

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
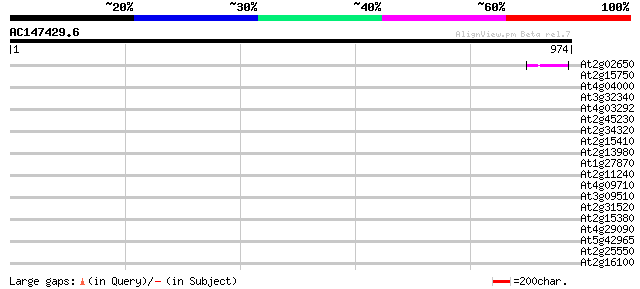
Query= AC147429.6 - phase: 0 /pseudo
(974 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At2g02650 putative reverse transcriptase 44 4e-04
At2g15750 putative non-LTR retroelement reverse transcriptase 38 0.032
At4g04000 putative reverse transcriptase 37 0.054
At3g32340 hypothetical protein 37 0.071
At4g03292 putative protein 35 0.21
At2g45230 putative non-LTR retroelement reverse transcriptase 35 0.21
At2g34320 putative non-LTR retroelement reverse transcriptase 34 0.35
At2g15410 putative retroelement pol polyprotein 34 0.35
At2g13980 pseudogene 33 0.78
At1g27870 hypothetical protein 33 0.78
At2g11240 pseudogene 33 1.0
At4g09710 RNA-directed DNA polymerase -like protein 32 2.3
At3g09510 putative non-LTR reverse transcriptase 32 2.3
At2g31520 putative non-LTR retroelement reverse transcriptase 32 2.3
At2g15380 putative non-LTR retroelement reverse transcriptase 32 2.3
At4g29090 putative protein 31 3.0
At5g42965 putative protein 30 5.1
At2g25550 putative non-LTR retroelement reverse transcriptase 30 5.1
At2g16100 hypothetical protein 30 8.7
>At2g02650 putative reverse transcriptase
Length = 365
Score = 43.9 bits (102), Expect = 4e-04
Identities = 25/73 (34%), Positives = 36/73 (49%), Gaps = 2/73 (2%)
Query: 897 YGFRQVQFEADNESLIQRIKVGIDVDRSYLGLFLSEIHQLQLNFDICHFSFIPRDGNHTA 956
+G R V FE+D++SL+ I G D S LG + +I L C F+ R+ N A
Sbjct: 274 HGLRYVWFESDSKSLVTLINNG--EDHSLLGTLIYDIRHWMLKLPYCSLEFVNRERNSAA 331
Query: 957 HSLAQLAHSEPNL 969
+LA H+ L
Sbjct: 332 DALASHVHARDPL 344
>At2g15750 putative non-LTR retroelement reverse transcriptase
Length = 280
Score = 37.7 bits (86), Expect = 0.032
Identities = 21/64 (32%), Positives = 34/64 (52%), Gaps = 2/64 (3%)
Query: 898 GFRQVQFEADNESLIQRIKVGIDVDRSYLGLFLSEIHQLQLNFDICHFSFIPRDGNHTAH 957
G +++ F +D+ +L++ I L L +IH L ++FD C F FI R N A
Sbjct: 208 GIKRISFASDSLTLVKAINRKFLTKE--LHRILHDIHSLSISFDACSFFFISRKHNSEAD 265
Query: 958 SLAQ 961
+LA+
Sbjct: 266 ALAK 269
>At4g04000 putative reverse transcriptase
Length = 1077
Score = 37.0 bits (84), Expect = 0.054
Identities = 20/64 (31%), Positives = 36/64 (56%), Gaps = 2/64 (3%)
Query: 900 RQVQFEADNESLIQRIKVGIDVDRSYLGLFLSEIHQLQLNFDICHFSFIPRDGNHTAHSL 959
+ + F +D++ +++ + + V L L +I L +FD C F+FI R+ NH A +L
Sbjct: 1009 KAISFASDSQIIVKALNQKLQVKE--LHGILHDILSLSSSFDACTFTFINREFNHQADAL 1066
Query: 960 AQLA 963
A+ A
Sbjct: 1067 AKRA 1070
>At3g32340 hypothetical protein
Length = 871
