

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
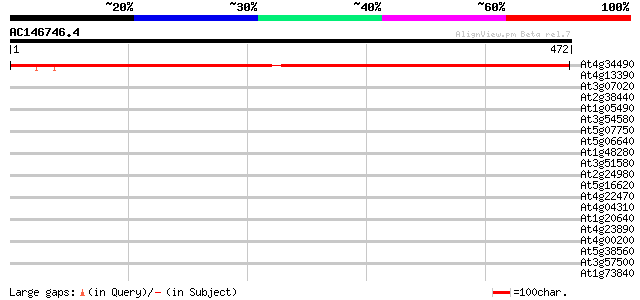
Query= AC146746.4 - phase: 0
(472 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At4g34490 putative cyclase associated protein CAP 672 0.0
At4g13390 extensin-like protein 36 0.052
At3g07020 UDP-glucose:sterol glucosyltransferase 35 0.069
At2g38440 unknown protein 34 0.20
At1g05490 hypothetical protein 33 0.34
At3g54580 extensin precursor -like protein 32 0.58
At5g07750 putative protein 32 0.76
At5g06640 putative protein 32 0.76
At1g48280 unknown protein 32 0.76
At3g51580 unknown protein 32 0.99
At2g24980 unknown protein 32 0.99
At5g16620 translocon Tic40-like protein 31 1.7
At4g22470 extensin - like protein 31 1.7
At4g04310 putative transposon protein 31 1.7
At1g20640 unknown protein 31 1.7
At4g23890 unknown protein 30 2.2
At4g00200 putative transcription factor 30 2.2
At5g38560 putative protein 30 2.9
At3g57500 hypothetical protein 30 2.9
At1g73840 proline-rich like protein precursor 30 2.9
>At4g34490 putative cyclase associated protein CAP
Length = 476
Score = 672 bits (1733), Expect = 0.0
Identities = 344/482 (71%), Positives = 395/482 (81%), Gaps = 18/482 (3%)
Query: 1 MDETLIKRLESAVTRLEALST---GIHHSSSASDASDAA------SDPSVVAFVDLIDQY 51
M+E LIKRLE+AVTRLE +S+ G+ S D S AA SDPS++A+ DLI Q
Sbjct: 1 MEEDLIKRLEAAVTRLEGISSNGGGVVSLSRGGDFSSAAGIDIASSDPSILAYEDLISQC 60
Query: 52 VSRLSKAADIIGGQVLDVTNRVKEAFSVQKELLIKLKQTQKPDPAGLAEFLKPLNEVIMK 111
V R AA+ IGG VLDVT V EAF+ QKELL+++KQTQKPD AGLA FLKPLN+V MK
Sbjct: 61 VGRALTAAEKIGGPVLDVTKIVAEAFASQKELLVRIKQTQKPDLAGLAGFLKPLNDVTMK 120
Query: 112 ASSLTEGRRSDFFNHLKAAVDSLSALAWIAFTGKDCGMSMPIAHVEESWQMAEFYSNKVL 171
A+++TEG+RSDFFNHLKAA DSLSALAWIAFTGKDCGMSMPIAHVEESWQMAEFY+NKVL
Sbjct: 121 ANAMTEGKRSDFFNHLKAACDSLSALAWIAFTGKDCGMSMPIAHVEESWQMAEFYNNKVL 180
Query: 172 VEYRNKDPNHVEWVKALKELYLPGLRDYVKSFHPLGPVWSQTGK-VFAPSKVNASAAPAA 230
VEYRNKD +HVEW KALKELYLPGLR+YVKS +PLGPVW+ +GK AP+K
Sbjct: 181 VEYRNKDADHVEWAKALKELYLPGLREYVKSHYPLGPVWNASGKPASAPAK-------GP 233
Query: 231 PSAPPPPPASLFSSESSQ-ASSSKPKVGMSAVFQEIGTGNVTAGLRKVTDDMKTKNRADR 289
P AP PPPA LFS+ESS+ +SSS K GMSAVFQ++ +G VT+GLRKVTDDMKTKNRADR
Sbjct: 234 PGAPAPPPAPLFSAESSKPSSSSNQKQGMSAVFQQLSSGAVTSGLRKVTDDMKTKNRADR 293
Query: 290 SGVVGNSVKESQAAPRAFSKVGPPKLELQMGRKWVVENQIDQKSLVIEDCDSKQSVYVYG 349
