

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126784.7 - phase: 0
(339 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
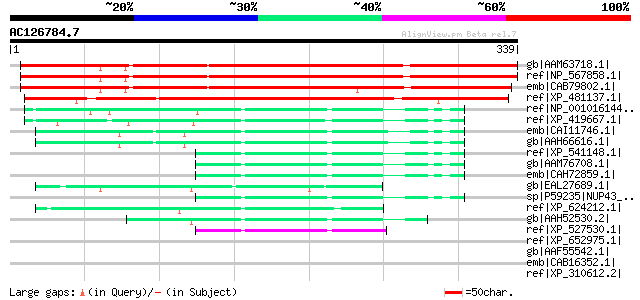
Score E
Sequences producing significant alignments: (bits) Value
gb|AAM63718.1| unknown [Arabidopsis thaliana] 378 e-103
ref|NP_567858.1| WD-40 repeat protein family [Arabidopsis thaliana] 376 e-103
emb|CAB79802.1| hypothetical protein [Arabidopsis thaliana] gi|2... 358 1e-97
ref|XP_481137.1| putative Nucleoporin Nup43 [Oryza sativa (japon... 269 8e-71
ref|NP_001016144.1| hypothetical protein LOC548898 [Xenopus trop... 64 8e-09
ref|XP_419667.1| PREDICTED: similar to Nucleoporin Nup43 (p42) [... 61 5e-08
emb|CAI11746.1| novel protein (zgc:77032) [Danio rerio] 59 2e-07
gb|AAH66616.1| Hypothetical protein zgc:77032 [Danio rerio] gi|4... 59 2e-07
ref|XP_541148.1| PREDICTED: hypothetical protein XP_541148 [Cani... 57 8e-07
gb|AAM76708.1| nucleoporin Nup43 [Homo sapiens] gi|40787641|gb|A... 55 2e-06
emb|CAH72859.1| nucleoporin 43kDa [Homo sapiens] 55 2e-06
gb|EAL27689.1| GA20514-PA [Drosophila pseudoobscura] 55 2e-06
sp|P59235|NUP43_MOUSE Nucleoporin Nup43 gi|26354845|dbj|BAC41049... 55 3e-06
ref|XP_624212.1| PREDICTED: similar to Nucleoporin Nup43 (p42) [... 54 5e-06
gb|AAH52530.2| Nup43 protein [Mus musculus] 54 9e-06
ref|XP_527530.1| PREDICTED: similar to Nucleoporin Nup43 (p42) [... 51 6e-05
ref|XP_652975.1| hypothetical protein 98.t00014 [Entamoeba histo... 44 0.009
gb|AAF55542.1| CG7671-PA [Drosophila melanogaster] gi|24647938|r... 43 0.015
emb|CAB16352.1| SPAC1A6.02 [Schizosaccharomyces pombe] gi|749090... 41 0.045
ref|XP_310612.2| ENSANGP00000019321 [Anopheles gambiae str. PEST... 41 0.059
>gb|AAM63718.1| unknown [Arabidopsis thaliana]
Length = 358
Score = 378 bits (971), Expect = e-103
Identities = 202/348 (58%), Positives = 234/348 (67%), Gaps = 23/348 (6%)
Query: 8 EIHRFPQYKYIDAVRWLPVLSAFNRFAVLATSDFDSNLSSIEIHSFKPNPLS-------F 60
++HR PQ KY+D VRWLP SA NRF A+ D D + SSIEI S PNP
Sbjct: 6 QVHRIPQSKYVDGVRWLPQASALNRFFATASYDADCDSSSIEIQSLDPNPRGNHNTNPLI 65
Query: 61 EFQSSWTSPSPISSIK-------SSQFLQKSIIASSTSSGSLHFLFADSTDARLVSEVSV 113
E SSWTSPS +SS++ F K +++++TSSGSLH L D + + EV
Sbjct: 66 ESLSSWTSPSRVSSLEVAGNGGGGGSF--KPMVSAATSSGSLHVLMIDLVEGAAIEEVYA 123
Query: 114 PENELHLEGKC-CIDLMDGGVECVTVGDDGRINLVT-VGDSNLNYRRLFDSGGLVSYTSV 171
E E G+ +D +GG ECVTVG+DGR+N+V V L YR++FD GLV+Y +V
Sbjct: 124 AEGERFHVGRVEGVDWREGG-ECVTVGEDGRVNVVKIVNGEGLRYRKVFDGNGLVAYRAV 182
Query: 172 KWASPVEFATGGYGFGLHWWDQRKPGGPVSQFKGNWDKNLNSGIVHSIDIHPSRKHTCLA 231
KWASP EF TGGYGFGL WDQRK G VSQ KGNW + S IVHSIDIHPSRKHTC+A
Sbjct: 183 KWASPTEFVTGGYGFGLQLWDQRKSGEAVSQLKGNWFQGKTSAIVHSIDIHPSRKHTCIA 242
Query: 232 AGSLGTVLAWDLRMQQQPIILSGTGAGDGAGNTAVQSISESEVWEVQYDRCIKSNTSSTR 291
GS GTV AWDLR QQPI+LSG GA + N +SESEVWEVQYD KSN SS+R
Sbjct: 243 GGSSGTVFAWDLRWPQQPIVLSGVGASENINN----PLSESEVWEVQYDSYTKSNVSSSR 298
