

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147179.2 + phase: 0
(185 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
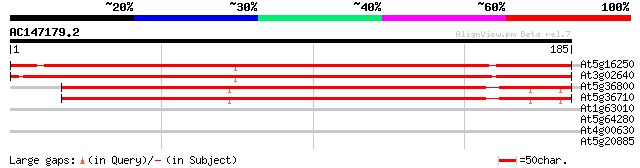
Score E
Sequences producing significant alignments: (bits) Value
At5g16250 unknown protein 255 8e-69
At3g02640 Unknown protein (F16B3.27) 243 3e-65
At5g36800 putative protein 200 3e-52
At5g36710 putative protein 200 3e-52
At1g63010 tetracycline resistance efflux protein like protein 32 0.25
At5g64280 2-oxoglutarate/malate translocator 29 1.3
At4g00630 putative potassium/H+ antiporter 27 6.3
At5g20885 unknown protein 27 8.2
>At5g16250 unknown protein
Length = 183
Score = 255 bits (652), Expect = 8e-69
Identities = 128/187 (68%), Positives = 149/187 (79%), Gaps = 6/187 (3%)
Query: 1 MGLSMNNPPATITPRHYNIHKLFLFCNYVLLGAASSCIFLTLSLRLVPSVCGFFLILLHI 60
MG ++ P + HY+ HK+FLF NY+LLGAASSCIFLTLSLRL+PS+CGF LILLH
Sbjct: 1 MGFISSSSP--VEESHYHTHKIFLFSNYILLGAASSCIFLTLSLRLIPSICGFLLILLHA 58
Query: 61 FTIAGAVSGCAAV--GANRWYSAHMVATVLTAIFQGSVSVLVFTRTSDFLGELQSYVREE 118
TIA AVSGCAA G NRWY+AHMVATVLTAIFQGSVSVL+FT TS FLG L+SYVREE
Sbjct: 59 TTIAAAVSGCAAASCGRNRWYAAHMVATVLTAIFQGSVSVLIFTNTSKFLGSLKSYVREE 118
Query: 119 DGSVILKLSGGLAILIFCLEWVVLTLAFFLKYYACVEGGNTCRTVVLGSAKVQQDEDLKD 178
D +VILKL GGL I+IFCL+W+VL AFFLKYYA V+GG+ + + KVQ +E+ KD
Sbjct: 119 DAAVILKLGGGLCIVIFCLDWIVLVCAFFLKYYAYVDGGD--GVAMKRTGKVQSEENPKD 176
Query: 179 WPWPFQV 185
WPWPFQV
Sbjct: 177 WPWPFQV 183
>At3g02640 Unknown protein (F16B3.27)
Length = 185
Score = 243 bits (621), Expect = 3e-65
Identities = 126/187 (67%), Positives = 147/187 (78%), Gaps = 4/187 (2%)
Query: 1 MGLSMNNPPATITPRHYNIHKLFLFCNYVLLGAASSCIFLTLSLRLVPSVCGFFLILLHI 60
MGL + P +I HY HKLFL NYVLLGA+SSCIFLTLSLRL+PS+CGFFLILLH
Sbjct: 1 MGL-IPQPQESIQESHYYTHKLFLTANYVLLGASSSCIFLTLSLRLIPSLCGFFLILLHA 59
Query: 61 FTIAGAVSGCAAV--GANRWYSAHMVATVLTAIFQGSVSVLVFTRTSDFLGELQSYVREE 118
TIA AVSGCAA G NRWY+AHM+ATVLTAIFQGSVSVL+FT TS+FL L SYVRE+
Sbjct: 60 TTIAAAVSGCAAASYGKNRWYAAHMIATVLTAIFQGSVSVLIFTNTSNFLESLNSYVREK 119
Query: 119 DGSVILKLSGGLAILIFCLEWVVLTLAFFLKYYACVEGGNTCRTVVLGSAKVQQDEDLKD 178
+ S+ILKL+GGL ++IFCLEW+VL LAFFLKYYA V+G N + + KVQ +E LK+
Sbjct: 120 EASMILKLAGGLCVVIFCLEWIVLVLAFFLKYYAYVDGDNN-GVAMKRTGKVQSEETLKN 178
Query: 179 WPWPFQV 185
PW FQV
Sbjct: 179 SPWAFQV 185
>At5g36800 putative protein
Length = 183
Score = 200 bits (509), Expect = 3e-52
Identities = 109/177 (61%), Positives = 131/177 (73%), Gaps = 13/177 (7%)
Query: 18 NIHKLFLFCNYVLLGAASSCIFLTLSLRLVPSVCGFFLILLHIFTIAGAVSGCA------ 71
N H +FL CNY+LLG+ASSCIFLT+SLRL PS+ G LI L+ TIA AVSGC+
Sbjct: 9 NTHNIFLLCNYILLGSASSCIFLTISLRLFPSLSGLSLIFLYTLTIATAVSGCSIFASST 68
Query: 72 -AVGANRWYSAHMVATVLTAIFQGSVSVLVFTRTSDFLGELQSYVREEDGSVILKLSGGL 130
A ++R Y +HMVATVLTAIFQG+VSVL+FTRT DFL L+SYVREEDG VILKLSGGL
Sbjct: 69 SATASDRLYGSHMVATVLTAIFQGAVSVLIFTRTGDFLRFLKSYVREEDGEVILKLSGGL 128
Query: 131 AILIFCLEWVVLTLAFFLKYYACVEGGNTCRTVVLGSAKV-QQDEDLKDWP-WPFQV 185
+L+FCLEW+VL LAF LKY ++ V KV +Q+EDLKDWP +PFQ+
Sbjct: 129 CVLMFCLEWIVLVLAFLLKYSDYLDES----VVDDDDFKVRRQEEDLKDWPSYPFQL 181
>At5g36710 putative protein
Length = 218
Score = 200 bits (509), Expect = 3e-52
