

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
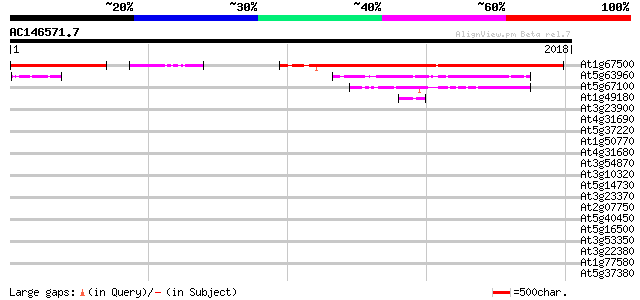
Query= AC146571.7 + phase: 0 /pseudo
(2018 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At1g67500 putative DNA polymerase zeta catalytic subunit 1252 0.0
At5g63960 DNA polymerase III catalytic subunit 283 7e-76
At5g67100 DNA polymerase alpha 1 128 4e-29
At1g49180 similar to MAP/ERK kinase kinase 3 gi|4505153 46 2e-04
At3g23900 unknown protein 38 0.053
At4g31690 putative protein 37 0.15
At5g37220 putative protein 35 0.45
At1g50770 hypothetical protein 35 0.58
At4g31680 putative protein 34 0.99
At3g54870 kinesin-like protein 34 0.99
At3g10320 unknown protein 34 0.99
At5g14730 unknown protein 33 1.3
At3g23370 hypothetical protein 33 1.3
At2g07750 putative RNA helicase 33 1.3
At5g40450 unknown protein 33 1.7
At5g16500 protein kinase-like protein 33 1.7
At3g53350 unknown protein 33 2.2
At3g22380 hypothetical protein 33 2.2
At1g77580 unknown protein 33 2.2
At5g37380 unknown protein 32 2.9
>At1g67500 putative DNA polymerase zeta catalytic subunit
Length = 1871
Score = 1252 bits (3239), Expect = 0.0
Identities = 656/1038 (63%), Positives = 793/1038 (76%), Gaps = 42/1038 (4%)
Query: 970 EDDEMPGNALDV-FLPISARNSQKQKKPWNKCVTIETPRSSGTKGVSTYYQNDGSHLYLL 1028
+ DE GN+ D+ F P+ +++++KK + + ++ G+ ++ NDGS+LYLL
Sbjct: 857 QHDEYEGNSNDIPFFPLE--DNKEEKKHFFQGTSL---------GIPLHHLNDGSNLYLL 905
Query: 1029 TPNILPPSAGSVQRWLFCDERDQEPDAEDQDVPKCTSGPLRHTPDQMRQEPGAKDKDISK 1088
TP PPS SV +W+ D+ D D+E Q PLR + GA D++
Sbjct: 906 TPAFSPPSVDSVLQWISNDKGDSNIDSEKQ--------PLRDN----HNDRGASFTDLAS 953
Query: 1089 CASGPTLRPELHQD------------TEKKLPCINEGQTERIKAHMDHSQDISQISGPGE 1136
++ ++ + Q TE ++ +G + SQ++SQISGP
Sbjct: 954 ASNVVSVSEHVEQHNNLFVNSESNAYTESEIDLKPKGTFLNLNLQASVSQELSQISGPDG 1013
Query: 1137 KSSFTPLSQIGFQDPASAGHGQQLALLSIEVLAESRGDLLPDPQFDAVNIVALGFQKDCD 1196
KS TPLSQ+GF+DPAS G GQQL +LSIEV AESRGDL PDP+FD+VN++AL Q D
