

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC133139.1 - phase: 0 /pseudo
(365 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
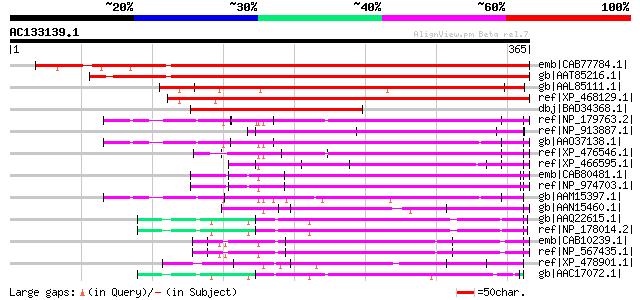
Score E
Sequences producing significant alignments: (bits) Value
emb|CAB77784.1| hypothetical protein [Arabidopsis thaliana] gi|1... 439 e-122
gb|AAT85216.1| unknown protein [Oryza sativa (japonica cultivar-... 365 1e-99
gb|AAL85111.1| unknown protein [Arabidopsis thaliana] gi|1453259... 295 1e-78
ref|XP_468129.1| mitochondrial transcription termination factor-... 267 3e-70
dbj|BAD34368.1| hypothetical protein [Oryza sativa (japonica cul... 166 8e-40
ref|NP_179763.2| mitochondrial transcription termination factor-... 94 6e-18
ref|NP_913887.1| unknown protein [Oryza sativa (japonica cultiva... 94 8e-18
gb|AAO37138.1| hypothetical protein [Arabidopsis thaliana] 91 4e-17
ref|XP_476546.1| hypothetical protein [Oryza sativa (japonica cu... 90 9e-17
ref|XP_466595.1| mitochondrial transcription termination factor-... 87 8e-16
emb|CAB80481.1| putative protein [Arabidopsis thaliana] gi|44671... 81 4e-14
ref|NP_974703.1| mitochondrial transcription termination factor-... 81 4e-14
gb|AAM15397.1| hypothetical protein [Arabidopsis thaliana] gi|44... 77 1e-12
gb|AAN15460.1| Unknown protein [Arabidopsis thaliana] gi|9758592... 76 1e-12
gb|AAQ22615.1| At1g78930 [Arabidopsis thaliana] 75 4e-12
ref|NP_178014.2| mitochondrial transcription termination factor-... 75 4e-12
emb|CAB10239.1| hypothetical protein [Arabidopsis thaliana] gi|7... 74 9e-12
ref|NP_567435.1| mitochondrial transcription termination factor-... 74 9e-12
ref|XP_478901.1| unknown protein [Oryza sativa (japonica cultiva... 73 1e-11
gb|AAC17072.1| Similar to hypothetical protein gb|Z97336 from A.... 70 1e-10
>emb|CAB77784.1| hypothetical protein [Arabidopsis thaliana]
gi|15236230|ref|NP_192208.1| mitochondrial transcription
termination factor family protein / mTERF family protein
[Arabidopsis thaliana] gi|3924606|gb|AAC79107.1|
hypothetical protein [Arabidopsis thaliana]
gi|7487645|pir||T01394 hypothetical protein T4I9.13 -
Arabidopsis thaliana
Length = 541
Score = 439 bits (1130), Expect = e-122
Identities = 238/361 (65%), Positives = 273/361 (74%), Gaps = 22/361 (6%)
Query: 19 MNFSGTITKPNFPF----PPTEMPILTFGHSKLTAATLALQLPCNYIRK--IRFITKLQC 72
+ F TKP F PP+ + + S T L CN+ ++ +QC
Sbjct: 3 IRFCNGFTKPGFLLVHFEPPSFFAVRSRSLSDSTYGNL-----CNHKKRPGTGIGLTVQC 57
Query: 73 SVAGRTDSYRT--------SAADSQGSKRDSGEIQRKRRGASSLYAGYARPSLSEMKKDK 124
++A R S R+ S+ S S RD + K R + SLY+ RPSL +M K+K
Sbjct: 58 AIANRRFSSRSLDSPRRERSSRSSSSSGRDRDRDKDKGRDSKSLYS---RPSLLDMNKEK 114
Query: 125 ATLRKVVYEFLRGIGIVPDELDGLELPVTVDVMKERVDFLHSLGLTIEDINNYPLVLGCS 184
A R VYEFLRGIGIVPDELDGLELPVT DVMKERV+FLH LGLTIEDINNYPLVLGCS
Sbjct: 115 AANRAKVYEFLRGIGIVPDELDGLELPVTADVMKERVEFLHKLGLTIEDINNYPLVLGCS 174
Query: 185 VKKNMVPVLDYLGKLGVRKSTITQFLRTYPQVLHASVVVDLVPVVKYLQGMDIKPDDIPR 244
VKKNMVPVLDYLGKLGVRKST T+FLR YPQVLH+SVV+DL PVVKYLQG+DIKP D+PR
Sbjct: 175 VKKNMVPVLDYLGKLGVRKSTFTEFLRRYPQVLHSSVVIDLAPVVKYLQGLDIKPSDVPR 234
Query: 245 VLERYPEVLGFKLEGTMSTSVAYLIGIGVGRRELGGILTRFPEILGMRVGRVIKPFVEYL 304
VLERYPEVLGFKLEGTMSTSVAYL+GIGV RRE+GGILTR+PEILGMRV R+IKP VEYL
Sbjct: 235 VLERYPEVLGFKLEGTMSTSVAYLVGIGVARREIGGILTRYPEILGMRVARIIKPLVEYL 294
Query: 305 ESLGIPRLAIARLIETQPYILGFDLDEKVKPNVKSLEEFNVRETSLASIIAQYPDIIGTD 364
E LGIPRLA ARLIE +P+ILGF+LD+ VKPNV+ L++FNVRETSL SIIAQYP+IIG D
Sbjct: 295 EVLGIPRLAAARLIEKRPHILGFELDDTVKPNVQILQDFNVRETSLPSIIAQYPEIIGID 354
Query: 365 L 365
L
Sbjct: 355 L 355
>gb|AAT85216.1| unknown protein [Oryza sativa (japonica cultivar-group)]
Length = 508
Score = 365 bits (937), Expect = 1e-99
Identities = 186/309 (60%), Positives = 237/309 (76%), Gaps = 7/309 (2%)
Query: 57 PCNYIRKIRFITKLQCSVAGRTDSYRTSAADSQGSKRDSGEIQRKRRGASSLYAGYARPS 116
P ++RK+ +T + A + S A S+G + +SSLYA RPS
Sbjct: 19 PSPHLRKLLRLT----ASASTSASSPPRAGCSRGPAHARPRPSPRPSPSSSLYA---RPS 71
Query: 117 LSEMKKDKATLRKVVYEFLRGIGIVPDELDGLELPVTVDVMKERVDFLHSLGLTIEDINN 176
L +M++ +A R V FL +G+ P EL GLELP TVDVM+ERV+FLHSL L+ ED+
Sbjct: 72 LLDMERGRAARRADVDAFLASLGVDPGELAGLELPATVDVMRERVEFLHSLDLSNEDLAA 131
Query: 177 YPLVLGCSVKKNMVPVLDYLGKLGVRKSTITQFLRTYPQVLHASVVVDLVPVVKYLQGMD 236
YPL LGCSV+KNMVPVLDYLGKLGVR+ + LR YPQVLHASVVVDL PVVKYLQGMD
Sbjct: 132 YPLALGCSVRKNMVPVLDYLGKLGVRQDALPDLLRRYPQVLHASVVVDLAPVVKYLQGMD 191
Query: 237 IKPDDIPRVLERYPEVLGFKLEGTMSTSVAYLIGIGVGRRELGGILTRFPEILGMRVGRV 296
++P D+PRVLERYPE+LGFKLEGTMSTS+AYL+GIGV RR++G ++TRFPE+LGMRVG++
Sbjct: 192 VRPHDVPRVLERYPELLGFKLEGTMSTSIAYLVGIGVARRQVGSVITRFPEVLGMRVGKI 251
Query: 297 IKPFVEYLESLGIPRLAIARLIETQPYILGFDLDEKVKPNVKSLEEFNVRETSLASIIAQ 356
IKPFVE+LE +G+ RLAIAR+IE +PY+LGF L++KVKPN+++L EF VR+ +LA I+AQ
Sbjct: 252 IKPFVEHLEGIGLQRLAIARIIEKKPYVLGFGLEDKVKPNIEALLEFGVRKEALAFIVAQ 311
Query: 357 YPDIIGTDL 365
YPDI+G +L
Sbjct: 312 YPDILGIEL 320
>gb|AAL85111.1| unknown protein [Arabidopsis thaliana] gi|14532592|gb|AAK64024.1|
unknown protein [Arabidopsis thaliana]
gi|3212859|gb|AAC23410.1| expressed protein [Arabidopsis
thaliana] gi|7486477|pir||T00682 hypothetical protein
At2g44020 [imported] - Arabidopsis thaliana
gi|18406426|ref|NP_566005.1| mitochondrial transcription
termination factor-related / mTERF-related [Arabidopsis
thaliana]
Length = 507
Score = 295 bits (755), Expect = 1e-78
Identities = 140/261 (53%), Positives = 203/261 (77%), Gaps = 4/261 (1%)
Query: 106 SSLYAGYARPSLS----EMKKDKATLRKVVYEFLRGIGIVPDELDGLELPVTVDVMKERV 161
SS + Y P+++ + KK+K R + ++L+G+GI+ DEL+ +ELP T++VM ERV
Sbjct: 57 SSKFPEYEMPTVTWGVIQGKKEKLVNRVKICDYLKGLGIITDELESIELPSTIEVMCERV 116
Query: 162 DFLHSLGLTIEDINNYPLVLGCSVKKNMVPVLDYLGKLGVRKSTITQFLRTYPQVLHASV 221
+FL LGLTI+DIN YPL+LGCSV+KN++PVL YL K+G+ +S + +F++ YPQVLHASV
Sbjct: 117 EFLQKLGLTIDDINEYPLMLGCSVRKNLIPVLAYLEKIGISRSKLGEFVKNYPQVLHASV 176
Query: 222 VVDLVPVVKYLQGMDIKPDDIPRVLERYPEVLGFKLEGTMSTSVAYLIGIGVGRRELGGI 281
VV+L PVVK+L+G+D++ D+ VL +YPE+LGFKLEGTMSTSVAYL+ IGV R++G +
Sbjct: 177 VVELAPVVKFLRGLDVEKQDLGYVLMKYPELLGFKLEGTMSTSVAYLVSIGVSPRDIGPM 236
Query: 282 LTRFPEILGMRVGRVIKPFVEYLESLGIPRLAIARLIETQPYILGFDLDEKVKPNVKSLE 341
+T++P +LGMRVG +IKP V+YL S+G+P+ +AR++E + YI+G++L+E VKPNV L
Sbjct: 237 VTQYPYLLGMRVGTMIKPLVDYLISIGLPKKIVARMLEKRSYIVGYNLEETVKPNVDCLI 296
Query: 342 EFNVRETSLASIIAQYPDIIG 362
F V++ L +IAQYP I+G
Sbjct: 297 SFGVKKELLPLLIAQYPQILG 317
Score = 67.0 bits (162), Expect = 9e-10
Identities = 59/268 (22%), Positives = 112/268 (41%), Gaps = 57/268 (21%)
Query: 131 VYEFLRGIGIVPDELD----------GLELPVTVDVMKERVDFLHSLGLTIEDIN----N 176
V +FLRG+ + +L G +L T M V +L S+G++ DI
Sbjct: 183 VVKFLRGLDVEKQDLGYVLMKYPELLGFKLEGT---MSTSVAYLVSIGVSPRDIGPMVTQ 239
Query: 177 YPLVLGCSVKKNMVPVLDYLGKLGVRKSTITQFLRTYPQVLHASVVVDLVPVVKYLQGMD 236
YP +LG V + P++DYL +G+ K + + L ++ ++ + P V L
Sbjct: 240 YPYLLGMRVGTMIKPLVDYLISIGLPKKIVARMLEKRSYIVGYNLEETVKPNVDCLISFG 299
