/80 (49%), Gaps = 8/80 (10%)
Query: 707 LGSDRRYNRYWLFLGPCNIDDPGHRRVYFESSEDGHWEVIDTEEALCALLSVLDDRGKRE 766
LG DR YNRYW F + R++ E+S+ W +E L AL+ L+ +G+RE
Sbjct: 608 LGKDRDYNRYWWFRS--------NGRIFVENSDSEEWGYYTAKEELDALMGSLNRKGERE 659
Query: 767 ALLIESLERRQTSLCRSMSR 786
L LE +C ++ +
Sbjct: 660 LSLYTQLEIFYDRICSTLQK 679
>ref|XP_546949.1| PREDICTED: similar to zonadhesin [Canis familiaris]
Length = 2810
Score = 46.2 bits (108), Expect = 0.006
Identities = 25/90 (27%), Positives = 39/90 (42%)
Query: 103 SAIQSHPNCNMKAPVERHGMGKGLATNPNCKMKAPVKRHGMGKGLATNPNCNMKAPVKRH 162
S I S P AP E+H +A P + P K+H + +A P AP ++H
Sbjct: 682 SEIASVPTEKPMAPTEKHTFPSEIAAVPTKQPMTPTKKHAVPSEIAAAPTKKPMAPTEKH 741
Query: 163 GMGKGLAANPNSNMKAPVKRHGMGKGLMTI 192
+ +AA P AP ++H + T+
Sbjct: 742 TVPSEVAAVPTEKNMAPTEKHTFPSEIATV 771
Score = 38.1 bits (87), Expect = 1.6
Identities = 18/65 (27%), Positives = 30/65 (45%)
Query: 119 RHGMGKGLATNPNCKMKAPVKRHGMGKGLATNPNCNMKAPVKRHGMGKGLAANPNSNMKA 178
+H + +A+ P K AP ++H +A P P K+H + +AA P A
Sbjct: 677 KHTVPSEIASVPTEKPMAPTEKHTFPSEIAAVPTKQPMTPTKKHAVPSEIAAAPTKKPMA 736
Query: 179 PVKRH 183
P ++H
Sbjct: 737 PTEKH 741
>ref|XP_640302.1| hypothetical protein DDB0216395 [Dictyostelium discoideum]
gi|60468316|gb|EAL66324.1| DDT domain-containing protein
[Dictyostelium discoideum]
Length = 885
Score = 45.4 bits (106), Expect = 0.010
Identities = 37/129 (28%), Positives = 60/129 (45%), Gaps = 30/129 (23%)
Query: 646 GAKIKRSVNPIEQMQCTKEVHMNSHACPVDSSLLVSKFHIQEASLEKRKVSAYSHPIQSV 705
GAK K+ + P+ + + K L K I + LEK YS ++S
Sbjct: 738 GAKQKKPITPVVEKEIEK---------------LEKKLEILDEKLEK-----YSTRLES- 776
Query: 706 FLGSDRRYNRYWLFLGPCNIDDPGHRRVYFESSEDGHWEVIDTEEALCALLSVLDDRGKR 765
LG DR Y YW + + ++Y E+ E+G WE +++ L L+ LD+RG R
Sbjct: 777 -LGRDRNYRNYWYWYQLPS-------KIYVEN-ENGGWEYYSSKKELDDLIKYLDNRGIR 827
Query: 766 EALLIESLE 774
E L+ +++
Sbjct: 828 ERKLLFNIK 836
>dbj|BAC70063.1| hypothetical protein [Streptomyces avermitilis MA-4680]
gi|29828894|ref|NP_823528.1| hypothetical protein
SAV2352 [Streptomyces avermitilis MA-4680]
Length = 376
Score = 43.9 bits (102), Expect = 0.029
Identities = 29/107 (27%), Positives = 40/107 (37%), Gaps = 6/107 (5%)
Query: 81 PKNYNSACQQDQEPVKRRKASKSAIQSHPNCNMKAPVERHGMGKGLATNPNCKMKAPVKR 140
P+ A + P K+ A K+A + AP ++ GK A K AP K+
Sbjct: 244 PRTAKKAAARKAAPAKKTPAKKAAAKK------TAPAKKSAPGKTAAKKAAAKKSAPAKK 297
Query: 141 HGMGKGLATNPNCNMKAPVKRHGMGKGLAANPNSNMKAPVKRHGMGK 187
GK A AP K+ GK A + AP K+ K
Sbjct: 298 SAPGKTAAKKAAAKKSAPAKKSAPGKTAAKKAAAKKTAPAKKSAAKK 344
Score = 35.8 bits (81), Expect = 7.8
Identities = 27/95 (28%), Positives = 35/95 (36%), Gaps = 2/95 (2%)
Query: 94 PVKRRKASKSAIQSHPNCNMKAPVERHGMGKGL-ATNPNCKMKAPVKRHGMGKGLATNPN 152
P R KA+ + + P KA + K A K AP K+ GK A
Sbjct: 230 PEAREKAAPAG-EEKPRTAKKAAARKAAPAKKTPAKKAAAKKTAPAKKSAPGKTAAKKAA 288
Query: 153 CNMKAPVKRHGMGKGLAANPNSNMKAPVKRHGMGK 187
AP K+ GK A + AP K+ GK
Sbjct: 289 AKKSAPAKKSAPGKTAAKKAAAKKSAPAKKSAPGK 323
