

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC119415.1 + phase: 0
(338 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
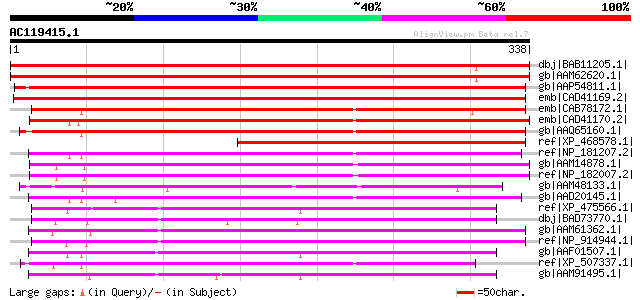
Score E
Sequences producing significant alignments: (bits) Value
dbj|BAB11205.1| flavanone 3-hydroxylase-like protein [Arabidopsi... 496 e-139
gb|AAM62620.1| flavanone 3-hydroxylase-like protein [Arabidopsis... 487 e-136
gb|AAP54811.1| unknown protein [Oryza sativa (japonica cultivar-... 463 e-129
emb|CAD41169.2| OSJNBa0064M23.14 [Oryza sativa (japonica cultiva... 463 e-129
emb|CAB78172.1| putative flavanone 3-beta-hydroxylase [Arabidops... 380 e-104
emb|CAD41170.2| OSJNBa0064M23.15 [Oryza sativa (japonica cultiva... 380 e-104
gb|AAQ65160.1| At4g10500 [Arabidopsis thaliana] gi|7267747|emb|C... 378 e-103
ref|XP_468578.1| Putative flavanone 3-hydroxylase [Oryza sativa ... 290 3e-77
ref|NP_181207.2| oxidoreductase, 2OG-Fe(II) oxygenase family pro... 254 3e-66
gb|AAM14878.1| putative flavonol synthase [Arabidopsis thaliana]... 241 2e-62
ref|NP_182007.2| oxidoreductase, 2OG-Fe(II) oxygenase family pro... 241 2e-62
gb|AAM48133.1| putative flavanone 3-hydroxylase [Saussurea medus... 240 5e-62
gb|AAD20145.1| putative giberellin beta-hydroxylase [Arabidopsis... 239 7e-62
ref|XP_475566.1| putative leucoanthocyanidin dioxygenase (EC 1.1... 233 6e-60
dbj|BAD73770.1| putative anthocyanidin synthase [Oryza sativa (j... 233 6e-60
gb|AAM61362.1| putative ethylene-forming enzyme [Arabidopsis tha... 229 7e-59
ref|NP_914944.1| putative ethylene-forming enzyme [Oryza sativa ... 229 1e-58
gb|AAF01507.1| putative leucoanthocyanidin dioxygenase [Arabidop... 226 6e-58
ref|XP_507337.1| PREDICTED P0562A06.31 gene product [Oryza sativ... 224 2e-57
gb|AAM91495.1| AT5g05600/MOP10_14 [Arabidopsis thaliana] gi|1017... 224 2e-57
>dbj|BAB11205.1| flavanone 3-hydroxylase-like protein [Arabidopsis thaliana]
gi|20148253|gb|AAM10017.1| flavanone 3-hydroxylase-like
protein [Arabidopsis thaliana]
gi|15238567|ref|NP_197841.1| oxidoreductase, 2OG-Fe(II)
oxygenase family protein [Arabidopsis thaliana]
gi|14423476|gb|AAK62420.1| flavanone 3-hydroxylase-like
protein [Arabidopsis thaliana]
Length = 341
Score = 496 bits (1277), Expect = e-139
Identities = 233/341 (68%), Positives = 280/341 (81%), Gaps = 3/341 (0%)
Query: 1 MDTKVLSSGIHYSKLPESYIRPESDRPCLSQVSEFENVPIIDLGSHNRTQIVQQIGEACS 60
M K++S+G ++ LPE+Y+RP SDRP LS+VS+ E+ P+IDL S +R+ ++QQI +AC+
Sbjct: 1 MAAKLISTGFRHTTLPENYVRPISDRPRLSEVSQLEDFPLIDLSSTDRSFLIQQIHQACA 60
Query: 61 SYGFFQVVNHGVPLEELKKTAEVAYDFFKLPVEEKMKLYSDDPTKTMRLSTSFNVNKEEV 120
+GFFQV+NHGV + + + VA +FF + +EEKMKLYSDDPTKT RLSTSFNV KEEV
Sbjct: 61 RFGFFQVINHGVNKQIIDEMVSVAREFFSMSMEEKMKLYSDDPTKTTRLSTSFNVKKEEV 120
Query: 121 HNWRDYLRLHCYPLDNYVPEWPSNPPSFKETVANYCKEVRELGLRIEEYISESLGLEKDY 180
+NWRDYLRLHCYP+ YV EWPSNPPSFKE V+ Y +EVRE+G +IEE ISESLGLEKDY
Sbjct: 121 NNWRDYLRLHCYPIHKYVNEWPSNPPSFKEIVSKYSREVREVGFKIEELISESLGLEKDY 180
Query: 181 LRNALGEQGQHMAVNYYPPCPQPELTYGLPGHTDPNALTILLQDLHVAGLQVLKDGKWLA 240
++ LGEQGQHMAVNYYPPCP+PELTYGLP HTDPNALTILLQD V GLQ+L DG+W A
Sbjct: 181 MKKVLGEQGQHMAVNYYPPCPEPELTYGLPAHTDPNALTILLQDTTVCGLQILIDGQWFA 240
Query: 241 INPIPDAFVINIGDQLQALSNGLYKSVWHRAIVNAEKPRLSVASFLCPDNEALICPAKPL 300
+NP PDAFVINIGDQLQALSNG+YKSVWHRA+ N E PRLSVASFLCP + A++ PAKPL
Sbjct: 241 VNPHPDAFVINIGDQLQALSNGVYKSVWHRAVTNTENPRLSVASFLCPADCAVMSPAKPL 300
Query: 301 TE---DGSGAVYRGFTYPEYYSKFWSRDLEKEHCLEFFKNN 338
E D + VY+ FTY EYY KFWSR+L++EHCLE F NN
Sbjct: 301 WEAEDDETKPVYKDFTYAEYYKKFWSRNLDQEHCLENFLNN 341
>gb|AAM62620.1| flavanone 3-hydroxylase-like protein [Arabidopsis thaliana]
Length = 341
Score = 487 bits (1253), Expect = e-136
Identities = 231/341 (67%), Positives = 277/341 (80%), Gaps = 3/341 (0%)
Query: 1 MDTKVLSSGIHYSKLPESYIRPESDRPCLSQVSEFENVPIIDLGSHNRTQIVQQIGEACS 60
M K++S+G ++ LPE+Y+RP SDRP LS+VS+ E+ P+IDL S +R+ ++QQI +AC+
Sbjct: 1 MAAKLISTGFRHTTLPENYVRPISDRPRLSEVSQLEDFPLIDLSSTDRSFLIQQIHQACA 60
Query: 61 SYGFFQVVNHGVPLEELKKTAEVAYDFFKLPVEEKMKLYSDDPTKTMRLSTSFNVNKEEV 120
+GFFQV NHGV + + + VA +FF + +EEKMKLYSDDPTKT RLSTSFNV KEEV
