

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137602.4 - phase: 0
(130 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
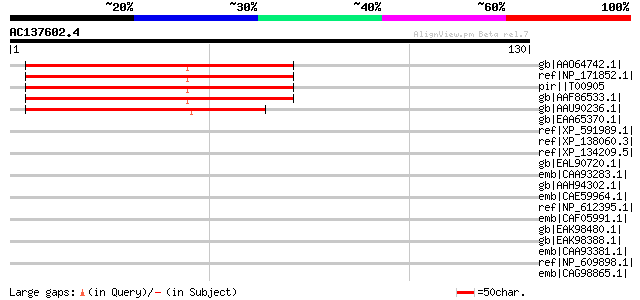
Score E
Sequences producing significant alignments: (bits) Value
gb|AAO64742.1| At1g03530 [Arabidopsis thaliana] 83 1e-15
ref|NP_171852.1| expressed protein [Arabidopsis thaliana] 83 1e-15
pir||T00905 hypothetical protein F21B7.19 - Arabidopsis thaliana 83 1e-15
gb|AAF86533.1| F21B7.15 [Arabidopsis thaliana] 83 1e-15
gb|AAU90236.1| unknown protein [Oryza sativa (japonica cultivar-... 78 4e-14
gb|EAA65370.1| hypothetical protein AN0728.2 [Aspergillus nidula... 39 0.028
ref|XP_591989.1| PREDICTED: similar to hypothetical protein BC00... 38 0.062
ref|XP_138060.3| PREDICTED: similar to hypothetical protein BC00... 37 0.081
ref|XP_134209.5| PREDICTED: similar to hypothetical protein BC00... 37 0.081
gb|EAL90720.1| NAF1 domain family [Aspergillus fumigatus Af293] ... 37 0.11
emb|CAA93283.1| Hypothetical protein F25H8.2 [Caenorhabditis ele... 37 0.11
gb|AAH94302.1| Hypothetical protein LOC306387 [Rattus norvegicus... 37 0.14
emb|CAE59964.1| Hypothetical protein CBG03454 [Caenorhabditis br... 37 0.14
ref|NP_612395.1| hypothetical protein LOC92345 [Homo sapiens] gi... 36 0.18
emb|CAF05991.1| related to nuclear assembly factor NAF1 [Neurosp... 36 0.18
gb|EAK98480.1| hypothetical protein CaO19.8124 [Candida albicans... 35 0.40
gb|EAK98388.1| hypothetical protein CaO19.494 [Candida albicans ... 35 0.40
emb|CAA93381.1| N1888 [Saccharomyces cerevisiae] gi|6324205|ref|... 35 0.53
ref|NP_609898.1| CG10341-PA [Drosophila melanogaster] gi|2294677... 34 0.69
emb|CAG98865.1| unnamed protein product [Kluyveromyces lactis NR... 34 0.90
>gb|AAO64742.1| At1g03530 [Arabidopsis thaliana]
Length = 801
Score = 83.2 bits (204), Expect = 1e-15
Identities = 39/71 (54%), Positives = 54/71 (75%), Gaps = 4/71 (5%)
Query: 5 PLIEGSILWITERQTSLGLIDEIFGQVKNPYYAVRYNSE----KGIREGTLISFVAEFVN 60
PL EGSILWITE++T LGL+DEIFG VK PYY VR+NSE +G+ +GT +SFVA+F
Sbjct: 405 PLTEGSILWITEKRTPLGLVDEIFGPVKCPYYIVRFNSESEVPEGVCQGTPVSFVADFAQ 464
Query: 61 YVMHPVSMMKR 71
++++ + K+
Sbjct: 465 HILNIKELQKK 475
>ref|NP_171852.1| expressed protein [Arabidopsis thaliana]
Length = 797
Score = 83.2 bits (204), Expect = 1e-15
Identities = 39/71 (54%), Positives = 54/71 (75%), Gaps = 4/71 (5%)
Query: 5 PLIEGSILWITERQTSLGLIDEIFGQVKNPYYAVRYNSE----KGIREGTLISFVAEFVN 60
PL EGSILWITE++T LGL+DEIFG VK PYY VR+NSE +G+ +GT +SFVA+F
Sbjct: 401 PLTEGSILWITEKRTPLGLVDEIFGPVKCPYYIVRFNSESEVPEGVCQGTPVSFVADFAQ 460
Query: 61 YVMHPVSMMKR 71
++++ + K+
Sbjct: 461 HILNIKELQKK 471
>pir||T00905 hypothetical protein F21B7.19 - Arabidopsis thaliana
Length = 681
Score = 83.2 bits (204), Expect = 1e-15
Identities = 39/71 (54%), Positives = 54/71 (75%), Gaps = 4/71 (5%)
Query: 5 PLIEGSILWITERQTSLGLIDEIFGQVKNPYYAVRYNSE----KGIREGTLISFVAEFVN 60
PL EGSILWITE++T LGL+DEIFG VK PYY VR+NSE +G+ +GT +SFVA+F