Query: 292 ILPAMICSEDGILGVIEQGEEPIELLAEPCAINSFDIDRHNPLVSGCS 339
ILP M CSEDGILG+IEQGEEPIELLAEPCAINSFDIDR NP CS
Sbjct: 299 ILPVMTCSEDGILGIIEQGEEPIELLAEPCAINSFDIDRQNPQDVICS 346
>ref|NP_567858.1| WD-40 repeat protein family [Arabidopsis thaliana]
Length = 361
Score = 376 bits (966), Expect = e-103
Identities = 201/348 (57%), Positives = 233/348 (66%), Gaps = 23/348 (6%)
Query: 8 EIHRFPQYKYIDAVRWLPVLSAFNRFAVLATSDFDSNLSSIEIHSFKPNPLS-------F 60
++HR PQ KY+D VRWLP SA NRF A+ D D + SSIEI S PNP
Sbjct: 9 QVHRIPQSKYVDGVRWLPQASALNRFFATASYDADCDSSSIEIQSLDPNPRGNHNTNPLI 68
Query: 61 EFQSSWTSPSPISSIK-------SSQFLQKSIIASSTSSGSLHFLFADSTDARLVSEVSV 113
E SSWTSPS +SS++ F K +++++TSSGSLH L D + + E
Sbjct: 69 ESLSSWTSPSRVSSLEVAGNGGGGGSF--KPMVSAATSSGSLHVLMIDLVEGAAIEEFYA 126
Query: 114 PENELHLEGKC-CIDLMDGGVECVTVGDDGRINLVT-VGDSNLNYRRLFDSGGLVSYTSV 171
E E G+ +D +GG ECVTVG+DGR+N+V V L YR++FD GLV+Y +V
Sbjct: 127 AEGERFHVGRVEGVDWREGG-ECVTVGEDGRVNVVKIVNGEGLRYRKVFDGNGLVAYRAV 185
Query: 172 KWASPVEFATGGYGFGLHWWDQRKPGGPVSQFKGNWDKNLNSGIVHSIDIHPSRKHTCLA 231
KWASP EF TGGYGFGL WDQRK G VSQ KGNW + S IVHSIDIHPSRKHTC+A
Sbjct: 186 KWASPTEFVTGGYGFGLQLWDQRKSGEAVSQLKGNWFQGKTSAIVHSIDIHPSRKHTCIA 245
Query: 232 AGSLGTVLAWDLRMQQQPIILSGTGAGDGAGNTAVQSISESEVWEVQYDRCIKSNTSSTR 291
GS GTV AWDLR QQPI+LSG GA + N +SESEVWEVQYD KSN SS+R
Sbjct: 246 GGSSGTVFAWDLRWPQQPIVLSGVGASENINN----PLSESEVWEVQYDSYTKSNVSSSR 301
Query: 292 ILPAMICSEDGILGVIEQGEEPIELLAEPCAINSFDIDRHNPLVSGCS 339
ILP M CSEDGILG+IEQGEEPIELLAEPCAINSFDIDR NP CS
Sbjct: 302 ILPVMTCSEDGILGIIEQGEEPIELLAEPCAINSFDIDRQNPQDVICS 349
>emb|CAB79802.1| hypothetical protein [Arabidopsis thaliana]
gi|2980782|emb|CAA18209.1| hypothetical protein
[Arabidopsis thaliana] gi|25372649|pir||A85361
hypothetical protein AT4g30840 [imported] - Arabidopsis
thaliana
Length = 384
Score = 358 bits (920), Expect = 1e-97
Identities = 199/373 (53%), Positives = 231/373 (61%), Gaps = 52/373 (13%)
Query: 8 EIHRFPQYKYIDAVRWLPVLSAFNRFAVLATSDFDSNLSSIEIHSFKPNPLS-------F 60
++HR PQ KY+D VRWLP SA NRF A+ D D + SSIEI S PNP
Sbjct: 9 QVHRIPQSKYVDGVRWLPQASALNRFFATASYDADCDSSSIEIQSLDPNPRGNHNTNPLI 68
Query: 61 EFQSSWTSPSPISSIK-------SSQFLQKSIIASSTSSGSLHFLFADSTDARLVSEVSV 113
E SSWTSPS +SS++ F K +++++TSSGSLH L D + + E
Sbjct: 69 ESLSSWTSPSRVSSLEVAGNGGGGGSF--KPMVSAATSSGSLHVLMIDLVEGAAIEEFYA 126
Query: 114 PENELHLEGKC-CIDLMDGGVECVTVGDDGRINLVT-VGDSNLNYRRLFDSGGLVSYTSV 171
E E G+ +D +GG ECVTVG+DGR+N+V V L YR++FD GLV+Y +V
Sbjct: 127 AEGERFHVGRVEGVDWREGG-ECVTVGEDGRVNVVKIVNGEGLRYRKVFDGNGLVAYRAV 185
Query: 172 KWASPVEFATGGYGFGLHWWDQRKPGGPVSQFKGNWDKNLNSGIVHSIDIHPSRKHTCLA 231
KWASP EF TGGYGFGL WDQRK G VSQ KGNW + S IVHSIDIHPSRKHTC+
Sbjct: 186 KWASPTEFVTGGYGFGLQLWDQRKSGEAVSQLKGNWFQGKTSAIVHSIDIHPSRKHTCIF 245
Query: 232 -----------------------------AGSLGTVLAWDLRMQQQPIILSGTGAGDGAG 262
GS GTV AWDLR QQPI+LSG GA +
Sbjct: 246 EKRLSHQLMNMIIPSVILNLYQVFGYIAFGGSSGTVFAWDLRWPQQPIVLSGVGASENIN 305
Query: 263 NTAVQSISESEVWEVQYDRCIKSNTSSTRILPAMICSEDGILGVIEQGEEPIELLAEPCA 322