Sbjct: 61 RFGFFQVKNHGVNKQIIDEMVSVAREFFSMSMEEKMKLYSDDPTKTTRLSTSFNVKKEEV 120
Query: 121 HNWRDYLRLHCYPLDNYVPEWPSNPPSFKETVANYCKEVRELGLRIEEYISESLGLEKDY 180
+NWRDYLRLHCYP+ YV EWPSNPPSFKE V+ Y +EVRE+G +IEE ISESLGLEKDY
Sbjct: 121 NNWRDYLRLHCYPIHKYVNEWPSNPPSFKEIVSKYSREVREVGFKIEELISESLGLEKDY 180
Query: 181 LRNALGEQGQHMAVNYYPPCPQPELTYGLPGHTDPNALTILLQDLHVAGLQVLKDGKWLA 240
++ LGEQGQHMAVNYYPPCP+PELTYGLP HTDPNALTILLQD V GLQ+L DG+W A
Sbjct: 181 MKKVLGEQGQHMAVNYYPPCPEPELTYGLPAHTDPNALTILLQDTTVCGLQILIDGQWFA 240
Query: 241 INPIPDAFVINIGDQLQALSNGLYKSVWHRAIVNAEKPRLSVASFLCPDNEALICPAKPL 300
+NP PDAFVINIGDQLQALSNG+YKSVW RA+ N E PRLSVASFLCP + A++ PAKPL
Sbjct: 241 VNPHPDAFVINIGDQLQALSNGVYKSVWRRAVTNTENPRLSVASFLCPADCAVMSPAKPL 300
Query: 301 TE---DGSGAVYRGFTYPEYYSKFWSRDLEKEHCLEFFKNN 338
E D + VY+ FTY EYY KFWSR+L++EH LE F NN
Sbjct: 301 WEAEDDETKPVYKDFTYAEYYKKFWSRNLDQEHFLENFLNN 341
>gb|AAP54811.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|21717150|gb|AAM76343.1| unknown protein [Oryza sativa
(japonica cultivar-group)] gi|37536444|ref|NP_922524.1|
unknown protein [Oryza sativa (japonica cultivar-group)]
gi|18057095|gb|AAL58118.1| putative flavanone
3-hydroxylase [Oryza sativa (japonica cultivar-group)]
Length = 342
Score = 463 bits (1192), Expect = e-129
Identities = 216/333 (64%), Positives = 256/333 (76%), Gaps = 1/333 (0%)
Query: 4 KVLSSGIHYSKLPESYIRPESDRPCLSQVSEFENVPIIDLGSHNRTQIVQQIGEACSSYG 63
++LS+ +H +P Y+RPES RP L V +P++DL S +R +V +G+AC ++G
Sbjct: 10 QLLSTAVH-DTMPGKYVRPESQRPRLDLVVSDARIPVVDLASPDRAAVVSAVGDACRTHG 68
Query: 64 FFQVVNHGVPLEELKKTAEVAYDFFKLPVEEKMKLYSDDPTKTMRLSTSFNVNKEEVHNW 123
FFQVVNHG+ + EV +FF+LP EEK KLYSDDP K +RLSTSFNV KE VHNW
Sbjct: 69 FFQVVNHGIDAALIASVMEVGREFFRLPAEEKAKLYSDDPAKKIRLSTSFNVRKETVHNW 128
Query: 124 RDYLRLHCYPLDNYVPEWPSNPPSFKETVANYCKEVRELGLRIEEYISESLGLEKDYLRN 183
RDYLRLHCYPL +VP+WPSNPPSFKE + YC EVRELG R+ E ISESLGLE Y+R
Sbjct: 129 RDYLRLHCYPLHQFVPDWPSNPPSFKEIIGTYCTEVRELGFRLYEAISESLGLEGGYMRE 188
Query: 184 ALGEQGQHMAVNYYPPCPQPELTYGLPGHTDPNALTILLQDLHVAGLQVLKDGKWLAINP 243
LGEQ QHMAVNYYP CP+PELTYGLP HTDPNALTILL D VAGLQVL DGKW+A+NP
Sbjct: 189 TLGEQEQHMAVNYYPQCPEPELTYGLPAHTDPNALTILLMDDQVAGLQVLNDGKWIAVNP 248
Query: 244 IPDAFVINIGDQLQALSNGLYKSVWHRAIVNAEKPRLSVASFLCPDNEALICPAKPLTED 303
P A VINIGDQLQALSNG Y+SVWHRA+VN+++ R+SVASFLCP N + PAK L D
Sbjct: 249 QPGALVINIGDQLQALSNGKYRSVWHRAVVNSDRERMSVASFLCPCNSVELGPAKKLITD 308
Query: 304 GSGAVYRGFTYPEYYSKFWSRDLEKEHCLEFFK 336
S AVYR +TY EYY KFWSR+L++EHCLE F+
Sbjct: 309 DSPAVYRNYTYDEYYKKFWSRNLDQEHCLELFR 341
>emb|CAD41169.2| OSJNBa0064M23.14 [Oryza sativa (japonica cultivar-group)]
gi|50928227|ref|XP_473641.1| OSJNBa0064M23.14 [Oryza
sativa (japonica cultivar-group)]
Length = 340
Score = 463 bits (1191), Expect = e-129
Identities = 214/335 (63%), Positives = 266/335 (78%), Gaps = 1/335 (0%)
Query: 3 TKVLSSGIHYSKLPESYIRPESDRPCLSQVSEFENVPIIDLGSHNRTQIVQQIGEACSSY 62
T++LS+ H LPE Y RPESDRP L++V+ N+P+IDL S ++ +++ +I +AC +Y
Sbjct: 4 TQLLSTVEHRETLPEGYARPESDRPRLAEVATDSNIPLIDLASPDKPRVIAEIAQACRTY 63
Query: 63 GFFQVVNHGVPLEELKKTAEVAYDFFKLPVEEKMKLYSDDPTKTMRLSTSFNVNKEEVHN 122
GFFQV NHG+ E L+K VA +FF+LP EEK KLYSD+P+K +RLSTSFNV KE VHN
Sbjct: 64 GFFQVTNHGIAEELLEKVMAVALEFFRLPPEEKEKLYSDEPSKKIRLSTSFNVRKETVHN 123
Query: 123 WRDYLRLHCYPLDNYVPEWPSNPPSFKETVANYCKEVRELGLRIEEYISESLGLEKDYLR 182
WRDYLRLHC+PL+ +VPEWPSNP FKE ++ YC+EVR+LGLR+ IS SLGLE+DY+
Sbjct: 124 WRDYLRLHCHPLEEFVPEWPSNPAQFKEIMSTYCREVRQLGLRLLGAISVSLGLEEDYIE 183
Query: 183 NALGEQGQHMAVNYYPPCPQPELTYGLPGHTDPNALTILLQDLHVAGLQVLKDG-KWLAI 241
LGEQ QHMAVNYYP CP+P+LTYGLP HTDPNALTILL D HVAGLQVL+DG +W+ +
Sbjct: 184 KVLGEQEQHMAVNYYPRCPEPDLTYGLPKHTDPNALTILLPDPHVAGLQVLRDGDQWIVV 243
Query: 242 NPIPDAFVINIGDQLQALSNGLYKSVWHRAIVNAEKPRLSVASFLCPDNEALICPAKPLT 301
NP P+A V+N+GDQ+QALSN YKSVWHRA+VN + R+SVASF+CP N A+I PA+ L
Sbjct: 244 NPRPNALVVNLGDQIQALSNDAYKSVWHRAVVNPVQERMSVASFMCPCNSAVISPARKLV 303
Query: 302 EDGSGAVYRGFTYPEYYSKFWSRDLEKEHCLEFFK 336
DG VYR FTY EYY KFWSR+L++EHCLE FK
Sbjct: 304 ADGDAPVYRSFTYDEYYKKFWSRNLDQEHCLELFK 338
>emb|CAB78172.1| putative flavanone 3-beta-hydroxylase [Arabidopsis thaliana]
gi|4539409|emb|CAB40042.1| putative flavanone