SG V KE++ + AFSK GPPK+ELQMGRKW VENQI +K LVI +CDSKQSVY+YG
Sbjct: 294 SGAVSAVEKETRTSKPAFSKTGPPKMELQMGRKWAVENQIGKKDLVISECDSKQSVYIYG 353
Query: 350 CKNSVLQIQGKVNNITIDNCKKTGVVFKDVVAAFEVVNSNGVEVQCQGSAPTISVDNTSG 409
CK+SVLQIQGKVNNITID C K GVVF DVVAAFE+VN N VEVQCQGSAPT+SVDNT+G
Sbjct: 354 CKDSVLQIQGKVNNITIDKCTKVGVVFTDVVAAFEIVNCNNVEVQCQGSAPTVSVDNTTG 413
Query: 410 CQIYLSKDSLETSISTAKSSEINVLVPNVESDGDWVEHSLPQQYIHLFKEGRFETTPASH 469
CQ+YL+KDSLET+I+TAKSSEINV+VP DGDWVEH+LPQQY H+F EG+FETTP SH
Sbjct: 414 CQLYLNKDSLETAITTAKSSEINVMVPGATPDGDWVEHALPQQYNHVFTEGKFETTPVSH 473
Query: 470 SG 471
SG
Sbjct: 474 SG 475
>At4g13390 extensin-like protein
Length = 429
Score = 35.8 bits (81), Expect = 0.052
Identities = 37/132 (28%), Positives = 53/132 (40%), Gaps = 12/132 (9%)
Query: 133 SLSALAWIAFTGKDCGMSMPIAHVEESWQMAEFYSNKVLVEYRNKDPNHVEWVKALKELY 192
SL L + + + K S P ++ S +YS V+Y++ P +V Y
Sbjct: 132 SLPPLTYYSPSPKVIYNSPPPPYIYSSPPPPPYYSPSPKVDYKSPPPPYVYSSPPPPPYY 191
Query: 193 LPGLR--------DYVKSFHPLGPVWSQTGKVFAPSKVNASAAPAAPSAPPPPPASLFSS 244
P + YV SF P P +S + KV S AP S+PPPPP S
Sbjct: 192 SPSPKVEYKSPPPPYVYSFPPPPPYYSPSPKVGYKSP----PAPYVYSSPPPPPYYSPSP 247
Query: 245 ESSQASSSKPKV 256
+ + S P V
Sbjct: 248 KVNYKSPPPPYV 259
Score = 32.7 bits (73), Expect = 0.44
Identities = 29/99 (29%), Positives = 39/99 (39%), Gaps = 12/99 (12%)
Query: 150 SMPIAHVEESWQMAEFYSNKVLVEYRNKDPNHVEWVKALKELYLPGLR--------DYVK 201
S P +V S +YS VEY++ P +V Y P + YV
Sbjct: 175 SPPPPYVYSSPPPPPYYSPSPKVEYKSPPPPYVYSFPPPPPYYSPSPKVGYKSPPAPYVY 234
Query: 202 SFHPLGPVWSQTGKVFAPSKVNASAAPAAPSAPPPPPAS 240
S P P +S + KV + P S+PPPPP S
Sbjct: 235 SSPPPPPYYSPSPKV----NYKSPPPPYVYSSPPPPPYS 269
>At3g07020 UDP-glucose:sterol glucosyltransferase
Length = 637
Score = 35.4 bits (80), Expect = 0.069
Identities = 50/235 (21%), Positives = 91/235 (38%), Gaps = 30/235 (12%)
Query: 234 PPPPPASLFS-SESSQASSSKPKVGMSAVFQEIGTGNVTAGLRKVTDDMKTKNRADRSGV 292
P PA L S SS +SSS S +EI G +G+ V+++++T + + +
Sbjct: 2 PEISPAELAKVSSSSSSSSSSSSGRASVKIEEIEGGAAASGVVIVSEELETNPKTVVASI 61
Query: 293 VGNSVKESQ-AAPRAFSKVGPPKLELQMGRKWVVENQIDQKSLVIEDCDSKQSVYVYGCK 351
+V ES ++FS+V LE + +Q L
Sbjct: 62 ADETVAESSGTGNKSFSRVWTMPLEGSSSSDKAESSSTNQPRL----------------D 105
Query: 352 NSVLQIQGKVNNITIDNCKKTGVVFKDVVAAFEVVNSNGVEVQCQGSAPTISVDNTSGCQ 411
S + Q KV +I ++ K +F D ++A G +++ T+ D T +
