

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148218.5 - phase: 0
(164 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
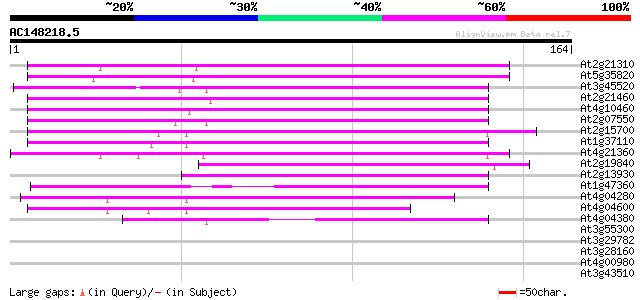
Score E
Sequences producing significant alignments: (bits) Value
At2g21310 putative retroelement pol polyprotein 87 3e-18
At5g35820 copia-like retrotransposable element 86 9e-18
At3g45520 copia-like polyprotein 83 8e-17
At2g21460 putative retroelement pol polyprotein 79 1e-15
At4g10460 putative retrotransposon 78 2e-15
At2g07550 putative retroelement pol polyprotein 77 3e-15
At2g15700 copia-like retroelement pol polyprotein 69 9e-13
At1g37110 63 6e-11
At4g21360 putative transposable element 62 1e-10
At2g19840 copia-like retroelement pol polyprotein 62 1e-10
At2g13930 putative retroelement pol polyprotein 58 2e-09
At1g47360 polyprotein, putative 56 8e-09
At4g04280 putative transposon protein 55 2e-08
At4g04600 putative polyprotein 52 2e-07
At4g04380 putative polyprotein 43 7e-05
At3g55300 putative protein 37 0.005
At3g29782 hypothetical protein 35 0.024
At3g28160 non-LTR reverse transcriptase, putative 34 0.040
At4g00980 unknown protein 32 0.15
At3g43510 unknown protein 31 0.26
>At2g21310 putative retroelement pol polyprotein
Length = 838
Score = 87.4 bits (215), Expect = 3e-18
Identities = 56/153 (36%), Positives = 83/153 (53%), Gaps = 12/153 (7%)
Query: 6 DIEKFTGDNDFRLWKVKMEA------VLIQQKCEKTLKGESSLPVTMSRAEKTE------ 53
++EK G+ D+ LWK K+ A +L + ++ ++ E S T S KTE
Sbjct: 7 EVEKLDGEGDYVLWKEKLLAHIELLGLLEGLEEDEAIEEEESTAETDSLLTKTEDKVLKE 66
Query: 54 MVDKARSAIVLCLGNKDLREATKETTATSMWSKFEYLYMTKSLAHRQVLKQQLYSFKMVK 113
KARS ++L LGN LR+ KE TA M + L+M KSL +R LKQ+LY +KM
Sbjct: 67 KRGKARSTVILSLGNHVLRKVIKEKTAAGMIRVLDKLFMAKSLPNRIYLKQRLYGYKMSD 126
Query: 114 SKVVVELLVEFNKIIGDFENIKLHLEDAGALMV 146
S + E + +F K+I D EN+K+ + D +V
Sbjct: 127 SMTIEENVNDFFKLISDLENVKVSVPDEDQAIV 159
>At5g35820 copia-like retrotransposable element
Length = 1342
Score = 85.9 bits (211), Expect = 9e-18
Identities = 56/161 (34%), Positives = 81/161 (49%), Gaps = 20/161 (12%)
Query: 6 DIEKFTGDNDFRLWKVKM-----------------EAVLIQQKCEKTLKGESSLPVTMSR 48
++EKF GD D+ LWK K+ EAV+ E + G S+
Sbjct: 7 EVEKFDGDGDYILWKEKLLAHMEMLGLLEGLGEEEEAVVEDSTTEISDGGNQDPETATSK 66
Query: 49 AEKT---EMVDKARSAIVLCLGNKDLREATKETTATSMWSKFEYLYMTKSLAHRQVLKQQ 105
