

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137077.1 - phase: 0
(1555 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
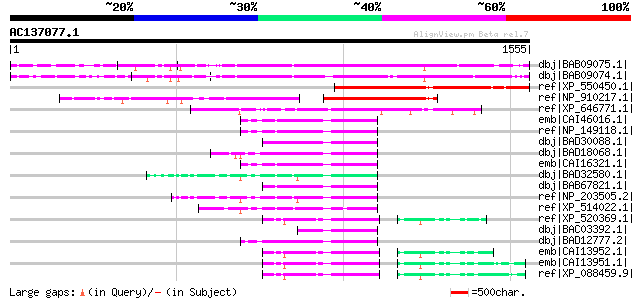
Score E
Sequences producing significant alignments: (bits) Value
dbj|BAB09075.1| unnamed protein product [Arabidopsis thaliana] g... 988 0.0
dbj|BAB09074.1| unnamed protein product [Arabidopsis thaliana] g... 901 0.0
ref|XP_550450.1| unknown protein [Oryza sativa (japonica cultiva... 518 e-145
ref|NP_910217.1| EST AU031036(E60617) corresponds to a region of... 351 1e-94
ref|XP_646771.1| hypothetical protein DDB0191321 [Dictyostelium ... 237 2e-60
emb|CAI46016.1| hypothetical protein [Homo sapiens] 142 1e-31
ref|NP_149118.1| regucalcin gene promotor region related protein... 142 1e-31
dbj|BAD30088.1| RGPR-p117 [Oryctolagus cuniculus] 140 4e-31
dbj|BAD18068.1| RGPR-p117 [Rattus norvegicus] 140 4e-31
emb|CAI16321.1| regucalcin gene promotor region related protein ... 138 2e-30
dbj|BAD32580.1| mKIAA1928 protein [Mus musculus] 138 2e-30
dbj|BAB67821.1| KIAA1928 protein [Homo sapiens] 137 2e-30
ref|NP_203505.2| regucalcin gene promotor region related protein... 137 4e-30
ref|XP_514022.1| PREDICTED: hypothetical protein XP_514022 [Pan ... 136 6e-30
ref|XP_520369.1| PREDICTED: hypothetical protein XP_520369 [Pan ... 131 2e-28
dbj|BAC03392.1| FLJ00305 protein [Homo sapiens] 131 2e-28
dbj|BAD12777.2| RGPR-p117 [Bos taurus] gi|46559376|ref|NP_996854... 129 1e-27
emb|CAI13952.1| RP11-413M3.10 [Homo sapiens] 128 1e-27
emb|CAI13951.1| RP11-413M3.10 [Homo sapiens] 128 1e-27
ref|XP_088459.9| PREDICTED: KIAA0310 [Homo sapiens] 128 1e-27
>dbj|BAB09075.1| unnamed protein product [Arabidopsis thaliana]
gi|15238109|ref|NP_199560.1| expressed protein
[Arabidopsis thaliana]
Length = 1361
Score = 988 bits (2554), Expect = 0.0
Identities = 613/1284 (47%), Positives = 761/1284 (58%), Gaps = 131/1284 (10%)
Query: 322 NTAVDWAAASDGKTEISYLQQAAQSVAGTLAETGTTESVSSWNQVSQG---NNGYLEH-- 376
N + + SD TE+ Q A TG + N S G + G L H
Sbjct: 159 NDGRGFGSYSDFFTELDATAGNLQGKADVAVATGGNLVANDTNNTSVGFEQHQGQLHHDS 218
Query: 377 ---------MVFDPQYPDWYYDTIAQEWRSLATYNSSVQSSVHGLQNGHTSTSTSSFNDD 427
++ YP W YD +W + +++S+ S +N ++ + N+
Sbjct: 219 ASGQYVDNSQSWENLYPGWKYDASTGQWFQVDGHDASMNSQ-ESYENSTSNWENVAANNS 277
Query: 428 NSLYSEYSQAGNHVSQGVGSQAVNGSWSGSHGVSQAGNYDGSHGVGSQAVNG-------- 479
+ Y S A S G+ +W+ VSQ N H V G
Sbjct: 278 DVAYQRQSTA----SAVAGTVENVSTWNQ---VSQVSNGYPEHMVFDSQYPGWYYDTIAQ 330
Query: 480 SWSGSHGVNQAGNYGGSQGVGSQAVNGSW---------SGSHGVNQAGNYGGSQGVGSQA 530
W NQA G Q Q NG+ S H V +Q Q+
Sbjct: 331 EWRSLDSYNQAFQTTG-QANDQQVQNGNSFTAVDHSRESNVHDVYDKNQILRTQKFDIQS 389
Query: 531 VNGSWSGSHGVNHQQGFDMYATEASTKIGNNTASSGNQQVHHSYGNQQVNTSSSFGSVAL 590
+GSW S+ +QQ +M+ E + G A+ + +S GNQQVN S G VA
Sbjct: 390 QHGSWDQSYYDKNQQATNMWQPENA---GAAEAAVTPASLSNSGGNQQVNNLYSTGPVA- 445
Query: 591 NNKGSFEPKAFVPHRDIAHQFNYQDTEFDNGTFAPKTFVPHGDIAQQFNYPNTKFDEQKQ 650
F+P +++G ++F+P Q N N +
Sbjct: 446 ---EQFKP-------------------YESGV---QSFIP-----QHMNVANVTQNGPMS 475
Query: 651 FSNVFAENQNSHSYSQQPIQGGLQYSYAPHAGRSSAGRPSHALVTFGFGGKLIIMKDPSA 710
FSN F Q S + Q Q +S P AGRSS GRP HALV FGFGGKLI+MKD S
Sbjct: 476 FSNGFYSRQESVDDAPQSFQSSQLFS--PSAGRSSDGRPPHALVNFGFGGKLILMKDDSG 533
Query: 711 L--TASYGSQDSVQGS-ISVLNLMEAVTGSNNSLTIGNATGDYFRALSQQSFPGPLVGGS 767
+S+GSQ GS ISVLNL E ++GS + ++G + YF L QQS PGPLVGG+
Sbjct: 534 SLQNSSFGSQKGTGGSSISVLNLAEVISGSASYSSLGENSLSYFSCLDQQSLPGPLVGGN 593
Query: 768 VGSKELYKWLDERIARCESPDMDYKKGERLRLLLSLLKIACQHYGKLRSPFGTDTILKEN 827
VGSK+L+KWLDERI CES MD+ +G+ L++LLSLL+I+CQ+YGKLRSPFG+D + KE
Sbjct: 594 VGSKDLHKWLDERILNCESSYMDFSRGKLLKMLLSLLRISCQYYGKLRSPFGSDALQKET 653
Query: 828 DAPESAVAKLFASAKVNGTEFTQYGMPSHCLQNFPSEEQMKAIASEMQNLLVSGKKMEAL 887
D+ E+AVAKLFA AK +G + Y S CLQ+ P E QM+ ASE+QNLL SG+KMEAL
Sbjct: 654 DSAEAAVAKLFAIAKEDGVQ-NGYAPISQCLQHLPPESQMQVTASEVQNLLASGRKMEAL 712
Query: 888 QRAQEGQLWGPALVLASQLGEQFYVDTVRQMALRQLVAGSPLRTLCLLIAGRPNDVFPTE 947
Q AQEG LWGPALV+A+QLG+QFYVDTV+QMALRQLV GSPLRTLCLL+AG+P +VF T
Sbjct: 713 QCAQEGHLWGPALVIAAQLGQQFYVDTVKQMALRQLVPGSPLRTLCLLVAGQPAEVFSTG 772
Query: 948 ETSISGHPGAVGMPQQSEQAGSNDMLEDWEENLAVITANRTKGDELVMMHLGDCLWKEKR 1007
TS PG+V +P Q Q G + ML+ WEENL +ITANRT DELV+ HLGDC+WKE+
Sbjct: 773 STSDISFPGSVNLPPQQPQFGCSSMLDSWEENLGIITANRTTDDELVITHLGDCMWKERG 832
Query: 1008 EITAAHICYLIAEVNFSSYSDATRLCLIGADHWTRPRTYASPEAIQRTELYEYSKLLGNS 1067
EI AAHICYLIA+ NF +YSD RLCL+GADHW PRTYASPEAIQRTELYEYSK LGNS
Sbjct: 833 EIIAAHICYLIADKNFDTYSDTARLCLVGADHWKYPRTYASPEAIQRTELYEYSKTLGNS 892
Query: 1068 QFVLHSFQPYKLIYAHMLAEVGKVSDSLKYCQAVLKSLKTGRAPEVETWKQLVLALEERI 1127
Q+ L +FQPYK++YAHMLAEVGK+S + KYCQAVLK LKTGR+PEVE WKQ V +LEERI
Sbjct: 893 QYTLLTFQPYKVMYAHMLAEVGKLSTAQKYCQAVLKCLKTGRSPEVEMWKQFVSSLEERI 952
Query: 1128 RTHQQGGYAANLAPAKLVGKLLNFFDSTAHRVVGGLPPPAPTSSQATVHGSE-QHYQHMA 1186
R HQQGGY ANL P KLVG LLNFF S HR VGG+PPPAP S++ + G+E QH Q A
Sbjct: 953 RIHQQGGYTANLHPEKLVGVLLNFFGSKTHRPVGGMPPPAPHSTKGNLQGNEYQHQQQEA 1012
Query: 1187 PRVSTSQSTMAMSSLVPSASLEPISEWTADNNRMAAKPNRSVSEPDIGRSPRQE------ 1240
+++ SQS MSSL+P AS+EP E RMA RSVSEPD GR+P QE
Sbjct: 1013 TKLAYSQSVNTMSSLMPPASVEPTHESGGSGRRMAVH-TRSVSEPDFGRTPIQEMADSSK 1071
Query: 1241 SPSPDAQGKVQVSG--GASRFSRFGFGSQLLQKTVGLVL-RSGKQAKLGEKNKFYYDEKL 1297
+ D K++ SG SRFSRFGFG + + TVG VL RS K+AKLG +N+FYYD+KL
Sbjct: 1072 EKAVDGVTKLKSSGSVAGSRFSRFGFG--IFKDTVGRVLARSSKEAKLGAENQFYYDDKL 1129
Query: 1298 KRWVEEGAEVPAEEAALPPPPPTTAAFQNGSADYNLKSALKTEGLTPNEFSSTRTSSPEL 1357
KRWVE G E PAEEAAL PPPPT AFQN S Y KS + +SS + E
Sbjct: 1130 KRWVERGVEPPAEEAAL-PPPPTIGAFQNNSLGYENKSDMIPSN---GNWSSGGPTPSEN 1185
Query: 1358 SPGMPPIPPSSNQFSARSRLGVRSRYVDTFNQNG-GSSANLFQSPSVQSVKPALPANAKF 1416
S G+PPI SNQFSAR R GVR+RYVDT+N G G+S + SPSVQ+ KP +PA AKF
Sbjct: 1186 SSGIPPISHGSNQFSARGRTGVRARYVDTYNPPGRGNSHTMIPSPSVQTAKPPIPAKAKF 1245
Query: 1417 FIP-APVPSSSEQNME-AIAESNLEDSAANENPSTSSTNDWSYHPPKHAQTMTMQRFPSA 1474
F+P AP S++Q ME A AE+ E+ +A+E ++S PP M MQR+PS
Sbjct: 1246 FVPAAPASFSNDQAMEPAAAETRQEEISADEVVASSGA------PP----PMMMQRYPSM 1295
Query: 1475 GNISKQG---QTDGNESHFSHSRRTASWSGSFNDSFSPPKMGEIKPSGAALGMPPSAFMP 1531
NI + G +G+ SRRTASWSG+FN SF+PP PS F P
Sbjct: 1296 DNIQRNGLGISVNGDNHQPPTSRRTASWSGNFNTSFTPP-------------TSPSTFKP 1342
Query: 1532 DPSSLMQGPTRSGSFGEDLQEVEL 1555
+ + S S GE+LQEVEL
Sbjct: 1343 -----VLLNSSSSSLGEELQEVEL 1361
Score = 229 bits (584), Expect = 6e-58
Identities = 179/520 (34%), Positives = 252/520 (48%), Gaps = 91/520 (17%)
Query: 1 MASNPPFQMEDQTDEDFFDKLVEDDDDVGPVKPVVIKSGN----DEGNDSDDANAFANLS 56
MAS F +EDQTDEDFFDKLV DD P + S D+ +DSDD AF+NLS
Sbjct: 1 MASASQFLLEDQTDEDFFDKLV--DDAYSPTEAQASSSVTELKFDDESDSDDIRAFSNLS 58
Query: 57 ISDVDAGAFDNSVVGESGVEVKEELGVVKSDVGLDGGGDDDGKEGNLLMGSSSVEC--DS 114
I G D ++ ++ +DV +G G++ + +V+
Sbjct: 59 IGKDPLGGGDGTL----------NEAILGNDVANEGASGSVGEDEPSSIAPEAVQFPHSD 108
Query: 115 KTELGKEEIGIGSEFTAVAPVGKSNEVASSGLMDIA-AVGESNEVALSGIKEKDWNCFSA 173
EL +E+ +EVA L + A NE + G+KE DW F A
