

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149579.7 - phase: 0 /pseudo
(367 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
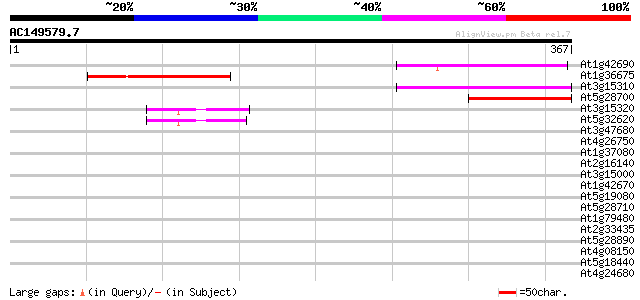
Score E
Sequences producing significant alignments: (bits) Value
At1g42690 unknown protein 86 3e-17
At1g36675 putative protein 84 9e-17
At3g15310 unknown protein 84 2e-16
At5g28700 putative protein 65 8e-11
At3g15320 hypothetical protein 49 6e-06
At5g32620 putative protein 45 5e-05
At3g47680 unknown protein 41 0.001
At4g26750 unknown protein 40 0.003
At1g37080 hypothetical protein 40 0.003
At2g16140 pseudogene; similar to MURA transposase of maize Muta... 39 0.003
At3g15000 unknown protein 39 0.006
At1g42670 unknown protein 39 0.006
At5g19080 unknown protein 37 0.013
At5g28710 putative protein 37 0.022
At1g79480 hypothetical protein 36 0.029
At2g33435 RRM-containing protein 36 0.038
At5g28890 putative protein 35 0.065
At4g08150 KNAT1 homeobox-like protein 35 0.065
At5g18440 putative protein 33 0.25
At4g24680 unknown protein 33 0.25
>At1g42690 unknown protein
Length = 333
Score = 85.9 bits (211), Expect = 3e-17
Identities = 48/119 (40%), Positives = 68/119 (56%), Gaps = 7/119 (5%)
Query: 254 DTYMVNRFIQRRKQIEEGSGSRSRK-------YLKRDHAGANQRLIDDYFANEPTYDDAM 306
D ++F Q Q E G+R R Y++R+ + +L++DYF PTY +
Sbjct: 11 DDMFDDKFDQCFDQALESYGNRQRVKPRKKKLYIERNREEGHIQLVNDYFTENPTYPPHI 70
Query: 307 FRRRYRMQKHVFLRIVGDLSSSDNYFTQRVDAANKEGISPLAKCTTSMQMLPYGVAADA 365
FRRR+RM K +F+RIV S+ YF QR DA + G S L K T +++ML YG+AADA
Sbjct: 71 FRRRFRMNKSLFMRIVERFSNEVPYFKQRRDATRRLGFSALQKSTAAIRMLAYGIAADA 129
>At1g36675 putative protein
Length = 268
Score = 84.3 bits (207), Expect = 9e-17
Identities = 42/93 (45%), Positives = 56/93 (60%), Gaps = 1/93 (1%)
Query: 52 SSSVPNFHPYYGSMMKYPSQTPPFNGSMPMENDNFHNVGATQYPEFSTQITPSGMAIADE 111
SS++ N PY+GS+M SQ P ++ S PM N+ NVGAT++PEFSTQ+ M+ E
Sbjct: 2 SSAITNHRPYHGSIMSPSSQAPSYS-STPMGNETNTNVGATEFPEFSTQMALGVMSGVHE 60
Query: 112 VTPEDSTPKSKRSKEPAWNTQQNLVLISAWIKY 144
P + S+ W T QNLVL+S WIKY
Sbjct: 61 TIPNEEDLTCNHSRSSKWTTDQNLVLLSRWIKY 93
Score = 32.7 bits (73), Expect = 0.32
Identities = 14/19 (73%), Positives = 16/19 (83%)
Query: 324 DLSSSDNYFTQRVDAANKE 342
DLSSSDNYFT+R +A KE
Sbjct: 175 DLSSSDNYFTKRFEATKKE 193
>At3g15310 unknown protein
Length = 415
Score = 83.6 bits (205), Expect = 2e-16
Identities = 45/114 (39%), Positives = 64/114 (55%)