Query: 237 IKPDDIPRVLERYPEVLGFKLEGTMSTS-------------------------------- 264
+K + +P ++ +YP++LG ++ MST
Sbjct: 300 VKKELLPLLIAQYPQILGLPVKAKMSTQQYFFSLKLKIDPEGFARVVEKMPQIVSLKQNV 359
Query: 265 ----VAYLIGIGVGRRELGGILTRFPEILGMRVGRVIKPFVEYLESLGIPRLAIARLIET 320
+ +L+G ++ ++ R P+IL RV + + Y +G P + L+E
Sbjct: 360 IMKPIEFLLGRAFQVEDIAKMVVRCPQILCSRVELMKNSYYFYKTEMGRP---MKELVEY 416
Query: 321 QPYILGFDLDEKVKPNVKSLEEFNVRET 348
Y + L+ ++KP + L+ +R +
Sbjct: 417 PEYFT-YSLESRIKPRYQKLQSKGIRSS 443
>ref|XP_468129.1| mitochondrial transcription termination factor-like [Oryza sativa
(japonica cultivar-group)] gi|51964458|ref|XP_507014.1|
PREDICTED OJ1311_D08.15 gene product [Oryza sativa
(japonica cultivar-group)] gi|47497486|dbj|BAD19540.1|
mitochondrial transcription termination factor-like
[Oryza sativa (japonica cultivar-group)]
Length = 485
Score = 267 bits (683), Expect = 3e-70
Identities = 133/261 (50%), Positives = 189/261 (71%), Gaps = 7/261 (2%)
Query: 112 YARPSLS----EMKKDKATLRKVVYEFLRGIGIVPD--ELDGLELPVTVDVMKERVDFLH 165
Y PS++ + +K++ R + +FLR G+ EL+ +ELP +++V++ER+DFL
Sbjct: 38 YEMPSVTWGVIQGRKERLVSRVLALDFLRSAGVSDPAGELEAVELPSSLEVLQERLDFLL 97
Query: 166 SLGLTIEDINNYPLVLGCSVKKNMVPVLDYLGKLGVRKSTITQFLRTYPQVLHASVVVDL 225
LGL+ +D++ YPL+L CS++KN +PVL YL KLGV ++ + F+R YP LHASV VDL
Sbjct: 98 RLGLSTDDLSAYPLLLACSLRKNAIPVLSYLEKLGVTRARLAAFVRAYPACLHASVAVDL 157
Query: 226 VPVVKYLQGMDIKPDDIPRVLERYPEVLGFKLEGTMSTSVAYLIGI-GVGRRELGGILTR 284
PVVK L+G+D+ D+PRVLERYP++LG K +GT+STSVAYL+GI GV R++G ++T
Sbjct: 158 TPVVKSLRGLDVDRQDLPRVLERYPDILGLKPDGTISTSVAYLVGIVGVAPRDIGPMVTH 217
Query: 285 FPEILGMRVGRVIKPFVEYLESLGIPRLAIARLIETQPYILGFDLDEKVKPNVKSLEEFN 344
FP LGMRVG IKP EY+ SLG+P +AR++E +PYILG+DL+E VKPNV++L F
Sbjct: 218 FPFFLGMRVGTTIKPLCEYITSLGLPMRILARILEKRPYILGYDLEETVKPNVEALLSFG 277
Query: 345 VRETSLASIIAQYPDIIGTDL 365
+R+ L +IAQYP I+G L
Sbjct: 278 IRKEMLPLVIAQYPPILGLPL 298
Score = 38.9 bits (89), Expect = 0.25
Identities = 29/111 (26%), Positives = 55/111 (49%), Gaps = 10/111 (9%)
Query: 155 DVMKERVDFLHSLGLTIED----INNYPLVLGCSVKKNMVPVLDYLG-KLGVRKSTITQF 209
+ +K V+ L S G+ E I YP +LG +K + + KL +
Sbjct: 264 ETVKPNVEALLSFGIRKEMLPLVIAQYPPILGLPLKTKLAAQQYFFNLKLQIDPDAFACA 323
Query: 210 LRTYPQV--LHASVVVDLVPVVKYLQGMDIKPDDIPRVLERYPEVLGFKLE 258
+ PQ+ LH ++++ LV ++L+G I +D+ R++ R P++L ++E
Sbjct: 324 IEKLPQLVSLHQNIILKLV---EFLRGRGISNEDVARMVVRCPQILLLRME 371
>dbj|BAD34368.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|50726597|dbj|BAD34231.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 416
Score = 166 bits (421), Expect = 8e-40
Identities = 83/121 (68%), Positives = 96/121 (78%)
Query: 128 RKVVYEFLRGIGIVPDELDGLELPVTVDVMKERVDFLHSLGLTIEDINNYPLVLGCSVKK 187
R V FL +G+ P EL GLELP TVDVM+ERV+FL SLGL+ E + YPL LGCSV+K
Sbjct: 199 RADVDAFLASLGVDPGELAGLELPATVDVMRERVEFLQSLGLSNEGLAAYPLALGCSVRK 258
Query: 188 NMVPVLDYLGKLGVRKSTITQFLRTYPQVLHASVVVDLVPVVKYLQGMDIKPDDIPRVLE 247
NMVPVLDYLGKLGVR+ + LR YPQVLHASVVVDL PVVKYLQ MD++P ++PRVLE
Sbjct: 259 NMVPVLDYLGKLGVRQDALPDLLRRYPQVLHASVVVDLAPVVKYLQVMDVRPHEVPRVLE 318
Query: 248 R 248
R
Sbjct: 319 R 319
Score = 40.4 bits (93), Expect = 0.085
Identities = 26/104 (25%), Positives = 50/104 (48%), Gaps = 4/104 (3%)
Query: 229 VKYLQGMDIKPDDIPRVLERYPEVLGFKLEGTMSTSVAYLIGIGVGRRELGGILTRFPEI 288
V++LQ + + + L YP LG + M + YL +GV + L +L R+P++
Sbjct: 232 VEFLQSLGLSNEG----LAAYPLALGCSVRKNMVPVLDYLGKLGVRQDALPDLLRRYPQV 287
Query: 289 LGMRVGRVIKPFVEYLESLGIPRLAIARLIETQPYILGFDLDEK 332
L V + P V+YL+ + + + R++E ++ L +
Sbjct: 288 LHASVVVDLAPVVKYLQVMDVRPHEVPRVLERVEFLHSLGLSAR 331
Score = 38.5 bits (88), Expect = 0.32
Identities = 28/107 (26%), Positives = 50/107 (46%), Gaps = 7/107 (6%)
Query: 251 EVLGFKLEGT---MSTSVAYLIGIGVGRRELGGILTRFPEILGMRVGRVIKPFVEYLESL 307
E+ G +L T M V +L +G+ L +P LG V + + P ++YL L
Sbjct: 215 ELAGLELPATVDVMRERVEFLQSLGLSNEGLAA----YPLALGCSVRKNMVPVLDYLGKL 270
Query: 308 GIPRLAIARLIETQPYILGFDLDEKVKPNVKSLEEFNVRETSLASII 354
G+ + A+ L+ P +L + + P VK L+ +VR + ++
Sbjct: 271 GVRQDALPDLLRRYPQVLHASVVVDLAPVVKYLQVMDVRPHEVPRVL 317
Score = 38.1 bits (87), Expect = 0.42
Identities = 26/96 (27%), Positives = 48/96 (49%), Gaps = 11/96 (11%)
Query: 266 AYLIGIGVGRRELGGILTRFPEILGMRVGRVIKPFVEYLESLGIPRLAIARLIETQPYIL 325
A+L +GV EL G+ P + V++ VE+L+SLG+ +A P L
Sbjct: 204 AFLASLGVDPGELAGL--ELPATVD-----VMRERVEFLQSLGLSNEGLA----AYPLAL 252
Query: 326 GFDLDEKVKPNVKSLEEFNVRETSLASIIAQYPDII 361
G + + + P + L + VR+ +L ++ +YP ++
Sbjct: 253 GCSVRKNMVPVLDYLGKLGVRQDALPDLLRRYPQVL 288
>ref|NP_179763.2| mitochondrial transcription termination factor-related /
mTERF-related [Arabidopsis thaliana]
Length = 640
Score = 94.0 bits (232), Expect = 6e-18
Identities = 71/305 (23%), Positives = 141/305 (45%), Gaps = 21/305 (6%)
Query: 67 ITKLQCSVAGRTDSYRTSAADSQGSKRDSGEIQRKRRGASSLYAGYARPSLSEMKKDKAT 126
I K+ C G DS R + S GE + + R + +++++
Sbjct: 231 IAKIICMSKGNLDSIRIMI-EWLKSIHVKGEF---------IAVAFLRSGDNILQRNREE 280
Query: 127 LRKVVYEFLRGIGIVPDELDGLE------LPVTVDVMKERVDFLHSLGLTIED----INN 176
L ++V E+L G+ D + + L +++ +K RVDF +G+ D + +
Sbjct: 281 LNEIV-EYLESNGVRRDWMGYVVGRCPELLSFSMEEVKSRVDFFLKMGMNQNDFGTMVYD 339
Query: 177 YPLVLGCSVKKNMVPVLDYLGKLGVRKSTITQFLRTYPQVLHASVVVDLVPVVKYLQGMD 236
YP ++G + M ++YL + G+ + + L P ++ S+ P+VKY +
Sbjct: 340 YPKIIGFFSFQVMEKKINYLKEFGLSTEEVGRLLAYKPHLMGCSIEERWKPLVKYFYYLG 399
Query: 237 IKPDDIPRVLERYPEVLGFKLEGTMSTSVAYLIGIGVGRRELGGILTRFPEILGMRVGRV 296
I + + R+L P + LE T++ V +L +G+ +G +L +FP +L + +
Sbjct: 400 IPKEGMKRILVVKPILYCIDLEKTIAPKVRFLQEMGIPNEAIGNMLVKFPSLLTNSLYKK 459
Query: 297 IKPFVEYLESLGIPRLAIARLIETQPYILGFDLDEKVKPNVKSLEEFNVRETSLASIIAQ 356
I+P + L G+ + I ++I P +LG + K++PN++ +R L +IA
Sbjct: 460 IRPVIFLLTRAGVTQKDIGKVIAMDPALLGCSIGTKLEPNMRYYISLGIRFYQLGEMIAD 519
Query: 357 YPDII 361
+P ++
Sbjct: 520 FPMLL 524
Score = 85.1 bits (209), Expect = 3e-15
Identities = 49/195 (25%), Positives = 101/195 (51%), Gaps = 5/195 (2%)
Query: 156 VMKERVDFLHSLGLTIEDINNY----PLVLGCSVKKNMVPVLDYLGKLGVRKSTITQFLR 211
VM++++++L GL+ E++ P ++GCS+++ P++ Y LG+ K + + L
Sbjct: 351 VMEKKINYLKEFGLSTEEVGRLLAYKPHLMGCSIEERWKPLVKYFYYLGIPKEGMKRILV 410
Query: 212 TYPQVLHASVVVDLVPVVKYLQGMDIKPDDIPRVLERYPEVLGFKLEGTMSTSVAYLIGI 271
P + + + P V++LQ M I + I +L ++P +L L + + L
Sbjct: 411 VKPILYCIDLEKTIAPKVRFLQEMGIPNEAIGNMLVKFPSLLTNSLYKKIRPVIFLLTRA 470
Query: 272 GVGRRELGGILTRFPEILGMRVGRVIKPFVEYLESLGIPRLAIARLIETQPYILGFDLDE 331
GV ++++G ++ P +LG +G ++P + Y SLGI + +I P +L +++D
Sbjct: 471 GVTQKDIGKVIAMDPALLGCSIGTKLEPNMRYYISLGIRFYQLGEMIADFPMLLRYNVD- 529
Query: 332 KVKPNVKSLEEFNVR 346
++P + L +R
Sbjct: 530 NLRPKYRYLRRTMIR 544
Score = 81.6 bits (200), Expect = 3e-14
Identities = 42/180 (23%), Positives = 93/180 (51%), Gaps = 1/180 (0%)
Query: 186 KKNMVPVLDYLGKLGVRKSTITQFLRTYPQVLHASVVVDLVPVVKYLQGMDIKPDDIPRV 245
++ + +++YL GVR+ + + P++L S+ ++ V + M + +D +
Sbjct: 278 REELNEIVEYLESNGVRRDWMGYVVGRCPELLSFSME-EVKSRVDFFLKMGMNQNDFGTM 336
Query: 246 LERYPEVLGFKLEGTMSTSVAYLIGIGVGRRELGGILTRFPEILGMRVGRVIKPFVEYLE 305
