

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146788.10 - phase: 0
(496 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
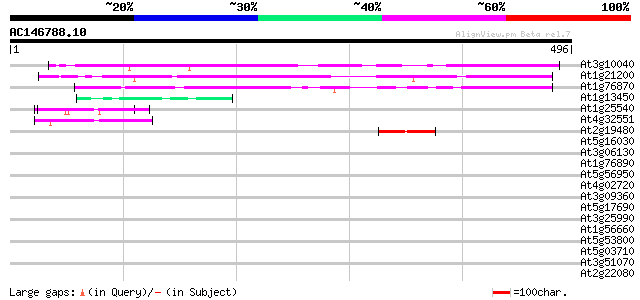
Score E
Sequences producing significant alignments: (bits) Value
At3g10040 unknown protein 382 e-106
At1g21200 unknown protein 272 3e-73
At1g76870 hypothetical protein 237 1e-62
At1g13450 hypothetical protein 47 3e-05
At1g25540 hypothetical protein 45 7e-05
At4g32551 Leunig protein 43 5e-04
At2g19480 putative nucleosome assembly protein 43 5e-04
At5g16030 unknown protein 42 0.001
At3g06130 unknown protein 42 0.001
At1g76890 unknown protein 41 0.002
At5g56950 nucleosome assembly protein 40 0.002
At4g02720 unknown protein 40 0.002
At3g09360 putative transcription factor 40 0.002
At5g17690 chromo domain protein 40 0.003
At3g25990 putative DNA-binding protein, GT-1 40 0.003
At1g56660 hypothetical protein 40 0.003
At5g53800 unknown protein 40 0.004
At5g03710 putative protein 40 0.004
At3g51070 putative protein 40 0.004
At2g22080 En/Spm-like transposon protein 40 0.004
>At3g10040 unknown protein
Length = 418
Score = 382 bits (981), Expect = e-106
Identities = 224/467 (47%), Positives = 276/467 (58%), Gaps = 84/467 (17%)
Query: 35 QNLPNQQQQNPHQLHHSQVVSYAPQHHDTDTNQHHHHSMKHGFPPFSSSKNKQQQQSSQM 94
QN PN Q N Q H P + T +Q +K +P S K
Sbjct: 8 QNPPNPQ--NSIQFQH-------PHPYTTSGDQQTQPPIKSLYPYASKPKQMSPISGGGC 58
Query: 95 SDEDEPNFPA-----EESSGGDPKRKISPWQRMKWTDTMVRLLIMAVYYIGDEAGSEGTD 149
DED + E+S+G D KRK+S W RMKWTDTMVRLLIMAV+YIGDEAG
Sbjct: 59 DDEDRGSGSGSGCNPEDSAGTDGKRKLSQWHRMKWTDTMVRLLIMAVFYIGDEAGLNDPV 118
Query: 150 PNKKKSSG---------LLQKKGKWKSVSRAMMEKGFYVSPQQCEDKFNDLNKRYKRVND 200
KKK+ G +LQKKGKWKSVSRAM+EKGF VSPQQCEDKFNDLNKRYKRVND
Sbjct: 119 DAKKKTGGGGGGGGGGGMLQKKGKWKSVSRAMVEKGFSVSPQQCEDKFNDLNKRYKRVND 178
Query: 201 ILGKGTACRVVENQGLLDSMD-LSPKMKDEVRKLLNSKHLFFREMCAYHNSCGHGGVATS 259
ILGKG ACRVVENQGLL+SMD L+PK+KDEV+KLLNSKHLFFREMCAYHNSCGH G
Sbjct: 179 ILGKGIACRVVENQGLLESMDHLTPKLKDEVKKLLNSKHLFFREMCAYHNSCGHLG---- 234
Query: 260 NVQHQVEGGSGTTTPPQNQPQQQHQHQHQQQNQQHCFHSSENGVGSLGVSRGEGLRMLKI 319
Q PQQ QQ+CFH++E G + R E
Sbjct: 235 -------------GHDQQPPQQNPISIPIPSQQQNCFHAAEAGKMARIAERVE------- 274
Query: 320 GSGYVEEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEENDGKPSRKRARKGGFS 379
VEEE + D ++SE +E E+E +RK+ R
Sbjct: 275 ----VEEEVESDMAEDSESEMEESEEEE---------------------TRKKRR----- 304
Query: 380 FPRSSSSTQLVNQMNNEISGVFQDGGKSTWEKKQWIRNRIMQLEEQKIGYESQAFQLEKQ 439
+ V ++ E + V +D GKS WEKK+WIR +++++EE+KIGYE + ++EKQ
Sbjct: 305 ------ISTAVKRLREEAASVVEDVGKSVWEKKEWIRRKMLEIEEKKIGYEWEGVEMEKQ 358
Query: 440 RLKWARYSSKKEREMERAKLENERRRLENERMVLLIRKKELELMHIQ 486
R+KW RY SKKEREME+AKL+N+RRRLE ERM+L++R+ E+EL +Q
Sbjct: 359 RVKWMRYRSKKEREMEKAKLDNQRRRLETERMILMLRRSEIELNELQ 405
>At1g21200 unknown protein
Length = 443
Score = 272 bits (696), Expect = 3e-73
Identities = 168/472 (35%), Positives = 252/472 (52%), Gaps = 80/472 (16%)
Query: 26 HQQQQQQKQQNLPNQQQQNPHQLHHSQVVSYAPQHHDTDTNQHHHHSMKHGFPPFSSSKN 85
HQ Q+ ++ PN + + L + V HH Q+ + SM S +
Sbjct: 31 HQDSMNQQHRHNPNSRPLH-EGLPFTMVTGQTCDHH-----QNQNMSM--------SEQQ 76
Query: 86 KQQQQSSQMSDEDEPNFPAEESSG----GDPKRKISPWQRMKWTDTMVRLLIMAVYYIGD 141
K +++ + +SD+DEP+F E G + K SPWQR+KWTD MV+LLI AV YIGD
Sbjct: 77 KAEREKNSVSDDDEPSFTEEGGDGVHNEANRSTKGSPWQRVKWTDKMVKLLITAVSYIGD 136
Query: 142 EAGSEGTDPNKKKSSGLLQKKGKWKSVSRAMMEKGFYVSPQQCEDKFNDLNKRYKRVNDI 201
++ D + ++ +LQKKGKWKSVS+ M E+G++VSPQQCEDKFNDLNKRYK++ND+
Sbjct: 137 DSS---IDSSSRRKFAVLQKKGKWKSVSKVMAERGYHVSPQQCEDKFNDLNKRYKKLNDM 193
Query: 202 LGKGTACRVVENQGLLDSMD-LSPKMKDEVRKLLNSKHLFFREMCAYHNSCGHGGVATSN 260
LG+GT+C+VVEN LLDS+ L+ K KD+VRK+++SKHLF+ EMC+YHN
Sbjct: 194 LGRGTSCQVVENPALLDSIGYLNDKEKDDVRKIMSSKHLFYEEMCSYHNGNRLHLPHDLA 253
Query: 261 VQHQVEGGSGTTTPPQNQPQQQHQHQHQQQNQQHCFHSSENGVGSLGVSRGEGLRMLKIG 320
+Q ++ + N ++HQ
Sbjct: 254 LQRSLQLALRSRDDHDNDDSRKHQ------------------------------------ 277
Query: 321 SGYVEEEDDEDEEDESEDFSDEGEDESGEG-CSKGH----------INDQDEEENDGKPS 369
+E+ DDED + + ++ + E G C H I E+ PS
Sbjct: 278 ---MEDLDDEDHDGDGDEHDEYEEQHYAYGDCRVNHYGGGGGPLKKIRPSLSHEDGDHPS 334