Sbjct: 109 ARELRDDEM--------------RSEVADMPLSETAKECTIVNEPGIPGVKELDWGSFDA 154
Query: 174 D-DANGVGGFGSYSDFFSELGDQSSDFPVISHDNLNSQVS-PVIEAHNVVLNSSVDYSQY 231
D N GFGSYSDFF+EL + NL + V N+V N
Sbjct: 155 DLSVNDGRGFGSYSDFFTELDATAG--------NLQGKADVAVATGGNLVAN-------- 198
Query: 232 QGVQGYDTSFDNHTGKQGDGLNTSVNYVQYQEGGGAYDASSNLHNNGQDLSSSQNWEDLY 291
D NTSV + Q+Q G +D++S GQ + +SQ+WE+LY
Sbjct: 199 ------------------DTNNTSVGFEQHQ-GQLHHDSAS-----GQYVDNSQSWENLY 234
Query: 292 PGWKYDHITGQWYQIEDYNATTTSQQTSEANTAVDWAAASDGKTEISYLQQAAQSVAGTL 351
PGWKYD TGQW+Q++ ++A+ SQ++ E N+ +W + ++++Y +Q+ S
Sbjct: 235 PGWKYDASTGQWFQVDGHDASMNSQESYE-NSTSNWENVAANNSDVAYQRQSTAS----- 288
Query: 352 AETGTTESVSSWNQVSQGNNGYLEHMVFDPQYPDWYYDTIAQEWRSLATYNSSVQSSVHG 411
A GT E+VS+WNQVSQ +NGY EHMVFD QYP WYYDTIAQEWRSL +YN + Q++
Sbjct: 289 AVAGTVENVSTWNQVSQVSNGYPEHMVFDSQYPGWYYDTIAQEWRSLDSYNQAFQTTGQA 348
Query: 412 ----LQNGHTSTSTSSFNDDNSLYSEYSQAGNHVSQGVGSQAVNGSWSGSH--GVSQAGN 465
+QNG++ T+ + N ++ Y + +Q Q+ +GSW S+ QA N
Sbjct: 349 NDQQVQNGNSFTAVDHSRESN-VHDVYDKNQILRTQKFDIQSQHGSWDQSYYDKNQQATN 407
Query: 466 --YDGSHGVGSQAVN-GSWSGSHGVNQAGNYGGSQGVGSQ 502
+ G AV S S S G Q N + V Q
Sbjct: 408 MWQPENAGAAEAAVTPASLSNSGGNQQVNNLYSTGPVAEQ 447
>dbj|BAB09074.1| unnamed protein product [Arabidopsis thaliana]
gi|15238106|ref|NP_199559.1| expressed protein
[Arabidopsis thaliana]
Length = 1350
Score = 901 bits (2328), Expect = 0.0
Identities = 555/1233 (45%), Positives = 719/1233 (58%), Gaps = 144/1233 (11%)
Query: 365 QVSQGNNGYLEH-MVFDPQYPDWYYDTIAQEWRSLATYNSSVQSSVHGLQNGHTSTSTSS 423
Q G+ Y+++ ++ YP W YD +W + +++V S + + S ++
Sbjct: 220 QHDSGSGQYVDNSQSWENLYPGWKYDASTGQWYQVDGQDATVNSQESYINSTGNWESVAA 279
Query: 424 FNDDNSLYSEYSQAGNHVSQGVGSQAVNGSWSGSHGVSQAGNYDGSHGVGSQAVNG---- 479
N D + + + S G+ +W+ VSQ GN H V G
Sbjct: 280 DNSDVAYLKQST-----TSAMAGTAESVSTWNQ---VSQVGNGYPEHMVFDAQYPGWYYD 331
Query: 480 ----SWSGSHGVNQAGNYGGSQGVGSQAV-----------NGSWSGSHGVNQAGNYGGSQ 524
W NQA + Q V N S S + VN +Q
Sbjct: 332 TIAQEWRSLDSYNQASQTTVTGQAHDQQVQNGHARTTTYHNNSQSSVYDVNNKNQTFKAQ 391
Query: 525 GVGSQAVNGSWSGSHGVNHQQGFDMYATEASTKIGNNTASSGNQQVHHSYGNQQVNTSSS 584
Q +GSW S+ N+QQ + T +G + + + GNQQVN S
Sbjct: 392 DFAIQGQHGSWDESYYANNQQAGN---TWQPVNVGKAEPAVTSDSLSRFGGNQQVNNLYS 448
Query: 585 FGSVALNNKGSFEPKAFVPHRDIAHQFNYQDTEFDNGTFAPKTFVPHGDIAQQFNYPNTK 644
SVA +F T ++F+P Q N +
Sbjct: 449 TESVA--------------------------EQFKPNTIGAQSFIP-----QHMNVASAT 477
Query: 645 FDEQKQFSNVFAENQNSHSYSQQPIQGGLQYSYAPHAGRSSAGRPSHALVTFGFGGKLII 704
+ FSN Q S ++Q+ Q +S P GRSS RP HALV+FGFGGKLI+
Sbjct: 478 QNGPLSFSNDLYNRQQSVDHAQKSFQNNQLFS--PSVGRSSDRRPPHALVSFGFGGKLIV 535
Query: 705 MKDP--SALTASYGSQDSVQGSISVLNLMEAVTGSNNSLTIGNATGDYFRALSQQSFPGP 762
MKD S S+GSQ SI+VLNL E ++GS + + G + YFR L QQS PGP
Sbjct: 536 MKDNNGSLQNTSFGSQGIGGSSITVLNLAEVISGSASYSSPGEDSLSYFRCLHQQSLPGP 595
Query: 763 LVGGSVGSKELYKWLDERIARCESPDMDYKKGERLRLLLSLLKIACQHYGKLRSPFGTDT 822
LVGG+VGSKEL+KW+DER+ CES +MD+ +G+ L++LLSLL+I+CQ+YGKLRSPFG+D
Sbjct: 596 LVGGNVGSKELHKWIDERLLHCESSNMDFSRGKLLKMLLSLLRISCQYYGKLRSPFGSDA 655
Query: 823 ILKENDAPESAVAKLFASAKVNGTEFTQYGMPSHCLQNFPSEEQMKAIASEMQNLLVSGK 882
KE D PE+AVAKLFA AK +G + Y S CLQ+ P E QM+ ASE+QNLL SG+
Sbjct: 656 SQKETDTPEAAVAKLFAFAKKDGIQ-NGYAPISQCLQHLPPESQMQVTASEVQNLLASGR 714
Query: 883 KMEALQRAQEGQLWGPALVLASQLGEQFYVDTVRQMALRQLVAGSPLRTLCLLIAGRPND 942
KMEALQ AQEG LWGPALV+A+QLG+QFYVDTV+QMALRQL+ GSPLRTLCLL+AG+P +
Sbjct: 715 KMEALQCAQEGHLWGPALVIAAQLGDQFYVDTVKQMALRQLIPGSPLRTLCLLVAGQPAE 774
Query: 943 VFPTEETSISGHPGAVGMPQQSEQAGSNDMLEDWEENLAVITANRTKGDELVMMHLGDCL 1002
V PT S+ ML++WEENL +ITANRT D+LV++HLGD +
Sbjct: 775 VCPT--------------------GSSSSMLDNWEENLGIITANRTTDDDLVIIHLGDSM 814
Query: 1003 WKEKREITAAHICYLIAEVNFSSYSDATRLCLIGADHWTRPRTYASPEAIQRTELYEYSK 1062
WKE+ EI AAHICYLIA+ NF YS++ RLCL+GADHW PRTYASP+AIQRTELYEYSK
Sbjct: 815 WKERGEIIAAHICYLIADKNFDPYSESARLCLVGADHWKCPRTYASPDAIQRTELYEYSK 874
Query: 1063 LLGNSQFVLHSFQPYKLIYAHMLAEVGKVSDSLKYCQAVLKSLKTGRAPEVETWKQLVLA 1122
LGNSQ++L FQPYK+IYAHMLAEVGK+S + KYCQAV++ LKT R+ EVE WKQ +
Sbjct: 875 TLGNSQYILLPFQPYKIIYAHMLAEVGKLSTAQKYCQAVIRCLKTSRSSEVEMWKQFASS 934
Query: 1123 LEERIRTHQQGGYAANLAPAKLVGKLLNFFDSTAHRVVGGLPPPAPTSSQATVHGSE-QH 1181
LEERIR+HQ+GG NLAPAKLVGKLLN + G+PPPAP S+ +E QH
Sbjct: 935 LEERIRSHQEGG---NLAPAKLVGKLLN--------SLWGMPPPAPHSTTGNPQVNEYQH 983
Query: 1182 YQHMAPRVSTSQSTMAMSSLVPSASLEPISEWTADNNRMAAKPNRSVSEPDIGRSPRQE- 1240
Q A ++S SQS MSSL+P AS+EP+ EW + MAA +RSVSEPD R+P Q+
Sbjct: 984 QQQEAAKLSYSQSANTMSSLMPPASIEPVHEWGGNGRTMAAH-SRSVSEPDFSRTPIQDQ 1042
Query: 1241 -------SPSPDAQGKVQVSGGASRFSRFGFGSQLLQKTVGLVL--RSGKQAKLGEKNKF 1291
+P Q K +SRFSRFG G +L+ TVG V RS +AKLG +N+F
Sbjct: 1043 TDSSKDKAPDGVTQVKSTRKVPSSRFSRFGIG--ILKNTVGKVFPSRSSNEAKLGNENQF 1100
Query: 1292 YYDEKLKRWVEEGAEVPAEEAALPPPPPTTAAFQNGSADYNLKSALKTEGLTPN--EFSS 1349
YYD+ LKRWVE G E PAEEAAL PPPPT+ F++ S + KS +K E ++P+ +SS
Sbjct: 1101 YYDDNLKRWVERGVEPPAEEAAL-PPPPTSVPFRSNSLGHENKSEIKNE-MSPSSGSWSS 1158
Query: 1350 TRTSSPELSPGMPPIPPSSNQFSARSRLGVRSRYVDTFNQNGGSSANLFQSPSVQSVKPA 1409
+ E SPG+PP+ SNQFSAR R+GVR+RYVDT+NQ S++++QSP VQS KP
Sbjct: 1159 GSPTPSENSPGIPPVSQGSNQFSARGRMGVRARYVDTYNQ---GSSSMYQSPPVQSSKPP 1215
Query: 1410 LPANAKFFIP-APVPSSSEQNMEAIA----ESNLEDSAANENPSTSSTNDWSYHPPKHAQ 1464
+PA AKFF+P AP +++Q ME+++ + N D A + + S+ P
Sbjct: 1216 IPAKAKFFVPAAPASFANDQVMESVSAETRQENSGDEAVVGSAGAPGPSQASFQSPT-PS 1274
Query: 1465 TMTMQRFPSAGNISKQGQTDGNESHF--SHSRRTASWSGSFNDSFSPPKMGEIKPSGAAL 1522
+ MQRFPS NI + G S SRRTASWSGS N S + P+ A
Sbjct: 1275 PIAMQRFPSVDNIRRSGSGTSLNGDLPQSVSRRTASWSGSVNSS------SFMSPTSA-- 1326
Query: 1523 GMPPSAFMPDPSSLMQGPTRSGSFGEDLQEVEL 1555
S F P P + + S S GE+LQEVEL
Sbjct: 1327 ----STFRPSPLN-----SSSSSLGEELQEVEL 1350
Score = 267 bits (683), Expect = 2e-69
Identities = 223/629 (35%), Positives = 297/629 (46%), Gaps = 109/629 (17%)
Query: 1 MASNPPFQMEDQTDEDFFDKLVEDDDDVGPVKPVVIKSGN-DEGNDSDDANAFANLSISD 59
MAS F ++DQTDEDFFDKLV DD P K D+G+DSDDA AFANLS+ D
Sbjct: 1 MASTADFLLDDQTDEDFFDKLV--DDSYTPTASSSAKELKFDDGSDSDDAKAFANLSVVD 58
Query: 60 VDAGAFDNSVVGESGVEVKEELGVVKSDVGLDGGGDDDGKEGNLLMGSSSVECDSKTELG 119
V+G+ V + E G G+D EG GS E S +
Sbjct: 59 --------DVLGDGDVALNEA-----------GLGNDVANEGT--SGSVGKEEPSSSIAP 97
Query: 120 KEEIGIGSEFTAVAPVGKSNEVASSGLMDIAAVG---ESNEVALSG---IKEKDWNCFSA 173
+ + S+ + V +V S + D+A ESN V SG +KE DW F A
Sbjct: 98 EAVQFVNSDANRLRDV----DVVRSEVDDMALTETGKESNIVDGSGSPGVKEVDWGSFYA 153
Query: 174 DDA-NGVGGFGSYSDFFSELGDQSSDFPVISHDNLNSQVS-PVIEAHNVVLNSSVDYSQY 231
D + N GGFGSYSDFF+EL + N+ Q V N+V N +++ S
Sbjct: 154 DSSVNDGGGFGSYSDFFTELDATAG--------NVQGQAEVAVATGGNLVANDTINTSV- 204
Query: 232 QGVQGYDTS--FDNHTGKQGDGLNTSVNYVQYQEGGGAYDASSNLHNNGQDLSSSQNWED 289
G D S F+ H G+ VQ+ G G Y + +SQ+WE+
Sbjct: 205 ----GLDNSAGFEQHQGQ-----------VQHDSGSGQY------------VDNSQSWEN 237
Query: 290 LYPGWKYDHITGQWYQIEDYNATTTSQQTSEANTAVDWAAASDGKTEISYLQQAAQSVAG 349
LYPGWKYD TGQWYQ++ +AT SQ+ S N+ +W + + ++++YL+Q+ S
Sbjct: 238 LYPGWKYDASTGQWYQVDGQDATVNSQE-SYINSTGNWESVAADNSDVAYLKQSTTS--- 293