+ YP+++GF M + YL G+ E+G +L P ++G + KP V+Y
Sbjct: 337 VYDYPKIIGFFSFQVMEKKINYLKEFGLSTEEVGRLLAYKPHLMGCSIEERWKPLVKYFY 396
Query: 306 SLGIPRLAIARLIETQPYILGFDLDEKVKPNVKSLEEFNVRETSLASIIAQYPDIIGTDL 365
LGIP+ + R++ +P + DL++ + P V+ L+E + ++ +++ ++P ++ L
Sbjct: 397 YLGIPKEGMKRILVVKPILYCIDLEKTIAPKVRFLQEMGIPNEAIGNMLVKFPSLLTNSL 456
Score = 69.7 bits (169), Expect = 1e-10
Identities = 47/213 (22%), Positives = 101/213 (47%), Gaps = 5/213 (2%)
Query: 157 MKERVDFLHSLGLTIE----DINNYPLVLGCSVKKNMVPVLDYLGKLGVRKSTITQFLRT 212
+ E V++L S G+ + + P +L S+++ V D+ K+G+ ++ +
Sbjct: 281 LNEIVEYLESNGVRRDWMGYVVGRCPELLSFSMEEVKSRV-DFFLKMGMNQNDFGTMVYD 339
Query: 213 YPQVLHASVVVDLVPVVKYLQGMDIKPDDIPRVLERYPEVLGFKLEGTMSTSVAYLIGIG 272
YP+++ + + YL+ + +++ R+L P ++G +E V Y +G
Sbjct: 340 YPKIIGFFSFQVMEKKINYLKEFGLSTEEVGRLLAYKPHLMGCSIEERWKPLVKYFYYLG 399
Query: 273 VGRRELGGILTRFPEILGMRVGRVIKPFVEYLESLGIPRLAIARLIETQPYILGFDLDEK 332
+ + + IL P + + + + I P V +L+ +GIP AI ++ P +L L +K
Sbjct: 400 IPKEGMKRILVVKPILYCIDLEKTIAPKVRFLQEMGIPNEAIGNMLVKFPSLLTNSLYKK 459
Query: 333 VKPNVKSLEEFNVRETSLASIIAQYPDIIGTDL 365
++P + L V + + +IA P ++G +
Sbjct: 460 IRPVIFLLTRAGVTQKDIGKVIAMDPALLGCSI 492
>ref|NP_913887.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|28201276|dbj|BAC56785.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
Length = 333
Score = 93.6 bits (231), Expect = 8e-18
Identities = 51/190 (26%), Positives = 103/190 (53%), Gaps = 2/190 (1%)
Query: 174 INNYPLVLGCSVKKNMVPVLDYLGKLGVRKSTITQFLRTYPQVLHASVVVDLVPVVKYLQ 233
+ P VL SV +VP + L L + + Q + +PQ+L SV L P++ + Q
Sbjct: 66 VTKCPKVLTLSVDDKLVPTVQCLTTLQAKPGEVAQAIVKFPQILFHSVEEKLCPLLAFFQ 125
Query: 234 GMDIKPDDIPRVLERYPEVLGFKLEGTMSTSVAYLIGIGVGRR-ELGGILTRFPEILGMR 292
+ I + ++L P ++ + +E S +V +L+G+G+ + +G I+ + P I+G
Sbjct: 126 TLGISEKQLAKLLMVNPRLISYSIEAKFSQTVNFLVGLGIDKEGMIGKIMAKEPYIMGYS 185
Query: 293 VGRVIKPFVEYLES-LGIPRLAIARLIETQPYILGFDLDEKVKPNVKSLEEFNVRETSLA 351
V + ++P E+L+S +G+ + R+I + P IL D+D+ ++PN+ L+ + +
Sbjct: 186 VDKRLRPTAEFLKSAVGLEGSNLQRVIMSFPDILSRDVDKILRPNLAFLQSCGFSKDQVM 245
Query: 352 SIIAQYPDII 361
+++A YP ++
Sbjct: 246 ALVAGYPPVL 255
Score = 69.7 bits (169), Expect = 1e-10
Identities = 40/177 (22%), Positives = 90/177 (50%), Gaps = 2/177 (1%)
Query: 168 GLTIEDINNYPLVLGCSVKKNMVPVLDYLGKLGVRKSTITQFLRTYPQVLHASVVVDLVP 227
G + I +P +L SV++ + P+L + LG+ + + + L P+++ S+
Sbjct: 96 GEVAQAIVKFPQILFHSVEEKLCPLLAFFQTLGISEKQLAKLLMVNPRLISYSIEAKFSQ 155
Query: 228 VVKYLQGMDI-KPDDIPRVLERYPEVLGFKLEGTMSTSVAYL-IGIGVGRRELGGILTRF 285
V +L G+ I K I +++ + P ++G+ ++ + + +L +G+ L ++ F
Sbjct: 156 TVNFLVGLGIDKEGMIGKIMAKEPYIMGYSVDKRLRPTAEFLKSAVGLEGSNLQRVIMSF 215
Query: 286 PEILGMRVGRVIKPFVEYLESLGIPRLAIARLIETQPYILGFDLDEKVKPNVKSLEE 342
P+IL V ++++P + +L+S G + + L+ P +L + ++P +K L E
Sbjct: 216 PDILSRDVDKILRPNLAFLQSCGFSKDQVMALVAGYPPVLIKSVKHCLEPRMKFLVE 272
Score = 48.1 bits (113), Expect = 4e-04
Identities = 28/119 (23%), Positives = 59/119 (49%), Gaps = 1/119 (0%)
Query: 245 VLERYPEVLGFKLEGTMSTSVAYLIGIGVGRRELGGILTRFPEILGMRVGRVIKPFVEYL 304
V+ + P+VL ++ + +V L + E+ + +FP+IL V + P + +
Sbjct: 65 VVTKCPKVLTLSVDDKLVPTVQCLTTLQAKPGEVAQAIVKFPQILFHSVEEKLCPLLAFF 124
Query: 305 ESLGIPRLAIARLIETQPYILGFDLDEKVKPNVKSLEEFNV-RETSLASIIAQYPDIIG 362
++LGI +A+L+ P ++ + ++ K V L + +E + I+A+ P I+G
Sbjct: 125 QTLGISEKQLAKLLMVNPRLISYSIEAKFSQTVNFLVGLGIDKEGMIGKIMAKEPYIMG 183
Score = 40.8 bits (94), Expect = 0.065
Identities = 26/96 (27%), Positives = 50/96 (52%), Gaps = 1/96 (1%)
Query: 267 YLIGI-GVGRRELGGILTRFPEILGMRVGRVIKPFVEYLESLGIPRLAIARLIETQPYIL 325
YL+ + + RR+L ++T+ P++L + V + P V+ L +L +A+ I P IL
Sbjct: 50 YLLNVVKIERRKLRYVVTKCPKVLTLSVDDKLVPTVQCLTTLQAKPGEVAQAIVKFPQIL 109
Query: 326 GFDLDEKVKPNVKSLEEFNVRETSLASIIAQYPDII 361
++EK+ P + + + E LA ++ P +I
Sbjct: 110 FHSVEEKLCPLLAFFQTLGISEKQLAKLLMVNPRLI 145
>gb|AAO37138.1| hypothetical protein [Arabidopsis thaliana]
Length = 429
Score = 91.3 bits (225), Expect = 4e-17
Identities = 72/306 (23%), Positives = 142/306 (45%), Gaps = 22/306 (7%)
Query: 67 ITKLQCSVAGRTDSYRTSAADSQGSKRDSGEIQRKRRGASSLYAGYARPSLSEMKKDKAT 126
I K+ C G DS R + S GE + + R + +++++
Sbjct: 94 IAKIICMSKGNLDSIRIMI-EWLKSIHVKGEF---------IAVAFLRSGDNILQRNREE 143
Query: 127 LRKVVYEFLRGIGIVPDELDGLE------LPVTVDVMKERVDFLHSLGLTIED----INN 176
L ++V E+L G+ D + + L +++ +K RVDF +G+ D + +
Sbjct: 144 LNEIV-EYLESNGVRRDWMGYVVGRCPELLSFSMEEVKSRVDFFLKMGMNQNDFGTMVYD 202
Query: 177 YPLVLGCSVKKNMVPVLDYLGKLGVRKSTITQFLRTYPQVLHASVVVDLVPVVKYLQGMD 236
YP ++G + M ++YL + G+ + + L P ++ S+ P+VKY +
Sbjct: 203 YPKIIGFFSFQVMEKKINYLKEFGLSTEEVGRLLAYKPHLMGCSIEERWKPLVKYFYYLG 262
Query: 237 IKPDDIPRVLERYPEVLGFKLEGTMSTSVAYLIGIGVGRRELGGILTRFPEILGMRVGRV 296
I + + R+L P + LE T++ V +L +G+ +G +L +FP +L + +
Sbjct: 263 IPKEGMKRILVVKPILYCIDLEKTIAPKVRFLQEMGIPNEAIGNMLVKFPSLLTNSLYKK 322
Query: 297 IKPFVEY-LESLGIPRLAIARLIETQPYILGFDLDEKVKPNVKSLEEFNVRETSLASIIA 355
I+P V + L G+ + I ++I P +LG + K++PN++ +R L +IA
Sbjct: 323 IRPVVIFLLTRAGVTQKDIGKVIAMDPALLGCSIGTKLEPNMRYYISLGIRFYQLGEMIA 382
Query: 356 QYPDII 361
+P ++
Sbjct: 383 DFPMLL 388
Score = 84.0 bits (206), Expect = 7e-15
Identities = 50/196 (25%), Positives = 103/196 (52%), Gaps = 6/196 (3%)
Query: 156 VMKERVDFLHSLGLTIEDINNY----PLVLGCSVKKNMVPVLDYLGKLGVRKSTITQFLR 211
VM++++++L GL+ E++ P ++GCS+++ P++ Y LG+ K + + L
Sbjct: 214 VMEKKINYLKEFGLSTEEVGRLLAYKPHLMGCSIEERWKPLVKYFYYLGIPKEGMKRILV 273
Query: 212 TYPQVLHASVVVDLVPVVKYLQGMDIKPDDIPRVLERYPEVLGFKLEGTMSTSVAYLIG- 270
P + + + P V++LQ M I + I +L ++P +L L + V +L+
Sbjct: 274 VKPILYCIDLEKTIAPKVRFLQEMGIPNEAIGNMLVKFPSLLTNSLYKKIRPVVIFLLTR 333
Query: 271 IGVGRRELGGILTRFPEILGMRVGRVIKPFVEYLESLGIPRLAIARLIETQPYILGFDLD 330
GV ++++G ++ P +LG +G ++P + Y SLGI + +I P +L +++D
Sbjct: 334 AGVTQKDIGKVIAMDPALLGCSIGTKLEPNMRYYISLGIRFYQLGEMIADFPMLLRYNVD 393
Query: 331 EKVKPNVKSLEEFNVR 346
++P + L +R
Sbjct: 394 -NLRPKYRYLRRTMIR 408
Score = 81.6 bits (200), Expect = 3e-14
Identities = 42/180 (23%), Positives = 93/180 (51%), Gaps = 1/180 (0%)
Query: 186 KKNMVPVLDYLGKLGVRKSTITQFLRTYPQVLHASVVVDLVPVVKYLQGMDIKPDDIPRV 245
++ + +++YL GVR+ + + P++L S+ ++ V + M + +D +
Sbjct: 141 REELNEIVEYLESNGVRRDWMGYVVGRCPELLSFSME-EVKSRVDFFLKMGMNQNDFGTM 199
Query: 246 LERYPEVLGFKLEGTMSTSVAYLIGIGVGRRELGGILTRFPEILGMRVGRVIKPFVEYLE 305
+ YP+++GF M + YL G+ E+G +L P ++G + KP V+Y
Sbjct: 200 VYDYPKIIGFFSFQVMEKKINYLKEFGLSTEEVGRLLAYKPHLMGCSIEERWKPLVKYFY 259
Query: 306 SLGIPRLAIARLIETQPYILGFDLDEKVKPNVKSLEEFNVRETSLASIIAQYPDIIGTDL 365
LGIP+ + R++ +P + DL++ + P V+ L+E + ++ +++ ++P ++ L
Sbjct: 260 YLGIPKEGMKRILVVKPILYCIDLEKTIAPKVRFLQEMGIPNEAIGNMLVKFPSLLTNSL 319
>ref|XP_476546.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|34394417|dbj|BAC83514.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 608
Score = 90.1 bits (222), Expect = 9e-17
Identities = 56/217 (25%), Positives = 107/217 (48%), Gaps = 5/217 (2%)
Query: 150 LPVTVDVMKERVDFLHSLGLTIED----INNYPLVLGCSVKKNMVPVLDYLGKLGVRKST 205