Query: 254 DTYMVNRFIQRRKQIEEGSGSRSRKYLKRDHAGANQRLIDDYFANEPTYDDAMFRRRYRM 313
D N F++ E +R R Y R + L +DYF++ + FRRR+RM
Sbjct: 22 DQQFENVFVELSNSEEVAKKNRKRAYFDRKREEGHVLLWNDYFSDNAIFPLQTFRRRFRM 81
Query: 314 QKHVFLRIVGDLSSSDNYFTQRVDAANKEGISPLAKCTTSMQMLPYGVAADAVD 367
+K +FLRIV LSS +F R DA + G SP+ KCT ++++L YG A+DAVD
Sbjct: 82 KKPLFLRIVDRLSSELMFFQHRRDATGRFGHSPIQKCTAAIRLLAYGYASDAVD 135
>At5g28700 putative protein
Length = 292
Score = 64.7 bits (156), Expect = 8e-11
Identities = 33/67 (49%), Positives = 43/67 (63%)
Query: 301 TYDDAMFRRRYRMQKHVFLRIVGDLSSSDNYFTQRVDAANKEGISPLAKCTTSMQMLPYG 360
TY +FR R+RM K +F+RIV S+ YF QR DA + S L K T +++ML YG
Sbjct: 33 TYPPHIFRHRFRMNKPLFMRIVERFSNEVPYFKQRRDATGRLDFSALQKSTAAIRMLAYG 92
Query: 361 VAADAVD 367
+AADAVD
Sbjct: 93 IAADAVD 99
>At3g15320 hypothetical protein
Length = 287
Score = 48.5 bits (114), Expect = 6e-06
Identities = 23/71 (32%), Positives = 41/71 (57%), Gaps = 9/71 (12%)
Query: 90 GATQYPEFSTQITPSGMAIA---DEVTPEDSTPKSKRSKEPAWNTQQNLVLISAWIKYGT 146
G+T+ P FS+Q++ G + DEV P+ S + K W ++++VL+SAW+
Sbjct: 7 GSTEVPAFSSQLSEEGSDLEGSEDEVKPKQSISRKK------WTAKEDIVLVSAWLNTSK 60
Query: 147 SSVVGRNQRGE 157
V+G +Q+G+
Sbjct: 61 DPVIGNDQQGQ 71
>At5g32620 putative protein
Length = 301
Score = 45.4 bits (106), Expect = 5e-05
Identities = 22/69 (31%), Positives = 39/69 (55%), Gaps = 9/69 (13%)
Query: 90 GATQYPEFSTQITPSGMAIA---DEVTPEDSTPKSKRSKEPAWNTQQNLVLISAWIKYGT 146
G+T+ P FS+Q++ G + DEV P+ S + K W ++++VL+SAW+
Sbjct: 30 GSTEVPAFSSQLSEEGSELEGSEDEVKPKQSISRKK------WTAKEDIVLVSAWLNPSK 83
Query: 147 SSVVGRNQR 155
V+G +Q+
Sbjct: 84 DPVIGNDQQ 92
>At3g47680 unknown protein
Length = 302
Score = 41.2 bits (95), Expect = 0.001
Identities = 19/49 (38%), Positives = 31/49 (62%), Gaps = 3/49 (6%)
Query: 110 DEVTPEDS--TPKSKRSKEPAWNTQQNLVLISAWIKYGTSSVVGRNQRG 156
D+V+P + TPK KR + W+ ++LVL+SAW+ +V+G Q+G
Sbjct: 30 DDVSPNQAAHTPKVKRERRK-WSAGEDLVLVSAWLNTSKDAVIGNEQKG 77
>At4g26750 unknown protein
Length = 421
Score = 39.7 bits (91), Expect = 0.003
Identities = 19/59 (32%), Positives = 35/59 (59%), Gaps = 3/59 (5%)
Query: 13 YSNYPFNYENPNNYPNPNQFFNQR--PQNIPNFGFPPNFNQSSSVPNFHPYYGSMMKYP 69
YS+ P++ + P + P P + + P ++PNF P+F++ SS+P+ P+Y S + P
Sbjct: 260 YSHQPYHQDPPKHMPPPQNYSSHEPSPNSLPNFQSYPSFSE-SSLPSTSPHYPSHYQNP 317
Score = 28.1 bits (61), Expect = 7.9
Identities = 23/74 (31%), Positives = 32/74 (43%), Gaps = 12/74 (16%)