Query: 350 TLAETGTTESVSSWNQVSQGNNGYLEHMVFDPQYPDWYYDTIAQEWRSLATYNSSVQSSV 409
A GT ESVS+WNQVSQ NGY EHMVFD QYP WYYDTIAQEWRSL +YN + Q++V
Sbjct: 294 --AMAGTAESVSTWNQVSQVGNGYPEHMVFDAQYPGWYYDTIAQEWRSLDSYNQASQTTV 351
Query: 410 HG------LQNGHTSTSTSSFNDDNSLYSEYSQAGNHVSQGVGSQAVNGSWSGSHGVS-- 461
G +QNGH T+T N +S+Y ++ +Q Q +GSW S+ +
Sbjct: 352 TGQAHDQQVQNGHARTTTYHNNSQSSVYDVNNKNQTFKAQDFAIQGQHGSWDESYYANNQ 411
Query: 462 QAGNYDGSHGVGS---QAVNGSWSGSHGVNQAGNYGGSQGVGSQAVNGSWSGSHGVNQAG 518
QAGN VG + S S G Q N ++ V Q + + Q
Sbjct: 412 QAGNTWQPVNVGKAEPAVTSDSLSRFGGNQQVNNLYSTESVAEQFKPNTIGAQSFIPQHM 471
Query: 519 NYGGSQGVGSQAVNGSWSGSHGV-NHQQGFDMYATEASTKIGNNTASSGNQQVHHSYGNQ 577
N V S NG S S+ + N QQ D A NN S V S +
Sbjct: 472 N------VASATQNGPLSFSNDLYNRQQSVD----HAQKSFQNNQLFS--PSVGRSSDRR 519
Query: 578 QVNTSSSFG-----SVALNNKGSFEPKAF 601
+ SFG V +N GS + +F
Sbjct: 520 PPHALVSFGFGGKLIVMKDNNGSLQNTSF 548
>ref|XP_550450.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|55295836|dbj|BAD67704.1| unknown protein [Oryza sativa
(japonica cultivar-group)]
Length = 577
Score = 518 bits (1335), Expect = e-145
Identities = 303/597 (50%), Positives = 379/597 (62%), Gaps = 33/597 (5%)
Query: 972 MLEDWEENLAVITANRTKGDELVMMHLGDCLWKEKREITAAHICYLIAEVNFSSYSDATR 1031
ML+DW+ENLA+ITANRTKGD+LV+ HLGDCLWKEK E+ AAH CYL AE+N SYS+++R
Sbjct: 1 MLDDWQENLAIITANRTKGDDLVITHLGDCLWKEKNEVAAAHSCYLAAELNIESYSESSR 60
Query: 1032 LCLIGADHWTRPRTYASPEAIQRTELYEYSKLLGNSQFVLHSFQPYKLIYAHMLAEVGKV 1091
+CLIGADH PRT+ SPEAIQRTE+YEY+K+LGNSQ++L FQPYKLIYA+ML EVGKV
Sbjct: 61 MCLIGADHLRCPRTFTSPEAIQRTEVYEYAKVLGNSQYILLPFQPYKLIYAYMLVEVGKV 120
Query: 1092 SDSLKYCQAVLKSLK-TGRAPEVETWKQLVLALEERIRTHQQGGYAANLAPAKLVGKLLN 1150
SDSL+YCQA LK LK +GRAPE+E WKQL +LEERIRT+QQGGY NLAPAKLVGKL
Sbjct: 121 SDSLRYCQACLKVLKASGRAPELEAWKQLFSSLEERIRTYQQGGYGTNLAPAKLVGKLFT 180
Query: 1151 FFDSTAHRVVGGLPPPAPTSSQATVHGSEQHYQHMAPRVSTSQSTMAMSSLVPSASLEPI 1210
D + R++G P Q ++ + + A +Q MAMSSL+ S S + +
Sbjct: 181 SLDKSLSRMMGTQPSALSPVPQGSLTERDSYSAPAATNFVNNQPVMAMSSLMSSVSEQSM 240
Query: 1211 SEWTADNN-RMAAKPNRSVSEPDIGRSPRQESPSPDAQGKVQVSGGASRFSRFGFGSQLL 1269
SE + + NRSVSEPD GR+P Q + +AQ SG S SRFG+ LL
Sbjct: 241 SEMSGNTGPDRKVTHNRSVSEPDFGRTPNQGAGLDNAQ---STSGSGS--SRFGW---LL 292
Query: 1270 QKTVGLVLRSGKQAKLGEKNKFYYDEKLKRWVEEGAEVPAEEAALPPPPPTTAAFQNGSA 1329
QKT+GLV RS QAKLGE+NKFYYDEKLKRWVEEGA++PAEE L PPPPT A FQNG
Sbjct: 293 QKTMGLVSRSHHQAKLGEQNKFYYDEKLKRWVEEGADIPAEEPPL-PPPPTKALFQNGIP 351
Query: 1330 DYNLKSALKTEGLTPNEFSSTRTSSPE-LSPGMPPIPPSSNQFSARSRLGVRSRYVDTFN 1388
D + + + T N FS R +P S GMPP+PPS NQFSAR R+GVRSRYVDTFN
Sbjct: 352 DQS-SNGPGSVSYTANGFSEARPLNPSGPSSGMPPMPPSQNQFSARGRMGVRSRYVDTFN 410
Query: 1389 QNGGSSAN-LFQSPSVQSVKPALPANAKFFIPAPVPSSSEQNMEAIAESNLEDSAANENP 1447
+ G S+ + P+ S+ P + A FF+P P +SEQ + + + +
Sbjct: 411 KGGASATGPSYNKPATPSMNPL--SGATFFVPTPATVASEQIPDPTVNVHQDQPS----- 463
Query: 1448 STSSTNDWSYHPPKHAQTM----TMQRFPSAGNISKQGQTDGNESHFSHSRRTASWSGSF 1503
ST + + S PP Q++ +QR+PS NI + GN S FS S R ASWSG++
Sbjct: 464 STIALRESSASPPPSVQSIPVQSNIQRYPSMDNIMTPSGS-GNGSSFSRS-RAASWSGAY 521
Query: 1504 NDSFSPPKMGEIKPSGAALGMPPSAFMPDPSSLMQGPTRS-----GSFGEDLQEVEL 1555
++ S + P G M S + + S GEDL EVEL
Sbjct: 522 SEQLSGNAVSR-SPDGQRTMMQSPLIPGQKQSHSRSSSNSSLQFNNGLGEDLHEVEL 577
>ref|NP_910217.1| EST AU031036(E60617) corresponds to a region of the predicted
gene.~Similar to Human mRNA for KIAA0310 gene. (AB002308)
[Oryza sativa (japonica cultivar-group)]
Length = 1172
Score = 351 bits (900), Expect = 1e-94
Identities = 193/347 (55%), Positives = 241/347 (68%), Gaps = 12/347 (3%)
Query: 939 RPNDVFPTEETSISGHPGAVGMPQQSEQA-GSNDMLEDWEENLAVITANRTKGDELVMMH 997
+P DVF E + G+ G + +PQ+ +A S ML+DW+ENLA+ITANRTKGD+LV+ H
Sbjct: 803 QPADVF-NAENPVDGNYGNLHIPQRPVEAVNSKSMLDDWQENLAIITANRTKGDDLVITH 861
Query: 998 LGDCLWKEKREITAAHICYLIAEVNFSSYSDATRLCLIGADHWTRPRTYASPEAIQRTEL 1057
LGDCLWKEK E+ AAH CYL AE+N SYS+++R+CLIGADH PRT+ SPEAIQRTE+
Sbjct: 862 LGDCLWKEKNEVAAAHSCYLAAELNIESYSESSRMCLIGADHLRCPRTFTSPEAIQRTEV 921
Query: 1058 YEYSKLLGNSQFVLHSFQPYKLIYAHMLAEVGKVSDSLKYCQAVLKSLK-TGRAPEVETW 1116
YEY+K+LGNSQ++L FQPYKLIYA+ML EVGKVSDSL+YCQA LK LK +GRAPE+E W
Sbjct: 922 YEYAKVLGNSQYILLPFQPYKLIYAYMLVEVGKVSDSLRYCQACLKVLKASGRAPELEAW 981
Query: 1117 KQLVLALEERIRTHQQGGYAANLAPAKLVGKLLNFFDSTAHRVVGGLPPPAPTSSQATVH 1176
KQL +LEERIRT+QQGGY NLAPAKLVGKL D + R++G P Q ++
Sbjct: 982 KQLFSSLEERIRTYQQGGYGTNLAPAKLVGKLFTSLDKSLSRMMGTQPSALSPVPQGSLT 1041
Query: 1177 GSEQHYQHMAPRVSTSQSTMAMSSLVPSASLEPISEWTADNN-RMAAKPNRSVSEPDIGR 1235
+ + A +Q MAMSSL+ S S + +SE + + NRSVSEPD GR
Sbjct: 1042 ERDSYSAPAATNFVNNQPVMAMSSLMSSVSEQSMSEMSGNTGPDRKVTHNRSVSEPDFGR 1101
Query: 1236 SPRQESPSPDAQGKVQVSGGASRFSRFGFGSQLLQKTVGLVLRSGKQ 1282
+P Q + +AQ SG S SRFG+ LLQKT+GLV RS Q
Sbjct: 1102 TPNQGAGLDNAQ---STSGSGS--SRFGW---LLQKTMGLVSRSHHQ 1140
Score = 274 bits (700), Expect = 2e-71
Identities = 220/765 (28%), Positives = 355/765 (45%), Gaps = 94/765 (12%)
Query: 148 DIAAVGESNEVALSGIKEKDWNCFSADDANGVGGFGSYSDFFSELGDQSSDFPVISHDNL 207
+ A+ G + +V + +K+ W+ F+ + +GVGG + +F +GD + +H
Sbjct: 89 EAASPGSAKDVH-TAVKQVQWSAFATN--SGVGGDDPFGEF---MGDDAFFGGNNTHTMT 142
Query: 208 NSQVSPVIEAHNVVLNSSVDYSQYQGVQGYDTSFDNHTGKQGDGL-----NTSVNYVQYQ 262
Q P+ + S++ +Q D D+ G NTSV
Sbjct: 143 GDQ--PIQASLVPPTTSTIGSMDHQFSNQVDNIADSQPGWPATAAEFMDHNTSVQ--SDS 198
Query: 263 EGGGAYDASSNLHNNGQDLSSSQNWEDLYPGWKYDHITGQWYQIEDYNATTTSQQTSEAN 322
A D++S + + +E LYPGWKYD T QWYQ++ ++ T+Q ++
Sbjct: 199 TTAAAVDSTS---------TDPKYFESLYPGWKYDEATQQWYQVD---SSYTAQGNADNL 246
Query: 323 TAVDWAAASDGKTE---------ISYLQQAAQSVAGTLAETGTTESVSSW--NQVSQGNN 371
A+ + + +SYL ++Q+ T+AE G+T + +SW N+ + G
Sbjct: 247 GALPVVGGDNVHQQQQQQQQQFTVSYLHDSSQAGLETIAEEGSTMA-TSWTPNESNTGAV 305
Query: 372 GYLEHMVFDPQYPDWYYDTIAQEWRSLATYNSS-VQSSVHGLQN-GHTSTSTSSFNDDNS 429
Y +MVF +YP WY+DT Q+W SL +Y VQ+ + G+ T + +S
Sbjct: 306 EYPSNMVFYAEYPGWYFDTNTQQWLSLESYQQGGVQAETTAAASAGYAGTGHNVAQTSDS 365
Query: 430 LYSEYSQAGNHVSQGVGSQAVNGSWSGSHG-----VSQAGNYDGSHG-------VGSQAV 477
+YS G +G +++ S+ GS+ +Q N + + + A
Sbjct: 366 YTGDYSHQGQQQHGSLGDNSLSDSFYGSNQHTENQTAQQANVESLESSKYYHADINTYAH 425
Query: 478 NGSWSGSHGVNQAGNYGGSQGVGSQAVNGSWSGSHG------VNQAGNYGGSQGVGSQAV 531
+ S S +QA G Q+V + S G + +G G QA
Sbjct: 426 STSQYTSSEDHQASYKGFGSSTSHQSVYKGFEPSVGHQSTFTSHHSGYNGSESSTVQQAA 485
Query: 532 NGSWSGSHGVNHQQGFDMYATEASTKIGNNTASSGNQQVHHSYGNQQVNTSSSFGSVALN 591
+ + S G + +GF+ Y SG+Q + Y T
Sbjct: 486 HQGFKPSTGNQNYKGFEPY--------------SGHQLGYKGYDYSTGQTGHQ------- 524
Query: 592 NKGSFEPKAFVPHRDIAHQFNYQDTEFDNGTFAPKTFVPHGDIAQQFNYPNTKFDEQKQF 651
F P +A+Q NG P+ VP ++ Q ++ D
Sbjct: 525 ---EFGPSTDSQANHVAYQQLPSHYSSFNGAAKPQDSVPTANMPQMQTRADS--DGCMNL 579
Query: 652 SNVFAENQNSHSYSQQPIQGG----LQYSYAPHAGRSSAGRPSHALVTFGFGGKLIIMKD 707
N + +S +++QQ Q+ Y+ H RSSAGRP HALVTFGFGGKL+++++
Sbjct: 580 PNNYLSTGSSVNFAQQQFISSNALPQQFGYSSHEQRSSAGRPPHALVTFGFGGKLVVVRE 639
Query: 708 PSALTASY--GSQDSVQGSISVLNLMEAVTG--SNNSLTIGNATGDYFRALSQQSFPGPL 763
+++ ++ G+Q + G++S+LN+ E V+ + S+ G+A YF AL +Q PGPL