L +++D ++ RV F +G+ D + +YP LG + M + YL + G+
Sbjct: 279 LNLSMDELETRVRFYTDMGMNDNDFGTMVYDYPKALGFFSLEEMNSKVQYLKEFGLSTDE 338
Query: 206 ITQFLRTYPQVLHASVVVDLVPVVKYLQGMDIKPDDIPRVLERYPEVLGFKLEGTMSTSV 265
+ + + PQ++ S+ P+VKYL ++I D + R+L P + LE ++ V
Sbjct: 339 LGKLMAFKPQLMACSIEERWKPLVKYLYHLNISRDGMKRMLVVQPTIFCLDLETVIAPKV 398
Query: 266 AYLIGIGVGRRELGGILTRFPEILGMRVGRVIKPFVEYL-ESLGIPRLAIARLIETQPYI 324
+L IGV +GG+L +FP +L + + I+P V +L + + I ++I P +
Sbjct: 399 QFLQDIGVRSDAVGGVLVKFPPVLTYSLYKKIRPVVIFLMTKAAVKQEDIGKVIALDPQL 458
Query: 325 LGFDLDEKVKPNVKSLEEFNVRETSLASIIAQYPDII 361
LG + K++ +VK L + L ++ +P ++
Sbjct: 459 LGCSIVRKLEVSVKYLRSLGIYHFVLGQMVTDFPTLL 495
Score = 85.1 bits (209), Expect = 3e-15
Identities = 48/170 (28%), Positives = 89/170 (52%), Gaps = 1/170 (0%)
Query: 192 VLDYLGKLGVRKSTITQFLRTYPQVLHASVVVDLVPVVKYLQGMDIKPDDIPRVLERYPE 251
++ Y+ GVRK I + PQ+L+ S+ +L V++ M + +D ++ YP+
Sbjct: 254 IIYYMESCGVRKDWIGHVVGRCPQLLNLSMD-ELETRVRFYTDMGMNDNDFGTMVYDYPK 312
Query: 252 VLGFKLEGTMSTSVAYLIGIGVGRRELGGILTRFPEILGMRVGRVIKPFVEYLESLGIPR 311
LGF M++ V YL G+ ELG ++ P+++ + KP V+YL L I R
Sbjct: 313 ALGFFSLEEMNSKVQYLKEFGLSTDELGKLMAFKPQLMACSIEERWKPLVKYLYHLNISR 372
Query: 312 LAIARLIETQPYILGFDLDEKVKPNVKSLEEFNVRETSLASIIAQYPDII 361
+ R++ QP I DL+ + P V+ L++ VR ++ ++ ++P ++
Sbjct: 373 DGMKRMLVVQPTIFCLDLETVIAPKVQFLQDIGVRSDAVGGVLVKFPPVL 422
Score = 83.2 bits (204), Expect = 1e-14
Identities = 50/222 (22%), Positives = 112/222 (49%), Gaps = 17/222 (7%)
Query: 130 VVYEFLRGIGIVPDELDGLELPVTVDVMKERVDFLHSLGLTIEDINNY----PLVLGCSV 185
+VY++ + +G +++ M +V +L GL+ +++ P ++ CS+
Sbjct: 306 MVYDYPKALGFF-----------SLEEMNSKVQYLKEFGLSTDELGKLMAFKPQLMACSI 354
Query: 186 KKNMVPVLDYLGKLGVRKSTITQFLRTYPQVLHASVVVDLVPVVKYLQGMDIKPDDIPRV 245
++ P++ YL L + + + + L P + + + P V++LQ + ++ D + V
Sbjct: 355 EERWKPLVKYLYHLNISRDGMKRMLVVQPTIFCLDLETVIAPKVQFLQDIGVRSDAVGGV 414
Query: 246 LERYPEVLGFKLEGTMSTSVAYLIG-IGVGRRELGGILTRFPEILGMRVGRVIKPFVEYL 304
L ++P VL + L + V +L+ V + ++G ++ P++LG + R ++ V+YL
Sbjct: 415 LVKFPPVLTYSLYKKIRPVVIFLMTKAAVKQEDIGKVIALDPQLLGCSIVRKLEVSVKYL 474
Query: 305 ESLGIPRLAIARLIETQPYILGFDLDEKVKPNVKSLEEFNVR 346
SLGI + +++ P +L +++D ++P + L VR
Sbjct: 475 RSLGIYHFVLGQMVTDFPTLLRYNVD-VLRPKYQYLRRVMVR 515
Score = 59.7 bits (143), Expect = 1e-07
Identities = 32/142 (22%), Positives = 70/142 (48%), Gaps = 1/142 (0%)
Query: 224 DLVPVVKYLQGMDIKPDDIPRVLERYPEVLGFKLEGTMSTSVAYLIGIGVGRRELGGILT 283
+L ++ Y++ ++ D I V+ R P++L ++ + T V + +G+ + G ++
Sbjct: 250 ELEEIIYYMESCGVRKDWIGHVVGRCPQLLNLSMD-ELETRVRFYTDMGMNDNDFGTMVY 308
Query: 284 RFPEILGMRVGRVIKPFVEYLESLGIPRLAIARLIETQPYILGFDLDEKVKPNVKSLEEF 343
+P+ LG + V+YL+ G+ + +L+ +P ++ ++E+ KP VK L
Sbjct: 309 DYPKALGFFSLEEMNSKVQYLKEFGLSTDELGKLMAFKPQLMACSIEERWKPLVKYLYHL 368
Query: 344 NVRETSLASIIAQYPDIIGTDL 365
N+ + ++ P I DL
Sbjct: 369 NISRDGMKRMLVVQPTIFCLDL 390
>ref|XP_466595.1| mitochondrial transcription termination factor-like protein [Oryza
sativa (japonica cultivar-group)]
gi|47848306|dbj|BAD22170.1| mitochondrial transcription
termination factor-like protein [Oryza sativa (japonica
cultivar-group)] gi|47497302|dbj|BAD19344.1|
mitochondrial transcription termination factor-like
protein [Oryza sativa (japonica cultivar-group)]
Length = 271
Score = 87.0 bits (214), Expect = 8e-16
Identities = 49/162 (30%), Positives = 89/162 (54%), Gaps = 4/162 (2%)
Query: 174 INNYPLVLGCSVKKNMVPVLDYLGKLGVRKSTITQFLRTYPQVLHASVVVDLVPVVKYLQ 233
+ +P +V + + P++ L +LGV +S I ++ PQ+ S+ +L P++ YL+
Sbjct: 10 VRKFPAFAYYNVDRKIKPLVALLLELGVPRSNIPGIIKKRPQLCGISLSDNLKPMMTYLE 69
Query: 234 GMDIKPDDIPRVLERYPEVLGFKLEGTMSTSVAYLIGIGVGRRELGGILTRFPEILGMRV 293
+ I D +VL R+P +L + + + T+V++L +GV + +G ILTR P I+ V
Sbjct: 70 NVGINKDKWSKVLSRFPALLTYSRQ-KVETTVSFLTELGVPKENIGKILTRCPHIMSYSV 128
Query: 294 GRVIKPFVEYLESLGIPRLAIARLIETQPYILGFDLDEKVKP 335
++P EY +S+G A LI+ P G +++ K+KP
Sbjct: 129 NDNLRPTAEYFQSIGAD---AASLIQKSPQAFGLNIEAKLKP 167
Score = 68.6 bits (166), Expect = 3e-10
Identities = 37/160 (23%), Positives = 82/160 (51%), Gaps = 4/160 (2%)
Query: 206 ITQFLRTYPQVLHASVVVDLVPVVKYLQGMDIKPDDIPRVLERYPEVLGFKLEGTMSTSV 265
I +R +P + +V + P+V L + + +IP ++++ P++ G L + +
Sbjct: 6 IKNVVRKFPAFAYYNVDRKIKPLVALLLELGVPRSNIPGIIKKRPQLCGISLSDNLKPMM 65
Query: 266 AYLIGIGVGRRELGGILTRFPEILGMRVGRVIKPFVEYLESLGIPRLAIARLIETQPYIL 325
YL +G+ + + +L+RFP +L +V + V +L LG+P+ I +++ P+I+
Sbjct: 66 TYLENVGINKDKWSKVLSRFPALLTYSRQKV-ETTVSFLTELGVPKENIGKILTRCPHIM 124
Query: 326 GFDLDEKVKPNVKSLEEFNVRETSLASIIAQYPDIIGTDL 365
+ +++ ++P + E F AS+I + P G ++
Sbjct: 125 SYSVNDNLRP---TAEYFQSIGADAASLIQKSPQAFGLNI 161
Score = 63.5 bits (153), Expect = 9e-09
Identities = 44/196 (22%), Positives = 91/196 (45%), Gaps = 12/196 (6%)
Query: 155 DVMKERVDFLHSLGLTIED----INNYPLVLGCSVKKNMVPVLDYLGKLGVRKSTITQFL 210
D +K + +L ++G+ + ++ +P +L S +K + + +L +LGV K I + L
Sbjct: 59 DNLKPMMTYLENVGINKDKWSKVLSRFPALLTYSRQK-VETTVSFLTELGVPKENIGKIL 117
Query: 211 RTYPQVLHASVVVDLVPVVKYLQGMDIKPDDIPRVLERYPEVLGFKLEGTMSTSVAYLIG 270
P ++ SV +L P +Y Q + D ++++ P+ G +E + + +
Sbjct: 118 TRCPHIMSYSVNDNLRPTAEYFQSIGA---DAASLIQKSPQAFGLNIEAKLKPITEFFLE 174
Query: 271 IGVGRRELGGILTRFPEILGMRVGRVIKPFVEYLESLGIPRLAIARLIETQPYILGFDLD 330
E+G + RF I + + + P EY ++G PR + + P G+ L+
Sbjct: 175 RDFTMEEIGTMANRFGIIHTLSMEDNLLPKYEYFLTMGYPRNELVKF----PQYFGYSLE 230
Query: 331 EKVKPNVKSLEEFNVR 346
+++KP + + VR
Sbjct: 231 QRIKPRYARMIDCGVR 246
Score = 59.3 bits (142), Expect = 2e-07
Identities = 31/122 (25%), Positives = 68/122 (55%), Gaps = 1/122 (0%)
Query: 240 DDIPRVLERYPEVLGFKLEGTMSTSVAYLIGIGVGRRELGGILTRFPEILGMRVGRVIKP 299
++I V+ ++P + ++ + VA L+ +GV R + GI+ + P++ G+ + +KP
Sbjct: 4 EEIKNVVRKFPAFAYYNVDRKIKPLVALLLELGVPRSNIPGIIKKRPQLCGISLSDNLKP 63
Query: 300 FVEYLESLGIPRLAIARLIETQPYILGFDLDEKVKPNVKSLEEFNVRETSLASIIAQYPD 359
+ YLE++GI + ++++ P +L + +KV+ V L E V + ++ I+ + P
Sbjct: 64 MMTYLENVGINKDKWSKVLSRFPALLTYS-RQKVETTVSFLTELGVPKENIGKILTRCPH 122
Query: 360 II 361
I+
Sbjct: 123 IM 124
>emb|CAB80481.1| putative protein [Arabidopsis thaliana] gi|4467122|emb|CAB37556.1|
putative protein [Arabidopsis thaliana]
gi|23308295|gb|AAN18117.1| At4g38160/F20D10_280
[Arabidopsis thaliana] gi|20147357|gb|AAM10391.1|
AT4g38160/F20D10_280 [Arabidopsis thaliana]
gi|15233700|ref|NP_195529.1| mitochondrial transcription
termination factor-related / mTERF-related [Arabidopsis
thaliana] gi|7485857|pir||T05643 hypothetical protein
F20D10.280 - Arabidopsis thaliana
Length = 333
Score = 81.3 bits (199), Expect = 4e-14
Identities = 46/215 (21%), Positives = 114/215 (52%), Gaps = 9/215 (4%)
Query: 155 DVMKERVDFLHSLGLTIED------INNYPLVLGCSVKKNMVPVLDYLGKLGVRKSTITQ 208
DV E D+L ++ + I++ ++ P +L + + ++P+++ L LG +
Sbjct: 39 DVASENWDYLSNI-VGIQERKLPYIVSRCPKILTLRLDERLIPMVECLSSLGRNPREVAS 97
Query: 209 FLRTYPQVLHASVVVDLVPVVKYLQGMDIKPDDIPRVLERYPEVLGFKLEGTMSTSVAYL 268
+ +P +L SV L P++ + Q + + + +++ P ++ + ++ ++ V++L
Sbjct: 98 AITKFPPILSHSVEEKLCPLLAFFQALGVPETQLGKMILFNPRLISYSIDTKLTVIVSFL 157