Query: 370 RKRARK-GGFSFPRSSSSTQLVNQMNNEISGVFQDGGKSTWEKKQWIRNRIMQLEEQKIG 428
+ + S P+ S VNQ E G++ +KQW+ +R +QLEEQK+
Sbjct: 335 HVNSLECNKVSLPQIPFSQADVNQGGAE-------SGRAGSVQKQWMESRTLQLEEQKLQ 387
Query: 429 YESQAFQLEKQRLKWARYSSKKEREMERAKLENERRRLENERMVLLIRKKEL 480
+ + +LEKQR +W R+S K+++E+ER ++ENER +LEN+RM L ++++EL
Sbjct: 388 IQVELLELEKQRFRWQRFSKKRDQELERMRMENERMKLENDRMGLELKQREL 439
>At1g76870 hypothetical protein
Length = 385
Score = 237 bits (605), Expect = 1e-62
Identities = 148/432 (34%), Positives = 232/432 (53%), Gaps = 81/432 (18%)
Query: 58 PQHHDTDTNQHHHHSMKH-GFPPFSSSK-NKQQQQSSQMSDEDEPNFPAEESSGGDPKRK 115
P + + QHH +S + GF ++ N + MS++DE S G + ++
Sbjct: 22 PNAINQNQKQHHPNSRQDSGFNNTMDTRHNNVDRGKKSMSEDDE--LCLLSSDGQNKSKE 79
Query: 116 ISPWQRMKWTDTMVRLLIMAVYYIGDEAGSEGTDPNKKKSSGLLQKKGKWKSVSRAMMEK 175
SPWQR+KW D MV+L+I A+ YIG+++GS+ K +LQKKGKW+SVS+ M E+
Sbjct: 80 NSPWQRVKWMDKMVKLMITALSYIGEDSGSD-------KKFAVLQKKGKWRSVSKVMDER 132
Query: 176 GFYVSPQQCEDKFNDLNKRYKRVNDILGKGTACRVVENQGLLDSMD-LSPKMKDEVRKLL 234
G++VSPQQCEDKFNDLNKRYK++N++LG+GT+C VVEN LLD +D L+ K KDEVR+++
Sbjct: 133 GYHVSPQQCEDKFNDLNKRYKKLNEMLGRGTSCEVVENPSLLDKIDYLNEKEKDEVRRIM 192
Query: 235 NSKHLFFREMCAYHNSCGHGGVATSNVQHQVEGGSGTTTPPQNQPQQQHQH------QHQ 288
+SKHLF+ EMC+YHN N H P + Q+ H +
Sbjct: 193 SSKHLFYEEMCSYHN---------GNRLHL----------PHDPAVQRSLHLITLGSRDD 233
Query: 289 QQNQQHCFHSSENGVGSLGVSRGEGLRMLKIGSGYVEEEDDEDEEDESEDFSDEGEDESG 348
N +H H +E+ ++DD+ EED SD
Sbjct: 234 HDNDEHGKHQNED-----------------------LDDDDDYEEDHDGALSDR------ 264
Query: 349 EGCSKGHINDQDEEENDGKPSRKRARKGGFSFPRSSSSTQLVNQMNNEISGVFQDGGKST 408
+ E+ G P++ G+ P S VN+ G+ D K+
Sbjct: 265 ---PLKRLRQSQSHEDVGHPNK------GYDVPCLPRSQADVNR------GISLDSRKAA 309
Query: 409 WEKKQWIRNRIMQLEEQKIGYESQAFQLEKQRLKWARYSSKKEREMERAKLENERRRLEN 468
++Q I ++ ++LE +K+ +++ +LE+Q+ KW +S ++E+++ + ++ENER +LEN
Sbjct: 310 GLQRQQIESKSLELEGRKLQIQAEMMELERQQFKWEVFSKRREQKLAKMRMENERMKLEN 369
Query: 469 ERMVLLIRKKEL 480
ERM L +++ EL
Sbjct: 370 ERMSLELKRIEL 381
>At1g13450 hypothetical protein
Length = 406
Score = 46.6 bits (109), Expect = 3e-05
Identities = 38/138 (27%), Positives = 52/138 (37%), Gaps = 27/138 (19%)
Query: 60 HHDTDTNQHHHHSMKHGFPPFSSSKNKQQQQSSQMSDEDEPNFPAEESSGGDPKRKISPW 119
+ D HHHH+ N QS + + ESSG D + K
Sbjct: 38 NESVDLQSHHHHN----------HHNHHLHQS-----QPQQQILLGESSGEDHEVKAPKK 82
Query: 120 QRMKWTDTMVRLLIMAVYYIGDEAGSEGTDPNKKKSSGLLQKKGKWKSVSRAMMEKGFYV 179
+ W R LIM G +G K + L W+ +S M EKGF
Sbjct: 83 RAETWVQDETRSLIMF------RRGMDGLFNTSKSNKHL------WEQISSKMREKGFDR 130
Query: 180 SPQQCEDKFNDLNKRYKR 197
SP C DK+ +L K +K+
Sbjct: 131 SPTMCTDKWRNLLKEFKK 148
>At1g25540 hypothetical protein
Length = 809
Score = 45.4 bits (106), Expect = 7e-05
Identities = 34/100 (34%), Positives = 43/100 (43%), Gaps = 17/100 (17%)
Query: 23 IPLHQQQQQQKQQNLPNQQQQNPHQL----HH---SQVVSYAPQHHDTDTNQHHHHSMKH 75
+P QQQQQQ Q QQQQ HQL HH Q Q H QHHH +
Sbjct: 687 MPQLQQQQQQHQ-----QQQQQQHQLSQLQHHQQQQQQQQQQQQQHQLTQLQHHHQQQQQ 741
Query: 76 GFP-----PFSSSKNKQQQQSSQMSDEDEPNFPAEESSGG 110
P +S N+ QQQ+S ++ + P + GG
Sbjct: 742 ASPLNQMQQQTSPLNQMQQQTSPLNQMQQQQQPQQMVMGG 781
Score = 44.7 bits (104), Expect = 1e-04
Identities = 29/100 (29%), Positives = 47/100 (47%), Gaps = 3/100 (3%)
Query: 25 LHQQQQQQKQQNLPNQQQQNPHQLHHSQVVSYAPQHHDTDTNQHHHHSMKHGFPPFSSSK 84
LHQQQQQQ+Q QQQQ+ Q Q+ QH QH ++H +
Sbjct: 662 LHQQQQQQQQIQQQQQQQQHLQQQQMPQLQQQQQQHQQQQQQQHQLSQLQH--HQQQQQQ 719
Query: 85 NKQQQQSSQMSDEDEPNFPAEESSG-GDPKRKISPWQRMK 123
+QQQQ Q++ + +++S +++ SP +M+
Sbjct: 720 QQQQQQQHQLTQLQHHHQQQQQASPLNQMQQQTSPLNQMQ 759
Score = 38.5 bits (88), Expect = 0.009
Identities = 26/97 (26%), Positives = 39/97 (39%), Gaps = 1/97 (1%)
Query: 3 EANAMFSTITPSGLLGVVDNIPLHQQQQQQKQQNLPNQQQQNPHQLHHSQVVSYAPQHHD 62
+A + + P ++ IP +QQQQQQ+Q + QQQQ Q Q Q
Sbjct: 631 KACRLIGMLFPGDMVVFKPQIP-NQQQQQQQQLHQQQQQQQQIQQQQQQQQHLQQQQMPQ 689
Query: 63 TDTNQHHHHSMKHGFPPFSSSKNKQQQQSSQMSDEDE 99
Q H + S ++ QQQQ Q + +
Sbjct: 690 LQQQQQQHQQQQQQQHQLSQLQHHQQQQQQQQQQQQQ 726
Score = 37.7 bits (86), Expect = 0.015
Identities = 27/90 (30%), Positives = 35/90 (38%), Gaps = 8/90 (8%)