Sbjct: 640 TISMSTNFDSGNQGNPCGTVSILNVSEIVSDRVDHPSIPSGSALS-YFHALCRQPIPGPL 698
Query: 764 VGGSVGSKELYKWLDERIARCESPDMDYKKGERLRLLLSLLKIACQHYGKLRSPFGTDTI 823
VGGS +K++ KWLDE +S +++ G+ +LL+SLLKI CQHYGKLRSPFG+D
Sbjct: 699 VGGSAAAKDVNKWLDEITGGYDSSIREFQGGDDQKLLISLLKILCQHYGKLRSPFGSDPS 758
Query: 824 LKENDAPESAVAKLFASAKVNGTEFTQYGMPSHCLQNFPSEEQMK 868
+ D PE AV KLF+S K +G +YG HC++N PSE Q++
Sbjct: 759 QEGIDGPEMAVTKLFSSCKSSGAHKGEYGAIVHCMKNIPSENQIQ 803
>ref|XP_646771.1| hypothetical protein DDB0191321 [Dictyostelium discoideum]
gi|60474012|gb|EAL71949.1| hypothetical protein
DDB0191321 [Dictyostelium discoideum]
Length = 1992
Score = 237 bits (605), Expect = 2e-60
Identities = 243/948 (25%), Positives = 392/948 (40%), Gaps = 127/948 (13%)
Query: 541 VNHQQGFDMYATEASTKIGNNTASSGNQQVHHSYGNQQV-----NTSSSFGS---VALNN 592
+N++ + ++ + NN G+ + S NQQ+ SS FG + ++
Sbjct: 1035 LNNKNNTSLDSSNGAIPFFNNNVGIGSPSIPMSSSNQQIPPQYQQPSSQFGGNQVIRNHS 1094
Query: 593 KGSFEPKAFVPHRDIAHQFNYQDTEFDNGTFAPKTFVPHGDIAQQFNYPNTKFDEQKQFS 652
F + D + N + + NG + P QQ + +Q+Q
Sbjct: 1095 LPDFSNHFPISPNDQFNNNNNNNYQGYNGGNSMFNGPPPQQQQQQQQQQQQQQQQQQQQQ 1154
Query: 653 NVFAENQNSHSYSQQPIQGGLQYSYAP---HAGRSSA--------------GRPSHALVT 695
+ Q PI + H RS++ RP H +
Sbjct: 1155 QQQQQQQQQQQQGMMPINNNNNNNNNNNFFHHARSTSWNGNDAHQRGIPFYSRP-HPVFA 1213
Query: 696 FGFGGKLIIMKDPSAL--TASYGSQDSVQGSISVLNLMEAVTGSNNSLTIGNATGDYFRA 753
FGFGGK++ +K S + T G+ + + SI + + ++++ G N ++
Sbjct: 1214 FGFGGKVVSIKSNSGIQVTQPDGTVKTSKYSIQI-SPIKSLIGHNT----------FYNN 1262
Query: 754 LSQQSFPGPLVGGSVGSKE-LYKWLDERIARCESPDMDY---KKGERLRLLLSLLKIACQ 809
LS +FPGPL S SK+ + KW+ ER+ S +D K E +L LLKI C
Sbjct: 1263 LS--TFPGPLTSSS--SKDTINKWISERVHDIASGKIDQLCSKDSEASTILWELLKILCN 1318
Query: 810 HYGKLRSPFGTDTILKENDAPESAVAKLFASAKVNGTEFTQYGMPSHCLQNFP-SEEQMK 868
+YG L + +D + +VAK S + P+ +N S+
Sbjct: 1319 NYGNL---------MNSSDKDDKSVAKSIQSLLSDNKTTPTLFQPTQ--KNIGRSDADSS 1367
Query: 869 AIASEMQNLLVSGKKMEALQRAQEGQLWGPALVLASQLGEQFYVDTVRQMALRQLVAGSP 928
+ + +Q+L+ G + A Q A + +LW ALVLA QLG + Y T+ L GSP
Sbjct: 1368 LLLNRVQSLMTQGDIIAAHQFALDNELWSHALVLAHQLGAEVYGRTMSIFTNSSLPTGSP 1427
Query: 929 LRTLCLLIAGRPNDVFPTEETSISGHPGAVGMPQQSEQAGSNDMLEDWEENLAVITANRT 988
L+TL +L +G D+F I+ + + + L+ W+EN+A + N
Sbjct: 1428 LKTLYILYSGNTRDLFKDIPQGITNN-------NNNNNNNNGAFLDSWKENIATLLFNGK 1480
Query: 989 KGDELVMMHLGDCLWKEKREITAAHICYLIAEVNFSSYSD-ATRLCLIGADHWTRPRTYA 1047
V+ +GDC+W+ ++ I AAHICYL+A+ F ++ +R+ L+G DH + +
Sbjct: 1481 ANGCKVIGEIGDCIWQVQKRIEAAHICYLLADYQFGPINNPGSRIVLLGGDHKI-DKHFI 1539
Query: 1048 SPEAIQRTELYEYSKLLGNSQFVLHSFQPYKLIYAHMLAEVGKVSDSLKYCQAVLKSLKT 1107
+ E++QR+E+YE+SK+LGNSQ+ L YK IYA L ++G V SLKY + LK
Sbjct: 1540 TSESLQRSEIYEFSKILGNSQYSLPIIFVYKFIYATYLCDLGLVDLSLKYLNHIKHLLKN 1599
Query: 1108 GRAP------EVETWKQLVLALEERIRTHQQGGYAANLAPA--KLVG---KLLNFFDSTA 1156
+ +++T+ + + + + G + ++ K++G + + +
Sbjct: 1600 HQVSNPCFIYQLDTFAERITSFSSNNKLSASGSWIPSIFTKGFKILGGSNRQVPTVGANN 1659
Query: 1157 HRVVGGLPPPAPTSSQATVHGSEQHYQHMAPRVSTSQSTMAM------SSLVPSASLEPI 1210
+ PP T S + + Q P S S + S S +P
Sbjct: 1660 NNNEMISPPSIVTGSSTPISTPTKSQQQQPPSQPISASKLPFKKDEDDEDSYSSFSTKPK 1719
Query: 1211 SEWTADNNRMAAKPNRSVSEPDIGRSPRQESPSPDAQGKVQVSGGASRFSRFGFGSQLLQ 1270
+ T +N+ + S S D SP SPS D SG LL
Sbjct: 1720 ASSTNNNSSNKVDSDLS-STSDNTASP--SSPSKDENSNETKSG-------------LLA 1763
Query: 1271 KTVGLVLRSGKQAKLGEKNKFYYDEKLKRWVEEGAEVPAEEAALPPPPPTTAAFQ----- 1325
K+ L N FYY+E+L +WVE G E A A+ PPPPP T ++
Sbjct: 1764 GLFWRKKPKTKEINLDNNNDFYYNEELGKWVERGKEAEAIAASQPPPPPPTESYSPSHTD 1823
Query: 1326 --------NGSADYNLKSALKTEGLT--PNEFSSTRTSSPELSPGMPPIPPSSNQFSARS 1375
G + + S L + T P+ + S P PP N++S S
Sbjct: 1824 GMSTPPPFGGGGNASPPSVLSSSPSTRLPSGGGGGGSGSGGAPSSDPSQPPGVNKYSLVS 1883
Query: 1376 RLGV----RSRYVDTFNQ-------NGGSSANLFQSPSVQSVKPALPA 1412
G + RYVDT NQ N SS +L P + P +PA
Sbjct: 1884 PGGPNTRRKPRYVDTLNQTVYTGGVNNSSSQSLPPKPPTSTFTPFMPA 1931
>emb|CAI46016.1| hypothetical protein [Homo sapiens]
Length = 1061
Score = 142 bits (357), Expect = 1e-31
Identities = 124/421 (29%), Positives = 194/421 (45%), Gaps = 56/421 (13%)
Query: 691 HALVTFGFGGKLIIMKDPSALTASYGSQDSVQGSISVLNLMEAVTGSNNSLTIGNATGDY 750
H V+FG GG+L+ + PS+ T Q ++ L+ ME + +
Sbjct: 280 HVPVSFGPGGQLVHV-GPSSPTDG-------QAALVELHSMEVILNDSEEQE-------- 323
Query: 751 FRALSQQSFPGPLVGGSVGSKELYKWLDERIAR-CESPDMDYKKGERLRLLLSLLKIACQ 809
+SF GPL+ V ++ + ++ A+ C+S + + LL LL + C+
Sbjct: 324 ----EMRSFSGPLIREDVHKVDIMTFCQQKAAQSCKSETLGSRDSA---LLWQLLVLLCR 376
Query: 810 HYGKLRSPFGTDTILKENDAPESAVAKLFASAKVNGTE----FTQYGMPSHCLQNFP-SE 864
G + + ++++ E + + +N T+ G P+ P S
Sbjct: 377 QNGSMVGSDIAELLMQDCKKLEKYKRQPPVANLINLTDEDWPVLSSGTPNLLTGEIPPSV 436
Query: 865 EQMKAIASEMQNLLVSGKKMEALQRAQEGQLWGPALVLASQLGEQFYVDTVRQMALRQLV 924
E I + LL G+K EAL+ A + LWG AL L+S++ Q Y V L
Sbjct: 437 ETPAQIVEKFTRLLYYGRKKEALEWAMKNHLWGHALFLSSKMDPQTY-SWVMSGFTSTLA 495
Query: 925 AGSPLRTLCLLIAGRPNDVFPTEETSISGHPGAVGMPQQSEQAGSNDMLEDWEENLAVIT 984
PL+TL L++GR +PQ + G DW +LAVI
Sbjct: 496 LNDPLQTLFQLMSGR--------------------IPQATTCCGEKQW-GDWRPHLAVIL 534
Query: 985 ANRTKGDEL---VMMHLGDCLWKEKREITAAHICYLIAEVNFSSYSDAT-RLCLIGADHW 1040
+N+ EL ++ +GD L K + AAH CYL+A V F Y+ T L L+G+ H
Sbjct: 535 SNQAGDPELYQRAIVAIGDTL-AGKGLVEAAHFCYLMAHVPFGHYTVKTDHLVLLGSSHS 593
Query: 1041 TRPRTYASPEAIQRTELYEYSKLLGNSQFVLHSFQPYKLIYAHMLAEVGKVSDSLKYCQA 1100
+A+ EAIQRTE++EY ++LG + + SFQ YKL+YA LA+ G VS +L YC+A
Sbjct: 594 QEFLKFATTEAIQRTEIFEYCQMLGRPKSFIPSFQVYKLLYASRLADYGLVSQALHYCEA 653
Query: 1101 V 1101
+
Sbjct: 654 I 654
>ref|NP_149118.1| regucalcin gene promotor region related protein [Homo sapiens]
gi|14517637|dbj|BAB61035.1| RGPR-p117 [Homo sapiens]
Length = 1060
Score = 142 bits (357), Expect = 1e-31
Identities = 124/421 (29%), Positives = 194/421 (45%), Gaps = 56/421 (13%)
Query: 691 HALVTFGFGGKLIIMKDPSALTASYGSQDSVQGSISVLNLMEAVTGSNNSLTIGNATGDY 750
H V+FG GG+L+ + PS+ T Q ++ L+ ME + +
Sbjct: 279 HVPVSFGPGGQLVHV-GPSSPTDG-------QAALVELHSMEVILNDSEEQE-------- 322
Query: 751 FRALSQQSFPGPLVGGSVGSKELYKWLDERIAR-CESPDMDYKKGERLRLLLSLLKIACQ 809
+SF GPL+ V ++ + ++ A+ C+S + + LL LL + C+
Sbjct: 323 ----EMRSFSGPLIREDVHKVDIMTFCQQKAAQSCKSETLGSRDSA---LLWQLLVLLCR 375
Query: 810 HYGKLRSPFGTDTILKENDAPESAVAKLFASAKVNGTE----FTQYGMPSHCLQNFP-SE 864
G + + ++++ E + + +N T+ G P+ P S
Sbjct: 376 QNGSMVGSDIAELLMQDCKKLEKYKRQPPVANLINLTDEDWPVLSSGTPNLLTGEIPPSV 435
Query: 865 EQMKAIASEMQNLLVSGKKMEALQRAQEGQLWGPALVLASQLGEQFYVDTVRQMALRQLV 924
E I + LL G+K EAL+ A + LWG AL L+S++ Q Y V L
Sbjct: 436 ETPAQIVEKFTRLLYYGRKKEALEWAMKNHLWGHALFLSSKMDPQTY-SWVMSGFTSTLA 494
Query: 925 AGSPLRTLCLLIAGRPNDVFPTEETSISGHPGAVGMPQQSEQAGSNDMLEDWEENLAVIT 984
PL+TL L++GR +PQ + G DW +LAVI
Sbjct: 495 LNDPLQTLFQLMSGR--------------------IPQAATCCGEKQW-GDWRPHLAVIL 533
Query: 985 ANRTKGDEL---VMMHLGDCLWKEKREITAAHICYLIAEVNFSSYSDAT-RLCLIGADHW 1040
+N+ EL ++ +GD L K + AAH CYL+A V F Y+ T L L+G+ H
Sbjct: 534 SNQAGDPELYQRAIVAIGDTL-AGKGLVEAAHFCYLMAHVPFGHYTVKTDHLVLLGSSHS 592
Query: 1041 TRPRTYASPEAIQRTELYEYSKLLGNSQFVLHSFQPYKLIYAHMLAEVGKVSDSLKYCQA 1100