Query: 269 IGIGVGRREL-GGILTRFPEILGMRVGRVIKPFVEYLES-LGIPRLAIARLIETQPYILG 326
+G+ + + G +L + P ++G V + ++P E+L+S +G+ I ++ P +L
Sbjct: 158 ASLGLDQDGMIGKVLVKNPFLMGYSVDKRLRPTTEFLKSSVGLSEDGIKSVVMNFPQLLC 217
Query: 327 FDLDEKVKPNVKSLEEFNVRETSLASIIAQYPDII 361
D+++ +KPN L+E ++ +A+++ YP I+
Sbjct: 218 RDVNKILKPNYDYLKECGFGDSQIATMVTGYPQIL 252
Score = 72.8 bits (177), Expect = 2e-11
Identities = 56/238 (23%), Positives = 114/238 (47%), Gaps = 17/238 (7%)
Query: 128 RKVVYEFLRGIGIVPDELDGLELPVTVDVMKERVDFLHSLGLTIED----INNYPLVLGC 183
RK+ Y R I+ LD +P+ V+ L SLG + I +P +L
Sbjct: 57 RKLPYIVSRCPKILTLRLDERLIPM--------VECLSSLGRNPREVASAITKFPPILSH 108
Query: 184 SVKKNMVPVLDYLGKLGVRKSTITQFLRTYPQVLHASVVVDLVPVVKYLQGMDIKPDD-I 242
SV++ + P+L + LGV ++ + + + P+++ S+ L +V +L + + D I
Sbjct: 109 SVEEKLCPLLAFFQALGVPETQLGKMILFNPRLISYSIDTKLTVIVSFLASLGLDQDGMI 168
Query: 243 PRVLERYPEVLGFKLEGTMSTSVAYL-IGIGVGRRELGGILTRFPEILGMRVGRVIKPFV 301
+VL + P ++G+ ++ + + +L +G+ + ++ FP++L V +++KP
Sbjct: 169 GKVLVKNPFLMGYSVDKRLRPTTEFLKSSVGLSEDGIKSVVMNFPQLLCRDVNKILKPNY 228
Query: 302 EYLESLGIPRLAIARLIETQPYILGFDLDEKVKPNVKSLEEFNVRETSLASIIAQYPD 359
+YL+ G IA ++ P IL + ++P ++ L + R +A YP+
Sbjct: 229 DYLKECGFGDSQIATMVTGYPQILIKSVKNSLQPRIRFLVQVMGRG---MDEVASYPE 283
Score = 71.2 bits (173), Expect = 5e-11
Identities = 35/175 (20%), Positives = 91/175 (52%), Gaps = 3/175 (1%)
Query: 194 DYLGKL-GVRKSTITQFLRTYPQVLHASVVVDLVPVVKYLQGMDIKPDDIPRVLERYPEV 252
DYL + G+++ + + P++L + L+P+V+ L + P ++ + ++P +
Sbjct: 46 DYLSNIVGIQERKLPYIVSRCPKILTLRLDERLIPMVECLSSLGRNPREVASAITKFPPI 105
Query: 253 LGFKLEGTMSTSVAYLIGIGVGRRELGGILTRFPEILGMRVGRVIKPFVEYLESLGIPRL 312
L +E + +A+ +GV +LG ++ P ++ + + V +L SLG+ +
Sbjct: 106 LSHSVEEKLCPLLAFFQALGVPETQLGKMILFNPRLISYSIDTKLTVIVSFLASLGLDQD 165
Query: 313 A-IARLIETQPYILGFDLDEKVKPNVKSLE-EFNVRETSLASIIAQYPDIIGTDL 365
I +++ P+++G+ +D++++P + L+ + E + S++ +P ++ D+
Sbjct: 166 GMIGKVLVKNPFLMGYSVDKRLRPTTEFLKSSVGLSEDGIKSVVMNFPQLLCRDV 220
>ref|NP_974703.1| mitochondrial transcription termination factor-related /
mTERF-related [Arabidopsis thaliana]
Length = 363
Score = 81.3 bits (199), Expect = 4e-14
Identities = 46/215 (21%), Positives = 114/215 (52%), Gaps = 9/215 (4%)
Query: 155 DVMKERVDFLHSLGLTIED------INNYPLVLGCSVKKNMVPVLDYLGKLGVRKSTITQ 208
DV E D+L ++ + I++ ++ P +L + + ++P+++ L LG +
Sbjct: 39 DVASENWDYLSNI-VGIQERKLPYIVSRCPKILTLRLDERLIPMVECLSSLGRNPREVAS 97
Query: 209 FLRTYPQVLHASVVVDLVPVVKYLQGMDIKPDDIPRVLERYPEVLGFKLEGTMSTSVAYL 268
+ +P +L SV L P++ + Q + + + +++ P ++ + ++ ++ V++L
Sbjct: 98 AITKFPPILSHSVEEKLCPLLAFFQALGVPETQLGKMILFNPRLISYSIDTKLTVIVSFL 157
Query: 269 IGIGVGRREL-GGILTRFPEILGMRVGRVIKPFVEYLES-LGIPRLAIARLIETQPYILG 326
+G+ + + G +L + P ++G V + ++P E+L+S +G+ I ++ P +L
Sbjct: 158 ASLGLDQDGMIGKVLVKNPFLMGYSVDKRLRPTTEFLKSSVGLSEDGIKSVVMNFPQLLC 217
Query: 327 FDLDEKVKPNVKSLEEFNVRETSLASIIAQYPDII 361
D+++ +KPN L+E ++ +A+++ YP I+
Sbjct: 218 RDVNKILKPNYDYLKECGFGDSQIATMVTGYPQIL 252
Score = 72.8 bits (177), Expect = 2e-11
Identities = 56/238 (23%), Positives = 114/238 (47%), Gaps = 17/238 (7%)
Query: 128 RKVVYEFLRGIGIVPDELDGLELPVTVDVMKERVDFLHSLGLTIED----INNYPLVLGC 183
RK+ Y R I+ LD +P+ V+ L SLG + I +P +L
Sbjct: 57 RKLPYIVSRCPKILTLRLDERLIPM--------VECLSSLGRNPREVASAITKFPPILSH 108
Query: 184 SVKKNMVPVLDYLGKLGVRKSTITQFLRTYPQVLHASVVVDLVPVVKYLQGMDIKPDD-I 242
SV++ + P+L + LGV ++ + + + P+++ S+ L +V +L + + D I
Sbjct: 109 SVEEKLCPLLAFFQALGVPETQLGKMILFNPRLISYSIDTKLTVIVSFLASLGLDQDGMI 168
Query: 243 PRVLERYPEVLGFKLEGTMSTSVAYL-IGIGVGRRELGGILTRFPEILGMRVGRVIKPFV 301
+VL + P ++G+ ++ + + +L +G+ + ++ FP++L V +++KP
Sbjct: 169 GKVLVKNPFLMGYSVDKRLRPTTEFLKSSVGLSEDGIKSVVMNFPQLLCRDVNKILKPNY 228
Query: 302 EYLESLGIPRLAIARLIETQPYILGFDLDEKVKPNVKSLEEFNVRETSLASIIAQYPD 359
+YL+ G IA ++ P IL + ++P ++ L + R +A YP+
Sbjct: 229 DYLKECGFGDSQIATMVTGYPQILIKSVKNSLQPRIRFLVQVMGRG---MDEVASYPE 283
Score = 71.2 bits (173), Expect = 5e-11
Identities = 35/175 (20%), Positives = 91/175 (52%), Gaps = 3/175 (1%)
Query: 194 DYLGKL-GVRKSTITQFLRTYPQVLHASVVVDLVPVVKYLQGMDIKPDDIPRVLERYPEV 252
DYL + G+++ + + P++L + L+P+V+ L + P ++ + ++P +
Sbjct: 46 DYLSNIVGIQERKLPYIVSRCPKILTLRLDERLIPMVECLSSLGRNPREVASAITKFPPI 105
Query: 253 LGFKLEGTMSTSVAYLIGIGVGRRELGGILTRFPEILGMRVGRVIKPFVEYLESLGIPRL 312
L +E + +A+ +GV +LG ++ P ++ + + V +L SLG+ +
Sbjct: 106 LSHSVEEKLCPLLAFFQALGVPETQLGKMILFNPRLISYSIDTKLTVIVSFLASLGLDQD 165
Query: 313 A-IARLIETQPYILGFDLDEKVKPNVKSLE-EFNVRETSLASIIAQYPDIIGTDL 365
I +++ P+++G+ +D++++P + L+ + E + S++ +P ++ D+
Sbjct: 166 GMIGKVLVKNPFLMGYSVDKRLRPTTEFLKSSVGLSEDGIKSVVMNFPQLLCRDV 220
>gb|AAM15397.1| hypothetical protein [Arabidopsis thaliana]
gi|4417266|gb|AAD20391.1| hypothetical protein
[Arabidopsis thaliana] gi|25412055|pir||C84604
hypothetical protein At2g21710 [imported] - Arabidopsis
thaliana
Length = 673
Score = 76.6 bits (187), Expect = 1e-12
Identities = 52/207 (25%), Positives = 105/207 (50%), Gaps = 14/207 (6%)
Query: 152 VTVDVMKERVDFLHSLGLTIEDINNY----PLVLGCSVKKNMVPVLDYLGKLGVRKSTIT 207
+ V ++ ++++L GL+ E++ P ++GCS+++ P++ Y LG+ K +
Sbjct: 373 IIVLLVLNQINYLKEFGLSTEEVGRLLAYKPHLMGCSIEERWKPLVKYFYYLGIPKEGMK 432
Query: 208 QFLRTYPQVLH--------ASVVVDLVPVVKYLQGMDIKPDDIPRVLERYPEVLGFKLEG 259
+ L P +L+ A VV+L V++LQ M I + I +L ++P +L L
Sbjct: 433 RILVVKP-ILYCIDLEKTIAPKVVELRYNVRFLQEMGIPNEAIGNMLVKFPSLLTNSLYK 491
Query: 260 TMSTSVAYLIGIGVGRRELGGILTRFPEILGMRVGRVIKPFVEYLESLGIPRLAIARLIE 319
+ + L GV ++++G ++ P +LG +G ++P + Y SLGI + +I
Sbjct: 492 KIRPVIFLLTRAGVTQKDIGKVIAMDPALLGCSIGTKLEPNMRYYISLGIRFYQLGEMIA 551
Query: 320 TQPYILGFDLDEKVKPNVKSLEEFNVR 346
P +L +++D ++P + L +R
Sbjct: 552 DFPMLLRYNVD-NLRPKYRYLRRTMIR 577
Score = 75.5 bits (184), Expect = 2e-12
Identities = 72/338 (21%), Positives = 144/338 (42%), Gaps = 54/338 (15%)
Query: 67 ITKLQCSVAGRTDSYRTSAADSQGSKRDSGEIQRKRRGASSLYAGYARPSLSEMKKDKAT 126
I K+ C G DS R + S GE + + R + +++++
Sbjct: 231 IAKIICMSKGNLDSIRIMI-EWLKSIHVKGEF---------IAVAFLRSGDNILQRNREE 280
Query: 127 LRKVVYEFLRGIGIVPDELDGLE------LPVTVDVMKERVDFLHSLGLTIED----INN 176
L ++V E+L G+ D + + L +++ +K RVDF +G+ D + +
Sbjct: 281 LNEIV-EYLESNGVRRDWMGYVVGRCPELLSFSMEEVKSRVDFFLKMGMNQNDFGTMVYD 339
Query: 177 YPLVLGCS----VKKNMVPVL----------------------DYLGKLGVRKSTITQFL 210
YP ++G ++K ++ L +YL + G+ + + L
Sbjct: 340 YPKIIGFFSFQVMEKKVLKALFNTPALRLSFKFIIVLLVLNQINYLKEFGLSTEEVGRLL 399
Query: 211 RTYPQVLHASVVVDLVPVVKYLQGMDIKPDDIPRVLERYPEVLGFKLEGTMSTSVA---- 266
P ++ S+ P+VKY + I + + R+L P + LE T++ V
Sbjct: 400 AYKPHLMGCSIEERWKPLVKYFYYLGIPKEGMKRILVVKPILYCIDLEKTIAPKVVELRY 459
Query: 267 ---YLIGIGVGRRELGGILTRFPEILGMRVGRVIKPFVEYLESLGIPRLAIARLIETQPY 323
+L +G+ +G +L +FP +L + + I+P + L G+ + I ++I P
Sbjct: 460 NVRFLQEMGIPNEAIGNMLVKFPSLLTNSLYKKIRPVIFLLTRAGVTQKDIGKVIAMDPA 519
Query: 324 ILGFDLDEKVKPNVKSLEEFNVRETSLASIIAQYPDII 361
+LG + K++PN++ +R L +IA +P ++
Sbjct: 520 LLGCSIGTKLEPNMRYYISLGIRFYQLGEMIADFPMLL 557