3-beta-hydroxylase [Arabidopsis thaliana]
gi|28827438|gb|AAO50563.1| putative flavanone
3-beta-hydroxylase [Arabidopsis thaliana]
gi|28393112|gb|AAO41989.1| putative flavanone
3-beta-hydroxylase [Arabidopsis thaliana]
gi|4115913|gb|AAD03424.1| contains similarity to
Iron/Ascorbate family of oxidoreductases (Pfam: PF00671,
Score=307.1, E=2.2e-88, N=1) [Arabidopsis thaliana]
gi|15235124|ref|NP_192787.1| oxidoreductase, 2OG-Fe(II)
oxygenase family protein [Arabidopsis thaliana]
gi|7433238|pir||T04184 hypothetical protein F7L13.70 -
Arabidopsis thaliana
Length = 348
Score = 380 bits (977), Expect = e-104
Identities = 179/328 (54%), Positives = 237/328 (71%), Gaps = 7/328 (2%)
Query: 15 LPESYIRPESDRPCLSQV-SEFENVPIIDLGS---HNRTQIVQQIGEACSSYGFFQVVNH 70
+P +Y+RP SDRP +S+V + +++P+IDL NR I+ Q ACSS GFFQ+ NH
Sbjct: 18 VPSNYVRPVSDRPKMSEVQTSGDSIPLIDLHDLHGPNRADIINQFAHACSSCGFFQIKNH 77
Query: 71 GVPLEELKKTAEVAYDFFKLPVEEKMKLYSDDPTKTMRLSTSFNVNKEEVHNWRDYLRLH 130
GVP E +KK A +FF+ E++K YS D KT RLSTSFNV+KE+V NWRD+LRLH
Sbjct: 78 GVPEETIKKMMNAAREFFRQSESERVKHYSADTKKTTRLSTSFNVSKEKVSNWRDFLRLH 137
Query: 131 CYPLDNYVPEWPSNPPSFKETVANYCKEVRELGLRIEEYISESLGLEKDYLRNALGEQGQ 190
CYP+++++ EWPS P SF+E A Y VR L L + E ISESLGL KD + N +G+ GQ
Sbjct: 138 CYPIEDFINEWPSTPISFREVTAEYATSVRALVLTLLEAISESLGLAKDRVSNTIGKHGQ 197
Query: 191 HMAVNYYPPCPQPELTYGLPGHTDPNALTILLQDLHVAGLQVLKDGKWLAINPIPDAFVI 250
HMA+NYYP CPQPELTYGLPGH D N +T+LLQD V+GLQV KDGKW+A+NP+P+ F++
Sbjct: 198 HMAINYYPRCPQPELTYGLPGHKDANLITVLLQD-EVSGLQVFKDGKWIAVNPVPNTFIV 256
Query: 251 NIGDQLQALSNGLYKSVWHRAIVNAEKPRLSVASFLCPDNEALICPAKPL--TEDGSGAV 308
N+GDQ+Q +SN YKSV HRA+VN++ R+S+ +F CP +A+I PA+ L E+ S A+
Sbjct: 257 NLGDQMQVISNEKYKSVLHRAVVNSDMERISIPTFYCPSEDAVISPAQELINEEEDSPAI 316
Query: 309 YRGFTYPEYYSKFWSRDLEKEHCLEFFK 336
YR FTY EY+ KFW + E C++ FK
Sbjct: 317 YRNFTYAEYFEKFWDTAFDTESCIDSFK 344
>emb|CAD41170.2| OSJNBa0064M23.15 [Oryza sativa (japonica cultivar-group)]
gi|50928229|ref|XP_473642.1| OSJNBa0064M23.15 [Oryza
sativa (japonica cultivar-group)]
Length = 352
Score = 380 bits (975), Expect = e-104
Identities = 182/331 (54%), Positives = 237/331 (70%), Gaps = 7/331 (2%)
Query: 14 KLPESYIRPESDRPCLSQVSEFEN--VPIIDL---GSHNRTQIVQQIGEACSSYGFFQVV 68
++P S+IRP DRP L V +P+IDL +R ++V+ IG AC + GFF V
Sbjct: 19 QVPSSHIRPVGDRPDLDNVDHESGAGIPVIDLKQLDGPDRRKVVEAIGSACETDGFFMVK 78
Query: 69 NHGVPLEELKKTAEVAYDFFKLPVEEKMKLYSDDPTKTMRLSTSFNVNKEEVHNWRDYLR 128
NHG+P E ++ VA +FF +P E++K YSDDP K +RLSTSFNV E+V NWRD+LR
Sbjct: 79 NHGIPEEVVEGMLRVAREFFHMPESERLKCYSDDPKKAIRLSTSFNVRTEKVSNWRDFLR 138
Query: 129 LHCYPLDNYVPEWPSNPPSFKETVANYCKEVRELGLRIEEYISESLGLEKDYLRNALGEQ 188
LHCYPL++++ +WPSNPPSF++ V Y +E R L LR+ E ISESLGLE+ ++ +A+G Q
Sbjct: 139 LHCYPLESFIDQWPSNPPSFRQVVGTYSREARALALRLLEAISESLGLERGHMVSAMGRQ 198
Query: 189 GQHMAVNYYPPCPQPELTYGLPGHTDPNALTILLQDLHVAGLQVLKDGKWLAINPIPDAF 248
QHMAVNYYPPCPQPELTYGLPGH DPNA+T+LLQD V+GLQV ++G+W+A+NP+PDA
Sbjct: 199 AQHMAVNYYPPCPQPELTYGLPGHKDPNAITLLLQD-GVSGLQVQRNGRWVAVNPVPDAL 257
Query: 249 VINIGDQLQALSNGLYKSVWHRAIVNAEKPRLSVASFLCPDNEALICPAKPLTEDG-SGA 307
VINIGDQ+QALSN YKSV HR IVN+E R+SV +F CP +A+I PA L +
Sbjct: 258 VINIGDQIQALSNDRYKSVLHRVIVNSESERISVPTFYCPSPDAVIAPAGALVDGALHPL 317
Query: 308 VYRGFTYPEYYSKFWSRDLEKEHCLEFFKNN 338
YR F Y YY +FW+ L+ CL+ F+ N
Sbjct: 318 AYRPFKYQAYYDEFWNMGLQSASCLDRFRPN 348
>gb|AAQ65160.1| At4g10500 [Arabidopsis thaliana] gi|7267747|emb|CAB78173.1|
putative Fe(II)/ascorbate oxidase [Arabidopsis thaliana]
gi|4539410|emb|CAB40043.1| putative Fe(II)/ascorbate
oxidase [Arabidopsis thaliana] gi|4115914|gb|AAD03425.1|
contains similarity to Iron/Ascorbate family of
oxidoreductases (Pfam: PF00671, Score=297.8, E=1.3e-85,
N=1) [Arabidopsis thaliana] gi|15235126|ref|NP_192788.1|
oxidoreductase, 2OG-Fe(II) oxygenase family protein
[Arabidopsis thaliana] gi|51972019|dbj|BAD44674.1|
putative Fe(II)/ascorbate oxidase [Arabidopsis thaliana]
gi|51971553|dbj|BAD44441.1| putative Fe(II)/ascorbate
oxidase [Arabidopsis thaliana] gi|7433239|pir||T04185
hypothetical protein F7L13.80 - Arabidopsis thaliana
Length = 349
Score = 378 bits (971), Expect = e-103