Sbjct: 285 PLTEGSILWITEKRTPLGLVDEIFGPVKCPYYIVRFNSESEVPEGVCQGTPVSFVADFAQ 344
Query: 61 YVMHPVSMMKR 71
++++ + K+
Sbjct: 345 HILNIKELQKK 355
>gb|AAF86533.1| F21B7.15 [Arabidopsis thaliana]
Length = 801
Score = 83.2 bits (204), Expect = 1e-15
Identities = 39/71 (54%), Positives = 54/71 (75%), Gaps = 4/71 (5%)
Query: 5 PLIEGSILWITERQTSLGLIDEIFGQVKNPYYAVRYNSE----KGIREGTLISFVAEFVN 60
PL EGSILWITE++T LGL+DEIFG VK PYY VR+NSE +G+ +GT +SFVA+F
Sbjct: 405 PLTEGSILWITEKRTPLGLVDEIFGPVKCPYYIVRFNSESEVPEGVCQGTPVSFVADFAQ 464
Query: 61 YVMHPVSMMKR 71
++++ + K+
Sbjct: 465 HILNIKELQKK 475
>gb|AAU90236.1| unknown protein [Oryza sativa (japonica cultivar-group)]
Length = 681
Score = 78.2 bits (191), Expect = 4e-14
Identities = 37/64 (57%), Positives = 48/64 (74%), Gaps = 4/64 (6%)
Query: 5 PLIEGSILWITERQTSLGLIDEIFGQVKNPYYAVRYNSEK----GIREGTLISFVAEFVN 60
PL EGSILWITE + LG++DE+FG VKNPYY VRYNS + I GT +SFVAEF +
Sbjct: 291 PLNEGSILWITESRIPLGIVDELFGPVKNPYYLVRYNSAEEVPADISAGTAVSFVAEFAD 350
Query: 61 YVMH 64
++++
Sbjct: 351 HILN 354
>gb|EAA65370.1| hypothetical protein AN0728.2 [Aspergillus nidulans FGSC A4]
gi|67516893|ref|XP_658332.1| hypothetical protein
AN0728_2 [Aspergillus nidulans FGSC A4]
gi|49085496|ref|XP_404865.1| hypothetical protein
AN0728.2 [Aspergillus nidulans FGSC A4]
Length = 680
Score = 38.9 bits (89), Expect = 0.028
Identities = 19/59 (32%), Positives = 37/59 (62%), Gaps = 6/59 (10%)
Query: 9 GSILWITERQTSLGLIDEIFGQVKNPYYAVRYNS-----EKGIREGTLISFVAEFVNYV 62
GS+L + +R+ + G++ E G+V+NP YAVR+ + + G+ GT++ +V + +V
Sbjct: 318 GSLLCLEDRRVA-GVVSETLGRVENPLYAVRFATTADVEKHGLSRGTVVYYVVDHSTFV 375
>ref|XP_591989.1| PREDICTED: similar to hypothetical protein BC008207, partial [Bos
taurus]
Length = 193
Score = 37.7 bits (86), Expect = 0.062
Identities = 18/42 (42%), Positives = 30/42 (70%), Gaps = 1/42 (2%)
Query: 5 PLIEGSILWITERQTSLGLIDEIFGQVKNPYYAVRYNSEKGI 46
P+ E ++++ T+RQ + G I EIFG V +P+Y +R+NS + I
Sbjct: 10 PVNEETVIFKTDRQAA-GKIFEIFGPVAHPFYVLRFNSSEHI 50
>ref|XP_138060.3| PREDICTED: similar to hypothetical protein BC008207 [Mus musculus]
Length = 535
Score = 37.4 bits (85), Expect = 0.081
Identities = 22/66 (33%), Positives = 39/66 (58%), Gaps = 9/66 (13%)
Query: 5 PLIEGSILWITERQTSLGLIDEIFGQVKNPYYAVRYNS-----EKGIREGTLISF---VA 56
P+ E ++++ ++RQ + G I EIFG V +P+Y +R+NS KGI+ + F +
Sbjct: 268 PVNEDTVIFKSDRQAA-GKIFEIFGPVAHPFYVLRFNSSDHIESKGIKINDTMYFAPSMK 326
Query: 57 EFVNYV 62
+F Y+
Sbjct: 327 DFTQYI 332
>ref|XP_134209.5| PREDICTED: similar to hypothetical protein BC008207 [Mus musculus]
Length = 729
Score = 37.4 bits (85), Expect = 0.081
Identities = 22/66 (33%), Positives = 39/66 (58%), Gaps = 9/66 (13%)
Query: 5 PLIEGSILWITERQTSLGLIDEIFGQVKNPYYAVRYNS-----EKGIREGTLISF---VA 56
P+ E ++++ ++RQ + G I EIFG V +P+Y +R+NS KGI+ + F +
Sbjct: 455 PVNEDTVIFKSDRQAA-GKIFEIFGPVAHPFYVLRFNSSDHIESKGIKINDTMYFAPSMK 513
Query: 57 EFVNYV 62
+F Y+
Sbjct: 514 DFTQYI 519