Query: 25 LHQQQQQQKQQNLPNQQQQNPHQL----HHSQVVSYAPQHHDTDTNQHHHHSMKHGFPPF 80
L QQQQ+QQ QQQQ HQL HH Q A + + M+ P
Sbjct: 710 LQHHQQQQQQQ----QQQQQQHQLTQLQHHHQQQQQASPLNQMQQQTSPLNQMQQQTSPL 765
Query: 81 SSSKNKQQQQSSQMSDEDEPNFPAEESSGG 110
+ + +QQ Q M + P GG
Sbjct: 766 NQMQQQQQPQQMVMGGQAFAQAPGRSQQGG 795
Score = 31.6 bits (70), Expect = 1.1
Identities = 18/85 (21%), Positives = 46/85 (53%), Gaps = 2/85 (2%)
Query: 412 KQWIRNRIMQLEEQKIGYESQAFQLEKQRLKWARYSSKKEREMERAKLENERRRLENERM 471
K I N+ Q ++Q + Q Q+++Q+ + ++ ++++ + ++++++ + ++
Sbjct: 648 KPQIPNQQQQQQQQLHQQQQQQQQIQQQQQQQQHLQQQQMPQLQQQQQQHQQQQQQQHQL 707
Query: 472 VLLIRKKELELMHIQQQQQQQHSST 496
L ++ + QQQQQQQH T
Sbjct: 708 SQLQHHQQQQQQ--QQQQQQQHQLT 730
Score = 29.6 bits (65), Expect = 4.0
Identities = 13/20 (65%), Positives = 13/20 (65%)
Query: 274 PPQNQPQQQHQHQHQQQNQQ 293
P Q Q QQQ HQ QQQ QQ
Sbjct: 652 PNQQQQQQQQLHQQQQQQQQ 671
Score = 28.5 bits (62), Expect = 8.9
Identities = 12/19 (63%), Positives = 13/19 (68%)
Query: 274 PPQNQPQQQHQHQHQQQNQ 292
P Q QQQHQ Q QQQ+Q
Sbjct: 688 PQLQQQQQQHQQQQQQQHQ 706
>At4g32551 Leunig protein
Length = 931
Score = 42.7 bits (99), Expect = 5e-04
Identities = 30/107 (28%), Positives = 46/107 (42%), Gaps = 6/107 (5%)
Query: 23 IPLHQQQQQQKQ--QNLPNQQQQNPHQLHHSQ-VVSYAPQHHDTDTNQHHHHSMKHGFPP 79
I +QQ QQ Q Q QQQQ Q+ Q ++ A Q QHHHH +
Sbjct: 84 IKAREQQLQQSQHPQVSQQQQQQQQQQIQMQQLLLQRAQQQQQQQQQQHHHHQQQQ---Q 140
Query: 80 FSSSKNKQQQQSSQMSDEDEPNFPAEESSGGDPKRKISPWQRMKWTD 126
+ +QQQQ Q P+ ++ S +++ +P Q+ + D
Sbjct: 141 QQQQQQQQQQQQQQQHQNQPPSQQQQQQSTPQHQQQPTPQQQPQRRD 187
Score = 33.1 bits (74), Expect = 0.36
Identities = 25/95 (26%), Positives = 36/95 (37%)
Query: 27 QQQQQQKQQNLPNQQQQNPHQLHHSQVVSYAPQHHDTDTNQHHHHSMKHGFPPFSSSKNK 86
QQQQQQ+QQ QQQQ+ +Q Q + H + G + S N
Sbjct: 139 QQQQQQQQQQQQQQQQQHQNQPPSQQQQQQSTPQHQQQPTPQQQPQRRDGSHLANGSANG 198
Query: 87 QQQQSSQMSDEDEPNFPAEESSGGDPKRKISPWQR 121
+S+ P + +S +R P QR
Sbjct: 199 LVGNNSEPVMRQNPGSGSSLASKAYEERVKMPTQR 233
Score = 32.7 bits (73), Expect = 0.47
Identities = 26/97 (26%), Positives = 35/97 (35%), Gaps = 23/97 (23%)
Query: 27 QQQQQQKQQNLP--------------NQQQQNPHQLHHSQVVSYAPQHHDTDTNQHHHHS 72
QQQQQQ+QQ + QQQQ+ H Q Q QH +
Sbjct: 101 QQQQQQQQQQIQMQQLLLQRAQQQQQQQQQQHHHHQQQQQQQQQQQQQQQQQQQQHQNQ- 159
Query: 73 MKHGFPPFSSSKNKQQQQSSQMSDEDEPNFPAEESSG 109
PP S+ +QQQ + Q + P + G
Sbjct: 160 -----PP---SQQQQQQSTPQHQQQPTPQQQPQRRDG 188
Score = 30.4 bits (67), Expect = 2.3
Identities = 13/19 (68%), Positives = 13/19 (68%)
Query: 276 QNQPQQQHQHQHQQQNQQH 294
Q Q QQQ Q Q QQQ QQH
Sbjct: 138 QQQQQQQQQQQQQQQQQQH 156
Score = 30.0 bits (66), Expect = 3.1
Identities = 18/70 (25%), Positives = 30/70 (42%), Gaps = 1/70 (1%)
Query: 27 QQQQQQKQQNLPNQQQQNPHQLHHSQVVSYAPQHHDTDTNQHHHHSMKHGFPPFSSSKNK 86
QQQQQQ+ + QQQQ Q Q Q+ Q + +H P + +
Sbjct: 125 QQQQQQQHHHHQQQQQQQQQQQQQQQQQQQQHQNQPPSQQQQQQSTPQHQQQP-TPQQQP 183
Query: 87 QQQQSSQMSD 96
Q++ S +++
Sbjct: 184 QRRDGSHLAN 193
Score = 29.6 bits (65), Expect = 4.0
Identities = 12/18 (66%), Positives = 12/18 (66%)
Query: 276 QNQPQQQHQHQHQQQNQQ 293
Q Q QQQ H HQQQ QQ
Sbjct: 124 QQQQQQQQHHHHQQQQQQ 141
Score = 29.6 bits (65), Expect = 4.0
Identities = 12/18 (66%), Positives = 12/18 (66%)
Query: 276 QNQPQQQHQHQHQQQNQQ 293
Q Q QQQH H QQQ QQ
Sbjct: 125 QQQQQQQHHHHQQQQQQQ 142
Score = 29.6 bits (65), Expect = 4.0
Identities = 12/18 (66%), Positives = 12/18 (66%)
Query: 276 QNQPQQQHQHQHQQQNQQ 293
Q Q QQQ QH H QQ QQ
Sbjct: 123 QQQQQQQQQHHHHQQQQQ 140
Score = 29.6 bits (65), Expect = 4.0
Identities = 12/18 (66%), Positives = 12/18 (66%)
Query: 276 QNQPQQQHQHQHQQQNQQ 293
Q Q QQ H HQ QQQ QQ
Sbjct: 126 QQQQQQHHHHQQQQQQQQ 143
Score = 29.3 bits (64), Expect = 5.2
Identities = 17/105 (16%), Positives = 52/105 (49%), Gaps = 14/105 (13%)
Query: 391 NQMNNEISGVFQDGGKSTWEKKQWIRNRIMQLEEQKIGYESQAFQLEKQRLKWARYSSKK 450
N+ ++E++ + + ++Q +++ Q+ +Q+ + Q Q+++ L+ A+ ++
Sbjct: 68 NEKHSEVAASYIETQMIKAREQQLQQSQHPQVSQQQQQQQQQQIQMQQLLLQRAQQQQQQ 127
Query: 451 EREMERAKLENERRRLENERMVLLIRKKELELMHIQQQQQQQHSS 495
+++ + ++++ + ++ QQQQQQQH +