Sbjct: 106 KSKTERQQKVTHILAEDAAK---IFDDKISA-------GKKLKLLNRIATVKHDGT--VE 153
Query: 412 IYLSKDSLETSISTAKSSEINVLVPNVESDGDWVEHSLPQQYIHLFKEGRFETTP 466
+ D++ I + N + + DG +++ P Q + L R + P
Sbjct: 154 FEVPADAIPQPIVVDRGESKNGVCADESIDGVDLQYIPPMQIVMLIVGTRGDVQP 208
>At2g38440 unknown protein
Length = 1421
Score = 33.9 bits (76), Expect = 0.20
Identities = 41/174 (23%), Positives = 71/174 (40%), Gaps = 22/174 (12%)
Query: 269 NVTAGLRKVTDDMKTKNRADRSGVVGNSVKESQ---AAPRAFSKVGPP-KLELQMGRKWV 324
+VT KV+ D+ + S V G + S ++PR S+ L +Q V
Sbjct: 495 HVTPEANKVSHDLNVQESVSSSNVDGQTSLSSNGTCSSPRPVSQNDQSCSLTVQSLASEV 554
Query: 325 VENQIDQKSL----------VIEDCDSKQSVYVYGCKNSVLQIQGKVNNITIDNCKKTGV 374
VE + L ++ DS +S + KNS L + + T + +
Sbjct: 555 VETSPELVRLDLMKGGNDGRKVDPFDSSKSCASFDAKNSDLPSETSSISSTSEGSRCDST 614
Query: 375 VFKDVVAAFEVVNSNGVEVQCQGSAPTISVDNTSGCQIYLSKDSLETSISTAKS 428
+ K+ + A +VNS G++P VD+ +G Q+ ++ ET+ A S
Sbjct: 615 IEKNCMVASNLVNS--------GTSPQAFVDSQTGKQLPIADTDFETNSIVACS 660
>At1g05490 hypothetical protein
Length = 731
Score = 33.1 bits (74), Expect = 0.34
Identities = 29/113 (25%), Positives = 49/113 (42%), Gaps = 5/113 (4%)
Query: 232 SAPPPPPASLFSSESSQASSSKPKVGMSAVFQEIGTGNVTAGLRKVTDDMKTKNRADRSG 291
S+ P +S SS SS +SSS S V + +G LRK + +K + +R
Sbjct: 279 SSEDPSSSSSSSSSSSSSSSSSSSDDESYVKEVVGDNRDDDDLRKASSPIKRVSLVERKA 338
Query: 292 VV-----GNSVKESQAAPRAFSKVGPPKLELQMGRKWVVENQIDQKSLVIEDC 339
+V G+S+ + + K+ + E + ++ VV Q S V+ C
Sbjct: 339 LVRYKRSGSSLTKPRERDNKIQKLNHREEEKKERQREVVRVVTKQPSNVVYTC 391
>At3g54580 extensin precursor -like protein
Length = 951
Score = 32.3 bits (72), Expect = 0.58
Identities = 31/102 (30%), Positives = 43/102 (41%), Gaps = 13/102 (12%)
Query: 165 FYSNKVLVEYRNKDPNHVEWVKALKELYLPGLRDYVKSFHPLGPVWSQTGKVF----APS 220
+YS V Y++ P +V + Y P + Y KS P P +S + KV+ P
Sbjct: 509 YYSPSPKVYYKSPPPPYV-YSSPPPPYYSPSPKVYYKS--PPPPYYSPSPKVYYKSPPPP 565
Query: 221 KVNASAAPAAPSAPP------PPPASLFSSESSQASSSKPKV 256
V +S P S P PPP ++SS S PKV
Sbjct: 566 YVYSSPPPPYYSPSPKVYYKSPPPPYVYSSPPPPYHSPSPKV 607
>At5g07750 putative protein
Length = 1289
Score = 32.0 bits (71), Expect = 0.76
Identities = 15/47 (31%), Positives = 24/47 (50%)
Query: 211 SQTGKVFAPSKVNASAAPAAPSAPPPPPASLFSSESSQASSSKPKVG 257
S + K+ ++++ +P +APPPPP FS+ S S P G
Sbjct: 895 SPSVKILPLHGISSAPSPPVKTAPPPPPPPPFSNAHSVLSPPPPSYG 941