Query: 14 SNYPFNYE-----NPNNYPNPNQFFNQRPQNIPNFGFPPNFNQSS------SVPNFHPYY 62
S+YP N P++ P P+ + +Q P PP N SS S+PNF Y
Sbjct: 237 SSYPSNDHLPPPTGPSDSPYPHPYSHQPYHQDPPKHMPPPQNYSSHEPSPNSLPNFQSYP 296
Query: 63 G-SMMKYPSQTPPF 75
S PS +P +
Sbjct: 297 SFSESSLPSTSPHY 310
>At1g37080 hypothetical protein
Length = 224
Score = 39.7 bits (91), Expect = 0.003
Identities = 27/102 (26%), Positives = 48/102 (46%), Gaps = 12/102 (11%)
Query: 57 NFHPYYGSMMKYPSQTP-PFNGSMPMENDNFHNVGATQYPEFSTQITPSGMAIADEVTPE 115
NF S + P++ P P+ P +G +++ F+TQ S + + T
Sbjct: 9 NFIDLLTSQHETPNRQPNPYETHSPSIK-----LGESEFLVFNTQWHESHLQGGNTTTKR 63
Query: 116 DSTPKSKRSKEPAWNTQQNLVLISAWIKYGTSSVVGRNQRGE 157
+T ++K W +++++VLIS W+ SVVG QR +
Sbjct: 64 TTTTRNK------WTSKEDIVLISTWLNTSKDSVVGNEQRAD 99
>At2g16140 pseudogene; similar to MURA transposase of maize Mutator
transposon
Length = 311
Score = 39.3 bits (90), Expect = 0.003
Identities = 20/61 (32%), Positives = 34/61 (54%), Gaps = 8/61 (13%)
Query: 95 PEFSTQITPSGMAIADEVTPEDSTPKSKRSKEPAWNTQQNLVLISAWIKYGTSSVVGRNQ 154
P FSTQ T D + E+ PK KR + W+ ++++VL+S+W+ +V+G Q
Sbjct: 34 PIFSTQCT-------DSPSLEEHVPKVKRERMK-WSAKEDMVLVSSWLNTSKDAVIGNEQ 85
Query: 155 R 155
+
Sbjct: 86 K 86
>At3g15000 unknown protein
Length = 395
Score = 38.5 bits (88), Expect = 0.006
Identities = 29/101 (28%), Positives = 42/101 (40%), Gaps = 7/101 (6%)
Query: 9 NNQNYSNYPFNYENP--NNY--PNPNQFFNQRPQNIPNFGFPPNFNQSSSVPNFHPYYGS 64
NN +P Y P NN P P Q + P PN+G P N P + G
Sbjct: 287 NNMGGPRHPPPYGAPPQNNMGGPRPPQNYGGTPP--PNYGGAPPANNMGGAPPPNYGGGP 344
Query: 65 MMKYPSQTPPFNGSMPMENDNFHNVGA-TQYPEFSTQITPS 104
+Y + PP G P +N+N+ G+ Q P++ P+
Sbjct: 345 PPQYGAVPPPQYGGAPPQNNNYQQQGSGMQQPQYQNNYPPN 385
>At1g42670 unknown protein
Length = 579
Score = 38.5 bits (88), Expect = 0.006
Identities = 23/77 (29%), Positives = 39/77 (49%), Gaps = 16/77 (20%)
Query: 89 VGATQYPEFSTQITPSGMAIADEVTPEDSTP-------KSKRSKEPA--WNTQQNLVLIS 139
+G+ P +ST+ + D+ ED P K K++K+P W++ +++VLIS
Sbjct: 298 LGSANVPVYSTEWS-------DDDPSEDELPIAGKKGKKGKKAKKPRRNWSSTEDVVLIS 350
Query: 140 AWIKYGTSSVVGRNQRG 156
W+ VVG Q+G
Sbjct: 351 GWLNTSKDPVVGNEQKG 367
>At5g19080 unknown protein
Length = 378
Score = 37.4 bits (85), Expect = 0.013
Identities = 32/100 (32%), Positives = 42/100 (42%), Gaps = 22/100 (22%)
Query: 2 DPNNNHFNNQNYSNYPFNYENPNNYPNPNQ----FFNQRPQNIPNFGFPPNFNQ--SSSV 55
D NNNH + + N P+ Y +P P Q + + P + P PP Q SSS
Sbjct: 11 DNNNNH--HHPHHNPPYYYSDPPPQQPPPQNGYSYSHNYPVSTPQLSLPPPPAQPPSSSQ 68