N +SESEVWEVQYD KSN SS+RILP M CSEDGILG+IEQGEEPIELLAEPCA
Sbjct: 306 NP----LSESEVWEVQYDSYTKSNVSSSRILPVMTCSEDGILGIIEQGEEPIELLAEPCA 361
Query: 323 INSFDIDRHNPLV 335
INSFDIDR NP V
Sbjct: 362 INSFDIDRQNPQV 374
>ref|XP_481137.1| putative Nucleoporin Nup43 [Oryza sativa (japonica cultivar-group)]
gi|37806397|dbj|BAC99935.1| putative Nucleoporin Nup43
[Oryza sativa (japonica cultivar-group)]
Length = 354
Score = 269 bits (688), Expect = 8e-71
Identities = 158/332 (47%), Positives = 202/332 (60%), Gaps = 20/332 (6%)
Query: 11 RFPQYKYIDAVRWLPVLSAFNRFAVLATSDFDS---NLSSIEIHSFKPNPLSFEFQSSWT 67
R P ID +RWLP S+ + +LA + D + SS +H L SS
Sbjct: 9 RHPHPFSIDLIRWLPCSSSSSSDRLLAAAVHDPAAPSSSSSHLHL-----LPLHDPSSPL 63
Query: 68 SPSPISSIKSSQFLQKSIIASSTSSGSLHFLFADSTDARLVSEVSVPENE-LHLEGKCCI 126
+ P+ S +S S++A++TSSGSLH L S DA + VSVP H+ +
Sbjct: 64 AALPLPSRAASLRCSPSVLAAATSSGSLHLL-PSSLDAAGSAGVSVPAGAGFHVGPVRGL 122
Query: 127 DLMDGGVECVTVGDDGRINLVTVG-DSNLNYRRLFDSGGLVSYTSVKWASPVEFATGGYG 185
D GG E VT G+DGR+++V G D + RRL+D G+ Y + +WAS EFATGG G
Sbjct: 123 DCGGGGEEWVTAGEDGRVHVVGGGGDGRVVARRLWDGKGMAGYEAARWASAAEFATGGAG 182
Query: 186 FGLHWWDQRKPGGPVSQFKGNWDKNLNSGIVHSIDIHPSRKHTCLAAGSLGTVLAWDLRM 245
G+ WWD+RK V+Q KG W + + +G+VHSIDIHPSRKH C+ GS GT+ AWDLR
Sbjct: 183 CGVQWWDRRKGDAVVAQCKGVWGRGIVTGMVHSIDIHPSRKHICVVGGSSGTIFAWDLRW 242
Query: 246 QQQPIILSGTGAGDGAGNTAVQSISESEVWEVQYDRCIKS----NTSSTRILPAMICSED 301
QQPI LSG G N Q +SESEVWEV +D +S +++STRILP M+CSED
Sbjct: 243 PQQPIPLSGLGL-----NGTAQPVSESEVWEVLFDNYTQSTDIISSASTRILPVMMCSED 297
Query: 302 GILGVIEQGEEPIELLAEPCAINSFDIDRHNP 333
GIL V+EQ E P+ELLAEPCAINSFDID NP
Sbjct: 298 GILAVVEQDERPLELLAEPCAINSFDIDPENP 329
>ref|NP_001016144.1| hypothetical protein LOC548898 [Xenopus tropicalis]
Length = 375
Score = 63.5 bits (153), Expect = 8e-09
Identities = 70/304 (23%), Positives = 119/304 (39%), Gaps = 38/304 (12%)
Query: 11 RFPQYKYIDAVRWLPV-LSAFNRFAVLATSDFDSNLSSIEIHS---FKPNPLSFEFQSS- 65
+F +K I RW PV S+ + V AT +D+ + + + + F L E+Q
Sbjct: 8 KFVSHK-ISRTRWRPVSASSLQQPDVFATGSWDNEENKVCVWATSDFGAISLEEEYQGDP 66
Query: 66 --WTSPSPISSIKSSQFLQKSIIASSTSSGSLHFLFADSTDARLVSEVSVPENELHLEGK 123
I + QFL + I +++S+G++ + L + H+
Sbjct: 67 KQLCDIRHIGDVMDMQFLDQERIVTASSTGTVTIFRHHENNQTLSVNQKWEQAHYHVGSN 126
Query: 124 C---CIDLMDGGVECVTVGDDGRINLVTVGDSNLNYRRLFDSGGLVSYTSVKWASPVEFA 180
C ++ E V+VG+DGRIN ++ R D + V + E
Sbjct: 127 MRAPCTGIVCSSPEIVSVGEDGRINCFRAESRDVV--RTIDDADSSTMHGVTFLRTTEIL 184
Query: 181 TGGYGFGLHWWDQRKPGGPVSQFKGNWDKNLNSGIVHSIDIHPSRKHTCLAAGSLGTVLA 240
T L WD RK G +Q + + +H +D HP+++H G G +
Sbjct: 185 TVNSVGQLKLWDLRKQGNDPTQIFSVTGERVP---LHCVDRHPNQQHVVATGGQDGMLCI 241
Query: 241 WDLRMQQQPIILSGTGAGDGAGNTAVQSISESEVWEVQYDRCIKSNTSSTRILPAMICSE 300
WD+R + P+ ++ + ++E+WEV + SN CSE
Sbjct: 242 WDVRHGKMPM--------------SLLNAHKAEMWEVHFH---PSNPDH-----LFTCSE 279
Query: 301 DGIL 304
DG L
Sbjct: 280 DGSL 283
>ref|XP_419667.1| PREDICTED: similar to Nucleoporin Nup43 (p42) [Gallus gallus]