Score = 29.3 bits (64), Expect = 4.9
Identities = 16/53 (30%), Positives = 25/53 (46%), Gaps = 4/53 (7%)
Query: 205 PLGPVWSQT--GKVFAPSKVNASAAPAAPSAPPPPPASLFS--SESSQASSSK 253
P P W + P+ + S AP + PPPPP + +S +SS +S+
Sbjct: 733 PPSPPWKSVYASALAIPAICSTSQAPTSSPTPPPPPPAYYSVGQKSSDLQTSQ 785
Score = 28.9 bits (63), Expect = 6.4
Identities = 17/42 (40%), Positives = 22/42 (51%), Gaps = 5/42 (11%)
Query: 218 APSKVNASAAPAAPSAPPPPPASLFSS-----ESSQASSSKP 254
+P + +S PA+P PPP SL S SSQA +S P
Sbjct: 607 SPKETPSSLPPASPHQAPPPLPSLTSEAKTVLHSSQAVASPP 648
>At5g06640 putative protein
Length = 689
Score = 32.0 bits (71), Expect = 0.76
Identities = 31/109 (28%), Positives = 43/109 (39%), Gaps = 18/109 (16%)
Query: 165 FYSNKVLVEYRNKDPNHVEWVKALKELYLPGLRDYVKSFHPLGPVWSQTGKVFAPSKVNA 224
+YS VEY++ P +V + Y P + Y KS P S ++PS
Sbjct: 291 YYSPSPKVEYKSPPPPYV-YSSPPPPYYSPSPKVYYKSPPPPYVYSSPPPPYYSPSPKVD 349
Query: 225 SAAPAAP---SAPPPP--------------PASLFSSESSQASSSKPKV 256
+P P S+PPPP P ++SS Q S PKV
Sbjct: 350 YKSPPPPYVYSSPPPPYYSPSPKVDYKSPPPPYVYSSPPPQYYSPSPKV 398
>At1g48280 unknown protein
Length = 558
Score = 32.0 bits (71), Expect = 0.76
Identities = 17/74 (22%), Positives = 38/74 (50%)
Query: 221 KVNASAAPAAPSAPPPPPASLFSSESSQASSSKPKVGMSAVFQEIGTGNVTAGLRKVTDD 280
K + +++P AP PPPPP ++A+ ++ +S +FQ + + + L + +
Sbjct: 248 KRDENSSPFAPPTPPPPPPPPPPRPLAKAARAQKSPPVSQLFQLLNKQDNSRNLSQSVNG 307
Query: 281 MKTKNRADRSGVVG 294
K++ + + +VG
Sbjct: 308 NKSQVNSAHNSIVG 321
>At3g51580 unknown protein
Length = 390
Score = 31.6 bits (70), Expect = 0.99
Identities = 34/150 (22%), Positives = 61/150 (40%), Gaps = 11/150 (7%)
Query: 231 PSAPPPPPASLFSSESS---QASSSKPKVGMSAVFQEIGTGNVTAGLRKVTDDMKTKNRA 287
P +PPPPPA+L S+ S A+ + P + + E G + L K K +
Sbjct: 107 PMSPPPPPANLTDSQDSGKLPANMAPPPKSLESGKNETEPGKESPPLAKDPAKGKDDKGS 166
Query: 288 DRSGVVGNSVKESQAAPRAFSKVGPPKLELQMG-RKWVV----ENQIDQKSLVIEDCDSK 342
S V V +S R + + L + G W++ E + K+ ++ ++
Sbjct: 167 SESASVDTCVGKSNIC-RTENSLVACTLSIDKGAANWLILVQNEGETSLKAKIVLPVNAL 225
Query: 343 QSVYV--YGCKNSVLQIQGKVNNITIDNCK 370
Q + + + + + I G N I +D K
Sbjct: 226 QELTLPKHQSQKVNISISGDTNKIILDTGK 255
>At2g24980 unknown protein
Length = 559
Score = 31.6 bits (70), Expect = 0.99
Identities = 32/109 (29%), Positives = 45/109 (40%), Gaps = 18/109 (16%)
Query: 165 FYSNKVLVEYRNKDPNHVEWVKALKELYLPGLRDYVKS-------FHPLGPVWSQTGKVF 217