Score = 36.6 bits (83), Expect = 0.071
Identities = 23/68 (33%), Positives = 35/68 (50%), Gaps = 2/68 (2%)
Query: 896 DYGFRQVQFEADNESLIQRIKVGIDVDRSYLGLFLSEIHQLQLNFDICHFSFIPRDGNHT 955
+ GF ++F +D+ LI+ ++V L +I L LNF + F+F PR N
Sbjct: 25 ELGFSHIKFASDSSQLIKA--QNLEVQSKELHEIHHDILALSLNFVVDSFNFTPRSSNLI 82
Query: 956 AHSLAQLA 963
A SLA+ A
Sbjct: 83 ADSLAKEA 90
>At4g03292 putative protein
Length = 142
Score = 35.0 bits (79), Expect = 0.21
Identities = 20/60 (33%), Positives = 26/60 (43%), Gaps = 2/60 (3%)
Query: 897 YGFRQVQFEADNESLIQRIKVGIDVDRSYLGLFLSEIHQLQLNFDICHFSFIPRDGNHTA 956
+G R V FE DN L+ + D D LG L +I L C F+ R+ N A
Sbjct: 64 HGLRYVWFETDNRDLVTLL--NNDEDHRLLGPLLCDIRYWMLKLPYCSIDFVNRERNSAA 121
>At2g45230 putative non-LTR retroelement reverse transcriptase
Length = 1374
Score = 35.0 bits (79), Expect = 0.21
Identities = 20/68 (29%), Positives = 37/68 (54%), Gaps = 2/68 (2%)
Query: 894 LKDYGFRQVQFEADNESLIQRIKVGIDVDRSYLGLFLSEIHQLQLNFDICHFSFIPRDGN 953
L + +R+V FE+D++ L+ I+ +D+ L + +I L +F+ F F R+GN
Sbjct: 1280 LSRFNYRRVIFESDSQYLVSLIQNEMDIPS--LAPRIQDIRNLLRHFEEVKFQFTRREGN 1337
Query: 954 HTAHSLAQ 961
+ A A+
Sbjct: 1338 NVADRTAR 1345
>At2g34320 putative non-LTR retroelement reverse transcriptase
Length = 292
Score = 34.3 bits (77), Expect = 0.35
Identities = 28/104 (26%), Positives = 49/104 (46%), Gaps = 3/104 (2%)
Query: 865 LWLQEPGLPLDLKRLLKQKLLRS**PCALLKDYGFRQVQFEADNESLIQRIKVGIDVDRS 924
LW+ LP K +L+ +L + + ++++ FE+D ++L+ + D
Sbjct: 174 LWMGARALPRT-KNVLEAELEALRWAVLTMSRFNYKRIIFESDAQALVNLLNS--DDFWP 230
Query: 925 YLGLFLSEIHQLQLNFDICHFSFIPRDGNHTAHSLAQLAHSEPN 968
L L +I QL +F+ F F PR GN A +A+ + S N
Sbjct: 231 TLQPALEDIQQLLHHFEEVKFEFTPRGGNKVADRIARESISFSN 274
>At2g15410 putative retroelement pol polyprotein
Length = 1787
Score = 34.3 bits (77), Expect = 0.35
Identities = 18/71 (25%), Positives = 36/71 (50%), Gaps = 1/71 (1%)
Query: 900 RQVQFEADNESLIQRIKVGIDVDRSYLGLFLSEIHQLQLNFDICHFSFIPRDGNHTAHSL 959
+++Q D++ + + + + +L + +L NF++ + IPR N A +L
Sbjct: 1215 KKIQAYCDSQLVASQFSGNYEAKNERMDAYLKVVRELSYNFEVFELTKIPRSDNAPADAL 1274
Query: 960 AQLAH-SEPNL 969
A LA S+P+L
Sbjct: 1275 AVLASTSDPDL 1285
>At2g13980 pseudogene
Length = 171
Score = 33.1 bits (74), Expect = 0.78
Identities = 21/65 (32%), Positives = 30/65 (45%), Gaps = 2/65 (3%)
Query: 898 GFRQVQFEADNESLIQRIKVGIDVDRSYLGLFLSEIHQLQLNFDICHFSFIPRDGNHTAH 957
G+ +V E D E+L + ++L L +I F FSF+ R GN AH
Sbjct: 82 GYLRVIMEGDCETLTNLVSGSSS--HNHLANLLDDIRIWAKKFSNVQFSFVRRGGNKVAH 139
Query: 958 SLAQL 962
LA+L
Sbjct: 140 ELAKL 144