Identities = 185/335 (55%), Positives = 240/335 (71%), Gaps = 9/335 (2%)
Query: 7 SSGIHYSKLPESYIRPESDRPCLSQV-SEFENVPIIDLGS---HNRTQIVQQIGEACSSY 62
+S +H +P +Y+RP SDRP LS+V S +++P+IDL NR IVQQ+ ACS+Y
Sbjct: 15 ASSVH---IPSNYVRPISDRPNLSEVESSGDSIPLIDLRDLHGPNRAVIVQQLASACSTY 71
Query: 63 GFFQVVNHGVPLEELKKTAEVAYDFFKLPVEEKMKLYSDDPTKTMRLSTSFNVNKEEVHN 122
GFFQ+ NHGVP + K VA +FF P E++K YS DPTKT RLSTSFNV ++V N
Sbjct: 72 GFFQIKNHGVPDTTVNKMQTVAREFFHQPESERVKHYSADPTKTTRLSTSFNVGADKVLN 131
Query: 123 WRDYLRLHCYPLDNYVPEWPSNPPSFKETVANYCKEVRELGLRIEEYISESLGLEKDYLR 182
WRD+LRLHC+P+++++ EWPS+P SF+E A Y VR L LR+ E ISESLGLE D++
Sbjct: 132 WRDFLRLHCFPIEDFIEEWPSSPISFREVTAEYATSVRALVLRLLEAISESLGLESDHIS 191
Query: 183 NALGEQGQHMAVNYYPPCPQPELTYGLPGHTDPNALTILLQDLHVAGLQVLKDGKWLAIN 242
N LG+ QHMA NYYPPCP+PELTYGLPGH DP +T+LLQD V+GLQV KD KW+A++
Sbjct: 192 NILGKHAQHMAFNYYPPCPEPELTYGLPGHKDPTVITVLLQD-QVSGLQVFKDDKWVAVS 250
Query: 243 PIPDAFVINIGDQLQALSNGLYKSVWHRAIVNAEKPRLSVASFLCPDNEALICPAKPL-T 301
PIP+ F++NIGDQ+Q +SN YKSV HRA+VN E RLS+ +F P +A+I PA L
Sbjct: 251 PIPNTFIVNIGDQMQVISNDKYKSVLHRAVVNTENERLSIPTFYFPSTDAVIGPAHELVN 310
Query: 302 EDGSGAVYRGFTYPEYYSKFWSRDLEKEHCLEFFK 336
E S A+YR + + EY+ KFW+R L CL+ FK
Sbjct: 311 EQDSLAIYRTYPFVEYWDKFWNRSLATASCLDAFK 345
>ref|XP_468578.1| Putative flavanone 3-hydroxylase [Oryza sativa (japonica
cultivar-group)] gi|25446682|gb|AAN74829.1| Putative
flavanone 3-hydroxylase [Oryza sativa (japonica
cultivar-group)]
Length = 250
Score = 290 bits (743), Expect = 3e-77
Identities = 134/188 (71%), Positives = 159/188 (84%)
Query: 149 KETVANYCKEVRELGLRIEEYISESLGLEKDYLRNALGEQGQHMAVNYYPPCPQPELTYG 208
KE ++ YCKEVRELG R+ ISESLGLE+DY++ LGEQ QHMAVN+YP CP+PELT+G
Sbjct: 56 KEIISTYCKEVRELGFRLYGAISESLGLEQDYIKKVLGEQEQHMAVNFYPKCPEPELTFG 115
Query: 209 LPGHTDPNALTILLQDLHVAGLQVLKDGKWLAINPIPDAFVINIGDQLQALSNGLYKSVW 268
LP HTDPNALTILL D VAGLQVLK+G+W+A+NP P+A VINIGDQLQALSNG YKSVW
Sbjct: 116 LPAHTDPNALTILLMDQQVAGLQVLKEGRWIAVNPQPNALVINIGDQLQALSNGRYKSVW 175
Query: 269 HRAIVNAEKPRLSVASFLCPDNEALICPAKPLTEDGSGAVYRGFTYPEYYSKFWSRDLEK 328
HRA+VN++K R+SVASFLCP N+ LI PA+ L DGS AVYR +TY EYY KFWSR+L++
Sbjct: 176 HRAVVNSDKARMSVASFLCPCNDVLIGPAQKLITDGSPAVYRNYTYDEYYKKFWSRNLDQ 235
Query: 329 EHCLEFFK 336
EHCLE F+
Sbjct: 236 EHCLELFR 243
>ref|NP_181207.2| oxidoreductase, 2OG-Fe(II) oxygenase family protein [Arabidopsis
thaliana]
Length = 366
Score = 254 bits (648), Expect = 3e-66
Identities = 135/331 (40%), Positives = 199/331 (59%), Gaps = 11/331 (3%)
Query: 13 SKLPESYIRPESDRPCLSQVSEFEN------VPIIDLGS---HNRTQIVQQIGEACSSYG 63
+K+P YI PE DRP L++ + +P+ID NR +++ I EAC +YG
Sbjct: 30 TKVPTKYIWPEPDRPILTKSDKLIKPNKNLKLPLIDFAELLGPNRPHVLRTIAEACKTYG 89
Query: 64 FFQVVNHGVPLEELKKTAEVAYDFFKLPVEEKMKLYSDDPTKTMRLSTSFNVNKEEVHNW 123
FFQVVNHG+ + K +V FF+LP EE+ K S D + +R TSFN K+ V W
Sbjct: 90 FFQVVNHGMEGDVSKNMIDVCKRFFELPYEERSKYMSSDMSAPVRYGTSFNQIKDNVFCW 149
Query: 124 RDYLRLHCYPLDNYVPEWPSNPPSFKETVANYCKEVRELGLRIEEYISESLGLE-KDYLR 182
RD+L+L+ +PL +Y+P WPS+P F+ + A Y KE +E+ + + I ESL ++ D
Sbjct: 150 RDFLKLYAHPLPDYLPHWPSSPSDFRSSAATYAKETKEMFEMMVKAILESLEIDGSDEAA 209
Query: 183 NALGEQGQHMAVNYYPPCPQPELTYGLPGHTDPNALTILLQDLHVAGLQVLKDGKWLAIN 242
L E Q + VN YPPCP+PELT G+P H+D LT+LLQD V GLQ+L +W+ ++
Sbjct: 210 KELEEGSQVVVVNCYPPCPEPELTLGMPPHSDYGFLTLLLQD-EVEGLQILYRDEWVTVD 268
Query: 243 PIPDAFVINIGDQLQALSNGLYKSVWHRAIVNAEKPRLSVASFLCPDNEALICPAKPLTE 302
PIP +FV+N+GD L+ SNG YKSV HR +VN+ KPR+SVAS +++ P+ L +
Sbjct: 269 PIPGSFVVNVGDHLEIFSNGRYKSVLHRVLVNSTKPRISVASLHSFPLTSVVKPSPKLVD 328
Query: 303 DGSGAVYRGFTYPEYYSKFWSRDLEKEHCLE 333
+ + Y + + SR+ + ++ LE
Sbjct: 329 KHNPSQYMDTDFTTFLQYITSREPKWKNFLE 359
>gb|AAM14878.1| putative flavonol synthase [Arabidopsis thaliana]
gi|7433240|pir||T01606 probable flavonol synthase
[imported] - Arabidopsis thaliana
Length = 352
Score = 241 bits (615), Expect = 2e-62
Identities = 129/332 (38%), Positives = 192/332 (56%), Gaps = 9/332 (2%)
Query: 14 KLPESYIRPESDRPCL--SQVSEFENVPIIDLGSHN----RTQIVQQIGEACSSYGFFQV 67
++P+ Y+ P S RP L S + +P+IDL + R+ + +I AC +GFFQV
Sbjct: 21 QVPDRYVLPPSQRPALGSSLGTSETTLPVIDLSLLHQPFLRSLAIHEISMACKEFGFFQV 80