Sbjct: 128 QQQQHHHHQQQQQQQQQQQQQ--------------QQQQQQQHQN 158
Score = 29.3 bits (64), Expect = 5.2
Identities = 14/63 (22%), Positives = 38/63 (60%)
Query: 430 ESQAFQLEKQRLKWARYSSKKEREMERAKLENERRRLENERMVLLIRKKELELMHIQQQQ 489
E+Q + +Q+L+ +++ +++ ++ + + + ++L +R ++++ + H QQQQ
Sbjct: 80 ETQMIKAREQQLQQSQHPQVSQQQQQQQQQQIQMQQLLLQRAQQQQQQQQQQHHHHQQQQ 139
Query: 490 QQQ 492
QQQ
Sbjct: 140 QQQ 142
Score = 28.9 bits (63), Expect = 6.8
Identities = 20/52 (38%), Positives = 25/52 (47%), Gaps = 10/52 (19%)
Query: 262 QHQVEGGSGTTTPPQNQPQQQHQHQ-----HQQQNQQHCFHSSE--NGVGSL 306
Q+Q +GG PPQ QPQ Q +Q Q Q+ H H E G GS+
Sbjct: 453 QNQQQGGGN---PPQPQPQPQPLNQLALTNPQPQSSNHSIHQQEKLGGGGSI 501
Score = 28.5 bits (62), Expect = 8.9
Identities = 56/262 (21%), Positives = 93/262 (35%), Gaps = 45/262 (17%)
Query: 262 QHQVEGGSGTTTPPQNQPQQQHQHQHQQQ----------NQQHCFHSSENGV---GSLGV 308
Q Q + PP Q QQQ QHQQQ + H + S NG+ S V
Sbjct: 148 QQQQQQQQHQNQPPSQQQQQQSTPQHQQQPTPQQQPQRRDGSHLANGSANGLVGNNSEPV 207
Query: 309 SRGEGLRMLKIGSGYVEEEDDEDEEDESEDFS--DEGEDESGEGCSKGHINDQDEEENDG 366
R + S EE + ES D + D G+ H + G
Sbjct: 208 MRQNPGSGSSLASKAYEERVKMPTQRESLDEAAMKRFGDNVGQLLDPSHASILKSAAASG 267
Query: 367 KPSRK--RARKGGFSFPRSSSSTQLVN---QMNNEISGVF----------------QDGG 405
+P+ + + GG S + + QL + +EI+ V + G
Sbjct: 268 QPAGQVLHSTSGGMSPQVQTRNQQLPGSAVDIKSEINPVLTPRTAVPEGSLIGIPGSNQG 327
Query: 406 KSTWEKKQWIRNRIMQ-----LEEQKIGYESQAFQLEKQRLKWARYSSKKEREMERAKLE 460
+ K W Q L++QK +SQ+F +L +++ + + L
Sbjct: 328 SNNLTLKGWPLTGFDQLRSGLLQQQKPFMQSQSF----HQLNMLTPQHQQQLMLAQQNLN 383
Query: 461 NERRRLENERMVLLIRKKELEL 482
++ EN R+ +L+ + + L
Sbjct: 384 SQSVSEENRRLKMLLNNRSMTL 405
>At2g19480 putative nucleosome assembly protein
Length = 379
Score = 42.7 bits (99), Expect = 5e-04
Identities = 21/50 (42%), Positives = 31/50 (62%), Gaps = 1/50 (2%)
Query: 327 EDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEENDGKPSRKRARKG 376
EDD+DE DE +D DE +DE E +D++EE + GK S+K++ G
Sbjct: 307 EDDDDEIDEDDDEEDEEDDEDDEE-EDDEDDDEEEEADQGKKSKKKSSAG 355
>At5g16030 unknown protein
Length = 339
Score = 41.6 bits (96), Expect = 0.001
Identities = 21/58 (36%), Positives = 34/58 (58%), Gaps = 11/58 (18%)
Query: 326 EEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEENDGKPSRKRARKGGFSFPRS 383
EED+E+EE+E +D S+E + E ++DE+E + K +K+ G FS+ RS
Sbjct: 270 EEDEEEEEEEKQDMSEEDDKE-----------EEDEQEEEEKTKKKKRGPGCFSWVRS 316
Score = 30.8 bits (68), Expect = 1.8
Identities = 17/58 (29%), Positives = 28/58 (47%), Gaps = 11/58 (18%)
Query: 324 VEEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEENDGKPSRKRARKGGFSFP 381
+ EEDD++EEDE E+ + + G GC + +++ARK + FP
Sbjct: 283 MSEEDDKEEEDEQEEEEKTKKKKRGPGCFSW-----------VRSRQRQARKSKYIFP 329
>At3g06130 unknown protein
Length = 473
Score = 41.6 bits (96), Expect = 0.001
Identities = 43/181 (23%), Positives = 72/181 (39%), Gaps = 29/181 (16%)
Query: 241 FREMCAYHNSCGHGGVATSNVQHQVE---------GGSGTTTPPQNQPQQQHQHQHQQQN 291
F+ M H G GG +N + + GG G + P+ Q QH Q Q
Sbjct: 92 FKGMQIDHGKAGGGGGGNNNNNKKGQKNGGGGGGGGGGGNSNAPKMGQQLNPQHLQQLQQ 151
Query: 292 QQHCFHSSENGV-----GSLGVSRGEGLRMLKIGSGYVEEEDDEDEEDESEDFSDEGEDE 346
Q + + GS+ V++ + + +K D E+D+ +DFSDE +DE
Sbjct: 152 LQKMKGFQDLKLPPQLKGSVPVNKNQNQKGVKF---------DVPEDDDDDDFSDEFDDE 202
Query: 347 SGEGCSKGHINDQDEEENDGKPSRKRARKGGFSFPRSSSSTQLVNQMN-NEISGVFQDGG 405
+ +D+ ++E D P K + ++ + Q N N G ++GG
Sbjct: 203 FTD-----DDDDEFDDEFDDLPLPSNKMKPNMTMMPNAQQMMMNAQKNANLAGGPAKNGG 257
Query: 406 K 406
K
Sbjct: 258 K 258
>At1g76890 unknown protein
Length = 575
Score = 40.8 bits (94), Expect = 0.002
Identities = 29/109 (26%), Positives = 50/109 (45%), Gaps = 26/109 (23%)
Query: 106 ESSGGDPKRKISPWQRMKWTDTMVRLLIMAVYYIGDEAGSEGTDPNKKKSSGLLQKKGK- 164
ESSGG + + MK +T G+ AGS G + ++ LL+ + +
Sbjct: 10 ESSGGGVGGSVEEEKDMKMEET------------GEGAGSGGNRWPRPETLALLRIRSEM 57
Query: 165 -------------WKSVSRAMMEKGFYVSPQQCEDKFNDLNKRYKRVND 200
W+ +SR MME G+ S ++C++KF ++ K +KR +
Sbjct: 58 DKAFRDSTLKAPLWEEISRKMMELGYKRSSKKCKEKFENVYKYHKRTKE 106
Score = 38.5 bits (88), Expect = 0.009
Identities = 54/290 (18%), Positives = 96/290 (32%), Gaps = 80/290 (27%)