Length = 555
Score = 60.8 bits (146), Expect = 5e-08
Identities = 75/311 (24%), Positives = 119/311 (38%), Gaps = 49/311 (15%)
Query: 11 RFPQYKYIDAVRWLPVLSAF----NRFAVLATSDFDSNLSSIEIHSFKPNPLSFEFQSSW 66
RF K I RW P+ +A + FA + + D+ +S + L+ E+Q
Sbjct: 188 RFVSQK-ISKARWRPLPAAALQPPDLFATGSWDNEDNRISIWSVGDVGSAGLNGEYQGE- 245
Query: 67 TSPSPISSIKSS------QFLQKSIIASSTSSGSLHFLFADSTDARLVSEVSVPENELHL 120
P + I+ + QFL + I +S+GS+ + L + H
Sbjct: 246 --PQLLCDIRHNGDVMDMQFLDQERIVVGSSTGSVTVFRHHQNNQTLSASHRWENAHYHA 303
Query: 121 E------GKCCIDLMDGGVECVTVGDDGRINLVTVGDSNLNYRRLFDSGGLVSYTSVKWA 174
+ G C ++ E VTVG+DGRINL + R D+ + +V +
Sbjct: 304 DQYTACGGAACTGVICNNPEIVTVGEDGRINLYRADQKDAV--RTIDNADSSTLHAVTFL 361
Query: 175 SPVEFATGGYGFGLHWWDQRKPGG-PVSQFKGNWDKNLNSGIVHSIDIHPSRKHTCLAAG 233
E T L WD R+ P F D+ +H +D HP+++H G
Sbjct: 362 RTTEILTVNSIGQLKIWDFRQQRNEPAQIFSLTGDRVP----LHCVDRHPNQQHIVATGG 417
Query: 234 SLGTVLAWDLRMQQQPIILSGTGAGDGAGNTAVQSISESEVWEVQYDRCIKSNTSSTRIL 293
G + WD+R P+ ++ + E+E+WEV + SN
Sbjct: 418 QDGMLSIWDIRQGTMPV--------------SLLNAHEAEMWEVHFH---PSNPDH---- 456
Query: 294 PAMICSEDGIL 304
CSEDG L
Sbjct: 457 -LFTCSEDGSL 466
>emb|CAI11746.1| novel protein (zgc:77032) [Danio rerio]
Length = 372
Score = 59.3 bits (142), Expect = 2e-07
Identities = 68/296 (22%), Positives = 116/296 (38%), Gaps = 37/296 (12%)
Query: 18 IDAVRWLPVLSA-FNRFAVLATSDFDSNLSSIEIHSFKPNPLSFEFQSSWTSPSPI---- 72
I RW PV +A + V AT +D+ + + I S S P +
Sbjct: 13 ISKTRWRPVSAASLQQPEVFATGSWDNENNKVSIWSVGDRGGSTLDGDLDEEPQLLCDSQ 72
Query: 73 --SSIKSSQFLQKSIIASSTSSGSLHFLFADSTDARLVSEVSVPE--NELHLEGKCCIDL 128
+ QFL + + S++SSG++ +F +D + +S V E + + C +
Sbjct: 73 HEGDVMDLQFLDQDRLVSASSSGAVS-IFKLQSDCQALSLAHVWERAHRCSCDNAPCTAI 131
Query: 129 MDGGVECVTVGDDGRINLVTVGDSNLNYRRLFDSGGLVSYTSVKWASPVEFATGGYGFGL 188
+ E V+VG+DGR+ L + + R+ ++ + +V + E T L
Sbjct: 132 VCRSPEIVSVGEDGRVILYKADQAEVT--RVIENADSSTIHAVTFLRTTEVLTVNSIGQL 189
Query: 189 HWWDQRKPGGPVSQFKGNWDKNLNSGIVHSIDIHPSRKHTCLAAGSLGTVLAWDLRMQQQ 248
WD R+ +Q + +H +D HP+++H G G + WD+R
Sbjct: 190 KMWDFRQQSDSPAQILSLTGDRVP---LHCVDKHPNQQHIVATGGQDGMLCIWDVRQGNT 246
Query: 249 PIILSGTGAGDGAGNTAVQSISESEVWEVQYDRCIKSNTSSTRILPAMICSEDGIL 304
P L+ +E+WEV + SN + CSEDG L
Sbjct: 247 PFSLT--------------EAHSAEMWEVHFH---PSNPNH-----LFTCSEDGSL 280
>gb|AAH66616.1| Hypothetical protein zgc:77032 [Danio rerio]
gi|47086251|ref|NP_998057.1| hypothetical protein
zgc:77032 [Danio rerio]
Length = 372
Score = 58.9 bits (141), Expect = 2e-07
Identities = 68/296 (22%), Positives = 115/296 (37%), Gaps = 37/296 (12%)
Query: 18 IDAVRWLPVLSA-FNRFAVLATSDFDSNLSSIEIHSFKPNPLSFEFQSSWTSPSPI---- 72
I RW PV +A + V AT +D+ + + I S S P +
Sbjct: 13 ISKTRWRPVSAASLQQPEVFATGSWDNENNKVSIWSVGDRGGSTLDGDLDEEPQLLCDSQ 72
Query: 73 --SSIKSSQFLQKSIIASSTSSGSLHFLFADSTDARLVSEVSVPE--NELHLEGKCCIDL 128
+ QFL + + S++SSG++ +F +D + +S V E + C +
Sbjct: 73 HEGDVMDLQFLDQDRLVSASSSGAVS-IFKLQSDCQALSLAHVWERAHRCSCNNAPCTAI 131