Query: 68 VNHGVPLEELKKTAEVAYDFFKLPVEEKMKLYSDDPTKTMRLSTSFNVNKEEVHNWRDYL 127
+NHG+P + + A FF LPVEEKM L S + + +R TS N + + VH WRD++
Sbjct: 81 INHGIPSSVVNDALDAATQFFDLPVEEKMLLVSANVHEPVRYGTSLNHSTDRVHYWRDFI 140
Query: 128 RLHCYPLDNYVPEWPSNPPSFKETVANYCKEVRELGLRIEEYISESLGLEKDYLRNALGE 187
+ + +PL ++ WPSNPP +K+ V Y + L ++ E ISESLGLEK+YL+ + E
Sbjct: 141 KHYSHPLSKWIDMWPSNPPCYKDKVGKYAEATHLLHKQLIEAISESLGLEKNYLQEEIEE 200
Query: 188 QGQHMAVNYYPPCPQPELTYGLPGHTDPNALTILLQDLHVAGLQVLKDGK-WLAINPIPD 246
Q MAVN YP CP+PE+ G+P H+D ++LTILLQ GLQ++ K W+ + I
Sbjct: 201 GSQVMAVNCYPACPEPEMALGMPPHSDFSSLTILLQS--SKGLQIMDCNKNWVCVPYIEG 258
Query: 247 AFVINIGDQLQALSNGLYKSVWHRAIVNAEKPRLSVASFLCPDNEALICPAKPLTEDGSG 306
A ++ +GDQ++ +SNG+YKSV HR VN E RLS AS I PA L +
Sbjct: 259 ALIVQLGDQVEVMSNGIYKSVIHRVTVNKEVKRLSFASLHSLPLHKKISPAPKLVNPNNA 318
Query: 307 AVYRGFTYPEYYSKFWSRDLEKEHCLEFFKNN 338
Y F++ ++ + S D +E ++ K +
Sbjct: 319 PAYGEFSFNDFLNYISSNDFIQERFIDTIKKS 350
>ref|NP_182007.2| oxidoreductase, 2OG-Fe(II) oxygenase family protein [Arabidopsis
thaliana]
Length = 357
Score = 241 bits (615), Expect = 2e-62
Identities = 129/332 (38%), Positives = 192/332 (56%), Gaps = 9/332 (2%)
Query: 14 KLPESYIRPESDRPCL--SQVSEFENVPIIDLGSHN----RTQIVQQIGEACSSYGFFQV 67
++P+ Y+ P S RP L S + +P+IDL + R+ + +I AC +GFFQV
Sbjct: 26 QVPDRYVLPPSQRPALGSSLGTSETTLPVIDLSLLHQPFLRSLAIHEISMACKEFGFFQV 85
Query: 68 VNHGVPLEELKKTAEVAYDFFKLPVEEKMKLYSDDPTKTMRLSTSFNVNKEEVHNWRDYL 127
+NHG+P + + A FF LPVEEKM L S + + +R TS N + + VH WRD++
Sbjct: 86 INHGIPSSVVNDALDAATQFFDLPVEEKMLLVSANVHEPVRYGTSLNHSTDRVHYWRDFI 145
Query: 128 RLHCYPLDNYVPEWPSNPPSFKETVANYCKEVRELGLRIEEYISESLGLEKDYLRNALGE 187
+ + +PL ++ WPSNPP +K+ V Y + L ++ E ISESLGLEK+YL+ + E
Sbjct: 146 KHYSHPLSKWIDMWPSNPPCYKDKVGKYAEATHLLHKQLIEAISESLGLEKNYLQEEIEE 205
Query: 188 QGQHMAVNYYPPCPQPELTYGLPGHTDPNALTILLQDLHVAGLQVLKDGK-WLAINPIPD 246
Q MAVN YP CP+PE+ G+P H+D ++LTILLQ GLQ++ K W+ + I
Sbjct: 206 GSQVMAVNCYPACPEPEMALGMPPHSDFSSLTILLQS--SKGLQIMDCNKNWVCVPYIEG 263
Query: 247 AFVINIGDQLQALSNGLYKSVWHRAIVNAEKPRLSVASFLCPDNEALICPAKPLTEDGSG 306
A ++ +GDQ++ +SNG+YKSV HR VN E RLS AS I PA L +
Sbjct: 264 ALIVQLGDQVEVMSNGIYKSVIHRVTVNKEVKRLSFASLHSLPLHKKISPAPKLVNPNNA 323
Query: 307 AVYRGFTYPEYYSKFWSRDLEKEHCLEFFKNN 338
Y F++ ++ + S D +E ++ K +
Sbjct: 324 PAYGEFSFNDFLNYISSNDFIQERFIDTIKKS 355
>gb|AAM48133.1| putative flavanone 3-hydroxylase [Saussurea medusa]
gi|48431269|gb|AAT44124.1| F3H-like protein [Saussurea
medusa]
Length = 343
Score = 240 bits (612), Expect = 5e-62
Identities = 125/322 (38%), Positives = 191/322 (58%), Gaps = 14/322 (4%)
Query: 7 SSGIHYSKLPESYIRPESDRPCLSQVSEFENVPIIDLGSH---NRTQIVQQIGEACSSYG 63
S+G+ +P+ Y+ P RP VS +P+IDL + +R+ IVQQI +AC +G
Sbjct: 9 SNGVQ--SVPKDYVMPPERRPG-DFVSACNEIPVIDLQENPKIDRSDIVQQILKACQEFG 65
Query: 64 FFQVVNHGVPLEELKKTAEVAYDFFKLPVEEKMKLYSD--DPTKTMRLSTSFNVNKEEVH 121
FQV+NHGV + ++ + ++FF +PV++K++ Y + T ++ N KE+VH
Sbjct: 66 LFQVINHGVSEKMMEDMRVLYHEFFNMPVDDKLRGYFEPWSTTGCSLYTSGVNYAKEDVH 125
Query: 122 NWRDYLRLHCYPLDNYVPEWPSNPPSFKETVANYCKEVRELGLRIEEYISESLGLEKDYL 181
W+D LR C+PL+ + P WP P ++E V Y EVR++G +I + I E LGL + Y
Sbjct: 126 YWKDTLRHPCHPLEEHTPSWPEKPARYREEVGRYAVEVRKMGFKILDLIGEGLGLNEGYF 185
Query: 182 RNALGEQGQHMAVNYYPPCPQPELTYGLPGHTDPNALTILLQDLHVAGLQVLKDGKWLAI 241
Q Q MA+N+YP CP P L G+ GHTDPN +T L QD + GLQ+ KDGKW+ I
Sbjct: 186 NGV--NQYQTMAINHYPLCPDPGLAMGIDGHTDPNLITFLQQDQY--GLQMCKDGKWMGI 241
Query: 242 NPIPDAFVINIGDQLQALSNGLYKSVWHRAIVNAEKPRLSVASFLCPDN--EALICPAKP 299
+PIP+AFV+NIG QL+ +SNG KS HR + ++ R S+ +F P+ ++ PAK
Sbjct: 242 DPIPNAFVVNIGYQLEIISNGKLKSAEHRVVTSSTTARTSIVTFFGPNAGLPIVVEPAKE 301
Query: 300 LTEDGSGAVYRGFTYPEYYSKF 321
L S +++ + Y + + +
Sbjct: 302 LVTSTSPKMFKSYQYNNFIADY 323
>gb|AAD20145.1| putative giberellin beta-hydroxylase [Arabidopsis thaliana]
gi|25285698|pir||E84783 probable giberellin
beta-hydroxylase [imported] - Arabidopsis thaliana
Length = 392
Score = 239 bits (611), Expect = 7e-62
Identities = 135/357 (37%), Positives = 199/357 (54%), Gaps = 37/357 (10%)
Query: 13 SKLPESYIRPESDRPCLSQVSEFEN------VPIIDLGS---HNRTQIVQQIGEACSSYG 63
+K+P YI PE DRP L++ + +P+ID NR +++ I EAC +YG