+A+ EAIQRTE++EY ++LG + + SFQ YKL+YA LA+ G VS +L YC+A
Sbjct: 593 QEFLKFATTEAIQRTEIFEYCQMLGRPKSFIPSFQVYKLLYASRLADYGLVSQALHYCEA 652
Query: 1101 V 1101
+
Sbjct: 653 I 653
>dbj|BAD30088.1| RGPR-p117 [Oryctolagus cuniculus]
Length = 1045
Score = 140 bits (352), Expect = 4e-31
Identities = 111/354 (31%), Positives = 166/354 (46%), Gaps = 34/354 (9%)
Query: 757 QSFPGPLVGGSVGSKELYKWLDERIARCESPDMDYKKGERLRLLLSLLKIACQHYGKLRS 816
+SFPGPL+ V ++ + ++ A +S G LL LL + C+ G +
Sbjct: 321 RSFPGPLIREDVHKVDVMTFCQQKAA--QSRKSGTPGGRDSALLWQLLVLLCRQNGSMAG 378
Query: 817 PFGTDTILK-----ENDAPESAVAKLFASAKVNGTEFTQYGMPSHCLQNFPSEEQMKAIA 871
+ +L+ E + VA L A + + + PS E +
Sbjct: 379 SDIAELLLQDCKKLEKYKKQPPVANLINLADEDWPVLSSGPRNLLTGETPPSVETPAQVV 438
Query: 872 SEMQNLLVSGKKMEALQRAQEGQLWGPALVLASQLGEQFYVDTVRQMALRQLVAGSPLRT 931
+ LL G+K EAL+ A + LWG AL L+S++ + Y V L PL+T
Sbjct: 439 GKFTQLLYYGRKKEALEWAMKNHLWGHALFLSSKMEPRTY-KWVMSGFTSTLALNDPLQT 497
Query: 932 LCLLIAGRPNDVFPTEETSISGHPGAVGMPQQSEQAGSNDMLEDWEENLAVITANRTKGD 991
L L++GR +PQ + G DW +LAVI +N+
Sbjct: 498 LFQLLSGR--------------------IPQAATCCGDTQW-GDWRPHLAVILSNQAGDP 536
Query: 992 EL---VMMHLGDCLWKEKREITAAHICYLIAEVNFSSYS-DATRLCLIGADHWTRPRTYA 1047
EL V++ +GD L K + AAH CYL+A+V F Y+ +A RL L+G+ H +A
Sbjct: 537 ELRRRVIVSMGDTL-ASKGLVEAAHFCYLMAQVPFGHYTVNADRLALLGSSHSQEFPRFA 595
Query: 1048 SPEAIQRTELYEYSKLLGNSQFVLHSFQPYKLIYAHMLAEVGKVSDSLKYCQAV 1101
S EAIQRTE+ EY + LG ++ SFQ YKL+YA LA+ G S + YC+A+
Sbjct: 596 SAEAIQRTEVLEYCRTLGGRPGLIASFQVYKLLYASRLADYGLASQAFHYCEAI 649
>dbj|BAD18068.1| RGPR-p117 [Rattus norvegicus]
Length = 1057
Score = 140 bits (352), Expect = 4e-31
Identities = 141/530 (26%), Positives = 230/530 (42%), Gaps = 76/530 (14%)
Query: 601 FVPHRDIAHQFNYQDTEFDN-----GTFAPKTFVPHGDIAQQFNYPNTKFDEQKQFSNVF 655
F + D QF ++ DN G P F P +Q +K + +Q +
Sbjct: 163 FEANSDTQFQFTSKNPYNDNPASAFGLEQPGEFFPESGAQKQKPSLTSKSNLLQQHESGL 222
Query: 656 AENQNSHSYSQQPIQGGLQY------SYAP-HAGRSSAGRPS--------HALVTFGFGG 700
+ + S+ SQ + QY ++ P A +SA P H V+FG GG
Sbjct: 223 SSS--SYELSQYMTEAPEQYEPMVSATWRPIQADDTSATVPKAPMRFYVPHVSVSFGPGG 280
Query: 701 KLIIMKDPSALTASYGSQDSVQGSISVLNLMEAVTGSNNSLTIGNATGDYFRALSQQSFP 760
+LI + S S Q ++ ++ ME + D+ ++FP
Sbjct: 281 QLICV--------SPNSPADGQTALVEVHSMEVILN------------DFEDQEEMRAFP 320
Query: 761 GPLVGGSVGSKELYKWLDERIARCESPDMDYKKGERLRLLLSLLKIACQHYGKLRSPFGT 820
GPL+ + ++ + ++ A+C + + L L LL + C+ G +
Sbjct: 321 GPLIREDIHKVDIMTFCQKKAAQCLKSETPGSRDSAL--LWQLLVLLCRQNGSMVGSDIA 378
Query: 821 DTILKENDAPESAVAKLFASAKVNGTEFTQYGMPSHCLQNFPSEEQMKA-----IASEMQ 875
+ ++++ E + + +N T+ + S E + I +
Sbjct: 379 ELLMQDCKKLEKYKRQPPVANLINLTDEDWPVLSSGTRNLLTGEIPLNVDTPAQIVEKFT 438
Query: 876 NLLVSGKKMEALQRAQEGQLWGPALVLASQLGEQFYVDTVRQMALRQLVAGSPLRTLCLL 935
NLL G+K EAL+ A + LWG AL LAS++ + Y + V L PL+TL L
Sbjct: 439 NLLYYGRKKEALEWAMKNHLWGHALFLASKMDPRTY-NWVMSGFTTTLALNDPLQTLFQL 497
Query: 936 IAGRPNDVFPTEETSISGHPGAVGMPQQSEQAGSNDMLEDWEENLAVITANRTKGDEL-- 993
++GR +PQ + G DW +LAVI +N+ EL
Sbjct: 498 MSGR--------------------IPQAATCCGDKQW-GDWRPHLAVILSNQAGDAELYQ 536
Query: 994 -VMMHLGDCLWKEKREITAAHICYLIAEVNFSSYSDAT-RLCLIGADHWTRPRTYASPEA 1051
++ +GD L K + A+H CYL+A V F Y+ T L L+G++H +A+ EA
Sbjct: 537 RAIVSMGDTL-AGKGLVEASHFCYLMAHVPFGHYTVKTDHLALVGSNHSQEFLKFATIEA 595
Query: 1052 IQRTELYEYSKLLGNSQFVLHSFQPYKLIYAHMLAEVGKVSDSLKYCQAV 1101
IQRTE++EY ++LG + + SFQ YKL+YA LA+ G S +L YC+A+
Sbjct: 596 IQRTEIFEYCQMLGRPKSFIPSFQVYKLLYASRLADYGLASQALHYCEAI 645
>emb|CAI16321.1| regucalcin gene promotor region related protein (RGPR) [Homo sapiens]
gi|56204433|emb|CAI22848.1| regucalcin gene promotor
region related protein (RGPR) [Homo sapiens]
Length = 1061
Score = 138 bits (347), Expect = 2e-30
Identities = 122/421 (28%), Positives = 192/421 (44%), Gaps = 56/421 (13%)
Query: 691 HALVTFGFGGKLIIMKDPSALTASYGSQDSVQGSISVLNLMEAVTGSNNSLTIGNATGDY 750
H V+FG GG+L+ + PS+ T Q ++ L+ ME + +
Sbjct: 280 HVPVSFGPGGQLVHV-GPSSPTDG-------QAALVELHSMEVILNDSEEQE-------- 323
Query: 751 FRALSQQSFPGPLVGGSVGSKELYKWLDERIAR-CESPDMDYKKGERLRLLLSLLKIACQ 809
+SF GPL+ V ++ + ++ A+ C+S + + LL LL + C+
Sbjct: 324 ----EMRSFSGPLIREDVHKVDIMTFCQQKAAQSCKSETLGSRDSA---LLWQLLVLLCR 376
Query: 810 HYGKLRSPFGTDTILKENDAPESAVAKLFASAKVNGTE----FTQYGMPSHCLQNFP-SE 864
G + + ++++ E + + +N T+ G P+ P S
Sbjct: 377 QNGSMVGSDIAELLMQDCKKLEKYKRQPPVANLINLTDEDWPVLSSGTPNLLTGEIPPSV 436
Query: 865 EQMKAIASEMQNLLVSGKKMEALQRAQEGQLWGPALVLASQLGEQFYVDTVRQMALRQLV 924
E I + LL G+K L+ A + LWG AL L+S++ Q Y V L
Sbjct: 437 ETPAQIVEKFTRLLYYGRKKATLEWAMKNHLWGHALFLSSKMDPQTY-SWVMSGFTSTLA 495
Query: 925 AGSPLRTLCLLIAGRPNDVFPTEETSISGHPGAVGMPQQSEQAGSNDMLEDWEENLAVIT 984
PL+TL L++GR +PQ + G DW +LAVI
Sbjct: 496 LNDPLQTLFQLMSGR--------------------IPQAATCCGEKQW-GDWRPHLAVIL 534
Query: 985 ANRTKGDEL---VMMHLGDCLWKEKREITAAHICYLIAEVNFSSYSDAT-RLCLIGADHW 1040
+N+ EL ++ +GD L K + AAH CYL+A V F Y+ T L L+G+ H
Sbjct: 535 SNQAGDPELYQRAIVAIGDTL-AGKGLVEAAHFCYLMAHVPFGHYTVKTDHLVLLGSSHS 593
Query: 1041 TRPRTYASPEAIQRTELYEYSKLLGNSQFVLHSFQPYKLIYAHMLAEVGKVSDSLKYCQA 1100
+A+ EAIQRTE++EY ++LG + + SFQ YKL+YA LA+ G VS +L YC+A
Sbjct: 594 QEFLKFATTEAIQRTEIFEYCQMLGRPKSFIPSFQVYKLLYASRLADYGLVSQALHYCEA 653
Query: 1101 V 1101
+
Sbjct: 654 I 654
>dbj|BAD32580.1| mKIAA1928 protein [Mus musculus]
Length = 1064
Score = 138 bits (347), Expect = 2e-30
Identities = 172/726 (23%), Positives = 292/726 (39%), Gaps = 137/726 (18%)
Query: 411 GLQNGHTSTSTSSFNDDNSLYSEYSQAGNHVSQGVGSQAVNGSWSGSHGVSQAGNYDGSH 470
GLQ+GH L+S+YS H W +H S++
Sbjct: 35 GLQSGHYRPR---------LHSQYSGDKYH------------QWQDAHKNSKS------- 66
Query: 471 GVGSQAVNGSWSGSHGVNQAGNYGGSQGV-GSQAVNGSWSGSHGVNQAGNYGGSQGVGSQ 529
Q + SH V+++G + SQ V G+ + GS+ SH +++G Q +
Sbjct: 67 ---QQDLRDDHQQSHSVSRSGEW--SQPVSGADYLKGSYP-SHLYSRSGYGDPYQRYHTP 120
Query: 530 AVNGSWSGSHGVNHQQGFDMYATEASTKIGNNTASSGNQQV-HHSYGNQQV----NTSSS 584
++ +G + G E A G+ + H +G+Q+ +
Sbjct: 121 TPRDEYA--YGNYYYHGHPQLLPE------ERVARQGSPYIWHEDHGDQRYFGEHHREKH 172
Query: 585 FGSVALNNKGSFEPKAFVPHRDIAHQFNYQDTEFDNGTFAPKTFVPHGDIAQQFNYPNTK 644
G+ N+ F+ + P+RD + Q+ P F P + +Q +K
Sbjct: 173 NGTFGANSDTQFQFTSKNPYRDSPASVSGQEQ--------PGEFFPESEAQKQKPLLTSK 224
Query: 645 FDEQKQFSNVFAENQNSHSYSQQPIQGGLQYSYAPHAGRSSAGRP--------------- 689
+Q + +S SY Y P S+A RP
Sbjct: 225 SSLLQQHES----GLSSSSYELSQYMTAAPEEYEPMV--SAAWRPIQADDTSATVPKAPM 278
Query: 690 ----SHALVTFGFGGKLIIMKDPSALTASYGSQDSVQGSISVLNLMEAVTGSNNSLTIGN 745
H V+FG GG+L+ + S Q ++ ++ ME +
Sbjct: 279 RFYVPHVSVSFGPGGQLVCVPPNSPADG--------QTALVEVHSMEVLLN--------- 321
Query: 746 ATGDYFRALSQQSFPGPLVGGSVGSKELYKWLDERIARCESPDMDYKKGERLRLLLSLLK 805
D+ ++FPGPL+ + ++ + ++ +C + + LL LL
Sbjct: 322 ---DFEDQEEMRAFPGPLIREDIHKVDIMTFCQQKATQCLKSETPGSRDS--ALLWQLLV 376
Query: 806 IACQHYGKLRSPFGTDTILKENDAPESAVAKLFASAKVNGTEFTQYGMPSHCLQNF---- 861
+ C+ G + + ++++ E + + +N T+ + + S ++
Sbjct: 377 LLCRQNGSMVGSDIAELLMQDCKKLEKYKRQPPVANLINLTD-EDWPVLSSGTRDLLTGE 435
Query: 862 --PSEEQMKAIASEMQNLLVSGKKMEALQRAQEGQLWGPALVLASQLGEQFYVDTVRQMA 919
P+ + I + LL G+K EAL+ A + LWG AL LAS++ + Y + V