E E KARS I+L LGN LR+ K+ TA M + L+M KSL +R LKQ+
Sbjct: 67 LEDKILKEKRGKARSTIILSLGNNVLRKVIKQKTAAGMIKVLDQLFMAKSLPNRIYLKQR 126
Query: 106 LYSFKMVKSKVVVELLVEFNKIIGDFENIKLHLEDAGALMV 146
LY +KM ++ + E + +F K+I D EN+K+ + D +V
Sbjct: 127 LYGYKMSENMTMEENVNDFFKLISDLENVKVVVPDEDQAIV 167
>At3g45520 copia-like polyprotein
Length = 1363
Score = 82.8 bits (203), Expect = 8e-17
Identities = 54/155 (34%), Positives = 84/155 (53%), Gaps = 17/155 (10%)
Query: 2 GSKWDIEKFTGDNDFRLWKVKMEAVLIQQKCEKTLKGESSLPVTMSR------------A 49
G++ ++EKF G D+ +WK K+ A + L+ ES P+ R
Sbjct: 3 GARIEVEKFDGRGDYTMWKEKLLAHIDMLGLSAVLR-ESETPMGKERDSEKSDEDEKEER 61
Query: 50 EKTEMVD----KARSAIVLCLGNKDLREATKETTATSMWSKFEYLYMTKSLAHRQVLKQQ 105
EK E + KARS IVL + ++ LR+ KET+A +M + LYM+K+L +R LKQ+
Sbjct: 62 EKMEAFEEKKRKARSTIVLSVSDRVLRKIKKETSAAAMLEALDRLYMSKALPNRIYLKQK 121
Query: 106 LYSFKMVKSKVVVELLVEFNKIIGDFENIKLHLED 140
LYSFKM ++ + + EF I+ D EN+ + + D
Sbjct: 122 LYSFKMSENLSIEGNIDEFLHIVADLENLNVLVSD 156
>At2g21460 putative retroelement pol polyprotein
Length = 1333
Score = 79.0 bits (193), Expect = 1e-15
Identities = 52/150 (34%), Positives = 79/150 (52%), Gaps = 15/150 (10%)
Query: 6 DIEKFTGDNDFRLWKVKMEAVLIQQKCEKTLKGESSLPVTMSRAEKTEMVDK-------- 57
++EKF G D+ +WK K+ A L LK E L ++ + TE +K
Sbjct: 7 EVEKFDGRGDYTMWKEKLMAHLDILGLSVALKEEDDLVEKVAEMQLTEEEEKEEVLRREL 66
Query: 58 -------ARSAIVLCLGNKDLREATKETTATSMWSKFEYLYMTKSLAHRQVLKQQLYSFK 110
ARSAIVL + ++ LR+ KE +A +M + LYM+K+L +R KQ+LYSFK
Sbjct: 67 LEEKRRKARSAIVLSVTDRVLRKIKKEQSAAAMLGVLDKLYMSKALPNRIYQKQKLYSFK 126
Query: 111 MVKSKVVVELLVEFNKIIGDFENIKLHLED 140
M ++ + + EF +II D EN + + D
Sbjct: 127 MSENLSIEGNIDEFLRIIADLENTNVLVSD 156
>At4g10460 putative retrotransposon
Length = 1230
Score = 78.2 bits (191), Expect = 2e-15
Identities = 51/148 (34%), Positives = 78/148 (52%), Gaps = 13/148 (8%)
Query: 6 DIEKFTGDNDFRLWKVKMEAVLIQQKCEKTLKGESSLPVTMSRAEK-------------T 52
++EKF G D+ LWK K+ A + L+ S+ + E+
Sbjct: 7 EMEKFDGHGDYTLWKEKLMAHMDLLGLTVALRETQSVSDPLESEEEGKESEKGDKEALME 66
Query: 53 EMVDKARSAIVLCLGNKDLREATKETTATSMWSKFEYLYMTKSLAHRQVLKQQLYSFKMV 112
E KARS IVL + ++ LR++ KE TA SM + LYM+K+L +R LKQ+LYS+KM
Sbjct: 67 EKRQKARSTIVLSVSDQVLRKSKKEKTAPSMLEALDKLYMSKALPNRIYLKQKLYSYKMQ 126
Query: 113 KSKVVVELLVEFNKIIGDFENIKLHLED 140
++ V + EF ++I D EN + + D
Sbjct: 127 ENLSVEGNIDEFLRLIADLENTNVLVSD 154