Query: 129 MDGGVECVTVGDDGRINLVTVGDSNLNYRRLFDSGGLVSYTSVKWASPVEFATGGYGFGL 188
+ E V+VG+DGR+ L + + R+ ++ + +V + E T L
Sbjct: 132 VCRSPEIVSVGEDGRVILYKADQAEVT--RVIENADSSTIHAVTFLRTTEVLTVNSIGQL 189
Query: 189 HWWDQRKPGGPVSQFKGNWDKNLNSGIVHSIDIHPSRKHTCLAAGSLGTVLAWDLRMQQQ 248
WD R+ +Q + +H +D HP+++H G G + WD+R
Sbjct: 190 KMWDFRQQSDSPAQILSLTGDRMP---LHCVDKHPNQQHIVATGGQDGMLCIWDVRQGNT 246
Query: 249 PIILSGTGAGDGAGNTAVQSISESEVWEVQYDRCIKSNTSSTRILPAMICSEDGIL 304
P L+ +E+WEV + SN + CSEDG L
Sbjct: 247 PFSLT--------------EAHSAEMWEVHFH---PSNPNH-----LFTCSEDGSL 280
>ref|XP_541148.1| PREDICTED: hypothetical protein XP_541148 [Canis familiaris]
Length = 427
Score = 57.0 bits (136), Expect = 8e-07
Identities = 49/180 (27%), Positives = 70/180 (38%), Gaps = 27/180 (15%)
Query: 125 CIDLMDGGVECVTVGDDGRINLVTVGDSNLNYRRLFDSGGLVSYTSVKWASPVEFATGGY 184
C ++ E VTVG+DGRINL R D+ + +V + E T
Sbjct: 181 CTGVVCNSPEIVTVGEDGRINLFRADHKEAV--RTIDNADSSTLHAVTFLRTPEILTVNS 238
Query: 185 GFGLHWWDQRKPGGPVSQFKGNWDKNLNSGIVHSIDIHPSRKHTCLAAGSLGTVLAWDLR 244
L WD RK G SQ + +H +D HP+++H G G + WD+R
Sbjct: 239 IGQLKIWDFRKQGNEPSQILSLTGDRVP---LHCVDRHPNQQHVVATGGQDGMLSIWDVR 295
Query: 245 MQQQPIILSGTGAGDGAGNTAVQSISESEVWEVQYDRCIKSNTSSTRILPAMICSEDGIL 304
P+ ++ E+E+WEV + SN CSEDG L
Sbjct: 296 QGTMPV--------------SLLKAHEAEMWEVHFH---PSNPDH-----LFTCSEDGSL 333
>gb|AAM76708.1| nucleoporin Nup43 [Homo sapiens] gi|40787641|gb|AAH65028.1|
Nucleoporin 43kDa [Homo sapiens]
gi|27923819|sp|Q8NFH3|NUP43_HUMAN Nucleoporin Nup43
(p42) gi|38605733|ref|NP_942590.1| nucleoporin 43kDa
[Homo sapiens] gi|32189380|ref|NP_078923.3| nucleoporin
43kDa [Homo sapiens]
Length = 380
Score = 55.5 bits (132), Expect = 2e-06
Identities = 48/180 (26%), Positives = 70/180 (38%), Gaps = 27/180 (15%)
Query: 125 CIDLMDGGVECVTVGDDGRINLVTVGDSNLNYRRLFDSGGLVSYTSVKWASPVEFATGGY 184
C ++ E VTVG+DGRINL R D+ + +V + E T
Sbjct: 134 CTGVVCNNPEIVTVGEDGRINLFRADHKEAV--RTIDNADSSTLHAVTFLRTPEILTVNS 191
Query: 185 GFGLHWWDQRKPGGPVSQFKGNWDKNLNSGIVHSIDIHPSRKHTCLAAGSLGTVLAWDLR 244
L WD R+ G SQ + +H +D HP+++H G G + WD+R
Sbjct: 192 IGQLKIWDFRQQGNEPSQILSLTGDRVP---LHCVDRHPNQQHVVATGGQDGMLSIWDVR 248
Query: 245 MQQQPIILSGTGAGDGAGNTAVQSISESEVWEVQYDRCIKSNTSSTRILPAMICSEDGIL 304
P+ ++ E+E+WEV + SN CSEDG L
Sbjct: 249 QGTMPV--------------SLLKAHEAEMWEVHFH---PSNPEH-----LFTCSEDGSL 286
>emb|CAH72859.1| nucleoporin 43kDa [Homo sapiens]
Length = 441
Score = 55.5 bits (132), Expect = 2e-06
Identities = 48/180 (26%), Positives = 70/180 (38%), Gaps = 27/180 (15%)
Query: 125 CIDLMDGGVECVTVGDDGRINLVTVGDSNLNYRRLFDSGGLVSYTSVKWASPVEFATGGY 184
C ++ E VTVG+DGRINL R D+ + +V + E T
Sbjct: 195 CTGVVCNNPEIVTVGEDGRINLFRADHKEAV--RTIDNADSSTLHAVTFLRTPEILTVNS 252
Query: 185 GFGLHWWDQRKPGGPVSQFKGNWDKNLNSGIVHSIDIHPSRKHTCLAAGSLGTVLAWDLR 244
L WD R+ G SQ + +H +D HP+++H G G + WD+R
Sbjct: 253 IGQLKIWDFRQQGNEPSQILSLTGDRVP---LHCVDRHPNQQHVVATGGQDGMLSIWDVR 309
Query: 245 MQQQPIILSGTGAGDGAGNTAVQSISESEVWEVQYDRCIKSNTSSTRILPAMICSEDGIL 304