Sbjct: 436 IPPNVDTPAQIVEKFTKLLYYGRKKEALEWAMKNHLWGHALFLASKMDPRTY-NWVMSGF 494
Query: 920 LRQLVAGSPLRTLCLLIAGRPNDVFPTEETSISGHPGAVGMPQQSEQAGSNDMLEDWEEN 979
L PL+TL L++GR +PQ + G DW +
Sbjct: 495 TSTLALNDPLQTLFQLMSGR--------------------IPQAATVCGDKQW-GDWRPH 533
Query: 980 LAVITANRTKGDEL---VMMHLGDCLWKEKREITAAHICYLIAEVNFSSYSDAT-RLCLI 1035
LAVI +N+ EL ++ +GD L K + A+H CYL+A V F Y+ T L L+
Sbjct: 534 LAVILSNQAGDTELYQRAIVSMGDTL-AGKGLVEASHFCYLMAHVPFGHYTVKTDHLALV 592
Query: 1036 GADHWTRPRTYASPEAIQRTELYEYSKLLGNSQFVLHSFQPYKLIYAHMLAEVGKVSDSL 1095
G+ H +A+ EAIQRTE++EY ++LG + + SFQ YKL+YA LA+ G S +L
Sbjct: 593 GSSHSQEFMKFATIEAIQRTEIFEYCQMLGRPKSFIPSFQVYKLLYASRLADYGLASQAL 652
Query: 1096 KYCQAV 1101
YC+A+
Sbjct: 653 HYCEAI 658
>dbj|BAB67821.1| KIAA1928 protein [Homo sapiens]
Length = 757
Score = 137 bits (346), Expect = 2e-30
Identities = 110/355 (30%), Positives = 170/355 (46%), Gaps = 36/355 (10%)
Query: 757 QSFPGPLVGGSVGSKELYKWLDERIAR-CESPDMDYKKGERLRLLLSLLKIACQHYGKLR 815
+SF GPL+ V ++ + ++ A+ C+S + + LL LL + C+ G +
Sbjct: 22 RSFSGPLIREDVHKVDIMTFCQQKAAQSCKSETLGSRDSA---LLWQLLVLLCRQNGSMV 78
Query: 816 SPFGTDTILKENDAPESAVAKLFASAKVNGTE----FTQYGMPSHCLQNFP-SEEQMKAI 870
+ ++++ E + + +N T+ G P+ P S E I
Sbjct: 79 GSDIAELLMQDCKKLEKYKRQPPVANLINLTDEDWPVLSSGTPNLLTGEIPPSVETPAQI 138
Query: 871 ASEMQNLLVSGKKMEALQRAQEGQLWGPALVLASQLGEQFYVDTVRQMALRQLVAGSPLR 930
+ LL G+K EAL+ A + LWG AL L+S++ Q Y V L PL+
Sbjct: 139 VEKFTRLLYYGRKKEALEWAMKNHLWGHALFLSSKMDPQTY-SWVMSGFTSTLALNDPLQ 197
Query: 931 TLCLLIAGRPNDVFPTEETSISGHPGAVGMPQQSEQAGSNDMLEDWEENLAVITANRTKG 990
TL L++GR +PQ + G DW +LAVI +N+
Sbjct: 198 TLFQLMSGR--------------------IPQAATCCGEKQW-GDWRPHLAVILSNQAGD 236
Query: 991 DEL---VMMHLGDCLWKEKREITAAHICYLIAEVNFSSYSDAT-RLCLIGADHWTRPRTY 1046
EL ++ +GD L K + AAH CYL+A V F Y+ T L L+G+ H +
Sbjct: 237 PELYQRAIVAIGDTL-AGKGLVEAAHFCYLMAHVPFGHYTVKTDHLVLLGSSHSQEFLKF 295
Query: 1047 ASPEAIQRTELYEYSKLLGNSQFVLHSFQPYKLIYAHMLAEVGKVSDSLKYCQAV 1101
A+ EAIQRTE++EY ++LG + + SFQ YKL+YA LA+ G VS +L YC+A+
Sbjct: 296 ATTEAIQRTEIFEYCQMLGRPKSFIPSFQVYKLLYASRLADYGLVSQALHYCEAI 350
>ref|NP_203505.2| regucalcin gene promotor region related protein [Mus musculus]
gi|37589164|gb|AAH59194.1| Regucalcin gene promotor
region related protein [Mus musculus]
Length = 1051
Score = 137 bits (344), Expect = 4e-30
Identities = 158/653 (24%), Positives = 271/653 (41%), Gaps = 106/653 (16%)
Query: 484 SHGVNQAGNYGGSQGV-GSQAVNGSWSGSHGVNQAGNYGGSQGVGSQAVNGSWSGSHGVN 542
SH V+++G + SQ V G+ + GS+ SH +++G Q + ++ +G
Sbjct: 64 SHSVSRSGEW--SQPVSGADYLKGSYP-SHLYSRSGYGDPYQRYHTPTPRDEYA--YGNY 118
Query: 543 HQQGFDMYATEASTKIGNNTASSGNQQV-HHSYGNQQV----NTSSSFGSVALNNKGSFE 597
+ G E A G+ + H +G+Q+ + G+ N+ F+
Sbjct: 119 YYHGHPQLLPE------ERVARQGSPYIWHEDHGDQRYFGEHHREKHNGTFGANSDTQFQ 172
Query: 598 PKAFVPHRDIAHQFNYQDTEFDNGTFAPKTFVPHGDIAQQFNYPNTKFDEQKQFSNVFAE 657
+ P+RD + Q+ P F P + +Q +K +Q +
Sbjct: 173 FTSKNPYRDSPASVSGQEQ--------PGEFFPESEAQKQKPLLTSKSSLLQQHES---- 220
Query: 658 NQNSHSYSQQPIQGGLQYSYAPHAGRSSAGRP-------------------SHALVTFGF 698
+S SY Y P S+A RP H V+FG
Sbjct: 221 GLSSSSYELSQYMTAAPEEYEPMV--SAAWRPIQADDTSATVPKAPMRFYVPHVSVSFGP 278
Query: 699 GGKLIIMKDPSALTASYGSQDSVQGSISVLNLMEAVTGSNNSLTIGNATGDYFRALSQQS 758
GG+L+ + S Q ++ ++ ME + D+ ++
Sbjct: 279 GGQLVCVPPNSPADG--------QTALVEVHSMEVLLN------------DFEDQEEMRA 318
Query: 759 FPGPLVGGSVGSKELYKWLDERIARCESPDMDYKKGERLRLLLSLLKIACQHYGKLRSPF 818
FPGPL+ + ++ + ++ +C + + LL LL + C+ G +
Sbjct: 319 FPGPLIREDIHKVDIMTFCQQKATQCLKSETPGSRDS--ALLWQLLVLLCRQNGSMVGSD 376
Query: 819 GTDTILKENDAPESAVAKLFASAKVNGTEFTQYGMPSHCLQNF------PSEEQMKAIAS 872
+ ++++ E + + +N T+ + + S ++ P+ + I
Sbjct: 377 IAELLMQDCKKLEKYKRQPPVANLINLTD-EDWPVLSSGTRDLLTGEIPPNVDTPAQIVE 435
Query: 873 EMQNLLVSGKKMEALQRAQEGQLWGPALVLASQLGEQFYVDTVRQMALRQLVAGSPLRTL 932
+ LL G+K EAL+ A + LWG AL LAS++ + Y + V L PL+TL
Sbjct: 436 KFTKLLYYGRKKEALEWAMKNHLWGHALFLASKMDPRTY-NWVMSGFTSTLALNDPLQTL 494
Query: 933 CLLIAGRPNDVFPTEETSISGHPGAVGMPQQSEQAGSNDMLEDWEENLAVITANRTKGDE 992
L++GR +PQ + G DW +LAVI +N+ E
Sbjct: 495 FQLMSGR--------------------IPQAATVCGDKQW-GDWRPHLAVILSNQAGDTE 533
Query: 993 L---VMMHLGDCLWKEKREITAAHICYLIAEVNFSSYSDAT-RLCLIGADHWTRPRTYAS 1048
L ++ +GD L K + A+H CYL+A V F Y+ T L L+G+ H +A+
Sbjct: 534 LYQRAIVSMGDTL-AGKGLVEASHFCYLMAHVPFGHYTVKTDHLALVGSSHSQEFMKFAT 592
Query: 1049 PEAIQRTELYEYSKLLGNSQFVLHSFQPYKLIYAHMLAEVGKVSDSLKYCQAV 1101
EAIQRTE++EY ++LG + + SFQ YKL+YA LA+ G S +L YC+A+
Sbjct: 593 IEAIQRTEIFEYCQMLGRPKSFIPSFQVYKLLYASRLADYGLASQALHYCEAI 645
>ref|XP_514022.1| PREDICTED: hypothetical protein XP_514022 [Pan troglodytes]
Length = 1073
Score = 136 bits (342), Expect = 6e-30
Identities = 144/557 (25%), Positives = 238/557 (41%), Gaps = 84/557 (15%)
Query: 566 GNQQVHHSYGNQQVNTSSSFGSVALNNKGSFEPKAFVPHRD---IAHQFNYQDTEFDNGT 622
G + + SY + + ++GS + + + VP + I H+ +Y++ ++ +
Sbjct: 101 GYENSYQSYQSPTMREEYAYGSYYYHGHPQWLQEERVPRQQSPYIWHE-DYREQKYLDEH 159
Query: 623 FAPKTFVPHGDIAQQFNYPNTKFDEQKQFSNVFAENQNSHSYSQQP---IQGGLQYSYAP 679
P G ++ + N++ N ++ S+S + P G L
Sbjct: 160 HYENQHSPFGTNSETYFQSNSR--------NPCKDSPASNSGQEWPGELFPGSLLAEAQK 211
Query: 680 HAGRSSAGRPS-------HALVTFGFGGKLIIMKDPSALTASYGSQDSVQGSISVLNLME 732
+ SSAG + H V+FG GG+L+ + PS+ T Q ++ L+ ME
Sbjct: 212 NKDVSSAGPKAPMKFYIPHVPVSFGPGGQLVRV-GPSSPTDG-------QAALVELHSME 263
Query: 733 AVTGSNNSLTIGNATGDYFRALSQQSFPGPLVGGSVGSKELYKWLDERIAR-CESPDMDY 791
+ + +SF GPL+ V ++ + ++ A+ C+S +
Sbjct: 264 VILNDSEEQE------------EMRSFSGPLIREDVHKVDIMTFCQQKAAQSCKSETLGS 311
Query: 792 KKGERLRLLLSLLKIACQHYGKLRSPFGTDTILK-----ENDAPESAVAKLFASAKVNGT 846
+ LL LL + C+ G + + +++ E + VA L + +
Sbjct: 312 RDSA---LLWQLLVLLCRQNGSMVGSDIAELLMQDCKKLEKYKRQPPVANLISLTDEDWP 368
Query: 847 EFTQYGMPSHCLQNFP-SEEQMKAIASEMQNLLVSGKKMEALQRAQEGQLWGPALVLASQ 905
+ G P+ P S E I + LL G+K EAL+ A + LWG AL L+S+
Sbjct: 369 VLSS-GTPNLLTGEIPPSVETPAQIVEKFTRLLYYGRKKEALEWAMKNHLWGHALFLSSK 427
Query: 906 LGEQFYVDTVRQMALRQLVAGSPLRTLCLLIAGRPNDVFPTEETSISGHPGAVGMPQQSE 965
+ Y V L PL+TL L++GR +PQ +
Sbjct: 428 MDPWTY-SWVMSGFTSTLALNDPLQTLFQLMSGR--------------------IPQAAT 466
Query: 966 QAGSNDMLEDWEENLAVITANRTKGDELVMMHLGDCLWKEKREITAAHICYLIAEVNFSS 1025
G DW +LAVI +N+ EL G + AAH CYL+A V F
Sbjct: 467 CCGEKQW-GDWRPHLAVILSNQAGDPELYQPGKG--------LVEAAHFCYLVAHVPFGH 517
Query: 1026 YSDAT-RLCLIGADHWTRPRTYASPEAIQRTELYEYSKLLGNSQFVLHSFQPYKLIYAHM 1084
Y+ T L L+G+ H +A+ EAIQRTE++EY ++LG + + SFQ YKL+YA
Sbjct: 518 YTVKTDHLVLLGSSHSQEFLKFATTEAIQRTEIFEYCQMLGRPKSFIPSFQVYKLLYASR 577
Query: 1085 LAEVGKVSDSLKYCQAV 1101
LA+ G VS +L YC+A+
Sbjct: 578 LADYGLVSQALHYCEAI 594
>ref|XP_520369.1| PREDICTED: hypothetical protein XP_520369 [Pan troglodytes]
Length = 1445
Score = 131 bits (329), Expect = 2e-28
Identities = 105/368 (28%), Positives = 170/368 (45%), Gaps = 43/368 (11%)
Query: 757 QSFPGPLVGGSVGSKELYKWLDERIARCESPDMDYKKGERLRLLLSLLKIACQHYGKLRS 816
++FPGPL ++ + + RC + K E LL + + + C+ G +
Sbjct: 566 RAFPGPLAKDDTHKVDVINFAQNKAMRCLQNENLIDK-ESASLLWNFIVLLCRQNGTV-- 622