>At2g07550 putative retroelement pol polyprotein
Length = 1356
Score = 77.4 bits (189), Expect = 3e-15
Identities = 54/150 (36%), Positives = 81/150 (54%), Gaps = 15/150 (10%)
Query: 6 DIEKFTGDNDFRLWKVKMEAVLIQQKCEKTLKGESSLPVTMS-----------RAEKTEM 54
++EKF G D+ +WK K+ A + LK S S + EK E
Sbjct: 7 EVEKFDGRGDYTMWKEKLLAHMDILGLNTALKESESTGEKKSVLDESDEDYEEKLEKFEA 66
Query: 55 VD----KARSAIVLCLGNKDLREATKETTATSMWSKFEYLYMTKSLAHRQVLKQQLYSFK 110
++ KARSAIVL + ++ LR+ KE+TA +M + LYM+K+L +R KQ+LYSFK
Sbjct: 67 LEEKKKKARSAIVLSVTDRVLRKIKKESTAAAMLLALDKLYMSKALPNRIYPKQKLYSFK 126
Query: 111 MVKSKVVVELLVEFNKIIGDFENIKLHLED 140
M ++ V + EF +II D EN+ + + D
Sbjct: 127 MSENLSVEGNIDEFLQIITDLENMNVIISD 156
>At2g15700 copia-like retroelement pol polyprotein
Length = 1166
Score = 69.3 bits (168), Expect = 9e-13
Identities = 52/173 (30%), Positives = 83/173 (47%), Gaps = 24/173 (13%)
Query: 6 DIEKFTGDNDFRLWKVKMEAVLIQQKCEKTLKGESSL---PVTMSRAE------------ 50
++++F G DF LWKV+M A + L E L PV A
Sbjct: 8 EVDRFDGTGDFSLWKVRMLAHYGVLGLKGILNDEQLLRDPPVIEEEAAVAGRDYHVGDFE 67
Query: 51 --------KTEMVDKARSAIVLCLGNKDLREATKETTATSMWSKFEYLYMTKSLAHRQVL 102
K E +KA+ IVL +GN+ LR+ TA +MWS + LYM SL +R L
Sbjct: 68 LPSNVDLIKFEKSEKAKDLIVLNVGNQVLRKIKNCETAAAMWSTLKRLYMETSLPNRIYL 127
Query: 103 KQQLYSFKMVKSKVVVELLVEFNKIIGDFENIKLHL-EDAGALMVWCCLEDEE 154
+ + Y++KM S+ + + +F K++ D NI + + E+ A+++ L D +
Sbjct: 128 QLKFYTYKMTDSRSIDGNVDDFLKLVTDLNNIGVDVTEEVQAILLLSSLYDRK 180
>At1g37110
Length = 1356
Score = 63.2 bits (152), Expect = 6e-11
Identities = 40/156 (25%), Positives = 78/156 (49%), Gaps = 21/156 (13%)
Query: 6 DIEKFTGDNDFRLWKVKMEAVLIQQKCEKTLKGES---SLPVTMSRAE------------ 50
+I+ F GD DF LWK++++A L + TL S ++P+T S A+
Sbjct: 9 EIKVFNGDRDFSLWKIRIQAQLGVLGLKDTLTDFSLTKTVPLTKSEAKQESGDGESSGTK 68
Query: 51 ------KTEMVDKARSAIVLCLGNKDLREATKETTATSMWSKFEYLYMTKSLAHRQVLKQ 104
K E ++A++ I+ + + L + T +W+ YM SL +R +
Sbjct: 69 EVPDPVKIEQSEQAKNIIINHISDVVLLKVNHYATTADLWATLNKKYMETSLPNRIYTQL 128
Query: 105 QLYSFKMVKSKVVVELLVEFNKIIGDFENIKLHLED 140
+LYSFKMV + + + + EF +I+ + ++++ +++
Sbjct: 129 KLYSFKMVSTMTIDQNVDEFLRIVAELGSLEIQVDE 164
>At4g21360 putative transposable element
Length = 1308
Score = 62.0 bits (149), Expect = 1e-10
Identities = 47/171 (27%), Positives = 85/171 (49%), Gaps = 25/171 (14%)