>gb|AAN15460.1| Unknown protein [Arabidopsis thaliana] gi|9758592|dbj|BAB09225.1|
unnamed protein product [Arabidopsis thaliana]
gi|15240542|ref|NP_200369.1| mitochondrial transcription
termination factor family protein / mTERF family protein
[Arabidopsis thaliana] gi|17065230|gb|AAL32769.1|
Unknown protein [Arabidopsis thaliana]
Length = 496
Score = 76.3 bits (186), Expect = 1e-12
Identities = 57/223 (25%), Positives = 110/223 (48%), Gaps = 11/223 (4%)
Query: 150 LPVTVDVMKERVDFLHSLGLTIEDINNY----PLVLGCSVKKNMVPVLDYLGKLGVRKST 205
L + V +ER+D+L S+G+ DI P +L +V+ N+ + +L LG+ S
Sbjct: 212 LQINVFSAQERLDYLLSVGVKHRDIKRMLLRQPQILQYTVENNLKAHISFLMGLGIPNSK 271
Query: 206 ITQFLRTYPQVLHASVVVDLVPVVKYL-QGMDIKPDDIPRVLERYPEVLGFKLEGTMSTS 264
I Q + P + SV L P ++YL + + IK D+ +V++ P++L +L+ T +T
Sbjct: 272 IGQIVAATPSLFSYSVENSLRPTIRYLIEEVGIKETDVGKVVQLSPQILVQRLDITWNTR 331
Query: 265 VAYLI-GIGVGRRELGGILTRFPEILGMRVGRVIKPFVEYLESLGIPRLAIARLIETQPY 323
+L +G R + ++ + P++L + P + +L S+G+ I +++ +
Sbjct: 332 YMFLSKELGAPRDSVVKMVKKHPQLLHYSIDDGFLPRINFLRSIGMCNSDILKVLTSLTQ 391
Query: 324 ILGFDLDEKVKPNVKSL-EEFNVRETSLASIIAQYPDIIGTDL 365
+L L++ +KP L E N + I+ +YP + L
Sbjct: 392 VLSLSLEDNLKPKYMYLVNELN----NEVHILTKYPMYLSLSL 430
Score = 75.5 bits (184), Expect = 2e-12
Identities = 44/174 (25%), Positives = 86/174 (49%), Gaps = 3/174 (1%)
Query: 190 VPVLDYLGKLGVRKSTITQFLRTYPQVLHASVVVDLVPVVKYLQGMDIKPDDIPRVLERY 249
VP+LDYL G+++S Q + L +V + YL + +K DI R+L R
Sbjct: 185 VPLLDYLSTFGLKESHFVQMYERHMPSLQINVF-SAQERLDYLLSVGVKHRDIKRMLLRQ 243
Query: 250 PEVLGFKLEGTMSTSVAYLIGIGVGRRELGGILTRFPEILGMRVGRVIKPFVEYL-ESLG 308
P++L + +E + +++L+G+G+ ++G I+ P + V ++P + YL E +G
Sbjct: 244 PQILQYTVENNLKAHISFLMGLGIPNSKIGQIVAATPSLFSYSVENSLRPTIRYLIEEVG 303
Query: 309 IPRLAIARLIETQPYILGFDLDEKVKPNVKSL-EEFNVRETSLASIIAQYPDII 361
I + ++++ P IL LD L +E S+ ++ ++P ++
Sbjct: 304 IKETDVGKVVQLSPQILVQRLDITWNTRYMFLSKELGAPRDSVVKMVKKHPQLL 357
Score = 50.1 bits (118), Expect = 1e-04
Identities = 30/113 (26%), Positives = 57/113 (49%), Gaps = 9/113 (7%)
Query: 198 KLGVRKSTITQFLRTYPQVLHASVVVDLVPVVKYLQGMDIKPDDIPRVLERYPEVLGFKL 257
+LG + ++ + ++ +PQ+LH S+ +P + +L+ + + DI +VL +VL L
Sbjct: 338 ELGAPRDSVVKMVKKHPQLLHYSIDDGFLPRINFLRSIGMCNSDILKVLTSLTQVLSLSL 397
Query: 258 EGTMSTSVAYLIGIGVGRRELGG---ILTRFPEILGMRVGRVIKPFVEYLESL 307
E + YL+ EL ILT++P L + + + I+P +L L
Sbjct: 398 EDNLKPKYMYLV------NELNNEVHILTKYPMYLSLSLDQRIRPRHRFLVEL 444
>gb|AAQ22615.1| At1g78930 [Arabidopsis thaliana]
Length = 525
Score = 74.7 bits (182), Expect = 4e-12
Identities = 48/192 (25%), Positives = 96/192 (50%), Gaps = 7/192 (3%)
Query: 174 INNYPLVLGCSVKKNMVPVLDYLGKLGVRKSTITQFLRTYPQVLHASVVVDLVPVVKYLQ 233
I P +L C ++ + V+ +L LG +K + Q L P++ S+ L + +L
Sbjct: 326 IPKMPQLLLCKPQE-FLKVVCFLEDLGFQKEIVGQILCRCPEIFGCSIEKTLQKKLIFLT 384
Query: 234 GMDIKPDDIPRVLERYPEVLGFKLEGTMSTSVAYLIGIGVGRRELGGILTRFPEILGMRV 293
+ PR++++YPE L + + T+ + YL+ IG+ RE+ ++ +F ILG +
Sbjct: 385 RFGVSTTHFPRIIKKYPEFLIYDADKTVLPRLKYLMEIGISEREIAFMIRKFSPILGYSI 444
Query: 294 GRVIKPFVEYL-ESLGIPRLAIARLIETQPYILGFDLDEKVKPNVKSLEEFNVRETSLAS 352
+V++P E+L S+ P R + P + L++++KP + L+ N+ E +L
Sbjct: 445 DKVLRPKFEFLVNSMEKP----VREVIEYPRYFSYSLEKRIKPRFRVLKGRNI-ECTLQE 499
Query: 353 IIAQYPDIIGTD 364
++ + + D
Sbjct: 500 MLGKNDEEFAAD 511
Score = 54.3 bits (129), Expect = 6e-06
Identities = 59/292 (20%), Positives = 117/292 (39%), Gaps = 30/292 (10%)
Query: 91 SKRDSGEIQRKRRGASSLYAGYARPSLSEMKKDKATLRKVVYEFLRGIGI-------VPD 143
S + SGE +R G + + M K K KV FL +G+ +
Sbjct: 51 SWKGSGESERVEEGLGF------KEKVIYMVKQKGDGGKVA--FLESLGLSLSSAMYLAH 102
Query: 144 ELDGLELPVTVDVMKERVDFLHS----LGLTIEDINNYPLVLGCSVKKNMVPVLDYLGKL 199
+ LP+ +D +K + S GL + L L + +++ L + K+
Sbjct: 103 YVSSESLPILLDKVKYLKEIFFSGSDEKGLVGKYARRMMLYLSIPIDEDVQQTLSFFEKI 162
Query: 200 GVRKSTITQF----------LRTYPQVLHASVVVDLVPVVKYLQGMDIKPDDIPRVLERY 249
R+ + L ++P++L S D+ P+V++L+ + I + +VL Y
Sbjct: 163 EARRGGLDMLGSVDASFRFLLESFPRLLLLSEENDMKPMVEFLESIGIPKYCLGKVLLLY 222
Query: 250 PEVLGFKLEGTMSTSVAYLIGIGVGRRELGGILTRFPEILGMRVGRVIKPFVEYLESLGI 309
P ++ K E + + V ++ G +L ++P IL + + S +
Sbjct: 223 PPIMLGKTEEIKRRVATAMEKVSVVNKDSGKLLLKYPWILSPSIQENYSHIGSFFYSESV 282
Query: 310 PRLAIARLIETQPYILGFDLDEKVKPNVKSLEEFNVRETSLASIIAQYPDII 361
++ I I P +LG ++ VK ++ VR+ + +I + P ++
Sbjct: 283 LKMDIDHAIRRWPLLLGCSA-SNMEMMVKEFDKLGVRDKRMGKVIPKMPQLL 333
Score = 44.7 bits (104), Expect = 0.005
Identities = 46/216 (21%), Positives = 92/216 (42%), Gaps = 42/216 (19%)
Query: 160 RVDFLHSLGLTIEDINNYPLVLGCSVKKNMVPVL----DYLGKLGVRKSTITQFLRTYPQ 215
+V FL SLGL++ + L V +P+L YL ++ S + Y +
Sbjct: 83 KVAFLESLGLSLSSA----MYLAHYVSSESLPILLDKVKYLKEIFFSGSDEKGLVGKYAR 138
Query: 216 --VLHASVVVDLVPVVKYLQGMDIKPDDIPRVLERYPEVL----GFKLEGTMSTSVAYLI 269
+L+ S+ +D +D+ + L + ++ G + G++ S +L
Sbjct: 139 RMMLYLSIPID---------------EDVQQTLSFFEKIEARRGGLDMLGSVDASFRFL- 182
Query: 270 GIGVGRRELGGILTRFPEILGMRVGRVIKPFVEYLESLGIPRLAIARLIETQPYILGFDL 329
L FP +L + +KP VE+LES+GIP+ + +++ P I+
Sbjct: 183 ------------LESFPRLLLLSEENDMKPMVEFLESIGIPKYCLGKVLLLYPPIMLGKT 230
Query: 330 DEKVKPNVKSLEEFNVRETSLASIIAQYPDIIGTDL 365
+E + ++E+ +V ++ +YP I+ +
Sbjct: 231 EEIKRRVATAMEKVSVVNKDSGKLLLKYPWILSPSI 266
>ref|NP_178014.2| mitochondrial transcription termination factor-related /
mTERF-related [Arabidopsis thaliana]
Length = 591
Score = 74.7 bits (182), Expect = 4e-12
Identities = 48/192 (25%), Positives = 96/192 (50%), Gaps = 7/192 (3%)
Query: 174 INNYPLVLGCSVKKNMVPVLDYLGKLGVRKSTITQFLRTYPQVLHASVVVDLVPVVKYLQ 233
I P +L C ++ + V+ +L LG +K + Q L P++ S+ L + +L
Sbjct: 392 IPKMPQLLLCKPQE-FLKVVCFLEDLGFQKEIVGQILCRCPEIFGCSIEKTLQKKLIFLT 450
Query: 234 GMDIKPDDIPRVLERYPEVLGFKLEGTMSTSVAYLIGIGVGRRELGGILTRFPEILGMRV 293
+ PR++++YPE L + + T+ + YL+ IG+ RE+ ++ +F ILG +
Sbjct: 451 RFGVSTTHFPRIIKKYPEFLIYDADKTVLPRLKYLMEIGISEREIAFMIRKFSPILGYSI 510
Query: 294 GRVIKPFVEYL-ESLGIPRLAIARLIETQPYILGFDLDEKVKPNVKSLEEFNVRETSLAS 352
+V++P E+L S+ P R + P + L++++KP + L+ N+ E +L
Sbjct: 511 DKVLRPKFEFLVNSMEKP----VREVIEYPRYFSYSLEKRIKPRFRVLKGRNI-ECTLQE 565
Query: 353 IIAQYPDIIGTD 364
++ + + D
Sbjct: 566 MLGKNDEEFAAD 577
Score = 54.3 bits (129), Expect = 6e-06
Identities = 59/292 (20%), Positives = 117/292 (39%), Gaps = 30/292 (10%)
Query: 91 SKRDSGEIQRKRRGASSLYAGYARPSLSEMKKDKATLRKVVYEFLRGIGI-------VPD 143
S + SGE +R G + + M K K KV FL +G+ +
Sbjct: 117 SWKGSGESERVEEGLGF------KEKVIYMVKQKGDGGKVA--FLESLGLSLSSAMYLAH 168
Query: 144 ELDGLELPVTVDVMKERVDFLHS----LGLTIEDINNYPLVLGCSVKKNMVPVLDYLGKL 199
+ LP+ +D +K + S GL + L L + +++ L + K+
Sbjct: 169 YVSSESLPILLDKVKYLKEIFFSGSDEKGLVGKYARRMMLYLSIPIDEDVQQTLSFFEKI 228
Query: 200 GVRKSTITQF----------LRTYPQVLHASVVVDLVPVVKYLQGMDIKPDDIPRVLERY 249
R+ + L ++P++L S D+ P+V++L+ + I + +VL Y
Sbjct: 229 EARRGGLDMLGSVDASFRFLLESFPRLLLLSEENDMKPMVEFLESIGIPKYCLGKVLLLY 288
Query: 250 PEVLGFKLEGTMSTSVAYLIGIGVGRRELGGILTRFPEILGMRVGRVIKPFVEYLESLGI 309
P ++ K E + + V ++ G +L ++P IL + + S +
Sbjct: 289 PPIMLGKTEEIKRRVATAMEKVSVVNKDSGKLLLKYPWILSPSIQENYSHIGSFFYSESV 348