Sbjct: 30 TKVPTKYIWPEPDRPILTKSDKLIKPNKNLKLPLIDFAELLGPNRPHVLRTIAEACKTYG 89
Query: 64 FFQV--------------------------VNHGVPLEELKKTAEVAYDFFKLPVEEKMK 97
FFQV VNHG+ + K +V FF+LP EE+ K
Sbjct: 90 FFQVGSLNPKAHAVISRFLAKQVSYGNGQVVNHGMEGDVSKNMIDVCKRFFELPYEERSK 149
Query: 98 LYSDDPTKTMRLSTSFNVNKEEVHNWRDYLRLHCYPLDNYVPEWPSNPPSFKETVANYCK 157
S D + +R TSFN K+ V WRD+L+L+ +PL +Y+P WPS+P F+ + A Y K
Sbjct: 150 YMSSDMSAPVRYGTSFNQIKDNVFCWRDFLKLYAHPLPDYLPHWPSSPSDFRSSAATYAK 209
Query: 158 EVRELGLRIEEYISESLGLE-KDYLRNALGEQGQHMAVNYYPPCPQPELTYGLPGHTDPN 216
E +E+ + + I ESL ++ D L E Q + VN YPPCP+PELT G+P H+D
Sbjct: 210 ETKEMFEMMVKAILESLEIDGSDEAAKELEEGSQVVVVNCYPPCPEPELTLGMPPHSDYG 269
Query: 217 ALTILLQDLHVAGLQVLKDGKWLAINPIPDAFVINIGDQLQALSNGLYKSVWHRAIVNAE 276
LT+LLQD V GLQ+L +W+ ++PIP +FV+N+GD L+ SNG YKSV HR +VN+
Sbjct: 270 FLTLLLQD-EVEGLQILYRDEWVTVDPIPGSFVVNVGDHLEIFSNGRYKSVLHRVLVNST 328
Query: 277 KPRLSVASFLCPDNEALICPAKPLTEDGSGAVYRGFTYPEYYSKFWSRDLEKEHCLE 333
KPR+SVAS +++ P+ L + + + Y + + SR+ + ++ LE
Sbjct: 329 KPRISVASLHSFPLTSVVKPSPKLVDKHNPSQYMDTDFTTFLQYITSREPKWKNFLE 385
>ref|XP_475566.1| putative leucoanthocyanidin dioxygenase (EC 1.14.11.-) [Oryza
sativa (japonica cultivar-group)]
gi|46391159|gb|AAS90686.1| putative leucoanthocyanidin
dioxygenase [Oryza sativa (japonica cultivar-group)]
Length = 368
Score = 233 bits (594), Expect = 6e-60
Identities = 122/310 (39%), Positives = 179/310 (57%), Gaps = 9/310 (2%)
Query: 15 LPESYIRPESDRPCLSQVSEFE---NVPIIDLGSHNRTQIVQQIGEACSSYGFFQVVNHG 71
+P+ Y+RPE++RP S + N+P++D+ S T + EAC +GFFQ VNHG
Sbjct: 28 IPDRYVRPETERPSSSSEANAVANINIPVVDMSSSPGTAAAA-VAEACREWGFFQAVNHG 86
Query: 72 VPLEELKKTAEVAYDFFKLPVEEKMKLYSDDPTKTMRLSTSFNVNKEEVHNWRDYLRLHC 131
VP L++ V FF+ P+E K + Y + P + V+K + +W DY LH
Sbjct: 87 VPAALLRRARGVWRGFFQQPMEVKQR-YGNSPATYEGYGSRLGVDKGAILDWGDYYFLHV 145
Query: 132 YPLDNYVP-EWPSNPPSFKETVANYCKEVRELGLRIEEYISESLGLEKDYLRNALGEQ-- 188
P P +WP PP +ET Y +EVR L R+ ++ LG+E+ L+ A G +
Sbjct: 146 RPPHLLSPHKWPHLPPDLRETTTEYSEEVRRLCERLMAVMAVGLGVEEGRLQEAFGGREG 205
Query: 189 -GQHMAVNYYPPCPQPELTYGLPGHTDPNALTILLQDLHVAGLQVLKDGKWLAINPIPDA 247
G + VNYYP CPQP+LT GL H+DP +T+LL D V GLQV G W+ ++P+PDA
Sbjct: 206 AGVCVRVNYYPRCPQPDLTLGLSSHSDPGGMTVLLVDDRVKGLQVRHAGAWVTVDPVPDA 265
Query: 248 FVINIGDQLQALSNGLYKSVWHRAIVNAEKPRLSVASFLCPDNEALICPAKPLTEDGSGA 307
F+IN+GDQ+Q ++N LY+SV HR +VNA + RLS+A+F P ++ + P L
Sbjct: 266 FIINVGDQIQVVTNALYRSVEHRVVVNAAEERLSIATFYNPRSDLPVAPLPELVSPERPP 325
Query: 308 VYRGFTYPEY 317
+Y T+ +Y
Sbjct: 326 LYSPMTFDDY 335
>dbj|BAD73770.1| putative anthocyanidin synthase [Oryza sativa (japonica
cultivar-group)]
Length = 366
Score = 233 bits (594), Expect = 6e-60
Identities = 128/314 (40%), Positives = 181/314 (56%), Gaps = 12/314 (3%)
Query: 15 LPESYIRPESDRPC--LSQVSEFENVPIIDLGSHNRT---QIVQQIGEACSSYGFFQVVN 69
+P Y++P DRP S ++P++DLG+ Q+ + + AC +GFFQVVN
Sbjct: 26 IPRCYVKPPCDRPAPEADDASSGASIPVVDLGNGGDDEGGQLAEAVAAACRGWGFFQVVN 85
Query: 70 HGVPLEELKKTAEVAYDFFKLPVEEKMKLYSDDPTKTMRLSTSFNVNKEEVHNWRDYLRL 129
HGV E ++ E + FF+LP++EK K Y++ P + V K + +W DY L
Sbjct: 86 HGVRPELMRAAREAWHGFFRLPLQEKQK-YANSPRTYEGYGSRLGVEKGAILDWGDYYFL 144
Query: 130 HCYPLDNYVPE--WPSNPPSFKETVANYCKEVRELGLRIEEYISESLGLEKDYLRNALG- 186
P P WP+NP KE Y +EV +L R+ +S SLGL++ + A G
Sbjct: 145 VLSPDAAKSPAKYWPANPGICKEVSEEYGREVIKLCERLMRLLSASLGLDETRFQEAFGG 204
Query: 187 -EQGQHMAVNYYPPCPQPELTYGLPGHTDPNALTILLQDLHVAGLQVLK-DGKWLAINPI 244
+ G + NYYP CPQP+LT GL H+DP LT+LL D HV GLQV + DG W+ + P+
Sbjct: 205 ADCGAGLRANYYPRCPQPDLTLGLSAHSDPGILTVLLADDHVRGLQVRRRDGHWVTVQPL 264
Query: 245 PDAFVINIGDQLQALSNGLYKSVWHRAIVNAEKPRLSVASFLCPDNEALICPAKPL-TED 303
PDAF++N+GDQ++ LSN +YKSV HR IVNAE+ R+S+A F P + + PA L T +
Sbjct: 265 PDAFIVNVGDQIEILSNSMYKSVEHRVIVNAEEERISLALFYNPRGDVPVAPAPELVTPE 324
Query: 304 GSGAVYRGFTYPEY 317
YR T+ EY
Sbjct: 325 RPSLYYRPMTFDEY 338
>gb|AAM61362.1| putative ethylene-forming enzyme [Arabidopsis thaliana]
gi|9294689|dbj|BAB03055.1| unnamed protein product
[Arabidopsis thaliana] gi|29029088|gb|AAO64923.1|
At3g21420 [Arabidopsis thaliana]
gi|18402992|ref|NP_566685.1| oxidoreductase, 2OG-Fe(II)
oxygenase family protein [Arabidopsis thaliana]
Length = 364