>gb|EAL90720.1| NAF1 domain family [Aspergillus fumigatus Af293]
gi|44889979|emb|CAF32097.1| hypothetical protein,
conserved [Aspergillus fumigatus]
Length = 671
Score = 37.0 bits (84), Expect = 0.11
Identities = 21/62 (33%), Positives = 36/62 (57%), Gaps = 6/62 (9%)
Query: 6 LIEGSILWITERQTSLGLIDEIFGQVKNPYYAVRYNS-----EKGIREGTLISFVAEFVN 60
L GS+L + +R T G++ E G+V+ P YAVR+ + E+G+ +G + +V E
Sbjct: 316 LESGSLLCLEDR-TVAGVVSETLGRVEKPLYAVRFPNAAAIEERGLSKGKNVYYVEEHST 374
Query: 61 YV 62
+V
Sbjct: 375 FV 376
>emb|CAA93283.1| Hypothetical protein F25H8.2 [Caenorhabditis elegans]
gi|17540000|ref|NP_501790.1| dictyostelium discoideum .
Prespore-specific protein like (43.6 kD) (4K697)
[Caenorhabditis elegans] gi|7499846|pir||T21367
hypothetical protein F25H8.2 - Caenorhabditis elegans
Length = 390
Score = 37.0 bits (84), Expect = 0.11
Identities = 15/27 (55%), Positives = 21/27 (77%)
Query: 16 ERQTSLGLIDEIFGQVKNPYYAVRYNS 42
++ +LG I +IFGQVKNP Y +R+NS
Sbjct: 198 QKGNALGQIYDIFGQVKNPQYVIRFNS 224
>gb|AAH94302.1| Hypothetical protein LOC306387 [Rattus norvegicus]
gi|67078478|ref|NP_001019943.1| hypothetical protein
LOC306387 [Rattus norvegicus]
Length = 457
Score = 36.6 bits (83), Expect = 0.14
Identities = 17/42 (40%), Positives = 30/42 (70%), Gaps = 1/42 (2%)
Query: 5 PLIEGSILWITERQTSLGLIDEIFGQVKNPYYAVRYNSEKGI 46
P+ E ++++ ++RQ + G I EIFG V +P+Y +R+NS + I
Sbjct: 187 PVNEDTVIFKSDRQAA-GKIFEIFGPVAHPFYVLRFNSSEHI 227
>emb|CAE59964.1| Hypothetical protein CBG03454 [Caenorhabditis briggsae]
Length = 405
Score = 36.6 bits (83), Expect = 0.14
Identities = 13/29 (44%), Positives = 23/29 (78%)
Query: 16 ERQTSLGLIDEIFGQVKNPYYAVRYNSEK 44
E++ ++G + +IFGQVK+P Y +R+NS +
Sbjct: 198 EKRNAIGQVYDIFGQVKSPQYVIRFNSSE 226
>ref|NP_612395.1| hypothetical protein LOC92345 [Homo sapiens]
gi|14198291|gb|AAH08207.1| Hypothetical protein BC008207
[Homo sapiens]
Length = 494
Score = 36.2 bits (82), Expect = 0.18
Identities = 22/66 (33%), Positives = 39/66 (58%), Gaps = 9/66 (13%)
Query: 5 PLIEGSILWITERQTSLGLIDEIFGQVKNPYYAVRYNS-----EKGIREGTLISF---VA 56
P+ E ++++ ++RQ + G I EIFG V +P+Y +R+NS KGI+ + F +
Sbjct: 222 PVNEETVIFKSDRQAA-GKIFEIFGPVAHPFYVLRFNSSDHIESKGIKIKETMYFAPSMK 280
Query: 57 EFVNYV 62
+F Y+
Sbjct: 281 DFTQYI 286
>emb|CAF05991.1| related to nuclear assembly factor NAF1 [Neurospora crassa]
gi|32406390|ref|XP_323808.1| hypothetical protein
[Neurospora crassa] gi|28916943|gb|EAA26677.1|
hypothetical protein [Neurospora crassa]
Length = 701
Score = 36.2 bits (82), Expect = 0.18
Identities = 22/59 (37%), Positives = 32/59 (53%), Gaps = 6/59 (10%)
Query: 9 GSILWITERQTSLGLIDEIFGQVKNPYYAVRYNSEKGIRE-----GTLISFVAEFVNYV 62
GS+L E +T +G + EI G VK+P Y + ++ I+E GT I + E NYV
Sbjct: 271 GSVL-CKEDRTVIGALAEIIGNVKSPLYTCAFANQDEIKELGLEVGTQIFYSVEHANYV 328
>gb|EAK98480.1| hypothetical protein CaO19.8124 [Candida albicans SC5314]
Length = 612
Score = 35.0 bits (79), Expect = 0.40
Identities = 14/39 (35%), Positives = 28/39 (70%)