>At1g27870 hypothetical protein
Length = 213
Score = 33.1 bits (74), Expect = 0.78
Identities = 21/67 (31%), Positives = 36/67 (53%), Gaps = 6/67 (8%)
Query: 897 YGFRQVQFEADNESLIQRIKVGIDVDRSYLGL--FLSEIHQLQLNFDICHFSFIPRDGNH 954
+G+++V FE DN+ L I + + Y G+ ++ +IH F+ HF++I R N
Sbjct: 120 HGYKRVCFEGDNKMLFDLI----NGSKVYFGVHNWIRDIHWWLRKFESIHFNWIRRHHNT 175
Query: 955 TAHSLAQ 961
A LA+
Sbjct: 176 PADILAK 182
>At2g11240 pseudogene
Length = 1044
Score = 32.7 bits (73), Expect = 1.0
Identities = 21/68 (30%), Positives = 37/68 (53%), Gaps = 2/68 (2%)
Query: 896 DYGFRQVQFEADNESLIQRIKVGIDVDRSYLGLFLSEIHQLQLNFDICHFSFIPRDGNHT 955
D GF+++ +D++ L++ + G G+ +I L LNF+ FSF+ R+ N
Sbjct: 968 DLGFKKLVVASDSQQLVKVLN-GEPHPMELHGIVF-DISVLSLNFEENSFSFVKRENNSK 1025
Query: 956 AHSLAQLA 963
A +LA+ A
Sbjct: 1026 ADALAKAA 1033
>At4g09710 RNA-directed DNA polymerase -like protein
Length = 1274
Score = 31.6 bits (70), Expect = 2.3
Identities = 24/68 (35%), Positives = 34/68 (49%), Gaps = 2/68 (2%)
Query: 898 GFRQVQFEADNESLIQRIKVGIDVDRSYLGLFLSEIHQLQLNFDICHFSFIPRDGNHTAH 957
G RQ+ +D + LI + G + L L +I +L ++F F FIPR N A
Sbjct: 1199 GVRQLNVFSDCKELISLLNSGKSIVE--LRGLLHDIRELSVSFTHLCFFFIPRLSNVVAD 1256
Query: 958 SLAQLAHS 965
SLA+ A S
Sbjct: 1257 SLAKSALS 1264
>At3g09510 putative non-LTR reverse transcriptase
Length = 484
Score = 31.6 bits (70), Expect = 2.3
Identities = 25/64 (39%), Positives = 30/64 (46%), Gaps = 2/64 (3%)
Query: 898 GFRQVQFEADNESLIQRIKVGIDVDRSYLGLFLSEIHQLQLNFDICHFSFIPRDGNHTAH 957
G+ QV E D ++LI I GI S L L +I F F FI R GN AH
Sbjct: 395 GYTQVFMEGDCQTLINLIN-GISFHSS-LANHLEDISFWANKFASIQFGFIRRKGNKLAH 452
Query: 958 SLAQ 961
LA+
Sbjct: 453 VLAK 456
>At2g31520 putative non-LTR retroelement reverse transcriptase
Length = 1524
Score = 31.6 bits (70), Expect = 2.3
Identities = 25/64 (39%), Positives = 30/64 (46%), Gaps = 2/64 (3%)
Query: 898 GFRQVQFEADNESLIQRIKVGIDVDRSYLGLFLSEIHQLQLNFDICHFSFIPRDGNHTAH 957
G+ QV E D ++LI I GI S L L +I F F FI R GN AH
Sbjct: 1435 GYTQVFMEGDCQTLINLIN-GISFHSS-LANHLEDISFWANKFASIQFGFIRRKGNKLAH 1492
Query: 958 SLAQ 961
LA+
Sbjct: 1493 VLAK 1496
>At2g15380 putative non-LTR retroelement reverse transcriptase
Length = 1311
Score = 31.6 bits (70), Expect = 2.3
Identities = 18/62 (29%), Positives = 32/62 (51%), Gaps = 2/62 (3%)
Query: 902 VQFEADNESLIQRIKVGIDVDRSYLGLFLSEIHQLQLNFDICHFSFIPRDGNHTAHSLAQ 961
+ F +D+ +L++ + + L L +I L F C F+FIP++ N A +LA+
Sbjct: 1244 ISFVSDSLTLVKALNSKLQSKE--LHGILHDILALSSTFTCCSFNFIPKENNRQADALAK 1301
Query: 962 LA 963
A