P+ ++ E+E+WEV + SN CSEDG L
Sbjct: 310 QGTMPV--------------SLLKAHEAEMWEVHFH---PSNPEH-----LFTCSEDGSL 347
>gb|EAL27689.1| GA20514-PA [Drosophila pseudoobscura]
Length = 355
Score = 55.5 bits (132), Expect = 2e-06
Identities = 60/245 (24%), Positives = 97/245 (39%), Gaps = 19/245 (7%)
Query: 18 IDAVRWLPV-LSAFNRFAVLATSDFDSNLSSIEIHSFKPNPLS---FEFQSSWTSPSPIS 73
I AVRWLP L +RF T +D + + + ++ + N + +F I
Sbjct: 15 ISAVRWLPEQLMQSDRFV---TGSWDMDQNFVRLYRLQSNQYADTNVDFVPRCNDKVAIE 71
Query: 74 S-IKSSQFLQKSIIASSTSSGSLHFLFADSTDARLVSEVSVPENELHL-----EGKCCID 127
+ + +F K+ +A S + G L L + + LH + C D
Sbjct: 72 GDVTAMEFADKNTLAVSCADGHLSLLNMQRAVEEDQLQRTARSERLHYFKRTNKSSPCTD 131
Query: 128 LMDGGVECVTVGDDGRINLVTVGDSNLNYRRLFDSGGLVSYTSVKWASPVEFATGGYGFG 187
L G + TVG+DG +N++ SN+ + +S SV + S + T
Sbjct: 132 LSVYGGDIATVGEDGCVNIMNA--SNIKQVKRTIEADSMSLLSVCYISQQQLVTANRMGV 189
Query: 188 LHWWDQRKPGGP---VSQFKGNWDKNLNSGIVHSIDIHPSRKHTCLAAGSLGTVLAWDLR 244
+ D R +S G+ D+ S V ++ HP + H L GT+ WDLR
Sbjct: 190 IRILDARASTAAQEKISIMVGSKDEK-RSNFVSALGTHPMQHHILLCGTEEGTITVWDLR 248
Query: 245 MQQQP 249
Q P
Sbjct: 249 NLQFP 253
>sp|P59235|NUP43_MOUSE Nucleoporin Nup43 gi|26354845|dbj|BAC41049.1| unnamed protein
product [Mus musculus]
Length = 380
Score = 55.1 bits (131), Expect = 3e-06
Identities = 48/180 (26%), Positives = 69/180 (37%), Gaps = 27/180 (15%)
Query: 125 CIDLMDGGVECVTVGDDGRINLVTVGDSNLNYRRLFDSGGLVSYTSVKWASPVEFATGGY 184
C ++ E VTVG+DGRINL V R D+ + +V + E T
Sbjct: 134 CTGIVCDNPEIVTVGEDGRINLFRVDHKEAV--RTIDNADSSTLHAVTFLRTPEIVTVNS 191
Query: 185 GFGLHWWDQRKPGGPVSQFKGNWDKNLNSGIVHSIDIHPSRKHTCLAAGSLGTVLAWDLR 244
L WD R+ G Q + +H +D HP ++H G G + WD+R
Sbjct: 192 IGQLKIWDFRQQGSEPCQILSLTGDRVP---LHCVDRHPDQQHVVATGGQDGMLSIWDVR 248
Query: 245 MQQQPIILSGTGAGDGAGNTAVQSISESEVWEVQYDRCIKSNTSSTRILPAMICSEDGIL 304
P+ ++ E+E+WEV + SN CSEDG L
Sbjct: 249 QGTMPV--------------SLLKAHEAEMWEVHFH---PSNPDH-----LFTCSEDGSL 286
>ref|XP_624212.1| PREDICTED: similar to Nucleoporin Nup43 (p42) [Apis mellifera]
Length = 346
Score = 54.3 bits (129), Expect = 5e-06
Identities = 54/239 (22%), Positives = 97/239 (39%), Gaps = 14/239 (5%)
Query: 18 IDAVRWLPVLSAFNRFAVLATSDFDSNLSSIEIHSFKPNPLSFEFQSSWTSPSPISSIKS 77
I +RW F T +D ++ + +F+ N + + +S + + +
Sbjct: 14 ISKIRWKH--EDFEDATNFITGSWDDPVNKVTHWTFQMNDDGESYPAVVSSYAILGDVTE 71
Query: 78 SQFLQKSIIASSTSSGSLHFL-FADSTDARLVSEVS---VPENELHLEGKCCIDLMDGGV 133
+F+ K STS G++ L ++ ++ +S + + + C L
Sbjct: 72 IKFISKDFFVVSTSIGTVRLLQIHENPYSQFKEHMSWEFIHRFDNTNDYAPCTGLSTFEQ 131
Query: 134 ECVTVGDDGRINLVTVGDSNLNYRRLFDSGGLVSYTSVKWASPVEFATGGYGFGLHWWDQ 193
+ +VG+DGRINL+T G R+ D S + + E TG + WD
Sbjct: 132 DIASVGEDGRINLLTAGQKQPV--RVIDDADSCSLYCIDFLRHNEILTGNIRGHMKVWDL 189
Query: 194 RKPGG-PVSQFK-GNWDKNLNSGIVHSIDIHPSRKHTCLAAGSLGTVLAWDLRMQQQPI 250
R P + F + K + I H HP+++H +A G G++ WDLR P+
Sbjct: 190 RNDQDLPATTFMLSDQAKTEATSIAH----HPTQRHIVVAGGGDGSLTVWDLRHNTYPM 244
>gb|AAH52530.2| Nup43 protein [Mus musculus]
Length = 375