Query: 71 HSMKHGFPPFSSSKNKQQQQSSQMSDEDEPNFPAEES-----------SGGDPKRKISPW 119
H + G P N + Q Q + F ++E D +SP
Sbjct: 335 HKISGGQPQQPQQHNHKPSQRKQYQSDHSITFESKEPRAVLLDTTIKMGNYDNNHSVSP- 393
Query: 120 QRMKWTDTMVRLLIMAVYYIGDEAGSEGTDPNKKKSSGLLQKKGKWKSVSRAMMEKGFYV 179
+W T V LI + GT K W+ +S M G+
Sbjct: 394 SSSRWPKTEVEALIRIRKNLEANYQENGT------------KGPLWEEISAGMRRLGYNR 441
Query: 180 SPQQCEDKFNDLNKRYKRVNDILGKGTACRVVENQGLLDSMDLSPKMKDEVRKLLNSKHL 239
S ++C++K+ ++NK +K+V K ++ R L +
Sbjct: 442 SAKRCKEKWENINKYFKKV--------------------------KESNKKRPLDSKTCP 475
Query: 240 FFREMCAYHNSCGHGGVATSNVQHQVEGGSGTTTPPQNQPQQQHQHQHQQQNQQHCFHSS 299
+F ++ A +N + G P PQ+Q + Q + F +
Sbjct: 476 YFHQLEALYN------------ERNKSGAMPLPLPLMVTPQRQLLLSQETQTE---FETD 520
Query: 300 ENGVGSLGVSRGEGLRMLKIGSGYVEEEDDEDEEDESEDFSDEGEDESGE 349
+ K+G EEE + +E++ E+ EG++E+ E
Sbjct: 521 QRE---------------KVGDKEDEEEGESEEDEYDEEEEGEGDNETSE 555
>At5g56950 nucleosome assembly protein
Length = 374
Score = 40.4 bits (93), Expect = 0.002
Identities = 24/61 (39%), Positives = 33/61 (53%), Gaps = 9/61 (14%)
Query: 320 GSGYVEEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEENDGKPSRKRA---RKG 376
G + + DDED+ DE ED +E EDE E +D+DEEE K +K + +KG
Sbjct: 301 GEEFEIDNDDEDDIDEDEDEDEEDEDEDEEE------DDEDEEEEVSKTKKKPSVLHKKG 354
Query: 377 G 377
G
Sbjct: 355 G 355
>At4g02720 unknown protein
Length = 422
Score = 40.4 bits (93), Expect = 0.002
Identities = 34/136 (25%), Positives = 62/136 (45%), Gaps = 11/136 (8%)
Query: 325 EEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEENDGKPSRKRARKGGFSFPRSS 384
+ ED ++ DE +D + D++ +G + +D E E+DG SRKR +S
Sbjct: 70 QNEDSDENADEIQDKNGGERDDNSKGKERKGKSDS-ESESDGLRSRKR---------KSK 119
Query: 385 SSTQLVNQMNNEISGVFQDGGKSTWEKKQWIRNRIMQLEEQKIGYESQAFQLEKQRLKWA 444
SS + + S +G +S E++ R R ++K S++F+ ++ +
Sbjct: 120 SSRSKRRRKRSYDSDSESEGSESDSEEED-RRRRRKSSSKRKKSRSSRSFRKKRSHRRKT 178
Query: 445 RYSSKKEREMERAKLE 460
+YS E E +K E
Sbjct: 179 KYSDSDESSDEDSKAE 194
Score = 28.9 bits (63), Expect = 6.8
Identities = 22/97 (22%), Positives = 46/97 (46%), Gaps = 8/97 (8%)
Query: 345 DESGEGCSKGHIN--DQDEEENDGKPSRKRARKGGFSFPRSSSSTQLVNQMNNEISGVFQ 402
DES + SK I+ EEE+ S++R + S RS + ++ G +
Sbjct: 184 DESSDEDSKAEISASSSGEEEDTKSKSKRRKKSSDSSSKRSKGEK---TKSGSDSDGTEE 240
Query: 403 DGGKSTWEKKQWIRNRIMQLEEQKIGYESQAFQLEKQ 439
D S + + ++N ++L+E+++ + +L+K+
Sbjct: 241 D---SKMQVDETVKNTELELDEEELKKFKEMIELKKK 274
>At3g09360 putative transcription factor
Length = 600
Score = 40.4 bits (93), Expect = 0.002
Identities = 58/240 (24%), Positives = 98/240 (40%), Gaps = 46/240 (19%)
Query: 265 VEGGS-GTTTPPQNQPQQQHQHQHQQQNQQHCFHSSENGVGSLGVSRGEGLRMLKIGSGY 323
V GG G + PP Q ++ + + + + +E G+ SL + E L L+I G
Sbjct: 319 VSGGLVGGSNPPAFQRAEKERMEKAAREE------NEGGISSL--NHDEQLYHLRIYLGC 370
Query: 324 VEEEDDEDEE---------DESEDFSDEGEDESGEGCSKGHINDQDE---------EEND 365
V E+ ++D++ DES++FSD +DE G+IN+++E E N
Sbjct: 371 VAEKGEKDKDGAEEHADTSDESDNFSDISDDE-----VNGYINNEEETHYKTITWTEMNK 425
Query: 366 G---KPSRKRARKGGFSFPRSSSSTQLVNQMNNEISGVFQDGGKSTWEKKQWIRNRIMQL 422
+ + K A S +S++ D KS EK+Q
Sbjct: 426 DYLEEQAAKEAALKAASEALKASNSNCPEDARKAFEAAKADAAKSRKEKQQKKAEEAKNA 485
Query: 423 EEQKIGYESQAFQLEKQRLKWA-RY--------SSKKEREMERAKLEN--ERRRLENERM 471
E+ L+K+RL Y +S E+ +R+K E E+++ EN+ M
Sbjct: 486 APPATAVEAVRRTLDKKRLSSVINYDVLESLFDTSAPEKSPKRSKTETDIEKKKEENKEM 545
Score = 28.5 bits (62), Expect = 8.9
Identities = 16/45 (35%), Positives = 30/45 (66%), Gaps = 4/45 (8%)
Query: 321 SGYVEEEDDEDEEDES-EDFSDEGEDESGEGCSKGHINDQDEEEN 364
+G E+ED+EDEE+ + E + + + ++GE K + D++EEE+
Sbjct: 552 NGENEDEDEEDEEEGNVESYDMKTDFQNGE---KFYEEDEEEEED 593
>At5g17690 chromo domain protein
Length = 445
Score = 40.0 bits (92), Expect = 0.003
Identities = 23/53 (43%), Positives = 32/53 (59%), Gaps = 2/53 (3%)
Query: 299 SENGVGSLGVSRGEGLRMLKIGSGY-VEEEDDEDEEDESEDFSDEGEDESGEG 350
S++G G G GE + + +IG E+ D+E+EEDE ED + EDE GEG
Sbjct: 42 SDDGGGGGGGGSGESI-LREIGDDRPTEDGDEEEEEDEDEDDGGDEEDEEGEG 93
>At3g25990 putative DNA-binding protein, GT-1
Length = 372
Score = 40.0 bits (92), Expect = 0.003
Identities = 32/104 (30%), Positives = 48/104 (45%), Gaps = 14/104 (13%)