Query: 310 PRLAIARLIETQPYILGFDLDEKVKPNVKSLEEFNVRETSLASIIAQYPDII 361
++ I I P +LG ++ VK ++ VR+ + +I + P ++
Sbjct: 349 LKMDIDHAIRRWPLLLGCSA-SNMEMMVKEFDKLGVRDKRMGKVIPKMPQLL 399
Score = 44.7 bits (104), Expect = 0.005
Identities = 46/216 (21%), Positives = 92/216 (42%), Gaps = 42/216 (19%)
Query: 160 RVDFLHSLGLTIEDINNYPLVLGCSVKKNMVPVL----DYLGKLGVRKSTITQFLRTYPQ 215
+V FL SLGL++ + L V +P+L YL ++ S + Y +
Sbjct: 149 KVAFLESLGLSLSSA----MYLAHYVSSESLPILLDKVKYLKEIFFSGSDEKGLVGKYAR 204
Query: 216 --VLHASVVVDLVPVVKYLQGMDIKPDDIPRVLERYPEVL----GFKLEGTMSTSVAYLI 269
+L+ S+ +D +D+ + L + ++ G + G++ S +L
Sbjct: 205 RMMLYLSIPID---------------EDVQQTLSFFEKIEARRGGLDMLGSVDASFRFL- 248
Query: 270 GIGVGRRELGGILTRFPEILGMRVGRVIKPFVEYLESLGIPRLAIARLIETQPYILGFDL 329
L FP +L + +KP VE+LES+GIP+ + +++ P I+
Sbjct: 249 ------------LESFPRLLLLSEENDMKPMVEFLESIGIPKYCLGKVLLLYPPIMLGKT 296
Query: 330 DEKVKPNVKSLEEFNVRETSLASIIAQYPDIIGTDL 365
+E + ++E+ +V ++ +YP I+ +
Sbjct: 297 EEIKRRVATAMEKVSVVNKDSGKLLLKYPWILSPSI 332
>emb|CAB10239.1| hypothetical protein [Arabidopsis thaliana]
gi|7268166|emb|CAB78502.1| hypothetical protein
[Arabidopsis thaliana] gi|7485117|pir||E71408
hypothetical protein - Arabidopsis thaliana
Length = 590
Score = 73.6 bits (179), Expect = 9e-12
Identities = 43/171 (25%), Positives = 85/171 (49%), Gaps = 4/171 (2%)
Query: 195 YLGKLGVRKSTITQFLRTYPQVLHASVVVDLVPVVKYLQGMDIKPDDIPRVLERYPEVLG 254
YL LG+ I R + + S+ + PVV++L + I DIP +L + P++ G
Sbjct: 348 YLLDLGLNLEQIKTITRKFAAFPYYSLDGKIKPVVEFLLDLGIPKSDIPTILCKRPQICG 407
Query: 255 FKLEGTMSTSVAYLIGIGVGRRELGGILTRFPEILGMRVGRVIKPFVEYLESLGIPRLAI 314
L + ++A+L +G+ + + I++RFP IL ++ VE+L G+ I
Sbjct: 408 ISLTDNLKPTMAFLETLGIDKNQWAKIISRFPAILTYSRQKLTST-VEFLSQTGLTEEQI 466
Query: 315 ARLIETQPYILGFDLDEKVKPNVKSLEEFNVRETSLASIIAQYPDIIGTDL 365
R++ P I+ + +++K++P ++ NV +A ++ + P G +
Sbjct: 467 GRILTRCPNIMSYSVEDKLRPTMEYFRSLNV---DVAVLLHRCPQTFGLSI 514
Score = 63.2 bits (152), Expect = 1e-08
Identities = 54/227 (23%), Positives = 102/227 (44%), Gaps = 12/227 (5%)
Query: 140 IVPDELDGLEL-----PVTVDVMKERVDFLHSLGLTIEDINNYPLVLGCSVKKNMVPVLD 194
+V EL LE+ P + +E D L L ++ D P ++ S + P D
Sbjct: 258 LVGRELTTLEIRDSLIPYLEQLHEEHGDLLAELVVSFPDPPAEPRLVASSPVSVLPPRGD 317
Query: 195 YLGKLGVRKSTITQFLRTYPQVLHASVVVDLVPVVKYLQGMDIKPDDIPRVLERYPEVLG 254
RK LR +V L P YL + + + I + ++
Sbjct: 318 TDSAADTRK------LRAVSRVSELDTEGALRPQTLYLLDLGLNLEQIKTITRKFAAFPY 371
Query: 255 FKLEGTMSTSVAYLIGIGVGRRELGGILTRFPEILGMRVGRVIKPFVEYLESLGIPRLAI 314
+ L+G + V +L+ +G+ + ++ IL + P+I G+ + +KP + +LE+LGI +
Sbjct: 372 YSLDGKIKPVVEFLLDLGIPKSDIPTILCKRPQICGISLTDNLKPTMAFLETLGIDKNQW 431
Query: 315 ARLIETQPYILGFDLDEKVKPNVKSLEEFNVRETSLASIIAQYPDII 361
A++I P IL + +K+ V+ L + + E + I+ + P+I+
Sbjct: 432 AKIISRFPAILTYS-RQKLTSTVEFLSQTGLTEEQIGRILTRCPNIM 477
Score = 61.2 bits (147), Expect = 5e-08
Identities = 44/197 (22%), Positives = 92/197 (46%), Gaps = 21/197 (10%)
Query: 129 KVVYEFLRGIGIVPDELD----------GLELPVTVDVMKERVDFLHSLGLTIED----I 174
K V EFL +GI ++ G+ L D +K + FL +LG+ I
Sbjct: 379 KPVVEFLLDLGIPKSDIPTILCKRPQICGISL---TDNLKPTMAFLETLGIDKNQWAKII 435
Query: 175 NNYPLVLGCSVKKNMVPVLDYLGKLGVRKSTITQFLRTYPQVLHASVVVDLVPVVKYLQG 234
+ +P +L S +K + +++L + G+ + I + L P ++ SV L P ++Y +
Sbjct: 436 SRFPAILTYSRQK-LTSTVEFLSQTGLTEEQIGRILTRCPNIMSYSVEDKLRPTMEYFRS 494
Query: 235 MDIKPDDIPRVLERYPEVLGFKLEGTMSTSVAYLIGIGVGRRELGGILTRFPEILGMRVG 294
+++ D+ +L R P+ G +E + + + G G E+G +++R+ + +
Sbjct: 495 LNV---DVAVLLHRCPQTFGLSIESNLKPVTEFFLEKGFGLDEIGIMISRYGALYTFSLK 551
Query: 295 RVIKPFVEYLESLGIPR 311
+ P +Y +++ P+
Sbjct: 552 ENVMPKWDYFQTMDYPK 568
Score = 40.0 bits (92), Expect = 0.11
Identities = 31/122 (25%), Positives = 51/122 (41%), Gaps = 7/122 (5%)
Query: 161 VDFLHSLGLTIEDINNY----PLVLGCSVKKNMVPVLDYLGKLGVRKSTITQFLRTYPQV 216
V+FL GLT E I P ++ SV+ + P ++Y L V + L PQ
Sbjct: 453 VEFLSQTGLTEEQIGRILTRCPNIMSYSVEDKLRPTMEYFRSLNV---DVAVLLHRCPQT 509
Query: 217 LHASVVVDLVPVVKYLQGMDIKPDDIPRVLERYPEVLGFKLEGTMSTSVAYLIGIGVGRR 276
S+ +L PV ++ D+I ++ RY + F L+ + Y + +
Sbjct: 510 FGLSIESNLKPVTEFFLEKGFGLDEIGIMISRYGALYTFSLKENVMPKWDYFQTMDYPKS 569
Query: 277 EL 278
EL
Sbjct: 570 EL 571
>ref|NP_567435.1| mitochondrial transcription termination factor-related /
mTERF-related [Arabidopsis thaliana]
Length = 444
Score = 73.6 bits (179), Expect = 9e-12
Identities = 43/171 (25%), Positives = 85/171 (49%), Gaps = 4/171 (2%)
Query: 195 YLGKLGVRKSTITQFLRTYPQVLHASVVVDLVPVVKYLQGMDIKPDDIPRVLERYPEVLG 254
YL LG+ I R + + S+ + PVV++L + I DIP +L + P++ G
Sbjct: 202 YLLDLGLNLEQIKTITRKFAAFPYYSLDGKIKPVVEFLLDLGIPKSDIPTILCKRPQICG 261
Query: 255 FKLEGTMSTSVAYLIGIGVGRRELGGILTRFPEILGMRVGRVIKPFVEYLESLGIPRLAI 314
L + ++A+L +G+ + + I++RFP IL ++ VE+L G+ I
Sbjct: 262 ISLTDNLKPTMAFLETLGIDKNQWAKIISRFPAILTYSRQKLTST-VEFLSQTGLTEEQI 320
Query: 315 ARLIETQPYILGFDLDEKVKPNVKSLEEFNVRETSLASIIAQYPDIIGTDL 365
R++ P I+ + +++K++P ++ NV +A ++ + P G +
Sbjct: 321 GRILTRCPNIMSYSVEDKLRPTMEYFRSLNV---DVAVLLHRCPQTFGLSI 368
Score = 63.2 bits (152), Expect = 1e-08
Identities = 54/227 (23%), Positives = 102/227 (44%), Gaps = 12/227 (5%)
Query: 140 IVPDELDGLEL-----PVTVDVMKERVDFLHSLGLTIEDINNYPLVLGCSVKKNMVPVLD 194
+V EL LE+ P + +E D L L ++ D P ++ S + P D
Sbjct: 112 LVGRELTTLEIRDSLIPYLEQLHEEHGDLLAELVVSFPDPPAEPRLVASSPVSVLPPRGD 171
Query: 195 YLGKLGVRKSTITQFLRTYPQVLHASVVVDLVPVVKYLQGMDIKPDDIPRVLERYPEVLG 254
RK LR +V L P YL + + + I + ++
Sbjct: 172 TDSAADTRK------LRAVSRVSELDTEGALRPQTLYLLDLGLNLEQIKTITRKFAAFPY 225
Query: 255 FKLEGTMSTSVAYLIGIGVGRRELGGILTRFPEILGMRVGRVIKPFVEYLESLGIPRLAI 314
+ L+G + V +L+ +G+ + ++ IL + P+I G+ + +KP + +LE+LGI +
Sbjct: 226 YSLDGKIKPVVEFLLDLGIPKSDIPTILCKRPQICGISLTDNLKPTMAFLETLGIDKNQW 285
Query: 315 ARLIETQPYILGFDLDEKVKPNVKSLEEFNVRETSLASIIAQYPDII 361
A++I P IL + +K+ V+ L + + E + I+ + P+I+
Sbjct: 286 AKIISRFPAILTYS-RQKLTSTVEFLSQTGLTEEQIGRILTRCPNIM 331
Score = 61.2 bits (147), Expect = 5e-08
Identities = 44/197 (22%), Positives = 92/197 (46%), Gaps = 21/197 (10%)
Query: 129 KVVYEFLRGIGIVPDELD----------GLELPVTVDVMKERVDFLHSLGLTIED----I 174
K V EFL +GI ++ G+ L D +K + FL +LG+ I
Sbjct: 233 KPVVEFLLDLGIPKSDIPTILCKRPQICGISL---TDNLKPTMAFLETLGIDKNQWAKII 289
Query: 175 NNYPLVLGCSVKKNMVPVLDYLGKLGVRKSTITQFLRTYPQVLHASVVVDLVPVVKYLQG 234
+ +P +L S +K + +++L + G+ + I + L P ++ SV L P ++Y +
Sbjct: 290 SRFPAILTYSRQK-LTSTVEFLSQTGLTEEQIGRILTRCPNIMSYSVEDKLRPTMEYFRS 348
Query: 235 MDIKPDDIPRVLERYPEVLGFKLEGTMSTSVAYLIGIGVGRRELGGILTRFPEILGMRVG 294
+++ D+ +L R P+ G +E + + + G G E+G +++R+ + +
Sbjct: 349 LNV---DVAVLLHRCPQTFGLSIESNLKPVTEFFLEKGFGLDEIGIMISRYGALYTFSLK 405
Query: 295 RVIKPFVEYLESLGIPR 311
+ P +Y +++ P+
Sbjct: 406 ENVMPKWDYFQTMDYPK 422
Score = 40.0 bits (92), Expect = 0.11
Identities = 31/122 (25%), Positives = 51/122 (41%), Gaps = 7/122 (5%)
Query: 161 VDFLHSLGLTIEDINNY----PLVLGCSVKKNMVPVLDYLGKLGVRKSTITQFLRTYPQV 216
V+FL GLT E I P ++ SV+ + P ++Y L V + L PQ
Sbjct: 307 VEFLSQTGLTEEQIGRILTRCPNIMSYSVEDKLRPTMEYFRSLNV---DVAVLLHRCPQT 363
