

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
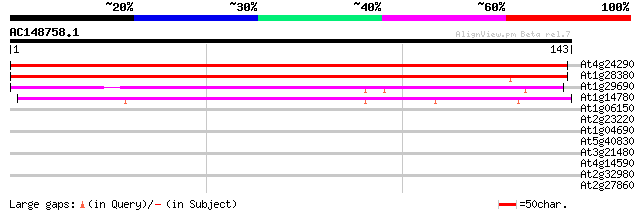
Query= AC148758.1 - phase: 0 /pseudo
(143 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At4g24290 unknown protein 182 7e-47
At1g28380 unknown protein 142 6e-35
At1g29690 unknown protein 109 4e-25
At1g14780 unknown protein 93 4e-20
At1g06150 unknown protein 28 1.3
At2g23220 putative cytochrome P450 28 2.2
At1g04690 putative K+ channel, beta subunit 28 2.2
At5g40830 unknown protein 27 3.7
At3g21480 unknown protein 27 3.7
At4g14590 hypothetical protein 27 4.9
At2g32980 unknown protein 26 6.4
At2g27860 putative dTDP-glucose 4-6-dehydratase 26 8.3
>At4g24290 unknown protein
Length = 606
Score = 182 bits (461), Expect = 7e-47
Identities = 89/142 (62%), Positives = 108/142 (75%)
Query: 1 MEDLHRFLEFQLPRQWAPVLGEIHLGSYRKHQVNTWLRFSILGPKLYINTTPVDVGNRPV 60
+E+LH+FLEFQLPRQWAPV E+ LG RK Q L+FS GPKLY+NTTPVDVG RP+
Sbjct: 325 IEELHQFLEFQLPRQWAPVFSELPLGPQRKQQSCASLQFSFFGPKLYVNTTPVDVGKRPI 384
Query: 61 TGLRLQLEGSRSNRLAIHLQHLASLPKSLPLADNANAYLSCDSYSCTFHKKVKRNCFSYV 120
TG+RL LEG RSNRLAIHLQHL+SLPK L D+ N + +S+ +++KV +S+V
Sbjct: 385 TGMRLYLEGRRSNRLAIHLQHLSSLPKIYQLEDDLNRSIRQESHDRRYYEKVNWKNYSHV 444
Query: 121 CTAPVESDDSLSIVTGAQLQVE 142
CT PVESDD LS+VTGAQL VE
Sbjct: 445 CTEPVESDDDLSVVTGAQLHVE 466
>At1g28380 unknown protein
Length = 612
Score = 142 bits (358), Expect = 6e-35
Identities = 76/147 (51%), Positives = 100/147 (67%), Gaps = 5/147 (3%)
Query: 1 MEDLHRFLEFQLPRQWAPVLGEIHLGSYRKHQVNTWLRFSILGPKLYINTTPVDVGNRPV 60
+E+LH+FLEFQLPRQWAPV G++ LG R Q + L+FS++GPKLY+NT+ VD G RPV
Sbjct: 331 IEELHQFLEFQLPRQWAPVYGDLPLGLRRSKQSSPSLQFSLMGPKLYVNTSKVDSGERPV 390
Query: 61 TGLRLQLEGSRSNRLAIHLQHLASLPKSLPLADNANAYLSCDSYSCTFHKKVKRNCFSYV 120
TGLR LEG + N LAIHLQHL++ P SL L+ + + ++ VK FS+V
Sbjct: 391 TGLRFFLEGKKGNHLAIHLQHLSACPPSLHLSHDDTYEPIEEPVEKGYYVPVKWGIFSHV 450
Query: 121 CTAPVE-----SDDSLSIVTGAQLQVE 142
CT PV+ SDD+ SIVT A L+V+
Sbjct: 451 CTYPVQYNGARSDDTASIVTKAWLEVK 477
>At1g29690 unknown protein
Length = 561
Score = 109 bits (273), Expect = 4e-25
Identities = 67/154 (43%), Positives = 93/154 (59%), Gaps = 17/154 (11%)
Query: 1 MEDLHRFLEFQLPRQWAPVLGEIHLGSYRKHQVNTWLRFSILGPKLYINTTPVDVGNRPV 60
+EDL FL++Q+ R WAP + RK V + L+FS++GPKL+I+ V VG +PV