+YS V Y++ P +V + Y P + Y KS P P +S + KV+
Sbjct: 411 YYSPSPKVYYKSPPPPYV-YSSPPPPYYSPSPKVYYKSPPPPYVYSSPPPPYYSPSPKVY 469
Query: 218 ----APSKVNASAAPAAPSAPP------PPPASLFSSESSQASSSKPKV 256
PS V +S P S P PPP+ ++SS S PKV
Sbjct: 470 YKSPPPSYVYSSPPPPYYSPSPKVYYKSPPPSYVYSSPPPPYYSPSPKV 518
>At5g16620 translocon Tic40-like protein
Length = 447
Score = 30.8 bits (68), Expect = 1.7
Identities = 22/75 (29%), Positives = 39/75 (51%), Gaps = 8/75 (10%)
Query: 216 VFAPSKVNASAAPAAPSAPPPPPASLFSSESSQASSSKPKVGMSAVFQEIGTGN-----V 270
+F+ S+ + + A+PS P PPP+S S+ S VG+SA+F + T N +
Sbjct: 78 IFSSSRDQQTTSVASPSVPVPPPSS--STIGSPLFWIGVGVGLSALFSYV-TSNLKKYAM 134
Query: 271 TAGLRKVTDDMKTKN 285
++ + + M T+N
Sbjct: 135 QTAMKTMMNQMNTQN 149
>At4g22470 extensin - like protein
Length = 375
Score = 30.8 bits (68), Expect = 1.7
Identities = 28/115 (24%), Positives = 44/115 (37%), Gaps = 7/115 (6%)
Query: 208 PVWSQTGKVFAPSKVNASAAPAAPSAPPPPPASLFSSESSQASS-----SKPKVGMSAVF 262
P S P A+ PA P P PPP S + + S P VG+ +
Sbjct: 120 PPLSPPPPAITPPPPLATTPPALPPKPLPPPLSPPQTTPPPPPAITPPLSPPLVGICS-- 177
Query: 263 QEIGTGNVTAGLRKVTDDMKTKNRADRSGVVGNSVKESQAAPRAFSKVGPPKLEL 317
+ + AG+ ++D + T RA+ + +V + A VG P+ L
Sbjct: 178 KNDTELKICAGILAISDGLLTTGRAEPCCSIIRNVSDLDAVTCFCKSVGAPRFSL 232
>At4g04310 putative transposon protein
Length = 1011
Score = 30.8 bits (68), Expect = 1.7
Identities = 34/123 (27%), Positives = 55/123 (44%), Gaps = 25/123 (20%)
Query: 304 PRAFSKVGPPKLELQMGRKWVVENQIDQKSLVIEDCDSKQSVYVYGCKNSVLQIQGKVNN 363
P+ PK+ + + VV Q Q SL + KQ CK+S L K
Sbjct: 351 PKKVKSTSQPKMVTE---EVVVHKQNRQPSLASSG-EQKQ------CKSSNLS---KSRG 397
Query: 364 ITIDNCKKTGVVFKDVVAAFEVVNSNGVEVQCQGSAPTIS-------VDNTSGC--QIYL 414
+T NC+K G + FE+ N++ ++ + S+P S ++N+S C ++L
Sbjct: 398 VTCYNCRKKGHLVATCPEKFELTNTS---LESKLSSPNSSSEVVSQGLENSSSCVTHLFL 454
Query: 415 SKD 417
SKD
Sbjct: 455 SKD 457
>At1g20640 unknown protein
Length = 844
Score = 30.8 bits (68), Expect = 1.7
Identities = 23/90 (25%), Positives = 41/90 (45%), Gaps = 12/90 (13%)
Query: 205 PLGPVWSQTGKVFAPSKVNASAAPAAPSAPPPPPASL---------FSSESSQASSSKPK 255
P+G ++ + + S+ + A P PPPPP L SS SSQ SS+ +
Sbjct: 628 PIGSFYANFPNLVSQSQEPSQQAKTTP--PPPPPVQLAKSPVSSYSHSSNSSQCCSSETQ 685
Query: 256 VGMSAVFQEIGTGNVTAGLRKVTDDMKTKN 285
+ A T +V L+K + +++ ++
Sbjct: 686 LNSGATTDPPST-DVGGALKKTSSEIELQS 714