Sbjct: 1302 AA 1303
>At4g29090 putative protein
Length = 575
Score = 31.2 bits (69), Expect = 3.0
Identities = 27/96 (28%), Positives = 45/96 (46%), Gaps = 3/96 (3%)
Query: 866 WLQEPGLPLDLKRLLKQKLLRS**PCALLKDYGFRQVQFEADNESLIQRIKVGIDVDRSY 925
W+ LP LK +L+ +L L + + V FE+D++ LI+ + D
Sbjct: 458 WMGARALP-KLKSVLEAELEAMRWAVLSLSRFQYNYVIFESDSQVLIEILNN--DEIWPS 514
Query: 926 LGLFLSEIHQLQLNFDICHFSFIPRDGNHTAHSLAQ 961
L + ++ +L F F FIPR+GN A +A+
Sbjct: 515 LKPTIQDLQRLLSQFTEVKFVFIPREGNTLAERVAR 550
>At5g42965 putative protein
Length = 129
Score = 30.4 bits (67), Expect = 5.1
Identities = 17/65 (26%), Positives = 33/65 (50%), Gaps = 2/65 (3%)
Query: 897 YGFRQVQFEADNESLIQRIKVGIDVDRSYLGLFLSEIHQLQLNFDICHFSFIPRDGNHTA 956
+ +++V FE+D++ L+ + G D D L + +I + F+ I R+GN A
Sbjct: 33 FNYKKVIFESDSQHLVSLL--GDDTDYPNLAPRIQDIRHIFQQFEEVKVMHIKREGNSVA 90
Query: 957 HSLAQ 961
+A+
Sbjct: 91 DRIAR 95
>At2g25550 putative non-LTR retroelement reverse transcriptase
Length = 1750
Score = 30.4 bits (67), Expect = 5.1
Identities = 24/64 (37%), Positives = 30/64 (46%), Gaps = 2/64 (3%)
Query: 898 GFRQVQFEADNESLIQRIKVGIDVDRSYLGLFLSEIHQLQLNFDICHFSFIPRDGNHTAH 957
G+ QV E D ++LI I GI S L L +I F F FI + GN AH
Sbjct: 1661 GYTQVFMEGDCQTLINLIN-GISFHSS-LANHLEDISFWANKFASIQFGFIRKKGNKLAH 1718
Query: 958 SLAQ 961
LA+
Sbjct: 1719 VLAK 1722
>At2g16100 hypothetical protein
Length = 250
Score = 29.6 bits (65), Expect = 8.7
Identities = 19/67 (28%), Positives = 34/67 (50%), Gaps = 6/67 (8%)
Query: 897 YGFRQVQFEADNESLIQRIKVGIDVDRSYLGL--FLSEIHQLQLNFDICHFSFIPRDGNH 954
+G+++V FE DN L I + + + G+ ++ +IH F+ HF++I R N
Sbjct: 157 HGYKRVCFEEDNNMLFDLI----NGSKVHFGVHNWIRDIHWWSRKFESIHFNWIRRHHNT 212
Query: 955 TAHSLAQ 961
A A+
Sbjct: 213 PADIFAK 219
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.363 0.165 0.602
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,023,627
Number of Sequences: 26719
Number of extensions: 655804
Number of successful extensions: 2450
Number of sequences better than 10.0: 19
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 14
Number of HSP's that attempted gapping in prelim test: 2438
Number of HSP's gapped (non-prelim): 24
length of query: 974
length of database: 11,318,596
effective HSP length: 109
effective length of query: 865
effective length of database: 8,406,225
effective search space: 7271384625
effective search space used: 7271384625
T: 11
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (22.0 bits)
S2: 65 (29.6 bits)
Medicago: description of AC147429.6