Query: 106 ESSGGDPKRKI-SPWQRMK-WTDTMVRLLIMAVYYIGDEAGSEGTDPNKKKSSGLLQKKG 163
ESSGG+ I +P +R + W R LI + + N KS+ K
Sbjct: 35 ESSGGEDHEIIKAPKKRAETWAQDETRTLISLRREMDNLF-------NTSKSN-----KH 82
Query: 164 KWKSVSRAMMEKGFYVSPQQCEDKFNDLNKRYKRVNDILGKGTA 207
W+ +S+ M EKGF SP C DK+ ++ K +K+ K T+
Sbjct: 83 LWEQISKKMREKGFDRSPSMCTDKWRNILKEFKKAKQHEDKATS 126
>At1g56660 hypothetical protein
Length = 522
Score = 40.0 bits (92), Expect = 0.003
Identities = 44/219 (20%), Positives = 95/219 (43%), Gaps = 20/219 (9%)
Query: 281 QQHQHQHQQQNQQHCFHSSENGVGSLGVSRGEGLRMLKIGSGYVEEEDDEDEEDESEDFS 340
++H+ +H++ ++ E G ++ E K SG E+ D+E + ED S
Sbjct: 109 EEHEKEHKKGKEKKHEELEEEKEGKKKKNKKE-----KDESGPEEKNKKADKEKKHEDVS 163
Query: 341 DEGEDESGEGCSKGHINDQDE---EENDGKPSRKRARKGGFSFPRSSSSTQLVNQMNNEI 397
E E+ E K ++DE EE KP +++ +K S S + + ++
Sbjct: 164 QEKEELEEEDGKKNKKKEKDESGTEEKKKKPKKEKKQK------EESKSNE-----DKKV 212
Query: 398 SGVFQDGGKSTWEKKQWIRNRIMQLEEQKIGYESQAFQLEKQRLKWARYSSKKEREMERA 457
G + G K EK+ + + +Q++ + +K++ + KK+ + E+
Sbjct: 213 KGKKEKGEKGDLEKEDEEKKKEHDETDQEMKEKDSKKNKKKEKDESCAEEKKKKPDKEK- 271
Query: 458 KLENERRRLENERMVLLIRKKELELMHIQQQQQQQHSST 496
K ++E E++++ K E + ++ ++H +T
Sbjct: 272 KEKDESTEKEDKKLKGKKGKGEKPEKEDEGKKTKEHDAT 310
Score = 32.7 bits (73), Expect = 0.47
Identities = 31/145 (21%), Positives = 67/145 (45%), Gaps = 10/145 (6%)
Query: 325 EEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEENDGKPSRKRARKGGFSFPRSS 384
EE + E ES+ +E E E +G K H ++E+E K ++K + G P
Sbjct: 93 EEGHGDLEVKESDVKVEEHEKEHKKGKEKKHEELEEEKEGKKKKNKKEKDESG---PEEK 149
Query: 385 SSTQLVNQMNNEISGVFQDGGKSTWEKKQWIRNRIMQLEEQKIGYESQAFQLEKQRLKWA 444
+ + + ++S K E++ +N+ + E+ + G E + + +K++ +
Sbjct: 150 NKKADKEKKHEDVS-----QEKEELEEEDGKKNK--KKEKDESGTEEKKKKPKKEKKQKE 202
Query: 445 RYSSKKEREMERAKLENERRRLENE 469
S ++++++ K + E+ LE E
Sbjct: 203 ESKSNEDKKVKGKKEKGEKGDLEKE 227
Score = 31.2 bits (69), Expect = 1.4
Identities = 81/452 (17%), Positives = 167/452 (36%), Gaps = 65/452 (14%)
Query: 60 HHDTDTNQHHHHSMKHGFPPFSSSKNKQQQQSSQMSDEDEPNFPAEESSGGDPKRKISPW 119
H D + + +H + K ++ + + + N ++ SG + K K +
Sbjct: 96 HGDLEVKESDVKVEEHEKEHKKGKEKKHEELEEEKEGKKKKNKKEKDESGPEEKNKKADK 155
Query: 120 QRMKWTDTMVRLLIMAVYYIGDEAGSEGTDPNKKKS---SGLLQKKGKWKSVSRAMMEKG 176
++ K D V +E E NKKK SG +KK K K +
Sbjct: 156 EK-KHED---------VSQEKEELEEEDGKKNKKKEKDESGTEEKKKKPKKEKK------ 199
Query: 177 FYVSPQQCEDKFNDLNKRYKRVNDILGKGTACRVVENQGLL-DSMDLSPKMKDEVRKLLN 235
Q+ E K N+ +K+ K + KG + E + D D K KD +
Sbjct: 200 -----QKEESKSNE-DKKVKGKKEKGEKGDLEKEDEEKKKEHDETDQEMKEKDSKKNKKK 253
Query: 236 SKHLFFREMCAYHNSCG------HGGVATSNVQHQVEGGSGTTTPPQNQPQQQHQHQHQQ 289
K E CA +T +++G G P+ + + + +H
Sbjct: 254 EKD----ESCAEEKKKKPDKEKKEKDESTEKEDKKLKGKKGKGEKPEKEDEGKKTKEHDA 309
Query: 290 QNQQHCFHSSENGVGSLGVSRGEGLRMLKIGSGYVEEEDDEDEEDE----------SEDF 339
Q+ ++++ G ++ + + + E+E + ++DE E
Sbjct: 310 TEQEMDDEAADHKEGKKKKNKDKAKKKETVIDEVCEKETKDKDDDEGETKQKKNKKKEKK 369
Query: 340 SDEGEDESGEGCSKGHINDQDEEENDGKPSRKRARKGGFSFPRSSSSTQLVNQMNNEISG 399
S++GE + E K + + + D K A K + T+ + +++ G
Sbjct: 370 SEKGEKDVKEDKKKENPLETEVMSRDIKLEEPEAEK------KEEDDTE--EKKKSKVEG 421
Query: 400 VFQDGGKSTWEKKQWIRNRIMQLEEQKIGYESQAFQLEKQRLKWARYSSKKERE------ 453
+ GK +KK +N+ +E K+ + + + + + +K +K+E++
Sbjct: 422 GESEEGKKK-KKKDKKKNKKKDTKEPKMTEDEEEKKDDSKDVKIEGSKAKEEKKDKDVKK 480
Query: 454 ----MERAKLENERRRLENERMVLLIRKKELE 481
+ KL+ + +++ + L+ K E+E
Sbjct: 481 KKGGNDIGKLKTKLAKIDEKIGALMEEKAEIE 512
Score = 28.5 bits (62), Expect = 8.9
Identities = 32/162 (19%), Positives = 66/162 (39%), Gaps = 18/162 (11%)
Query: 310 RGEGLRM-LKIGSGYVEEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEE-ENDGK 367
+GE + + +++ + +E+ + +E+ S + E + G+ D+D++ + DGK
Sbjct: 27 KGENVEVEMEVKAKSIEKVKAKKDEESSGKSKKDKEKKKGKNVDSEVKEDKDDDKKKDGK 86
Query: 368 PSRKRARKGGFSFPRSSSSTQLVNQMNNEISGVFQDGGKSTWEKKQWIRNRIMQLEEQKI 427
K+ +G S ++ G EKK +LEE+K
Sbjct: 87 MVSKKHEEGHGDLEVKESDVKVEEHEKEHKKGK---------EKKH------EELEEEKE 131
Query: 428 GYESQAFQLEKQRLKWARYSSKKEREMERAKLENERRRLENE 469
G + + + EK + K ++E + + E+ LE E