>ref|XP_395223.2| PREDICTED: similar to bromodomain adjacent to zinc finger domain,
1A isoform a [Apis mellifera]
Length = 1320
Score = 43.9 bits (102), Expect = 0.029
Identities = 35/115 (30%), Positives = 54/115 (46%), Gaps = 16/115 (13%)
Query: 680 VSKFHIQEASLEKRKVSAYSHPIQSVFLGSDRRYNRYWLFL--------------GPCNI 725
+ F I E L++ + ++ + + V+LGSDR Y RYW FL G C
Sbjct: 589 LKSFLIYENKLKELQQASRDNKMM-VYLGSDRAYRRYWRFLSIPGIFVENDEWWPGNCIS 647
Query: 726 DDPGHRRVYFESSEDGHWEVIDTEEALCALLSVLDDRGKREALLIESLERRQTSL 780
D V+++ SE W +E + AL++ L RG RE L ++ + TSL
Sbjct: 648 DGDKECPVHWKRSE-LKWSFFGKQEDIEALVNSLSKRGIREGELRNNIIQEMTSL 701
>ref|NP_702166.1| coatamer protein, beta subunit, putative [Plasmodium falciparum
3D7] gi|23497345|gb|AAN36890.1| coatamer protein, beta
subunit, putative [Plasmodium falciparum 3D7]
Length = 1394
Score = 43.5 bits (101), Expect = 0.037
Identities = 54/222 (24%), Positives = 96/222 (42%), Gaps = 31/222 (13%)
Query: 387 VENQEYSLDAWRRSLNPLTWIEILRQVL--VAAGFGSKQGAFQREGLGKELDILVNYG-- 442
+ EY + R L+ + +++IL ++ + G + ++ + I+ ++G
Sbjct: 111 ISPNEYVRGSTLRLLSKIKYLKILDPLMETITKNLGHRHSYVRKNAITCIHTIIKHHGVD 170
Query: 443 LCPGTLKCELFKILSERGNNGCKVSELAKSMQIAELNLSSTTEELESLIYST----LSSD 498
+ P +K E+ KIL G+ K S LA I L L +Y T L
Sbjct: 171 IIPNAVK-EVEKILFLEGDISTKRSALAMLTDIDPLTTLKYILSLNDQLYDTADVILLEV 229
Query: 499 ITLFEK-----ISSSAYRLRMSTVAKDDDDSQSDTEDSGSVDDELNDSDTCSSGDDFGSG 553
I LF+K + +Y L ++ KDDDD +D E++ DE +D GDD
Sbjct: 230 IHLFKKLYIPHVFDDSYLL-INDEDKDDDDENND-ENNDKESDEEDDH-----GDDHNDY 282
Query: 554 SIHSNIRKLRRH----------NSRKAKHNKLKVYTEIDESH 585
+I+SN+++ ++ ++ K N L Y +I+ H
Sbjct: 283 NINSNLKEEEKYKYNYNDPIYDENKNNKENNLLDYKKINNIH 324
>gb|AAN02801.1| putative late transcription factor [lumpy skin disease virus]
Length = 226
Score = 40.8 bits (94), Expect = 0.24
Identities = 46/190 (24%), Positives = 83/190 (43%), Gaps = 40/190 (21%)
Query: 460 GNNGCKVSELAKSMQIAELNLSSTTEELESLIYSTLSSDITLFEKISSS-----AYRLRM 514
GNNG + K+++ ++ STTE ++ + SD+ + SSS R +
Sbjct: 8 GNNG----DNFKTLEEIRAHVRSTTENMDKVADEIFPSDVKIPTTSSSSKQKKPVVRKKA 63
Query: 515 STVAK-----------DDDDSQSDTEDSGSVDDELNDSDTCSSGDDFG-------SGSIH 556
+T K +D D ++D ++ DDE ND+D + DD + +
Sbjct: 64 ATTTKKSKKIKEKTPIEDHDDENDNDN----DDENNDNDNNDNNDDINNIEADDKNNDTN 119
Query: 557 SNIRKLRRHNSRKAKHNKLKVYTEIDESHAGEVWLLGLMDSEYSDLKI--EEKLNALAAL 614
++ + ++ +K + K K T + H G++D+ SDLKI E + L L
Sbjct: 120 NSDTEENENDDKKEDNKKSKKSTNDNNQHDD-----GVIDN--SDLKIATESIIKGLKIL 172
Query: 615 TGLLSSGSSI 624
+SS S++
Sbjct: 173 NNKVSSVSTV 182
>ref|NP_659648.1| Late transcription factor VLTF-4 [Sheeppox virus]
Length = 228
Score = 40.8 bits (94), Expect = 0.24
Identities = 45/188 (23%), Positives = 81/188 (42%), Gaps = 34/188 (18%)
Query: 460 GNNGCKVSELAKSMQIAELNLSSTTEELESLIYSTLSSDITLFEKISSS-----AYRLRM 514
GNNG + K+++ ++ STTE ++ + SD+ + SSS R +
Sbjct: 8 GNNG----DNFKTLEEIRAHVRSTTENIDKVADEIFPSDVKIPTTSSSSKQKKPVVRKKA 63
Query: 515 STVAK-----------DDDDSQSDTEDSGSVDDELND-----SDTCSSGDDFGSGSIHSN 558
T K +D D ++D ++ DDE ND D + D+ + +++