Query: 1 MGSKWDIEKFTGDNDFRLWKVKMEA---------VLIQQKCEKTL------KGESSLPVT 45
M +K +I+ F GD DF LWK+++EA L KT+ K ES
Sbjct: 3 MSTKVEIKTFNGDRDFSLWKIRIEAQLGVLGLKPALSDFTLTKTILVVKSEKKESESEDD 62
Query: 46 MSRAEKTEMV---------DKARSAIVLCLGNKDLREATKETTATSMWSKFEYLYMTKSL 96
+ ++KTE V D+A++ I+ + + L + TA +W+ L+M SL
Sbjct: 63 ETDSKKTEEVPDPIKFEQSDQAKNFIINHITDTVLLKVQHCVTAAELWATLNKLFMETSL 122
Query: 97 AHRQVLKQQLYSFKMVKSKVVVELLVEFNKIIGDFENIKLHL-EDAGALMV 146
+R + +LYSFKMV + + + EF +I+ + ++++ + E+ A+++
Sbjct: 123 PNRIYTQLRLYSFKMVDNLSIDQNTDEFLRIVAELGSLQIQVGEEVQAILI 173
>At2g19840 copia-like retroelement pol polyprotein
Length = 1137
Score = 62.0 bits (149), Expect = 1e-10
Identities = 32/98 (32%), Positives = 56/98 (56%), Gaps = 1/98 (1%)
Query: 56 DKARSAIVLCLGNKDLREATKETTATSMWSKFEYLYMTKSLAHRQVLKQQLYSFKMVKSK 115
+KA I + +G+K LR TA W+ + LY+ KSL +R L+ ++Y+++M SK
Sbjct: 42 EKAMDMIFINVGDKVLRNIENSKTAAEAWATLDKLYLVKSLPNRVYLQLKVYNYRMQDSK 101
Query: 116 VVVELLVEFNKIIGDFENIKLHLED-AGALMVWCCLED 152
+ E + EF K+I D N+++ + D A+++ L D
Sbjct: 102 TLEENVDEFQKMISDLNNLQIQVPDEVQAILILSALPD 139
>At2g13930 putative retroelement pol polyprotein
Length = 1335
Score = 58.2 bits (139), Expect = 2e-09
Identities = 29/90 (32%), Positives = 51/90 (56%)
Query: 51 KTEMVDKARSAIVLCLGNKDLREATKETTATSMWSKFEYLYMTKSLAHRQVLKQQLYSFK 110
+ E DKA++ I L + +K LR+ TA W + L+M +SL HR + Y+FK
Sbjct: 42 RLERCDKAKNVIFLNVADKVLRKIELCKTAAEAWETLDRLFMIRSLPHRVYTQLSFYTFK 101
Query: 111 MVKSKVVVELLVEFNKIIGDFENIKLHLED 140
M ++K + E + +F KI+ D ++++ + D
Sbjct: 102 MQENKKIDENIDDFLKIVADLNHLQIDVTD 131
>At1g47360 polyprotein, putative
Length = 1182
Score = 56.2 bits (134), Expect = 8e-09
Identities = 37/134 (27%), Positives = 60/134 (44%), Gaps = 18/134 (13%)
Query: 7 IEKFTGDNDFRLWKVKMEAVLIQQKCEKTLKGESSLPVTMSRAEKTEMVDKARSAIVLCL 66
I F DF LWK ++ A L + + G++ P+T E E I LC
Sbjct: 6 IAVFDESGDFSLWKTRIMAHLSVIGLKDVVIGKTITPLTAEEEEDPE------KKIELC- 58
Query: 67 GNKDLREATKETTATSMWSKFEYLYMTKSLAHRQVLKQQLYSFKMVKSKVVVELLVEFNK 126
TA W + L+M +SL HR + Y+FKM ++K + E + +F K
Sbjct: 59 -----------QTAQEAWETLDRLFMIRSLPHRVYTQLSFYTFKMQENKKIDENIDDFLK 107
Query: 127 IIGDFENIKLHLED 140
I+ D ++++ + D
Sbjct: 108 IVADLNHLQIEVTD 121
>At4g04280 putative transposon protein
Length = 1104
Score = 54.7 bits (130), Expect = 2e-08
Identities = 41/151 (27%), Positives = 69/151 (45%), Gaps = 24/151 (15%)