Query: 56 P-----NFHPY---YGSMMKYPSQTPPF------NGSMPM 81
P ++ PY Y YP Q PP+ NG PM
Sbjct: 69 PPPSQISYRPYGQNYHQNQYYPQQAPPYFTGYHHNGFNPM 108
>At5g28710 putative protein
Length = 264
Score = 36.6 bits (83), Expect = 0.022
Identities = 22/74 (29%), Positives = 38/74 (50%), Gaps = 13/74 (17%)
Query: 89 VGATQYPEFSTQITPSGMAIADEVTPEDST----PKSKRSKEPA--WNTQQNLVLISAWI 142
+G+T P FST+ + D+ ED K K++K+P W++ +++VLI W+
Sbjct: 36 LGSTNVPVFSTEWS-------DDDPSEDEALIAGKKGKKAKKPRRNWSSIEDIVLICGWL 88
Query: 143 KYGTSSVVGRNQRG 156
+VG Q+G
Sbjct: 89 NTSKDPMVGNEQKG 102
>At1g79480 hypothetical protein
Length = 356
Score = 36.2 bits (82), Expect = 0.029
Identities = 37/120 (30%), Positives = 51/120 (41%), Gaps = 16/120 (13%)
Query: 2 DPNNNHFNNQNYSNY--PFNYENPNNYPNPNQFFNQRPQNIPNFGFPPNFNQSSSVPNFH 59
+PN+N ++ SN P + NPN+ PNP P++ N PN SSS PN +
Sbjct: 95 NPNSNPNPPESSSNPNPPDSSSNPNSNPNPPVTVPNPPESSSN----PNPPDSSSNPNSN 150
Query: 60 PYYGSMMKYPSQTPPFNGSMPMENDNFHNVGATQYPEFSTQITPSGMAIADEVTPEDSTP 119
P P+ PP P E+ + N PE S+ P I PE S+P
Sbjct: 151 PNPPESSSNPN--PPVTVPNPPESSSNPNP-----PESSSNPNP---PITIPYPPESSSP 200
Score = 31.6 bits (70), Expect = 0.72
Identities = 25/76 (32%), Positives = 34/76 (43%), Gaps = 16/76 (21%)
Query: 15 NYPFNYENPNNYPNPNQFFNQRPQNIPNFGFPPNFNQSSSVPNFHPYYGSMMKYPSQT-- 72
N P + NPN+ PNP P++ N PN SSS PN +P + P ++
Sbjct: 88 NPPDSSSNPNSNPNP-------PESSSN----PNPPDSSSNPNSNPNPPVTVPNPPESSS 136
Query: 73 ---PPFNGSMPMENDN 85
PP + S P N N
Sbjct: 137 NPNPPDSSSNPNSNPN 152
>At2g33435 RRM-containing protein
Length = 979
Score = 35.8 bits (81), Expect = 0.038
Identities = 29/85 (34%), Positives = 39/85 (45%), Gaps = 10/85 (11%)
Query: 265 RKQIEEGSGSRSRKYLKRDHAGANQRL-------IDDYFANEPTYDDAMFRRRYRMQKHV 317
RK+ + SGS K +K DHAGA Q L + DY A+E +D + RR K
Sbjct: 337 RKKEDATSGSTEEKSMKEDHAGAAQLLGHDIVEKVSDYHASEKGHDRSTKVRREERVKDS 396
Query: 318 FLRIVGDLSSSDNYFTQRVDAANKE 342
+ S S +RVD + KE
Sbjct: 397 SRKKEDATSGSRE---ERVDNSTKE 418
Score = 31.6 bits (70), Expect = 0.72
Identities = 18/46 (39%), Positives = 22/46 (47%), Gaps = 7/46 (15%)
Query: 265 RKQIEEGSGSRSRKYLKRDHAGANQ-------RLIDDYFANEPTYD 303
RK+ E S SR K +K DH GA Q ++ DY E YD
Sbjct: 462 RKKEEAISSSRGEKPIKEDHVGAAQLLGNDLVEMVSDYHETEKGYD 507
>At5g28890 putative protein
Length = 232
Score = 35.0 bits (79), Expect = 0.065
Identities = 20/77 (25%), Positives = 39/77 (49%), Gaps = 4/77 (5%)