>At4g23890 unknown protein
Length = 250
Score = 30.4 bits (67), Expect = 2.2
Identities = 13/25 (52%), Positives = 15/25 (60%)
Query: 231 PSAPPPPPASLFSSESSQASSSKPK 255
P PPPPPA S S+ A+ KPK
Sbjct: 138 PPPPPPPPAKQGSDASAVATDKKPK 162
>At4g00200 putative transcription factor
Length = 345
Score = 30.4 bits (67), Expect = 2.2
Identities = 17/48 (35%), Positives = 22/48 (45%), Gaps = 6/48 (12%)
Query: 219 PSKVNASAAPAA------PSAPPPPPASLFSSESSQASSSKPKVGMSA 260
P K APAA P +PPPP AS+FS + G+S+
Sbjct: 276 PRKQRVEHAPAAVMSVPPPPSPPPPAASVFSPTNPDREQPPSSFGISS 323
>At5g38560 putative protein
Length = 681
Score = 30.0 bits (66), Expect = 2.9
Identities = 17/44 (38%), Positives = 23/44 (51%), Gaps = 1/44 (2%)
Query: 205 PLGPVWSQTGKVFAPSKVNASA-APAAPSAPPPPPASLFSSESS 247
P G S G+ +P K + S P ++PPPPPA+ S SS
Sbjct: 141 PPGETPSPPGETPSPPKPSPSTPTPTTTTSPPPPPATSASPPSS 184
>At3g57500 hypothetical protein
Length = 139
Score = 30.0 bits (66), Expect = 2.9
Identities = 13/30 (43%), Positives = 16/30 (53%)
Query: 226 AAPAAPSAPPPPPASLFSSESSQASSSKPK 255
A AAPS PPPPP + S A+ + K
Sbjct: 109 AVSAAPSPPPPPPPATAEERSKPAAEEQTK 138
>At1g73840 proline-rich like protein precursor
Length = 388
Score = 30.0 bits (66), Expect = 2.9
Identities = 16/62 (25%), Positives = 26/62 (41%)
Query: 194 PGLRDYVKSFHPLGPVWSQTGKVFAPSKVNASAAPAAPSAPPPPPASLFSSESSQASSSK 253
P ++ ++ SF G V + + + P PPPPP+ +S SQ S
Sbjct: 94 PNVQAHMSSFQGGGSVHEPANTMQPQAPIRKHPTPQPMPMPPPPPSVSANSAQSQPRFSH 153
Query: 254 PK 255
P+
Sbjct: 154 PQ 155
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.312 0.128 0.363
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,444,649
Number of Sequences: 26719
Number of extensions: 446125
Number of successful extensions: 2977
Number of sequences better than 10.0: 56
Number of HSP's better than 10.0 without gapping: 23
Number of HSP's successfully gapped in prelim test: 34
Number of HSP's that attempted gapping in prelim test: 2660
Number of HSP's gapped (non-prelim): 262
length of query: 472
length of database: 11,318,596
effective HSP length: 103
effective length of query: 369
effective length of database: 8,566,539
effective search space: 3161052891
effective search space used: 3161052891
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 62 (28.5 bits)
Medicago: description of AC146746.4