Query: 217 LHASVVVDLVPVVKYLQGMDIKPDDIPRVLERYPEVLGFKLEGTMSTSVAYLIGIGVGRR 276
S+ +L PV ++ D+I ++ RY + F L+ + Y + +
Sbjct: 364 FGLSIESNLKPVTEFFLEKGFGLDEIGIMISRYGALYTFSLKENVMPKWDYFQTMDYPKS 423
Query: 277 EL 278
EL
Sbjct: 424 EL 425
>ref|XP_478901.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|27817836|dbj|BAC55604.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
Length = 503
Score = 73.2 bits (178), Expect = 1e-11
Identities = 63/273 (23%), Positives = 123/273 (44%), Gaps = 32/273 (11%)
Query: 108 LYAGYARPSLSEMKKDKATLRKVVYEFLRGIGIVPDELDGLELPVTVDVMKERVDFLHSL 167
LY+ R + + ++K L++ G P D + VD + + S
Sbjct: 100 LYSNNPRAPIKKPGREKPALKQ------NWEGRQPKTRDRCDTSKKVDALHAKSKASRST 153
Query: 168 GLTIEDINNYPLVLGCSVKKNM--------------VPVLDYLGKLGVRKSTITQFLRTY 213
GL DI+N + S+ +++ +P++DYL G+++S T +
Sbjct: 154 GLV--DIDNEVELKNESISRSLFQKLQEEYDFDDKWLPLIDYLCTFGLKESHFTNMYERH 211
Query: 214 P---QVLHASVVVDLVPVVKYLQGMDIKPDDIPRVLERYPEVLGFKLEGTMSTSVAYLIG 270
Q+ AS L ++L + +K D+ R+L R P++L + L + + VA+L+G
Sbjct: 212 MACFQISQASAEERL----EFLLSVGVKSKDMKRMLVRQPQILEYTLSN-LKSHVAFLVG 266
Query: 271 IGVGRRELGGILTRFPEILGMRVGRVIKPFVEYL-ESLGIPRLAIARLIETQPYILGFDL 329
IGV +G I++ P V + +KP + YL E +GI + ++++ P IL +
Sbjct: 267 IGVPSARIGQIISAAPSFFSYSVEQSLKPTIRYLIEEVGIEESDVGKVVQLSPQILVQRI 326
Query: 330 DEKVKPNVKSL-EEFNVRETSLASIIAQYPDII 361
D K L +E + ++ ++ ++P ++
Sbjct: 327 DSAWKSRFLFLSKELGAPKDNIVKMVTKHPQLL 359
Score = 63.5 bits (153), Expect = 9e-09
Identities = 42/184 (22%), Positives = 95/184 (50%), Gaps = 7/184 (3%)
Query: 158 KERVDFLHSLGLTIEDINNY----PLVLGCSVKKNMVPVLDYLGKLGVRKSTITQFLRTY 213
+ER++FL S+G+ +D+ P +L ++ N+ + +L +GV + I Q +
Sbjct: 223 EERLEFLLSVGVKSKDMKRMLVRQPQILEYTLS-NLKSHVAFLVGIGVPSARIGQIISAA 281
Query: 214 PQVLHASVVVDLVPVVKYL-QGMDIKPDDIPRVLERYPEVLGFKLEGTMSTSVAYLIG-I 271
P SV L P ++YL + + I+ D+ +V++ P++L +++ + +L +
Sbjct: 282 PSFFSYSVEQSLKPTIRYLIEEVGIEESDVGKVVQLSPQILVQRIDSAWKSRFLFLSKEL 341
Query: 272 GVGRRELGGILTRFPEILGMRVGRVIKPFVEYLESLGIPRLAIARLIETQPYILGFDLDE 331
G + + ++T+ P++L + I P + +L S+G+ + +++ + +L L+E
Sbjct: 342 GAPKDNIVKMVTKHPQLLHYSIEDGILPRINFLRSIGMRDTDVLKVLTSLTQVLSLSLEE 401
Query: 332 KVKP 335
+KP
Sbjct: 402 NLKP 405
Score = 50.8 bits (120), Expect = 6e-05
Identities = 29/110 (26%), Positives = 56/110 (50%), Gaps = 3/110 (2%)
Query: 198 KLGVRKSTITQFLRTYPQVLHASVVVDLVPVVKYLQGMDIKPDDIPRVLERYPEVLGFKL 257
+LG K I + + +PQ+LH S+ ++P + +L+ + ++ D+ +VL +VL L
Sbjct: 340 ELGAPKDNIVKMVTKHPQLLHYSIEDGILPRINFLRSIGMRDTDVLKVLTSLTQVLSLSL 399
Query: 258 EGTMSTSVAYLIGIGVGRRELGGILTRFPEILGMRVGRVIKPFVEYLESL 307
E + YL+ + LT++P L + + + I+P +L SL
Sbjct: 400 EENLKPKYLYLVN---DLKNDVQSLTKYPMYLSLSLDQRIRPRHRFLVSL 446
Score = 40.8 bits (94), Expect = 0.065
Identities = 25/101 (24%), Positives = 53/101 (51%), Gaps = 3/101 (2%)
Query: 171 IEDINNYPLVLGCSVKKNMVPVLDYLGKLGVRKSTITQFLRTYPQVLHASVVVDLVPVVK 230
++ + +P +L S++ ++P +++L +G+R + + + L + QVL S+ +L P
Sbjct: 349 VKMVTKHPQLLHYSIEDGILPRINFLRSIGMRDTDVLKVLTSLTQVLSLSLEENLKPKYL 408
Query: 231 YLQGMDIKPDDIPRVLERYPEVLGFKLEGTMSTSVAYLIGI 271
YL D+K D + L +YP L L+ + +L+ +
Sbjct: 409 YLVN-DLKND--VQSLTKYPMYLSLSLDQRIRPRHRFLVSL 446
>gb|AAC17072.1| Similar to hypothetical protein gb|Z97336 from A. thaliana. This
gene is probably cut off. [Arabidopsis thaliana]
gi|7487916|pir||T01062 hypothetical protein YUP8H12R.46
- Arabidopsis thaliana
Length = 600
Score = 69.7 bits (169), Expect = 1e-10
Identities = 52/194 (26%), Positives = 93/194 (47%), Gaps = 14/194 (7%)
Query: 174 INNYPLVLGCSVKKNMVPVLDYLGKLGVRKSTITQFLRTYPQVLHASVVVDLVPVVKYLQ 233
I +PL+LGCS NM ++ KLGVR + + + PQ+L + + VV +L+
Sbjct: 357 IRRWPLLLGCSAS-NMEMMVKEFDKLGVRDKRMGKVIPKMPQLLLCKPQ-EFLKVVCFLE 414
Query: 234 GMDIKPDDIPRVLERYPEVLGFKLEGTMSTSVAYLIGIGVGRRELGGILTRFPEILGMRV 293
+ + + + ++L R PE+ G +E T+ + +L GV I+ ++PE L
Sbjct: 415 DLGFQKEIVGQILCRCPEIFGCSIEKTLQKKLIFLTRFGVSTTHFPRIIKKYPEFLIYDA 474
Query: 294 GR---------VIKPFVEYLESLGIPRLAIARLIETQPYILGFDLDEKVKPNVKSLEEFN 344
+ + ++YL +GI IA +I ILG+ +D+ ++P + L N
Sbjct: 475 DKTKMTPNFVNICSYRLKYLMEIGISEREIAFMIRKFSPILGYSIDKVLRPKFEFL--VN 532
Query: 345 VRETSLASIIAQYP 358
E + +I +YP
Sbjct: 533 SMEKPVREVI-EYP 545
Score = 54.3 bits (129), Expect = 6e-06
Identities = 59/292 (20%), Positives = 117/292 (39%), Gaps = 30/292 (10%)
Query: 91 SKRDSGEIQRKRRGASSLYAGYARPSLSEMKKDKATLRKVVYEFLRGIGI-------VPD 143
S + SGE +R G + + M K K KV FL +G+ +
Sbjct: 117 SWKGSGESERVEEGLGF------KEKVIYMVKQKGDGGKVA--FLESLGLSLSSAMYLAH 168
Query: 144 ELDGLELPVTVDVMKERVDFLHS----LGLTIEDINNYPLVLGCSVKKNMVPVLDYLGKL 199
+ LP+ +D +K + S GL + L L + +++ L + K+
Sbjct: 169 YVSSESLPILLDKVKYLKEIFFSGSDEKGLVGKYARRMMLYLSIPIDEDVQQTLSFFEKI 228
Query: 200 GVRKSTITQF----------LRTYPQVLHASVVVDLVPVVKYLQGMDIKPDDIPRVLERY 249
R+ + L ++P++L S D+ P+V++L+ + I + +VL Y
Sbjct: 229 EARRGGLDMLGSVDASFRFLLESFPRLLLLSEENDMKPMVEFLESIGIPKYCLGKVLLLY 288
Query: 250 PEVLGFKLEGTMSTSVAYLIGIGVGRRELGGILTRFPEILGMRVGRVIKPFVEYLESLGI 309
P ++ K E + + V ++ G +L ++P IL + + S +
Sbjct: 289 PPIMLGKTEEIKRRVATAMEKVSVVNKDSGKLLLKYPWILSPSIQENYSHIGSFFYSESV 348
Query: 310 PRLAIARLIETQPYILGFDLDEKVKPNVKSLEEFNVRETSLASIIAQYPDII 361
++ I I P +LG ++ VK ++ VR+ + +I + P ++
Sbjct: 349 LKMDIDHAIRRWPLLLGCSA-SNMEMMVKEFDKLGVRDKRMGKVIPKMPQLL 399
Score = 44.7 bits (104), Expect = 0.005
Identities = 46/216 (21%), Positives = 92/216 (42%), Gaps = 42/216 (19%)
Query: 160 RVDFLHSLGLTIEDINNYPLVLGCSVKKNMVPVL----DYLGKLGVRKSTITQFLRTYPQ 215
+V FL SLGL++ + L V +P+L YL ++ S + Y +
Sbjct: 149 KVAFLESLGLSLSSA----MYLAHYVSSESLPILLDKVKYLKEIFFSGSDEKGLVGKYAR 204
Query: 216 --VLHASVVVDLVPVVKYLQGMDIKPDDIPRVLERYPEVL----GFKLEGTMSTSVAYLI 269
+L+ S+ +D +D+ + L + ++ G + G++ S +L
Sbjct: 205 RMMLYLSIPID---------------EDVQQTLSFFEKIEARRGGLDMLGSVDASFRFL- 248
Query: 270 GIGVGRRELGGILTRFPEILGMRVGRVIKPFVEYLESLGIPRLAIARLIETQPYILGFDL 329
L FP +L + +KP VE+LES+GIP+ + +++ P I+
Sbjct: 249 ------------LESFPRLLLLSEENDMKPMVEFLESIGIPKYCLGKVLLLYPPIMLGKT 296
Query: 330 DEKVKPNVKSLEEFNVRETSLASIIAQYPDIIGTDL 365
+E + ++E+ +V ++ +YP I+ +
Sbjct: 297 EEIKRRVATAMEKVSVVNKDSGKLLLKYPWILSPSI 332
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.322 0.142 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 606,467,664
Number of Sequences: 2540612
Number of extensions: 26051232
Number of successful extensions: 65515
Number of sequences better than 10.0: 100
Number of HSP's better than 10.0 without gapping: 54
Number of HSP's successfully gapped in prelim test: 46
Number of HSP's that attempted gapping in prelim test: 65001
Number of HSP's gapped (non-prelim): 316
length of query: 365
length of database: 863,360,394
effective HSP length: 129
effective length of query: 236
effective length of database: 535,621,446
effective search space: 126406661256
effective search space used: 126406661256
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 76 (33.9 bits)
Medicago: description of AC133139.1