Query: 817 PFGTD----------TILKENDAPESAVAKLFASAKVNGTEFTQYGMPS-HCLQNFPSE- 864
GTD T+ +P A F + V E + G L P+
Sbjct: 623 -VGTDIAELLLRDHRTVWLPGKSPNEANLIDFTNEAVEQVEEEESGEAQLSFLTGGPAAA 681
Query: 865 -EQMKAIASEMQNLLVSGKKMEALQRAQEGQLWGPALVLASQLGEQFYVDTVRQMALRQL 923
++ + LL+ G+K +AL+ A + LWG AL+LAS++ + + + + A L
Sbjct: 682 ASSLERETERFRELLLYGRKKDALESAMKNGLWGHALLLASKMDSRTHARVMTRFA-NSL 740
Query: 924 VAGSPLRTLCLLIAGRPNDVFPTEETSISGHPGAVGMPQQSEQAGSNDMLEDWEENLAVI 983
PL+T+ L++GR MP S ++ DW +LA++
Sbjct: 741 PINDPLQTVYQLMSGR--------------------MPAASTVCCGDEKWGDWRPHLAMV 780
Query: 984 TANRTKGDEL---VMMHLGDCLWKEKREITAAHICYLIAEVNFSSYSD-ATRLCLIGADH 1039
+N ++ M +GD L + + AAH CYL+A+ F Y+ T+L LIG++H
Sbjct: 781 LSNLNNNMDVESRTMATMGDTL-ASRGLLDAAHFCYLMAQAGFGVYTKKTTKLVLIGSNH 839
Query: 1040 WTRPRTYASPEAIQRTELYEYSKLLGNSQFVLHSFQPYKLIYAHMLAEVGKVSDSLKYCQ 1099
+A+ EAIQRTE YEY++ LG L SFQ +K IY+ LAE+G + + YC+
Sbjct: 840 SLPFLKFATNEAIQRTEAYEYAQSLGAQTCPLPSFQVFKFIYSCRLAEMGLATQAFHYCE 899
Query: 1100 AVLKSLKT 1107
A+ KS+ T
Sbjct: 900 AIAKSILT 907
Score = 55.5 bits (132), Expect = 1e-05
Identities = 70/289 (24%), Positives = 103/289 (35%), Gaps = 73/289 (25%)
Query: 1162 GLPPPAPTSSQATVHGSEQHYQHMAPRVSTSQSTMAMSSLVPSASLEPISEWTADNNRMA 1221
GLPP P Q +H+ + + SL +SE D
Sbjct: 1113 GLPPGVPPL---------QERRHLLQEARSPDPGIVPQEAPVGNSLSELSEENFDGKFAN 1163
Query: 1222 AKPNRSV----------------SEPDIGRSPRQESPSPDAQGKVQVSG---GASRFSRF 1262
P+R++ ++P + SP E+ P K + G S F R+
Sbjct: 1164 LTPSRTMPDSEAPPGWDRADSGPTQPPLSFSPAPETKRPGQAAKKETKEPKKGESWFFRW 1223
Query: 1263 GFGSQLLQKTVGLVLRSGKQAKLGEKNK-FYYDEKLKRWVEEGAEVPAEEAALPPPPPTT 1321
G + + + +KNK +DEK +WV P EE PPPPPT+
Sbjct: 1224 LPGKKKTEAYLP-----------DDKNKSIVWDEKKNQWVN--LNEPEEEKKAPPPPPTS 1270
Query: 1322 AAFQNGSADYNLKSALKTEGLTPNEFSSTRTSSPELSPGMPPIPPSSNQFSARSRLGVRS 1381
T ++P PG P P N +S R+ G R+
Sbjct: 1271 -------------------------MPKTVQAAPPALPGPPGAPV--NMYSRRAA-GTRA 1302
Query: 1382 RYVDTFNQNGGSSANLFQSPS--VQSVKPALPANAKFFIPAPVPSSSEQ 1428
RYVD N +G + +P+ V + P LP + F+P PV S Q
Sbjct: 1303 RYVDVLNPSGTQRSEPALAPADFVAPLAP-LPIPSNLFVPTPVSSVRPQ 1350
>dbj|BAC03392.1| FLJ00305 protein [Homo sapiens]
Length = 789
Score = 131 bits (329), Expect = 2e-28
Identities = 90/244 (36%), Positives = 126/244 (50%), Gaps = 27/244 (11%)
Query: 862 PSEEQMKAIASEMQNLLVSGKKMEALQRAQEGQLWGPALVLASQLGEQFYVDTVRQMALR 921
PS E I + LL G+K EAL+ A + LWG AL L+S++ Q Y V
Sbjct: 148 PSVETPAQIVEKFTRLLYYGRKKEALEWAMKNHLWGHALFLSSKMDPQTY-SWVMSGFTS 206
Query: 922 QLVAGSPLRTLCLLIAGRPNDVFPTEETSISGHPGAVGMPQQSEQAGSNDMLEDWEENLA 981
L PL+TL L++GR +PQ + G DW +LA
Sbjct: 207 TLALNDPLQTLFQLMSGR--------------------IPQAATCCGEKQW-GDWRPHLA 245
Query: 982 VITANRTKGDEL---VMMHLGDCLWKEKREITAAHICYLIAEVNFSSYSDAT-RLCLIGA 1037
VI +N+ EL ++ +GD L K + AAH CYL+A V F Y+ T L L+G+
Sbjct: 246 VILSNQAGDPELYQRAIVAIGDTL-AGKGLVEAAHFCYLMAHVPFGHYTVKTDHLVLLGS 304
Query: 1038 DHWTRPRTYASPEAIQRTELYEYSKLLGNSQFVLHSFQPYKLIYAHMLAEVGKVSDSLKY 1097
H +A+ EAIQRTE++EY ++LG + + SFQ YKL+YA LA+ G VS +L Y
Sbjct: 305 SHSQEFLKFATTEAIQRTEIFEYCQMLGRPKSFIPSFQVYKLLYASRLADYGLVSQALHY 364
Query: 1098 CQAV 1101
C+A+
Sbjct: 365 CEAI 368
>dbj|BAD12777.2| RGPR-p117 [Bos taurus] gi|46559376|ref|NP_996854.2| regucalcin gene
promotor region related protein [Bos taurus]
Length = 1052
Score = 129 bits (323), Expect = 1e-27
Identities = 117/420 (27%), Positives = 186/420 (43%), Gaps = 54/420 (12%)
Query: 691 HALVTFGFGGKLIIMKDPSALTASYGSQDSVQGSISVLNLMEAVTGSNNSLTIGNATGDY 750
H V+FG GG+L+ + S S Q ++ L+ ME + +
Sbjct: 279 HMPVSFGPGGQLVCV--------SPSSPSDGQTALVELHSMEVILNDSEEQE-------- 322
Query: 751 FRALSQQSFPGPLVGGSVGSKELYKWLDERIARCESPDMDYKKGERLRLLLSLLKIACQH 810
++F GPL+ V ++ + ++ A+ + + L L LL + C+
Sbjct: 323 ----EMRTFSGPLIREDVHKVDIMTFCQQKAAQSHKSETPGSRDSAL--LWQLLVLLCRQ 376
Query: 811 YGKLRSPFGTDTILKENDAPESAVAKLFASAKVNGTEFTQYGMPSHCL-----QNFPSEE 865
G + + ++++ E + + +N T+ + S + PS E
Sbjct: 377 NGSMVGSDIAELLMQDWKKLEKYKRQPPVANLINLTDEDWPVLSSGTRDLLTGEILPSVE 436
Query: 866 QMKAIASEMQNLLVSGKKMEALQRAQEGQLWGPALVLASQLGEQFYVDTVRQMALRQLVA 925
I + LL G+K EAL+ A + LWG AL LAS++ + + V L
Sbjct: 437 TPAQILEKFTKLLYYGRKKEALEWAMKNHLWGHALFLASKMDPRTH-SRVMSGFTSTLAL 495
Query: 926 GSPLRTLCLLIAGRPNDVFPTEETSISGHPGAVGMPQQSEQAGSNDMLEDWEENLAVITA 985
PL+TL L++GR +PQ + G DW +LAVI +
Sbjct: 496 NDPLQTLFQLMSGR--------------------IPQAATCCGDKQW-GDWRPHLAVILS 534
Query: 986 NRTKGDEL---VMMHLGDCLWKEKREITAAHICYLIAEVNFSSYSDATR-LCLIGADHWT 1041
N+ EL ++ +GD L K + AAH CYL+A V F Y+ T L L+G+ H
Sbjct: 535 NQGGDPELYQRTIVAMGDTL-AGKGLVEAAHFCYLMAHVPFGYYTVKTDYLALLGSSHSQ 593
Query: 1042 RPRTYASPEAIQRTELYEYSKLLGNSQFVLHSFQPYKLIYAHMLAEVGKVSDSLKYCQAV 1101
A+ EAIQRTE++EY ++LG + + SFQ YKL+YA LA+ G S +L YC+ V
Sbjct: 594 EFLKSATTEAIQRTEIFEYCQMLGRPKSFIPSFQVYKLLYASRLADYGLTSQALHYCEPV 653
>emb|CAI13952.1| RP11-413M3.10 [Homo sapiens]
Length = 2134
Score = 128 bits (322), Expect = 1e-27
Identities = 105/368 (28%), Positives = 171/368 (45%), Gaps = 44/368 (11%)
Query: 757 QSFPGPLVGGSVGSKELYKWLDERIARCESPDMDYKKGERLRLLLSLLKIACQHYGKLRS 816
++FPGPL ++ + + +C + K E LL + + + C+ G +
Sbjct: 1317 RAFPGPLAKDDTHKVDVINFAQNKAMKCLQNENLIDK-ESASLLWNFIVLLCRQNGTV-- 1373
Query: 817 PFGTD----------TILKENDAPESAVAKLFASAKVNGTEFTQYGMPS-HCLQNFPSE- 864
GTD T+ +P A F + V E + G L P+
Sbjct: 1374 -VGTDIAELLLRDHRTVWLPGKSPNEANLIDFTNEAVEQVEEEESGEAQLSFLTGGPAAA 1432
Query: 865 -EQMKAIASEMQNLLVSGKKMEALQRAQEGQLWGPALVLASQLGEQFYVDTVRQMALRQL 923
++ + LL+ G+K +AL+ A + LWG AL+LAS++ + + + + A L
Sbjct: 1433 ASSLERETERFRELLLYGRKKDALESAMKNGLWGHALLLASKMDSRTHARVMTRFA-NSL 1491
Query: 924 VAGSPLRTLCLLIAGRPNDVFPTEETSISGHPGAVGMPQQSEQAGSNDMLEDWEENLAVI 983
PL+T+ L++GR MP S G ++ DW +LA++
Sbjct: 1492 PINDPLQTVYQLMSGR--------------------MPAASTCCG-DEKWGDWRPHLAMV 1530
Query: 984 TANRTKGDEL---VMMHLGDCLWKEKREITAAHICYLIAEVNFSSYSD-ATRLCLIGADH 1039
+N ++ M +GD L + + AAH CYL+A+ F Y+ T+L LIG++H
Sbjct: 1531 LSNLNNNMDVESRTMATMGDTL-ASRGLLDAAHFCYLMAQAGFGVYTKKTTKLVLIGSNH 1589
Query: 1040 WTRPRTYASPEAIQRTELYEYSKLLGNSQFVLHSFQPYKLIYAHMLAEVGKVSDSLKYCQ 1099
+A+ EAIQRTE YEY++ LG L SFQ +K IY+ LAE+G + + YC+
Sbjct: 1590 SLPFLKFATNEAIQRTEAYEYAQSLGAETCPLPSFQVFKFIYSCRLAEMGLATQAFHYCE 1649
Query: 1100 AVLKSLKT 1107
A+ KS+ T
Sbjct: 1650 AIAKSILT 1657
Score = 55.5 bits (132), Expect = 1e-05
Identities = 76/312 (24%), Positives = 107/312 (33%), Gaps = 85/312 (27%)
Query: 1162 GLPPPAPTSSQATVHGSEQHYQHMAPRVSTSQSTMAMSSLVPSASLEPISEWTADNNRMA 1221
GLPP P Q +H+ + + SL +SE D
Sbjct: 1824 GLPPGVPPL---------QERRHLLQEARSPDPGIVPQEAPVGNSLSELSEENFDGKFAN 1874
Query: 1222 AKPNRSV----------------SEPDIGRSPRQESPSPDAQGKVQVSG---GASRFSRF 1262
P+R+V ++P + SP E+ P K + G S F R+
Sbjct: 1875 LTPSRTVPDSEAPPGWDRADSGPTQPPLSLSPAPETKRPGQAAKKETKEPKKGESWFFRW 1934
Query: 1263 GFGSQLLQKTVGLVLRSGKQAKLGEKNK-FYYDEKLKRWVEEGAEVPAEEAALPPPPPTT 1321
G + + + +KNK +DEK +WV P EE PPPPPT+
Sbjct: 1935 LPGKKKTEAYLP-----------DDKNKSIVWDEKKNQWVN--LNEPEEEKKAPPPPPTS 1981
Query: 1322 AAFQNGSADYNLKSALKTEGLTPNEFSSTRTSSPELSPGMPPIPPSSNQFSARSRLGVRS 1381