Score = 53.5 bits (127), Expect = 9e-06
Identities = 48/208 (23%), Positives = 76/208 (36%), Gaps = 26/208 (12%)
Query: 79 QFLQKSIIASSTSSGSLHFLFADSTDARLVSEVSVPENELHL-------EGKCCIDLMDG 131
QF + I +++S+G + + L P H C ++
Sbjct: 76 QFFDQERIVAASSTGCVTVFLHHPNNQTLSVNQQWPAAHYHTGPSSPSYSSAPCTGIVCD 135
Query: 132 GVECVTVGDDGRINLVTVGDSNLNYRRLFDSGGLVSYTSVKWASPVEFATGGYGFGLHWW 191
E VTVG+DGRINL V R D+ + +V + E T L W
Sbjct: 136 NPEIVTVGEDGRINLFRVDHKEAV--RTIDNADSSTLHAVTFLRTPEIVTVNSIGQLKIW 193
Query: 192 DQRKPGGPVSQFKGNWDKNLNSGIVHSIDIHPSRKHTCLAAGSLGTVLAWDLRMQQQPII 251
D R+ G Q + +H +D HP ++H G G + WD+R P+
Sbjct: 194 DFRQQGSEPCQILSLTGDRVP---LHCVDRHPDQQHVVATGGQDGMLSIWDVRQGTMPV- 249
Query: 252 LSGTGAGDGAGNTAVQSISESEVWEVQY 279
++ E+E+WEV +
Sbjct: 250 -------------SLLKAHEAEMWEVHF 264
>ref|XP_527530.1| PREDICTED: similar to Nucleoporin Nup43 (p42) [Pan troglodytes]
Length = 507
Score = 50.8 bits (120), Expect = 6e-05
Identities = 36/128 (28%), Positives = 53/128 (41%), Gaps = 5/128 (3%)
Query: 125 CIDLMDGGVECVTVGDDGRINLVTVGDSNLNYRRLFDSGGLVSYTSVKWASPVEFATGGY 184
C ++ E VTVG+DGRINL R D+ + +V + E T
Sbjct: 195 CTGVVCNNPEIVTVGEDGRINLFRADHKEAV--RTIDNADSSTLHAVTFLRTPEILTVNS 252
Query: 185 GFGLHWWDQRKPGGPVSQFKGNWDKNLNSGIVHSIDIHPSRKHTCLAAGSLGTVLAWDLR 244
L WD R+ G SQ + +H +D HP+++H G G + WD+R
Sbjct: 253 IGQLKIWDFRQQGNEPSQILSLTGDRVP---LHCVDRHPNQQHVVATGGQDGMLSIWDVR 309
Query: 245 MQQQPIIL 252
P+ L
Sbjct: 310 QGTMPVSL 317
>ref|XP_652975.1| hypothetical protein 98.t00014 [Entamoeba histolytica HM-1:IMSS]
gi|56469885|gb|EAL47589.1| hypothetical protein
98.t00014 [Entamoeba histolytica HM-1:IMSS]
Length = 685
Score = 43.5 bits (101), Expect = 0.009
Identities = 27/80 (33%), Positives = 41/80 (50%)
Query: 77 SSQFLQKSIIASSTSSGSLHFLFADSTDARLVSEVSVPENELHLEGKCCIDLMDGGVECV 136
SS+ L SI + +T S + LFA ST+ +++S + L C I L+ GG
Sbjct: 107 SSKKLDISITSPATFSTNHQLLFACSTNNKIISFTNKFTQNLIYSVNCGIQLISGGTTLA 166
Query: 137 TVGDDGRINLVTVGDSNLNY 156
T +DG IN+ + +NL Y
Sbjct: 167 TKYNDGSINVFKIESNNLTY 186
>gb|AAF55542.1| CG7671-PA [Drosophila melanogaster] gi|24647938|ref|NP_650716.2|
CG7671-PA [Drosophila melanogaster]
Length = 358
Score = 42.7 bits (99), Expect = 0.015
Identities = 53/247 (21%), Positives = 95/247 (38%), Gaps = 21/247 (8%)
Query: 18 IDAVRWLPV-LSAFNRFAVLATSDFDSNLSSIEIHSFKPNPLSFEFQSSWTS-PSPISSI 75
+ VRWLP L RF T +D + + + + + N S P + +
Sbjct: 15 VSQVRWLPEELQQSERFV---TGSWDMDQNFVRLWRLQSNQYVTATDSEVDQIPRCMDKV 71
Query: 76 K------SSQFLQKSIIASSTSSGSLHFLFADSTDARLVSEVSVPENELHL-----EGKC 124
+ + +F+ K +A S + G L + + LH E
Sbjct: 72 RMEDDVTAMEFVDKDTLAVSCADGHLSVFNVHRAVEEDQMQRTSRSGRLHCFQRSQEPAP 131
Query: 125 CIDLMDGGVECVTVGDDGRINLVTVGDSNLNYRRLFDSGGLVSYTSVKWASPVEFATGGY 184
C DL G + T G+DG +++V+V ++ +R ++ + + S+ + S E T
Sbjct: 132 CTDLSVYGTDIATAGEDGCVSIVSV-ENVRQVKRQIEADSMALF-SICYISQQELVTANR 189
Query: 185 GFGLHWWDQR--KPGGPVSQFKGNWDKNLNSGIVHSIDIHPSRKHTCLAAGSLGTVLAWD 242
+ D R P + F ++ S V S+ HP ++H L G++ WD