Sbjct: 132 G-KKKKNKKEKDESGPEEKNKKADKEKKHEDVSQEKEELEEE 172
>At5g53800 unknown protein
Length = 339
Score = 39.7 bits (91), Expect = 0.004
Identities = 35/150 (23%), Positives = 68/150 (45%), Gaps = 17/150 (11%)
Query: 326 EEDDEDEEDESEDFSDEGEDESGE-GCSKGHINDQDEE----------ENDGKPSRKRAR 374
EE D ES+ + +G D E G G ++++E ++D K SR R R
Sbjct: 40 EEKDVSRRRESKRRTKDGNDSGSESGLESGSESEKEERRRSRKDRGKRKSDRKSSRSRRR 99
Query: 375 KGGFSFPRSSSSTQLVNQMNNEISGVFQDGGKSTWEKKQWIRNRIMQLEEQKIGYESQAF 434
+ +S S S ++ ++ ++ +D E+++ R R + EE+K +
Sbjct: 100 RRDYSSSSSDSESESESEYSDSEESESED------ERRRRKRKRKEREEEEKERKRRRRE 153
Query: 435 QLEKQRLKWARYSSKKEREMERAKLENERR 464
+ +K+R K + KK +E ++ K E ++
Sbjct: 154 KDKKKRNKSDKDGDKKRKEKKKKKSEKVKK 183
>At5g03710 putative protein
Length = 81
Score = 39.7 bits (91), Expect = 0.004
Identities = 19/56 (33%), Positives = 35/56 (61%), Gaps = 1/56 (1%)
Query: 325 EEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEENDGKPSRKRARKGGFSF 380
EEE++E+EE+E E+ +E E+E E + +++EEE + + R+R + F+F
Sbjct: 23 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEDREREER-AFAF 77
Score = 38.1 bits (87), Expect = 0.011
Identities = 17/51 (33%), Positives = 31/51 (60%)
Query: 325 EEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEENDGKPSRKRARK 375
EEE++E+EE+E E+ +E E+E E + +++EEE + + R R+
Sbjct: 21 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEDRERE 71
Score = 36.6 bits (83), Expect = 0.033
Identities = 16/51 (31%), Positives = 31/51 (60%)
Query: 325 EEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEENDGKPSRKRARK 375
EEE++E+EE+E E+ +E E+E E + +++EEE + + + R+
Sbjct: 19 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEDRE 69
Score = 36.2 bits (82), Expect = 0.043
Identities = 16/44 (36%), Positives = 28/44 (63%)
Query: 322 GYVEEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEEND 365
G EEE++E+EE+E E+ +E E+E E + +++EEE +
Sbjct: 5 GEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 48
Score = 35.4 bits (80), Expect = 0.073
Identities = 15/41 (36%), Positives = 27/41 (65%)
Query: 325 EEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEEND 365
EEE++E+EE+E E+ +E E+E E + +++EEE +
Sbjct: 14 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 54
Score = 35.4 bits (80), Expect = 0.073
Identities = 15/41 (36%), Positives = 27/41 (65%)
Query: 325 EEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEEND 365
EEE++E+EE+E E+ +E E+E E + +++EEE +
Sbjct: 12 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 52
Score = 35.4 bits (80), Expect = 0.073
Identities = 15/41 (36%), Positives = 27/41 (65%)
Query: 325 EEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEEND 365
EEE++E+EE+E E+ +E E+E E + +++EEE +
Sbjct: 15 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 55
Score = 35.4 bits (80), Expect = 0.073
Identities = 15/41 (36%), Positives = 27/41 (65%)
Query: 325 EEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEEND 365
EEE++E+EE+E E+ +E E+E E + +++EEE +
Sbjct: 11 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 51
Score = 35.4 bits (80), Expect = 0.073
Identities = 15/41 (36%), Positives = 27/41 (65%)
Query: 325 EEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEEND 365
EEE++E+EE+E E+ +E E+E E + +++EEE +
Sbjct: 18 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 58
Score = 35.4 bits (80), Expect = 0.073
Identities = 15/41 (36%), Positives = 27/41 (65%)
Query: 325 EEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEEND 365
EEE++E+EE+E E+ +E E+E E + +++EEE +
Sbjct: 16 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 56
Score = 35.4 bits (80), Expect = 0.073
Identities = 15/41 (36%), Positives = 27/41 (65%)
Query: 325 EEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEEND 365
EEE++E+EE+E E+ +E E+E E + +++EEE +
Sbjct: 17 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 57
Score = 35.4 bits (80), Expect = 0.073
Identities = 15/41 (36%), Positives = 27/41 (65%)
Query: 325 EEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEEND 365
EEE++E+EE+E E+ +E E+E E + +++EEE +
Sbjct: 10 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 50