Sbjct: 307 IEDLQYFLDYQIARAWAPEQSNLQ----RKEPVCSSLQFSLMGPKLFISADQVTVGRKPV 362
Query: 61 TGLRLQLEGSRSNRLAIHLQHLASLPKSL-PLADN-----ANAYLSCDSYSCTFHKKVKR 114
TGLRL LEGS+ NRL+IHLQHL SLPK L P D+ A + + + + +K
Sbjct: 363 TGLRLSLEGSKQNRLSIHLQHLVSLPKILQPHWDSHVPIGAPKWQGPEEQDSRWFEPIKW 422
Query: 115 NCFSYVCTAPVESDDS-------LSIVTGAQLQV 141
FS+V T+P+E ++ + IVTGAQL V
Sbjct: 423 KNFSHVSTSPIEHTETHIGDLSGVHIVTGAQLGV 456
>At1g14780 unknown protein
Length = 627
Score = 93.2 bits (230), Expect = 4e-20
Identities = 60/159 (37%), Positives = 83/159 (51%), Gaps = 18/159 (11%)
Query: 3 DLHRFLEFQLPRQWAPVLGEIHLGSY-RKHQVNTWLRFSILGPKLYINTTPVDVGNRPVT 61
DL FL+F PR WAPV ++ G+ L + +GPKLY+NTTPV PVT
Sbjct: 334 DLQYFLDFSGPRAWAPVHNDLPFGAAPNMASAYPALHINFMGPKLYVNTTPVTSEKNPVT 393
Query: 62 GLRLQLEGSRSNRLAIHLQHLASLPKSL-PLADNANAYLSCDSYSCT--FHKKVKRNCFS 118
G+R LEG + NRLAIHLQHL + ++ + + + D + + + + FS
Sbjct: 394 GMRFFLEGKKCNRLAIHLQHLDNTRTTVGEKITDEHIWRGSDQITDNDRYFEPLNGKKFS 453
Query: 119 YVCTAPVESD--------------DSLSIVTGAQLQVEK 143
+VCT PV+ D D IVTGAQL+V+K
Sbjct: 454 HVCTVPVKYDPNWIKTTSNHKSQNDVAFIVTGAQLEVKK 492
>At1g06150 unknown protein
Length = 1288
Score = 28.5 bits (62), Expect = 1.3
Identities = 20/64 (31%), Positives = 30/64 (46%), Gaps = 1/64 (1%)
Query: 53 VDVGNRPVTGLRLQLEGSRSNRLAIHLQHLASLPKSLPLADNANAYLSCDSYSCTFHKKV 112
V VG+ V L + L H++HL L + PLAD+A+ + CD S + K+
Sbjct: 92 VAVGSCGVVQLGSLCKVEEDPALVTHIRHLF-LALTDPLADHASNLMQCDINSPSDRPKI 150
Query: 113 KRNC 116
C
Sbjct: 151 PSKC 154
>At2g23220 putative cytochrome P450
Length = 515
Score = 27.7 bits (60), Expect = 2.2
Identities = 18/56 (32%), Positives = 26/56 (46%)
Query: 58 RPVTGLRLQLEGSRSNRLAIHLQHLASLPKSLPLADNANAYLSCDSYSCTFHKKVK 113
R V G R EG+ + +A ++HL + A NA YL + F K+VK
Sbjct: 203 RMVAGKRFYGEGTEQDEVAQQVRHLMDEIVTSAGAGNAADYLPIMRWFTNFEKRVK 258
>At1g04690 putative K+ channel, beta subunit
Length = 328
Score = 27.7 bits (60), Expect = 2.2
Identities = 15/49 (30%), Positives = 25/49 (50%)
Query: 16 WAPVLGEIHLGSYRKHQVNTWLRFSILGPKLYINTTPVDVGNRPVTGLR 64
W+P+ + G Y K + + RF++ K N + VD R V+GL+
Sbjct: 207 WSPLASGVLTGKYNKGAIPSDSRFALENYKNLANRSLVDDVLRKVSGLK 255
>At5g40830 unknown protein
Length = 414
Score = 26.9 bits (58), Expect = 3.7
Identities = 19/64 (29%), Positives = 28/64 (43%), Gaps = 4/64 (6%)
Query: 77 IHLQHLASLPKSL--PLADNANAY--LSCDSYSCTFHKKVKRNCFSYVCTAPVESDDSLS 132