Query: 6 LIEGSILWITERQTSLGLIDEIFGQVKNPYYAVRYNSEK 44
+++ ++ E +T +G + EIFG+V+ P Y+V++NSE+
Sbjct: 320 ILQEKSIFCFEDRTVIGPLFEIFGRVQQPVYSVKFNSEE 358
>gb|EAK98388.1| hypothetical protein CaO19.494 [Candida albicans SC5314]
Length = 612
Score = 35.0 bits (79), Expect = 0.40
Identities = 14/39 (35%), Positives = 28/39 (70%)
Query: 6 LIEGSILWITERQTSLGLIDEIFGQVKNPYYAVRYNSEK 44
+++ ++ E +T +G + EIFG+V+ P Y+V++NSE+
Sbjct: 320 ILQEKSIFCFEDRTVIGPLFEIFGRVQQPVYSVKFNSEE 358
>emb|CAA93381.1| N1888 [Saccharomyces cerevisiae] gi|6324205|ref|NP_014275.1|
Protein required for the assembly of box H/ACA snoRNPs
and for pre-rRNA processing, forms a complex with Shq1p
and interacts with H/ACA snoRNP components Nhp2p and
Cbf5p; has similarity to Gar1p and other RNA-binding
proteins; Naf1p [Saccharomyces cerevisiae]
gi|1302056|emb|CAA96005.1| unnamed protein product
[Saccharomyces cerevisiae]
gi|1730772|sp|P53919|YNM4_YEAST Hypothetical 54.9 kDa
protein in SPC98-TOM70 intergenic region
Length = 492
Score = 34.7 bits (78), Expect = 0.53
Identities = 15/39 (38%), Positives = 26/39 (66%), Gaps = 1/39 (2%)
Query: 6 LIEGSILWITERQTSLGLIDEIFGQVKNPYYAVRYNSEK 44
L EGSI + +R T +G++ E+FG ++NP+Y ++ K
Sbjct: 164 LKEGSIFCLEDR-TLIGMLTEVFGPLQNPFYRIKLPDSK 201
>ref|NP_609898.1| CG10341-PA [Drosophila melanogaster] gi|22946771|gb|AAF53693.2|
CG10341-PA [Drosophila melanogaster]
gi|15291489|gb|AAK93013.1| GH23387p [Drosophila
melanogaster]
Length = 564
Score = 34.3 bits (77), Expect = 0.69
Identities = 14/37 (37%), Positives = 25/37 (66%)
Query: 10 SILWITERQTSLGLIDEIFGQVKNPYYAVRYNSEKGI 46
++L++ + + LG + ++ GQV +P Y VR+NS K I
Sbjct: 315 TVLFLEKGRKVLGEVFDVLGQVSDPLYCVRFNSNKQI 351
>emb|CAG98865.1| unnamed protein product [Kluyveromyces lactis NRRL Y-1140]
gi|50312251|ref|XP_456157.1| unnamed protein product
[Kluyveromyces lactis]
Length = 485
Score = 33.9 bits (76), Expect = 0.90
Identities = 17/41 (41%), Positives = 28/41 (67%), Gaps = 1/41 (2%)
Query: 6 LIEGSILWITERQTSLGLIDEIFGQVKNPYYAVRYNSEKGI 46
L EGSIL + E+ T +G + E+FG ++ P+Y V ++ +K I
Sbjct: 164 LKEGSILCLQEK-TLVGPVCEVFGPLQAPFYRVSFDKDKEI 203
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.322 0.138 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 214,773,971
Number of Sequences: 2540612
Number of extensions: 7529135
Number of successful extensions: 16518
Number of sequences better than 10.0: 84
Number of HSP's better than 10.0 without gapping: 25
Number of HSP's successfully gapped in prelim test: 59
Number of HSP's that attempted gapping in prelim test: 16487
Number of HSP's gapped (non-prelim): 85
length of query: 130
length of database: 863,360,394
effective HSP length: 106
effective length of query: 24
effective length of database: 594,055,522
effective search space: 14257332528
effective search space used: 14257332528
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 67 (30.4 bits)
Medicago: description of AC137602.4