Score = 229 bits (585), Expect = 7e-59
Identities = 126/337 (37%), Positives = 194/337 (57%), Gaps = 14/337 (4%)
Query: 13 SKLPESYIRPESDR----PCLSQVSEFENVPIIDLGSHNRTQI------VQQIGEACSSY 62
+K+PE +IR E +R L +P+IDL ++ + ++ +AC +
Sbjct: 26 NKVPERFIREEYERGVVVSSLKTHHLHHQIPVIDLSKLSKPDNDDFFFEILKLSQACEDW 85
Query: 63 GFFQVVNHGVPLEELKKTAEVAYDFFKLPVEEKMKLYSDDPTKTMRLSTSFNVNKEEVHN 122
GFFQV+NHG+ +E ++ EVA +FF +P+EEK K Y +P +F ++++ +
Sbjct: 86 GFFQVINHGIEVEVVEDIEEVASEFFDMPLEEKKK-YPMEPGTVQGYGQAFIFSEDQKLD 144
Query: 123 WRDYLRLHCYPLDNYVPE-WPSNPPSFKETVANYCKEVRELGLRIEEYISESLGLEKDYL 181
W + L +P P+ WPS P F E++ Y KE+REL R+ +YI+ SLGL+++
Sbjct: 145 WCNMFALGVHPPQIRNPKLWPSKPARFSESLEGYSKEIRELCKRLLKYIAISLGLKEERF 204
Query: 182 RNALGEQGQHMAVNYYPPCPQPELTYGLPGHTDPNALTILLQDLH-VAGLQVLKDGKWLA 240
GE Q + +NYYPPC P+L GL H+D +ALT+L Q + GLQ+LKD W+
Sbjct: 205 EEMFGEAVQAVRMNYYPPCSSPDLVLGLSPHSDGSALTVLQQSKNSCVGLQILKDNTWVP 264
Query: 241 INPIPDAFVINIGDQLQALSNGLYKSVWHRAIVNAEKPRLSVASFLCPDNEALICPAKPL 300
+ P+P+A VINIGD ++ LSNG YKSV HRA+ N EK RL++ +F P+ E I P L
Sbjct: 265 VKPLPNALVINIGDTIEVLSNGKYKSVEHRAVTNREKERLTIVTFYAPNYEVEIEPMSEL 324
Query: 301 TEDGSG-AVYRGFTYPEYYSKFWSRDLEKEHCLEFFK 336
+D + YR + + +Y + S L+ + L+F K
Sbjct: 325 VDDETNPCKYRSYNHGDYSYHYVSNKLQGKKSLDFAK 361
>ref|NP_914944.1| putative ethylene-forming enzyme [Oryza sativa (japonica
cultivar-group)] gi|15408799|dbj|BAB64195.1| putative
ethylene-forming enzyme [Oryza sativa (japonica
cultivar-group)]
Length = 366
Score = 229 bits (583), Expect = 1e-58
Identities = 129/329 (39%), Positives = 190/329 (57%), Gaps = 8/329 (2%)
Query: 15 LPESYIRPESDRPCLSQVSEF---ENVPIIDLGSHNR--TQIVQQIGEACSSYGFFQVVN 69
+PE YIR DRP + V + E +P+ID+G R + + AC +GFFQVVN
Sbjct: 38 VPERYIRDGDDRPDHAVVDDERAQERIPVIDVGELQRGSEDELDNLRLACEQWGFFQVVN 97
Query: 70 HGVPLEELKKTAEVAYDFFKLPVEEKMKLYSDDPTKTMRLSTSFNVNKEEVHNWRDYLRL 129
HGV E +++ + A +FF LP+EEK K Y +P +F + ++ +W + L L
Sbjct: 98 HGVEEETMEEMEKAAREFFMLPLEEKEK-YPMEPGGIQGYGHAFVFSDDQKLDWCNMLAL 156
Query: 130 HCYPLDNYVPE-WPSNPPSFKETVANYCKEVRELGLRIEEYISESLGLEKDYLRNALGEQ 188
P P WP+ P +F +T+ Y E+REL +R+ E+I+ +LGL L GE
Sbjct: 157 GVEPAFIRRPNLWPTTPANFSKTLEKYSVEIRELCVRLLEHIAAALGLAPARLNGMFGEA 216
Query: 189 GQHMAVNYYPPCPQPELTYGLPGHTDPNALTILLQDLHVAGLQVLK-DGKWLAINPIPDA 247
Q + +N+YPPCP+PEL GL H+D +A+T+L QD AGLQVL+ G W+A++P+P A
Sbjct: 217 VQAVRMNFYPPCPRPELVLGLSPHSDGSAVTVLQQDAAFAGLQVLRGGGGWVAVHPVPGA 276
Query: 248 FVINIGDQLQALSNGLYKSVWHRAIVNAEKPRLSVASFLCPDNEALICPAKPLTEDGSGA 307
V+N+GD L+ L+NG YKSV HRA+ + E R+SV +F P + + P L DG
Sbjct: 277 LVVNVGDTLEVLTNGRYKSVEHRAVASGEHDRMSVVTFYAPAYDVELGPLPELVADGEPR 336
Query: 308 VYRGFTYPEYYSKFWSRDLEKEHCLEFFK 336
YR + + EY + + L+ + LEF K
Sbjct: 337 RYRTYNHGEYSRHYVTSRLQGKKTLEFAK 365
>gb|AAF01507.1| putative leucoanthocyanidin dioxygenase [Arabidopsis thaliana]
gi|12321884|gb|AAG50980.1| leucoanthocyanidin
dioxygenase, putative; 41415-43854 [Arabidopsis
thaliana] gi|15229694|ref|NP_187728.1| oxidoreductase,
2OG-Fe(II) oxygenase family protein [Arabidopsis
thaliana]
Length = 400
Score = 226 bits (577), Expect = 6e-58
Identities = 130/315 (41%), Positives = 176/315 (55%), Gaps = 11/315 (3%)
Query: 13 SKLPESYIRPESDRPCLSQVSEFE-----NVPIIDLGS--HNRTQIVQQIGEACSSYGFF 65
+ LP+ YI+P S RP + + N+PIIDL S ++I EAC +GFF
Sbjct: 65 TSLPDRYIKPPSQRPQTTIIDHQPEVADINIPIIDLDSLFSGNEDDKKRISEACREWGFF 124
Query: 66 QVVNHGVPLEELKKTAEVAYDFFKLPVEEKMKLYSDDPTKTMRLSTSFNVNKEEVHNWRD 125
QV+NHGV E + E FF LPVE K ++YS+ P + V K + +W D
Sbjct: 125 QVINHGVKPELMDAARETWKSFFNLPVEAK-EVYSNSPRTYEGYGSRLGVEKGAILDWND 183
Query: 126 YLRLHCYPLD-NYVPEWPSNPPSFKETVANYCKEVRELGLRIEEYISESLGLEKDYLRNA 184
Y LH PL +WPS P + +E Y KE+ +LG R+ +S +LGL + L+ A
Sbjct: 184 YYYLHFLPLALKDFNKWPSLPSNIREMNDEYGKELVKLGGRLMTILSSNLGLRAEQLQEA 243
Query: 185 LGEQ--GQHMAVNYYPPCPQPELTYGLPGHTDPNALTILLQDLHVAGLQVLKDGKWLAIN 242
G + G + VNYYP CPQPEL GL H+DP +TILL D V GLQV W+ +N
Sbjct: 244 FGGEDVGACLRVNYYPKCPQPELALGLSPHSDPGGMTILLPDDQVVGLQVRHGDTWITVN 303
Query: 243 PIPDAFVINIGDQLQALSNGLYKSVWHRAIVNAEKPRLSVASFLCPDNEALICPAKPLTE 302
P+ AF++NIGDQ+Q LSN YKSV HR IVN+EK R+S+A F P ++ I P + L