Query: 4 KWDIEKFTGDNDFRLWKVKMEAVL-------------------IQQKCEKTLKGESSLPV 44
K +I+ F GD DF WK+++EA L + +K EK + E
Sbjct: 2 KVEIKTFNGDRDFSFWKIRIEAQLGVLGLKNSLTDFKLTKTVPVAKKEEKESEYEDDASD 61
Query: 45 TMSRAE-----KTEMVDKARSAIVLCLGNKDLREATKETTATSMWSKFEYLYMTKSLAHR 99
E K E ++A++ I+ + + L + TA +W+ L+M SL +R
Sbjct: 62 IKQATEEPDPIKFEQSEQAKNFIINHITDTVLLKVQHCKTAAEIWATLNKLFMETSLLNR 121
Query: 100 QVLKQQLYSFKMVKSKVVVELLVEFNKIIGD 130
+ +LYSFKMV + + + + EF +I+ D
Sbjct: 122 IYTQLKLYSFKMVDTLSIDQNVDEFLRILAD 152
>At4g04600 putative polyprotein
Length = 922
Score = 51.6 bits (122), Expect = 2e-07
Identities = 40/138 (28%), Positives = 62/138 (43%), Gaps = 26/138 (18%)
Query: 6 DIEKFTGDNDFRLWKVKMEAVL------------IQQKCEKTLKGE------------SS 41
+I+ F GDNDF LWK+++ A L K E K E SS
Sbjct: 8 EIKAFDGDNDFSLWKMRIMAQLGVLGLKGTLTDFALTKTEPLTKSEEKQVASGDESSDSS 67
Query: 42 LPVTMSRAE--KTEMVDKARSAIVLCLGNKDLREATKETTATSMWSKFEYLYMTKSLAHR 99
+T + K E ++A + I+ + + LR+ TA ++W LYM SL +R
Sbjct: 68 AVLTKEVPDPIKIEKSEQAINIIINHISDTVLRKVNHYKTAATLWELLNELYMETSLPNR 127
Query: 100 QVLKQQLYSFKMVKSKVV 117
+ + YSF+M+ SK +
Sbjct: 128 IYAQLKFYSFRMMTSKTI 145
>At4g04380 putative polyprotein
Length = 778
Score = 43.1 bits (100), Expect = 7e-05
Identities = 33/111 (29%), Positives = 52/111 (46%), Gaps = 17/111 (15%)
Query: 34 KTLKGESSLPVTMSRAEKTEMVD----KARSAIVLCLGNKDLREATKETTATSMWSKFEY 89
K E S EK E + K RS IVL + ++ L + +
Sbjct: 28 KDRDSEKSDEEEKEECEKVEAFEEKKRKTRSTIVLSVSDRVLEASDR------------- 74
Query: 90 LYMTKSLAHRQVLKQQLYSFKMVKSKVVVELLVEFNKIIGDFENIKLHLED 140
LYM+K+L ++ LKQ+LY FKM ++ + + EF I+ D EN+ + + D
Sbjct: 75 LYMSKALPNQIYLKQKLYRFKMSENLSMEGNIDEFLHIVADLENLNVLVSD 125
>At3g55300 putative protein
Length = 262
Score = 37.0 bits (84), Expect = 0.005
Identities = 19/54 (35%), Positives = 32/54 (59%)
Query: 87 FEYLYMTKSLAHRQVLKQQLYSFKMVKSKVVVELLVEFNKIIGDFENIKLHLED 140
+E Y TK+LA+ LK + SFKMV++K + E L K++ D ++ ++ D
Sbjct: 36 YEGNYQTKTLANMIYLKHKFTSFKMVENKSIEENLDTSLKLVADLASLDINTSD 89
>At3g29782 hypothetical protein
Length = 1130
Score = 34.7 bits (78), Expect = 0.024
Identities = 28/131 (21%), Positives = 55/131 (41%), Gaps = 8/131 (6%)
Query: 10 FTGDNDFRLWKVKMEAVLIQQKCEKTLKGE-------SSLPVTMSRAEKTEMVDKARSAI 62
F G + F+ W+ KM L+ E+ L + ++ T+ + D +
Sbjct: 71 FDGKSGFKTWQEKMRYYLVSINMERYLPEDPPIVPQGTTYVYTVGGMDTWAQGDYCCKGL 130