Score = 35.4 bits (80), Expect = 0.073
Identities = 15/41 (36%), Positives = 27/41 (65%)
Query: 325 EEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEEND 365
EEE++E+EE+E E+ +E E+E E + +++EEE +
Sbjct: 9 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 49
Score = 35.4 bits (80), Expect = 0.073
Identities = 15/41 (36%), Positives = 27/41 (65%)
Query: 325 EEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEEND 365
EEE++E+EE+E E+ +E E+E E + +++EEE +
Sbjct: 13 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 53
>At3g51070 putative protein
Length = 895
Score = 39.7 bits (91), Expect = 0.004
Identities = 34/136 (25%), Positives = 57/136 (41%), Gaps = 27/136 (19%)
Query: 262 QHQVEGGSGTTTPPQN-----QPQQQHQHQHQ--QQNQQHCFHSSENGVGSLGVSRGEGL 314
+ +E G G T+ QP++Q+ + QQN++ S ENG G
Sbjct: 221 EQPMETGQGETSETSKNEENGQPEEQNSGNEETGQQNEEKTTASEENGKGE--------- 271
Query: 315 RMLKIGSGYVEEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEENDGKPSRKRAR 374
+ +K +G EE +EE +++ +DE+ E Q EE D K + +
Sbjct: 272 KSMKDENGQQEEHTTAEEESGNKEEESTSKDENME---------QQEERKDEKKHEQGSE 322
Query: 375 KGGF--SFPRSSSSTQ 388
GF P+ S+ +Q
Sbjct: 323 ASGFGSGIPKESAESQ 338
Score = 36.2 bits (82), Expect = 0.043
Identities = 45/247 (18%), Positives = 92/247 (37%), Gaps = 22/247 (8%)
Query: 212 ENQGLLDSMDLSPKMKDEVRKLLNSKHLFFREMCAYHNSCGHGGVATSNVQHQVEGGSGT 271
+N G L D + K +DE RK K + + S + + G
Sbjct: 89 DNPGKLP--DDAVKSEDEQRKSAKEKSETTSSKTQTQETQQNNDDKISEEKEKDNGKENQ 146
Query: 272 TTPPQNQPQQQH--QHQHQQQNQQHCFHSSENGVGSLGVSRGEGLRMLKIGSGYVEEEDD 329
T + Q + + ++Q QQ + G+ G +G+G + G + ++
Sbjct: 147 TVQESEEGQMKKVVKEFEKEQKQQRDEDAGTQPKGTQGQEQGQGKEQPDVEQGNKQGQEQ 206
Query: 330 EDEEDESEDFSDEGEDESGEGCSKGHINDQDEEENDGKPSRKRARKGGFSFPRSSSSTQL 389
+ D + F+D + E +G ++ + E +G+P + +S +
Sbjct: 207 DSNTDVT--FTDATKQEQPMETGQGETSETSKNEENGQPEEQ------------NSGNEE 252
Query: 390 VNQMNNEISGVFQDGGKSTWEKKQWIRNRIMQLEEQKIGYESQAFQLEKQRLKWARYSSK 449
Q N E + ++ GK ++ +++ Q EE E + E+ K +
Sbjct: 253 TGQQNEEKTTASEENGKG----EKSMKDENGQQEEHTTAEEESGNKEEESTSKDENMEQQ 308
Query: 450 KEREMER 456
+ER+ E+
Sbjct: 309 EERKDEK 315
>At2g22080 En/Spm-like transposon protein
Length = 140
Score = 39.7 bits (91), Expect = 0.004
Identities = 16/48 (33%), Positives = 31/48 (64%)
Query: 325 EEEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEENDGKPSRKR 372
++ED+E++EDE++D +E +DE E + D+D+EE P +++
Sbjct: 92 DDEDEEEDEDENDDGGEEDDDEDAEVEEEEEEEDEDDEEALQPPKKRK 139
Score = 33.1 bits (74), Expect = 0.36
Identities = 13/42 (30%), Positives = 25/42 (58%)
Query: 326 EEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEEENDGK 367
E D D+E E +D +D+ +D++ E + D+DE ++ G+
Sbjct: 67 EAGDNDDEPEGDDGNDDEDDDNHENDDEDEEEDEDENDDGGE 108
Score = 32.7 bits (73), Expect = 0.47
Identities = 17/60 (28%), Positives = 36/60 (59%), Gaps = 11/60 (18%)
Query: 325 EEEDDEDEEDESEDFSDEGEDESGEGCSK-----GHINDQDEEENDG-----KPSRKRAR 374
++EDD++ E++ ED +E EDE+ +G + + +++EEE++ +P +KR +
Sbjct: 82 DDEDDDNHENDDED-EEEDEDENDDGGEEDDDEDAEVEEEEEEEDEDDEEALQPPKKRKK 140
Score = 28.9 bits (63), Expect = 6.8
Identities = 11/37 (29%), Positives = 22/37 (58%)
Query: 326 EEDDEDEEDESEDFSDEGEDESGEGCSKGHINDQDEE 362
++ ++DE+D++ + DE E+E + G D DE+
Sbjct: 78 DDGNDDEDDDNHENDDEDEEEDEDENDDGGEEDDDED 114
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.310 0.127 0.367
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,782,708
Number of Sequences: 26719
Number of extensions: 661723
Number of successful extensions: 11151
Number of sequences better than 10.0: 582
Number of HSP's better than 10.0 without gapping: 286
Number of HSP's successfully gapped in prelim test: 308
Number of HSP's that attempted gapping in prelim test: 5526
Number of HSP's gapped (non-prelim): 2754
length of query: 496
length of database: 11,318,596
effective HSP length: 103
effective length of query: 393
effective length of database: 8,566,539
effective search space: 3366649827
effective search space used: 3366649827
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 62 (28.5 bits)
Medicago: description of AC146788.10