Query: 79 MPMENDNFHNVGATQYPEFSTQITPSGMAIADEVTPEDSTPKSKRSKEPAWNTQQNLVLI 138
M F ++ +Q ++ + P ++DE + +TPK K+ W+ +++++LI
Sbjct: 1 MDSYTSGFMDLLQSQQESYNNENNPQ---LSDEDHEDVNTPKVAIPKKK-WSAKEDVILI 56
Query: 139 SAWIKYGTSSVVGRNQR 155
SAW+ VVG Q+
Sbjct: 57 SAWLNTSKDPVVGNEQK 73
>At4g08150 KNAT1 homeobox-like protein
Length = 398
Score = 35.0 bits (79), Expect = 0.065
Identities = 33/119 (27%), Positives = 49/119 (40%), Gaps = 26/119 (21%)
Query: 2 DPNNNHFNNQNYSNYPFNYENPNN---------YPNPNQFFNQRPQNI-------PNFGF 45
D NNN+ NN N SNY Y N NN +P+ + Q +N PN
Sbjct: 33 DTNNNN-NNNNSSNYGPGYNNTNNNNHHHQHMLFPHMSSLLPQTTENCFRSDHDQPNNNN 91
Query: 46 PPNFNQSSSVPNFHPYYGSMMKYPSQTPPFNGSMPMENDNFHNVGATQ-----YPEFST 99
P+ +S + +Y +M+ T N + NDN +V A + +P +ST
Sbjct: 92 NPSVKSEASSSRIN-HYSMLMRAIHNTQEANNN---NNDNVSDVEAMKAKIIAHPHYST 146
>At5g18440 putative protein
Length = 451
Score = 33.1 bits (74), Expect = 0.25
Identities = 15/33 (45%), Positives = 18/33 (54%), Gaps = 5/33 (15%)
Query: 24 NNYPNPN-----QFFNQRPQNIPNFGFPPNFNQ 51
+N P PN QFFN PQ +P F P + NQ
Sbjct: 49 HNMPMPNMPIHPQFFNNMPQQLPQFAMPNHINQ 81
>At4g24680 unknown protein
Length = 1480
Score = 33.1 bits (74), Expect = 0.25
Identities = 25/100 (25%), Positives = 40/100 (40%), Gaps = 16/100 (16%)
Query: 9 NNQNYSNYPFNYENPNNYPNPNQFFNQRPQNIPNFGFPPNFNQSSSVP-NFHP---YYGS 64
N+ N P++ + P + Q ++ Q+ PN FPP ++ P N H +YG
Sbjct: 312 NSWRRENQPYSEDAPRHCREEGQLDSRGSQSYPNANFPPRYDAWRGPPVNNHQGGGWYGG 371
Query: 65 MMKYPSQTPPFNGSMPMENDNFHNVGATQYPEFSTQITPS 104
Y PM FH +P + TQ+ P+
Sbjct: 372 NHPY---------GAPMGPGGFH---MDPFPFYPTQVPPA 399
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.333 0.143 0.475
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,676,818
Number of Sequences: 26719
Number of extensions: 396502
Number of successful extensions: 1894
Number of sequences better than 10.0: 71
Number of HSP's better than 10.0 without gapping: 23
Number of HSP's successfully gapped in prelim test: 49
Number of HSP's that attempted gapping in prelim test: 1771
Number of HSP's gapped (non-prelim): 128
length of query: 367
length of database: 11,318,596
effective HSP length: 101
effective length of query: 266
effective length of database: 8,619,977
effective search space: 2292913882
effective search space used: 2292913882
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.5 bits)
S2: 61 (28.1 bits)
Medicago: description of AC149579.7