T ++P PG P P N +S R+ G R+
Sbjct: 1982 -------------------------MPKTVQAAPPALPGPPGAPV--NMYSRRAA-GTRA 2013
Query: 1382 RYVDTFNQNGGSSANLFQSPSVQSVKPALPANAKF---FIPAPVPSSSEQNMEAIAESNL 1438
RYVD N +G Q +PAL A A F P P+PS+ E L
Sbjct: 2014 RYVDVLNPSG-----------TQRSEPAL-APADFVAPLAPLPIPSNLFVPTPDAEEPQL 2061
Query: 1439 EDSAANENPSTS 1450
D E P+ +
Sbjct: 2062 PDGTGREGPAAA 2073
>emb|CAI13951.1| RP11-413M3.10 [Homo sapiens]
Length = 1059
Score = 128 bits (322), Expect = 1e-27
Identities = 105/368 (28%), Positives = 171/368 (45%), Gaps = 44/368 (11%)
Query: 757 QSFPGPLVGGSVGSKELYKWLDERIARCESPDMDYKKGERLRLLLSLLKIACQHYGKLRS 816
++FPGPL ++ + + +C + K E LL + + + C+ G +
Sbjct: 217 RAFPGPLAKDDTHKVDVINFAQNKAMKCLQNENLIDK-ESASLLWNFIVLLCRQNGTV-- 273
Query: 817 PFGTD----------TILKENDAPESAVAKLFASAKVNGTEFTQYGMPS-HCLQNFPSE- 864
GTD T+ +P A F + V E + G L P+
Sbjct: 274 -VGTDIAELLLRDHRTVWLPGKSPNEANLIDFTNEAVEQVEEEESGEAQLSFLTGGPAAA 332
Query: 865 -EQMKAIASEMQNLLVSGKKMEALQRAQEGQLWGPALVLASQLGEQFYVDTVRQMALRQL 923
++ + LL+ G+K +AL+ A + LWG AL+LAS++ + + + + A L
Sbjct: 333 ASSLERETERFRELLLYGRKKDALESAMKNGLWGHALLLASKMDSRTHARVMTRFA-NSL 391
Query: 924 VAGSPLRTLCLLIAGRPNDVFPTEETSISGHPGAVGMPQQSEQAGSNDMLEDWEENLAVI 983
PL+T+ L++GR MP S G ++ DW +LA++
Sbjct: 392 PINDPLQTVYQLMSGR--------------------MPAASTCCG-DEKWGDWRPHLAMV 430
Query: 984 TANRTKGDEL---VMMHLGDCLWKEKREITAAHICYLIAEVNFSSYSD-ATRLCLIGADH 1039
+N ++ M +GD L + + AAH CYL+A+ F Y+ T+L LIG++H
Sbjct: 431 LSNLNNNMDVESRTMATMGDTL-ASRGLLDAAHFCYLMAQAGFGVYTKKTTKLVLIGSNH 489
Query: 1040 WTRPRTYASPEAIQRTELYEYSKLLGNSQFVLHSFQPYKLIYAHMLAEVGKVSDSLKYCQ 1099
+A+ EAIQRTE YEY++ LG L SFQ +K IY+ LAE+G + + YC+
Sbjct: 490 SLPFLKFATNEAIQRTEAYEYAQSLGAETCPLPSFQVFKFIYSCRLAEMGLATQAFHYCE 549
Query: 1100 AVLKSLKT 1107
A+ KS+ T
Sbjct: 550 AIAKSILT 557
Score = 57.8 bits (138), Expect = 3e-06
Identities = 100/409 (24%), Positives = 137/409 (33%), Gaps = 113/409 (27%)
Query: 1162 GLPPPAPTSSQATVHGSEQHYQHMAPRVSTSQSTMAMSSLVPSASLEPISEWTADNNRMA 1221
GLPP P Q +H+ + + SL +SE D
Sbjct: 724 GLPPGVPPL---------QERRHLLQEARSPDPGIVPQEAPVGNSLSELSEENFDGKFAN 774
Query: 1222 AKPNRSV----------------SEPDIGRSPRQESPSPDAQGKVQVSG---GASRFSRF 1262
P+R+V ++P + SP E+ P K + G S F R+
Sbjct: 775 LTPSRTVPDSEAPPGWDRADSGPTQPPLSLSPAPETKRPGQAAKKETKEPKKGESWFFRW 834
Query: 1263 GFGSQLLQKTVGLVLRSGKQAKLGEKNK-FYYDEKLKRWVEEGAEVPAEEAALPPPPPTT 1321
G + + + +KNK +DEK +WV P EE PPPPPT+
Sbjct: 835 LPGKKKTEAYLP-----------DDKNKSIVWDEKKNQWVN--LNEPEEEKKAPPPPPTS 881
Query: 1322 AAFQNGSADYNLKSALKTEGLTPNEFSSTRTSSPELSPGMPPIPPSSNQFSARSRLGVRS 1381
T ++P PG P P N +S R+ G R+
Sbjct: 882 -------------------------MPKTVQAAPPALPGPPGAPV--NMYSRRAA-GTRA 913
Query: 1382 RYVDTFNQNGGSSANLFQSPSVQSVKPALPANAKF---FIPAPVPSSSEQNMEAIAESNL 1438
RYVD N +G Q +PAL A A F P P+PS+ E L
Sbjct: 914 RYVDVLNPSG-----------TQRSEPAL-APADFVAPLAPLPIPSNLFVPTPDAEEPQL 961
Query: 1439 EDSAANENPSTSSTNDWSYHPPKHAQTMTMQRFPSAGNISKQGQTDGNESHFSHSRRTAS 1498
D E P+ + P+ A P +S G+E SR S
Sbjct: 962 PDGTGREGPAAAR----GLANPEPA--------PEPKVLSSAASLPGSE--LPSSRPEGS 1007
Query: 1499 WSGSFNDSFSPPKMGEIKPSGAALGMPPSAFMP--DPSSLMQGPTRSGS 1545
+ GE A G PPS MP +P+ L Q SGS
Sbjct: 1008 ------------QGGEAPGDLPAAGGPPSGAMPFYNPAQLAQACATSGS 1044
>ref|XP_088459.9| PREDICTED: KIAA0310 [Homo sapiens]
Length = 1984
Score = 128 bits (322), Expect = 1e-27
Identities = 105/368 (28%), Positives = 171/368 (45%), Gaps = 44/368 (11%)
Query: 757 QSFPGPLVGGSVGSKELYKWLDERIARCESPDMDYKKGERLRLLLSLLKIACQHYGKLRS 816
++FPGPL ++ + + +C + K E LL + + + C+ G +
Sbjct: 1122 RAFPGPLAKDDTHKVDVINFAQNKAMKCLQNENLIDK-ESASLLWNFIVLLCRQNGTV-- 1178
Query: 817 PFGTD----------TILKENDAPESAVAKLFASAKVNGTEFTQYGMPS-HCLQNFPSE- 864
GTD T+ +P A F + V E + G L P+
Sbjct: 1179 -VGTDIAELLLRDHRTVWLPGKSPNEANLIDFTNEAVEQVEEEESGEAQLSFLTGGPAAA 1237
Query: 865 -EQMKAIASEMQNLLVSGKKMEALQRAQEGQLWGPALVLASQLGEQFYVDTVRQMALRQL 923
++ + LL+ G+K +AL+ A + LWG AL+LAS++ + + + + A L
Sbjct: 1238 ASSLERETERFRELLLYGRKKDALESAMKNGLWGHALLLASKMDSRTHARVMTRFA-NSL 1296
Query: 924 VAGSPLRTLCLLIAGRPNDVFPTEETSISGHPGAVGMPQQSEQAGSNDMLEDWEENLAVI 983
PL+T+ L++GR MP S G ++ DW +LA++
Sbjct: 1297 PINDPLQTVYQLMSGR--------------------MPAASTCCG-DEKWGDWRPHLAMV 1335
Query: 984 TANRTKGDEL---VMMHLGDCLWKEKREITAAHICYLIAEVNFSSYSD-ATRLCLIGADH 1039
+N ++ M +GD L + + AAH CYL+A+ F Y+ T+L LIG++H
Sbjct: 1336 LSNLNNNMDVESRTMATMGDTL-ASRGLLDAAHFCYLMAQAGFGVYTKKTTKLVLIGSNH 1394
Query: 1040 WTRPRTYASPEAIQRTELYEYSKLLGNSQFVLHSFQPYKLIYAHMLAEVGKVSDSLKYCQ 1099
+A+ EAIQRTE YEY++ LG L SFQ +K IY+ LAE+G + + YC+
Sbjct: 1395 SLPFLKFATNEAIQRTEAYEYAQSLGAETCPLPSFQVFKFIYSCRLAEMGLATQAFHYCE 1454
Query: 1100 AVLKSLKT 1107
A+ KS+ T
Sbjct: 1455 AIAKSILT 1462
Score = 62.4 bits (150), Expect = 1e-07
Identities = 104/414 (25%), Positives = 143/414 (34%), Gaps = 103/414 (24%)
Query: 1162 GLPPPAPTSSQATVHGSEQHYQHMAPRVSTSQSTMAMSSLVPSASLEPISEWTADNNRMA 1221
GLPP P Q +H+ + + SL +SE D
Sbjct: 1629 GLPPGVPPL---------QERRHLLQEARSPDPGIVPQEAPVGNSLSELSEENFDGKFAN 1679
Query: 1222 AKPNRSV----------------SEPDIGRSPRQESPSPDAQGKVQVSG---GASRFSRF 1262
P+R+V ++P + SP E+ P K + G S F R+
Sbjct: 1680 LTPSRTVPDSEAPPGWDRADSGPTQPPLSLSPAPETKRPGQAAKKETKEPKKGESWFFRW 1739
Query: 1263 GFGSQLLQKTVGLVLRSGKQAKLGEKNK-FYYDEKLKRWVEEGAEVPAEEAALPPPPPTT 1321
G + + + +KNK +DEK +WV P EE PPPPPT+
Sbjct: 1740 LPGKKKTEAYLP-----------DDKNKSIVWDEKKNQWVN--LNEPEEEKKAPPPPPTS 1786
Query: 1322 AAFQNGSADYNLKSALKTEGLTPNEFSSTRTSSPELSPGMPPIPPSSNQFSARSRLGVRS 1381
T ++P PG P P N +S R+ G R+
Sbjct: 1787 -------------------------MPKTVQAAPPALPGPPGAPV--NMYSRRAA-GTRA 1818
Query: 1382 RYVDTFNQNGGSSANLFQSPSVQSVKPALPANAKF---FIPAPVPSSSEQNMEAIAESNL 1438
RYVD N +G Q +PAL A A F P P+PS+ E L
Sbjct: 1819 RYVDVLNPSG-----------TQRSEPAL-APADFVAPLAPLPIPSNLFVPTPDAEEPQL 1866
Query: 1439 EDSAANENPSTSS--TNDWSYHPPK---HAQTMTMQRFPSAGNISKQGQTDGNESHFSHS 1493
D E P+ + N PK A ++ PS+ QG S S
Sbjct: 1867 PDGTGREGPAAARGLANPEPAPEPKVLSSAASLPGSELPSSRPEGSQGGELSRCSSMSSL 1926
Query: 1494 RRTASWSGSFNDSFSPPKMGEIKPSGAALGMPPSAFMP--DPSSLMQGPTRSGS 1545
R S FN + G++ A G PPS MP +P+ L Q SGS
Sbjct: 1927 SREV--SQHFNQA-----PGDL----PAAGGPPSGAMPFYNPAQLAQACATSGS 1969
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.310 0.128 0.370
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,825,271,344
Number of Sequences: 2540612
Number of extensions: 129530699
Number of successful extensions: 448024
Number of sequences better than 10.0: 3674
Number of HSP's better than 10.0 without gapping: 789
Number of HSP's successfully gapped in prelim test: 3092
Number of HSP's that attempted gapping in prelim test: 372116
Number of HSP's gapped (non-prelim): 25832
length of query: 1555
length of database: 863,360,394
effective HSP length: 142
effective length of query: 1413
effective length of database: 502,593,490
effective search space: 710164601370
effective search space used: 710164601370
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 82 (36.2 bits)
Medicago: description of AC137077.1