Query: 63 VLCLGNKDLREA-TKETTATSMWSKFEYLYMTKSLAHRQVLKQQLYSFKMVKSKVVVELL 121
+L DL + +K ++ ++W E Y T ++ + +FKMV SK ++E +
Sbjct: 131 ILNRLLNDLFDLYSKAKSSKTLWLTLENKYKTDESRMQRFSTAKFLNFKMVDSKPIMEQV 190
Query: 122 VEFNKIIGDFE 132
+I + E
Sbjct: 191 EALQRICQEIE 201
>At3g28160 non-LTR reverse transcriptase, putative
Length = 630
Score = 33.9 bits (76), Expect = 0.040
Identities = 36/128 (28%), Positives = 61/128 (47%), Gaps = 24/128 (18%)
Query: 2 GSKWDIEKFTGDNDFRLWKVK----------MEAVLIQQKCEKTLKGESSLPVTMSRAEK 51
GSK D+EKF G D+ WKVK +E + + EK S + E+
Sbjct: 4 GSK-DLEKFDGFGDYISWKVKFLDHLKDLDLLEGLKEEVCLEKGSGNSGSDEGRVKDLEE 62
Query: 52 TEMVD----KARSAIVLCLGNKDLREATKETTATSMWSKFEYLYMTKSLAHRQVLKQQLY 107
++++ KAR+ I++ +G+ LR+ KE T + + +Y + HR+V K Y
Sbjct: 63 CKILEEKKRKARAFIIVSVGDHVLRKIMKERTTA---DEHKRIYEGE---HRRVFK---Y 113
Query: 108 SFKMVKSK 115
+ +V +K
Sbjct: 114 TVLVVSAK 121
>At4g00980 unknown protein
Length = 488
Score = 32.0 bits (71), Expect = 0.15
Identities = 15/70 (21%), Positives = 35/70 (49%)
Query: 72 REATKETTATSMWSKFEYLYMTKSLAHRQVLKQQLYSFKMVKSKVVVELLVEFNKIIGDF 131
R + K A +W + +++Y ++ ++ F+MV+ + ++E + FNKI
Sbjct: 275 RYSQKFKHAKELWDELKWVYQCDESKSKRSQVRKYIEFRMVEERPILEQVQVFNKIADSI 334
Query: 132 ENIKLHLEDA 141
+ + L++A
Sbjct: 335 VSAGMFLDEA 344
>At3g43510 unknown protein
Length = 336
Score = 31.2 bits (69), Expect = 0.26
Identities = 14/48 (29%), Positives = 27/48 (56%)
Query: 80 ATSMWSKFEYLYMTKSLAHRQVLKQQLYSFKMVKSKVVVELLVEFNKI 127
A +W + +++Y + + V ++ FKMV+ + V+E + EF KI
Sbjct: 156 AKELWEELKWVYRREESNSKMVQVRKYIDFKMVEERPVLEQVQEFIKI 203
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.317 0.131 0.381
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,549,732
Number of Sequences: 26719
Number of extensions: 127210
Number of successful extensions: 364
Number of sequences better than 10.0: 33
Number of HSP's better than 10.0 without gapping: 23
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 328
Number of HSP's gapped (non-prelim): 35
length of query: 164
length of database: 11,318,596
effective HSP length: 92
effective length of query: 72
effective length of database: 8,860,448
effective search space: 637952256
effective search space used: 637952256
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 56 (26.2 bits)
Medicago: description of AC148218.5