Sbjct: 190 MGVIRMLDARVATEAQPKTTFMVA-SQDDKSNFVSSLAKHPMQQHILLCGSEEGSITVWD 248
Query: 243 LRMQQQP 249
+R P
Sbjct: 249 MRNLSYP 255
>emb|CAB16352.1| SPAC1A6.02 [Schizosaccharomyces pombe] gi|7490901|pir||T38005
hypothetical protein SPAC1A6.02 - fission yeast
(Schizosaccharomyces pombe)
gi|51316564|sp|O13856|YEX2_SCHPO Hypothetical WD-repeat
protein C1A6.02 in chromosome I
Length = 361
Score = 41.2 bits (95), Expect = 0.045
Identities = 35/150 (23%), Positives = 63/150 (41%), Gaps = 10/150 (6%)
Query: 104 DARLVSEVSVPENELHLEGKCCIDLMDGGVECVTVGDDGRINLVTVGDSNLNYRRLFDSG 163
D +S V + H + I + + G E ++VG DG + + ++ + + D
Sbjct: 43 DVAQISLVEQWSTKRHKKSCRNISVNESGTEFISVGSDGVLKIADTSTGRVSSKWIVDKN 102
Query: 164 GLVS-YTSVKW-ASPVEFATGGYGFGLHWWDQRKPGGPVSQFKGNWDKNLNSGIVHSIDI 221
+S Y+ V+W + + FATG + WD+R GG + N I + I
Sbjct: 103 KEISPYSVVQWIENDMVFATGDDNGCVSVWDKRTEGGIIHTH--------NDHIDYISSI 154
Query: 222 HPSRKHTCLAAGSLGTVLAWDLRMQQQPII 251
P + +A G + D R ++PI+
Sbjct: 155 SPFEERYFVATSGDGVLSVIDARNFKKPIL 184
>ref|XP_310612.2| ENSANGP00000019321 [Anopheles gambiae str. PEST]
gi|55243318|gb|EAA06600.2| ENSANGP00000019321 [Anopheles
gambiae str. PEST]
Length = 292
Score = 40.8 bits (94), Expect = 0.059
Identities = 60/254 (23%), Positives = 104/254 (40%), Gaps = 22/254 (8%)
Query: 6 NSEIHRFPQYKYIDAVRWLPVLSAFNRFAVLATSDFDSNLSSIEIHSFKPNPLSFEFQSS 65
+ EI F + VRWLP + F V T + ++++ + + + L+ E +
Sbjct: 2 HDEIVSFLLVDKLSRVRWLPNQTEDEHFFV--TGSWGERVNTVRLWNLVHDRLTDEDDTG 59
Query: 66 WTS-PSPISS------IKSSQFLQKSIIASSTSSGSLHFLFADSTDARLVSEVSVPEN-- 116
P P + I +FL +A+ TS G+L L + ++RL + + N
Sbjct: 60 VPLVPQPTAKFGVTGDIVGLEFLDDKHLAAVTSEGTLSVLDLNR-ESRLSYDFTHTYNLH 118
Query: 117 ELHLEG---KCCIDLMDGGVECVTVGDDGRINLVTVGDSNLNYRRLFD-SGGLVSYTSVK 172
+LH G C + VT G+DG +N+V GD+ R + D GG V S
Sbjct: 119 DLHTNGGVRSACTGVSAFDQYLVTGGEDGTVNMVA-GDAGKVIRTIRDPDGGAVQCVSFI 177
Query: 173 WASPVEFATGGYGFG-LHWWDQRKPGG-PVSQFKGNWDKNLNSGIVHSIDIHPSRKHTCL 230
+ + G +G + +D R PV + +++ + I+ P K
Sbjct: 178 YP---DLVIVGQQYGVIDCYDTRDESSKPVFSVETCVEEDRDLNKPTCINHFPKNKQVVA 234
Query: 231 AAGSLGTVLAWDLR 244
GT++ WD+R
Sbjct: 235 IGLESGTIILWDIR 248
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.317 0.134 0.407
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 622,225,401
Number of Sequences: 2540612
Number of extensions: 27782202
Number of successful extensions: 64134
Number of sequences better than 10.0: 93
Number of HSP's better than 10.0 without gapping: 14
Number of HSP's successfully gapped in prelim test: 79
Number of HSP's that attempted gapping in prelim test: 64039
Number of HSP's gapped (non-prelim): 117
length of query: 339
length of database: 863,360,394
effective HSP length: 128
effective length of query: 211
effective length of database: 538,162,058
effective search space: 113552194238
effective search space used: 113552194238
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 75 (33.5 bits)
Medicago: description of AC126784.7