Identities = 109/177 (61%), Positives = 131/177 (73%), Gaps = 13/177 (7%)
Query: 18 NIHKLFLFCNYVLLGAASSCIFLTLSLRLVPSVCGFFLILLHIFTIAGAVSGCA------ 71
N H +FL CNY+LLG+ASSCIFLT+SLRL PS+ G LI L+ TIA AVSGC+
Sbjct: 44 NTHNIFLLCNYILLGSASSCIFLTISLRLFPSLSGLSLIFLYTLTIATAVSGCSIFASST 103
Query: 72 -AVGANRWYSAHMVATVLTAIFQGSVSVLVFTRTSDFLGELQSYVREEDGSVILKLSGGL 130
A ++R Y +HMVATVLTAIFQG+VSVL+FTRT DFL L+SYVREEDG VILKLSGGL
Sbjct: 104 SATASDRLYGSHMVATVLTAIFQGAVSVLIFTRTGDFLRFLKSYVREEDGEVILKLSGGL 163
Query: 131 AILIFCLEWVVLTLAFFLKYYACVEGGNTCRTVVLGSAKV-QQDEDLKDWP-WPFQV 185
+L+FCLEW+VL LAF LKY ++ V KV +Q+EDLKDWP +PFQ+
Sbjct: 164 CVLMFCLEWIVLVLAFLLKYSDYLDES----VVDDDDFKVRRQEEDLKDWPSYPFQL 216
>At1g63010 tetracycline resistance efflux protein like protein
Length = 699
Score = 31.6 bits (70), Expect = 0.25
Identities = 29/117 (24%), Positives = 53/117 (44%), Gaps = 5/117 (4%)
Query: 12 ITPRHYNIHKLFLFCNYVLLGAASSCIFLTLSLRLVPSVCGFFLILLHIFTIAGAVSGCA 71
+TP I + N +LL ++ +++ + +VP+ + + L T+ G V G
Sbjct: 233 VTPSEDTIDERSYHFNSLLLNLGNTFLYMVNTYIIVPTADDYSMSLGAAATVCGVVIGSM 292
Query: 72 AVGANRWYSAHMVATVLTAIFQGSVSVLVFTRTSDFLGELQSYVREEDGSVILKLSG 128
AV A + S + A + F+ LVF+ + F+G L + + S+ L L G
Sbjct: 293 AV-AQVFSSVYFSAWSNKSYFK----PLVFSSIALFIGNLMYALAYDANSIALLLLG 344
>At5g64280 2-oxoglutarate/malate translocator
Length = 549
Score = 29.3 bits (64), Expect = 1.3
Identities = 17/51 (33%), Positives = 27/51 (52%), Gaps = 2/51 (3%)
Query: 52 GFFLILLHIFTIAGAVSGCAAVGANRWYSAHMVATVLTAIFQGSVSVLVFT 102
G+ L+ + +FTI+G V G VGA W + A+++T S + FT
Sbjct: 109 GWQLLSIFLFTISGLVLGPLPVGA--WAFIGLTASIVTKTLPFSTAFAAFT 157
>At4g00630 putative potassium/H+ antiporter
Length = 617
Score = 26.9 bits (58), Expect = 6.3
Identities = 36/140 (25%), Positives = 56/140 (39%), Gaps = 19/140 (13%)
Query: 8 PPATITPRHYNIHKLFLFCNYVLLGAASSCIFLTLSLRLVPSVCGF----FLILLHIFTI 63
PP IT R F C L A+ SC++ L + P V G+ LI + +I
Sbjct: 7 PPTFITLRLMRRKLPFSMCCGYCLQASYSCLYFRKFLEVTP-VLGYLAAGILIGPYGLSI 65
Query: 64 AGAVSGCAAV----------GANRWYSAHMVATVLTAIF-QGSVSVLVFTRTSDFLGELQ 112
V G A+ S ++++ +F GS VLV T+ +G +
Sbjct: 66 IRNVHGTKAIAEFGVVFLLFNIGLELSVERLSSMKKYVFGLGSAQVLV---TAAVIGLIT 122
Query: 113 SYVREEDGSVILKLSGGLAI 132
YV + G + + GLA+
Sbjct: 123 HYVAGQAGPAAIVIGNGLAL 142
>At5g20885 unknown protein
Length = 176
Score = 26.6 bits (57), Expect = 8.2
Identities = 13/38 (34%), Positives = 23/38 (60%), Gaps = 3/38 (7%)
Query: 113 SYVREEDGSVILKLSGGLAILIFCLEWVVLTLAFFLKY 150
S+ E+ G ++ +L +A+LI L W+ LA+ L+Y
Sbjct: 2 SFFIEDSGLIVTQLLYKMAVLITVLRWI---LAWILRY 36
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.328 0.142 0.449
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,054,729
Number of Sequences: 26719
Number of extensions: 151102
Number of successful extensions: 473
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 461
Number of HSP's gapped (non-prelim): 8
length of query: 185
length of database: 11,318,596
effective HSP length: 93
effective length of query: 92
effective length of database: 8,833,729
effective search space: 812703068
effective search space used: 812703068
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 57 (26.6 bits)
Medicago: description of AC147179.2