Sbjct: 304 PLRHAFIVNIGDQIQILSNSKYKSVEHRVIVNSEKERVSLAFFYNPKSDIPIQPMQQLVT 363
Query: 303 DGSGAVYRGFTYPEY 317
+Y T+ +Y
Sbjct: 364 STMPPLYPPMTFDQY 378
>ref|XP_507337.1| PREDICTED P0562A06.31 gene product [Oryza sativa (japonica
cultivar-group)] gi|50948493|ref|XP_483774.1| putative
iron deficiency protein Ids3 [Oryza sativa (japonica
cultivar-group)] gi|45736159|dbj|BAD13205.1| putative
iron deficiency protein Ids3 [Oryza sativa (japonica
cultivar-group)] gi|45736113|dbj|BAD13144.1| putative
iron deficiency protein Ids3 [Oryza sativa (japonica
cultivar-group)]
Length = 383
Score = 224 bits (572), Expect = 2e-57
Identities = 124/305 (40%), Positives = 178/305 (57%), Gaps = 12/305 (3%)
Query: 8 SGIHYSKLPESYIRPESDRP-----CLSQVSEFENVPIIDLGS----HNRTQIVQQIGEA 58
SGI ++LP +Y+ P SDRP + +P++DL R +++ + A
Sbjct: 47 SGI--TRLPGNYVLPASDRPGQAAGAAAAAGGSVKLPVVDLSRLRVPSERGAVLRTLDAA 104
Query: 59 CSSYGFFQVVNHGVPLEELKKTAEVAYDFFKLPVEEKMKLYSDDPTKTMRLSTSFNVNKE 118
C YGFFQVVNHGV E + +VA FF+LP E+ + S D +R TSFN ++
Sbjct: 105 CREYGFFQVVNHGVGGEVVGGMLDVARRFFELPQPERERYMSADVRAPVRYGTSFNQVRD 164
Query: 119 EVHNWRDYLRLHCYPLDNYVPEWPSNPPSFKETVANYCKEVRELGLRIEEYISESLGLEK 178
V WRD+L+L C PL V WP++P +E + Y + + + + + E E+LG+
Sbjct: 165 AVLCWRDFLKLACMPLAAVVESWPTSPADLREVASRYAEANQRVFMEVMEAALEALGVGG 224
Query: 179 DYLRNALGEQGQHMAVNYYPPCPQPELTYGLPGHTDPNALTILLQDLHVAGLQVLKDGKW 238
+ L Q M VN YP CPQPELT G+P H+D LT++LQD VAGLQV+ G+W
Sbjct: 225 GGVMEDLAAGTQMMTVNCYPECPQPELTLGMPPHSDYGFLTLVLQD-EVAGLQVMHAGEW 283
Query: 239 LAINPIPDAFVINIGDQLQALSNGLYKSVWHRAIVNAEKPRLSVASFLCPDNEALICPAK 298
L ++P+P +FV+N+GD L+ LSNG Y+SV HR VN+ + R+SVASF E ++ PA
Sbjct: 284 LTVDPLPGSFVVNVGDHLEILSNGRYRSVLHRVKVNSRRLRVSVASFHSVAPERVVSPAP 343
Query: 299 PLTED 303
L +D
Sbjct: 344 ELIDD 348
>gb|AAM91495.1| AT5g05600/MOP10_14 [Arabidopsis thaliana]
gi|10178137|dbj|BAB11549.1| leucoanthocyanidin
dioxygenase-like protein [Arabidopsis thaliana]
gi|15239179|ref|NP_196179.1| oxidoreductase, 2OG-Fe(II)
oxygenase family protein [Arabidopsis thaliana]
gi|14532538|gb|AAK63997.1| AT5g05600/MOP10_14
[Arabidopsis thaliana]
Length = 371
Score = 224 bits (572), Expect = 2e-57
Identities = 130/316 (41%), Positives = 181/316 (57%), Gaps = 13/316 (4%)
Query: 13 SKLPESYIRPESDRPCLSQ-VSEFENVPIIDLGSHNRTQ------IVQQIGEACSSYGFF 65
S LP+ YI+P S RP ++ N+PIIDL + I+ +I EAC +GFF
Sbjct: 36 SSLPDRYIKPASLRPTTTEDAPTATNIPIIDLEGLFSEEGLSDDVIMARISEACRGWGFF 95
Query: 66 QVVNHGVPLEELKKTAEVAYDFFKLPVEEKMKLYSDDPTKTMRLSTSFNVNKEEVHNWRD 125
QVVNHGV E + E +FF +PV K + YS+ P + V K +W D
Sbjct: 96 QVVNHGVKPELMDAARENWREFFHMPVNAK-ETYSNSPRTYEGYGSRLGVEKGASLDWSD 154
Query: 126 YLRLHCYP--LDNYVPEWPSNPPSFKETVANYCKEVRELGLRIEEYISESLGLEKDYLRN 183
Y LH P L ++ +WPS PP+ +E + Y +E+ +L RI +S +LGL++D +
Sbjct: 155 YYFLHLLPHHLKDF-NKWPSFPPTIREVIDEYGEELVKLSGRIMRVLSTNLGLKEDKFQE 213
Query: 184 ALGEQ--GQHMAVNYYPPCPQPELTYGLPGHTDPNALTILLQDLHVAGLQVLKDGKWLAI 241
A G + G + VNYYP CP+PEL GL H+DP +TILL D V GLQV KD W+ +
Sbjct: 214 AFGGENIGACLRVNYYPKCPRPELALGLSPHSDPGGMTILLPDDQVFGLQVRKDDTWITV 273
Query: 242 NPIPDAFVINIGDQLQALSNGLYKSVWHRAIVNAEKPRLSVASFLCPDNEALICPAKPLT 301
P P AF++NIGDQ+Q LSN YKSV HR IVN++K R+S+A F P ++ I P + L
Sbjct: 274 KPHPHAFIVNIGDQIQILSNSTYKSVEHRVIVNSDKERVSLAFFYNPKSDIPIQPLQELV 333
Query: 302 EDGSGAVYRGFTYPEY 317
+ +Y T+ +Y
Sbjct: 334 STHNPPLYPPMTFDQY 349
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.317 0.137 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 643,178,527
Number of Sequences: 2540612
Number of extensions: 28716335
Number of successful extensions: 68273
Number of sequences better than 10.0: 1117
Number of HSP's better than 10.0 without gapping: 1005
Number of HSP's successfully gapped in prelim test: 112
Number of HSP's that attempted gapping in prelim test: 64820
Number of HSP's gapped (non-prelim): 1352
length of query: 338
length of database: 863,360,394
effective HSP length: 128
effective length of query: 210
effective length of database: 538,162,058
effective search space: 113014032180
effective search space used: 113014032180
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 75 (33.5 bits)
Medicago: description of AC119415.1