+H LA P SL P+ +++ + L C S+ C KK+ R+C A D
Sbjct: 183 VHKPGLALFPDSLWRPVGNSSVNWSGLGCKSFECLKGKKLSRDCVGCFDLATSHEKDRFV 242
Query: 133 IVTG 136
V G
Sbjct: 243 KVNG 246
>At3g21480 unknown protein
Length = 1045
Score = 26.9 bits (58), Expect = 3.7
Identities = 26/89 (29%), Positives = 31/89 (34%), Gaps = 9/89 (10%)
Query: 30 KHQVNTWLRFSILGPKLYINTTPVDVGNRPVTGLRLQLEGSRSNRLAIHLQHLASLPKSL 89
KHQ RF I T N T R LE S + + Q L S+
Sbjct: 831 KHQKKILARFDISEASSMKEATHFIADN--FTRTRNMLEAIASGKPVVTTQWLESI---- 884
Query: 90 PLADNANAYLSCDSYSCTFHKKVKRNCFS 118
D N Y+ D Y KK K CF+
Sbjct: 885 ---DQVNIYVDEDMYILRDSKKEKEFCFN 910
>At4g14590 hypothetical protein
Length = 508
Score = 26.6 bits (57), Expect = 4.9
Identities = 11/27 (40%), Positives = 21/27 (77%), Gaps = 1/27 (3%)
Query: 19 VLGEIHLGSYRKHQVNTWLRFSILGPK 45
+LG + LGS+++HQ+ +L+ +LGP+
Sbjct: 261 LLGNVKLGSHKRHQI-WFLKKFLLGPE 286
>At2g32980 unknown protein
Length = 296
Score = 26.2 bits (56), Expect = 6.4
Identities = 9/37 (24%), Positives = 21/37 (56%)
Query: 53 VDVGNRPVTGLRLQLEGSRSNRLAIHLQHLASLPKSL 89
+ V R + L+++L+G + ++ HL H+ + K +
Sbjct: 65 LSVVQRKIADLQVELQGRKDDKNVAHLTHVGEMQKKI 101
>At2g27860 putative dTDP-glucose 4-6-dehydratase
Length = 389
Score = 25.8 bits (55), Expect = 8.3
Identities = 21/87 (24%), Positives = 33/87 (37%), Gaps = 16/87 (18%)
Query: 51 TPVDVGNRP--------VTGLRLQLEGSRSNRLAIHLQHL--------ASLPKSLPLADN 94
TP D RP + L + S +N+ IH + LPK PL D+
Sbjct: 101 TPADYNTRPLDTIYSNFIDALPVVKYCSENNKRLIHFSTCEVYGKTIGSFLPKDHPLRDD 160
Query: 95 ANAYLSCDSYSCTFHKKVKRNCFSYVC 121
Y+ + S +++ +SY C
Sbjct: 161 PAFYVLKEDISPCIFGSIEKQRWSYAC 187
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.136 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,330,606
Number of Sequences: 26719
Number of extensions: 133016
Number of successful extensions: 245
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 232
Number of HSP's gapped (non-prelim): 12
length of query: 143
length of database: 11,318,596
effective HSP length: 89
effective length of query: 54
effective length of database: 8,940,605
effective search space: 482792670
effective search space used: 482792670
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 55 (25.8 bits)
Medicago: description of AC148758.1