Sbjct: 64 VTTTKKSKKIKEKTPIEDHDDENDNDNDNDNDDENNDDNDDNDDINNIEADYKNNDTNNS 123
Query: 559 IRKLRRHNSRKAKHNKLKVYTEIDESHAGEVWLLGLMDSEYSDLKI--EEKLNALAALTG 616
+ ++ +K + K K T + H G++D+ SDLKI E + L L
Sbjct: 124 DTEENENDDKKENNKKSKKITNDNNQHDD-----GVIDN--SDLKIATESIIKGLKILNN 176
Query: 617 LLSSGSSI 624
+SS S++
Sbjct: 177 KVSSVSTV 184
>ref|XP_578005.1| PREDICTED: similar to putative DNA-directed RNA polymerase,possible
RNA polymerase A/beta/A subunit, long PHYSPTS repeat at
C-terminus [Rattus norvegicus]
Length = 205
Score = 40.4 bits (93), Expect = 0.32
Identities = 37/154 (24%), Positives = 59/154 (38%), Gaps = 13/154 (8%)
Query: 76 SLPSGPKNYNSACQQDQEPVKRRKASKSAIQSHPNCNMKAPVERHGMGKGLATNPNCKMK 135
SL S P + S ++ +++ +++ S P AP+ H L + P
Sbjct: 27 SLHSAPISLRSVPMSLHSALRSLRSAPTSLHSAPMNLHNAPISLHSAPMNLHSAPTSLHS 86
Query: 136 APVKRHGMGKGLATNPNCNMKAPVKRHGMGKGLAANPNSNMKAPVKRHGMGKGLMTIWRA 195
AP+ H L + P AP+ H L + P S+ AP+ + L
Sbjct: 87 APISLHSAPMNLHSAPTSLHSAPISLHSAPMNLHSAPISHHSAPMSLRSVPMSL------ 140
Query: 196 TNHDARDLPISFGSVDKDVHL--TSNTKTPISVN 227
H A PIS S +H TS P+S++
Sbjct: 141 --HSA---PISLHSAPISLHSAPTSLHSAPMSLH 169
>gb|AAN02644.1| putative late transcription factor [lumpy skin disease virus]
gi|15149087|gb|AAK85037.1| LSDV076 putative late
transcription factor [lumpy skin disease virus]
gi|15150515|ref|NP_150510.1| LSDV076 putative late
transcription factor [lumpy skin disease virus]
Length = 223
Score = 39.7 bits (91), Expect = 0.54
Identities = 42/184 (22%), Positives = 77/184 (41%), Gaps = 31/184 (16%)
Query: 460 GNNGCKVSELAKSMQIAELNLSSTTEELESLIYSTLSSDITLFEKISSS----------- 508
GNNG + K+++ ++ STTE ++ + SD+ + SSS
Sbjct: 8 GNNG----DNFKTLEEIRAHVRSTTENMDKVADEIFPSDVKIPTTSSSSKQKKPVVRKKA 63
Query: 509 ------AYRLRMSTVAKDDDDSQSDTEDSGSVDDELNDSDTCSSGDDFGSGSIHSNIRKL 562
+ +++ T +D DD + D + D+ ND DD + + +S+ +
Sbjct: 64 VTTTKKSKKIKEKTPIEDHDDENDNDNDDENNDNNDNDDINNIEADDKNNDTNNSDTEE- 122
Query: 563 RRHNSRKAKHNKLKVYTEIDESHAGEVWLLGLMDSEYSDLKI--EEKLNALAALTGLLSS 620
++ +K + K K T + H +V + SDLKI E + L L +SS
Sbjct: 123 NENDDKKEDNKKSKKSTNDNNQHDDDV-------IDNSDLKIATESIIKGLKILNNKVSS 175
Query: 621 GSSI 624
S++
Sbjct: 176 VSTV 179
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.316 0.132 0.388
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,805,093,147
Number of Sequences: 2540612
Number of extensions: 77566588
Number of successful extensions: 251970
Number of sequences better than 10.0: 92
Number of HSP's better than 10.0 without gapping: 17
Number of HSP's successfully gapped in prelim test: 78
Number of HSP's that attempted gapping in prelim test: 250564
Number of HSP's gapped (non-prelim): 988
length of query: 1060
length of database: 863,360,394
effective HSP length: 139
effective length of query: 921
effective length of database: 510,215,326
effective search space: 469908315246
effective search space used: 469908315246
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 81 (35.8 bits)
Medicago: description of AC140023.16