Sbjct: 1014 KSGPTPLSQMGFRDPASMGAGQQLTILSIEVHAESRGDLRPDPRFDSVNVIALVVQNDDS 1073
Query: 1197 ATVEVLVLLHSKFVPCQRSLDGLSDCKVLNFTDEKHLLKEFTKIVSSSDPDILMGWEIQG 1256
EV VLL S QR++DGLS CK+ F +E+ L + F + + DPD+L+GW+IQG
Sbjct: 1074 FVAEVFVLLFSPDSIDQRNVDGLSGCKLSVFLEERQLFRYFIETLCKWDPDVLLGWDIQG 1133
Query: 1257 SSLGFLAERASHLGFGLLNDLSRTPSNSWINSQDIKTSEKSILEPDIPDTTSLDCCARES 1316
S+GFLAERA+ LG LN++SRTPS + N+ D K + L +PD + E
Sbjct: 1134 GSIGFLAERAAQLGIRFLNNISRTPSPTTTNNSDNKRKLGNNL---LPDPLVANPAQVEE 1190
Query: 1317 SIIEDEWGRTHASGVHVGGRIVLNLWRLIRGEVKLNLYSVEAVAEAVLRRKVPSLNHNVL 1376
+IEDEWGRTHASGVHVGGRIVLN WRLIRGEVKLN+Y++EAV+EAVLR+KVPS+ + VL
Sbjct: 1191 VVIEDEWGRTHASGVHVGGRIVLNAWRLIRGEVKLNMYTIEAVSEAVLRQKVPSIPYKVL 1250
Query: 1377 TKWFSRGPGQARYQCIKYIVDRAKLNLEILNQLDMVNRTSELARVFGIDFFSVLSRGSQY 1436
T+WFS GP ARY+CI+Y++ RA LNLEI++QLDM+NRTSELARVFGIDFFSVLSRGSQY
Sbjct: 1251 TEWFSSGPAGARYRCIEYVIRRANLNLEIMSQLDMINRTSELARVFGIDFFSVLSRGSQY 1310
Query: 1437 RVESMFLRLAHTQNYVAISPGNQQVASQPAMECLPLVMEPESGFYSDPVVVLDFQSLYPS 1496
RVESM LRLAHTQNY+AISPGNQQVASQPAMEC+PLVMEPES FY DPV+VLDFQSLYPS
Sbjct: 1311 RVESMLLRLAHTQNYLAISPGNQQVASQPAMECVPLVMEPESAFYDDPVIVLDFQSLYPS 1370
Query: 1497 MIIAYNLCFCTCLGKVVPSKTNTLGVSPFVPEQHILQDLKDQILLTPNGVMFVPSKIRRG 1556
MIIAYNLCF TCLGK+ K NTLGVS + + +LQDL +QIL TPN VM+VP ++RRG
Sbjct: 1371 MIIAYNLCFSTCLGKLAHLKMNTLGVSSYSLDLDVLQDL-NQILQTPNSVMYVPPEVRRG 1429
Query: 1557 VLPRLLEEILSTRIMVKQAMKKLSPSEQVLQRIFNARQLALKLISNVTYGYTAAGFSGRM 1616
+LPRLLEEILSTRIMVK+AMKKL+PSE VL RIFNARQLALKLI+NVTYGYTAAGFSGRM
Sbjct: 1430 ILPRLLEEILSTRIMVKKAMKKLTPSEAVLHRIFNARQLALKLIANVTYGYTAAGFSGRM 1489
Query: 1617 PCAELADSIVQCGRCTLEKAISFVNLHEKWNAKVIYGDTDSMFVLLKGRTAKEAFQIGSE 1676
PCAELADSIVQCGR TLEKAISFVN ++ WNA+V+YGDTDSMFVLLKGRT KEAF +G E
Sbjct: 1490 PCAELADSIVQCGRSTLEKAISFVNANDNWNARVVYGDTDSMFVLLKGRTVKEAFVVGQE 1549
Query: 1677 IASAITAMNPSPVTLKMEKVYHPCFLITKKRYVGYSYENPNQIEPVFDAKGIETVRRDTC 1736
IASAIT MNP PVTLKMEKVYHPCFL+TKKRYVGYSYE+PNQ EP+FDAKGIETVRRDTC
Sbjct: 1550 IASAITEMNPHPVTLKMEKVYHPCFLLTKKRYVGYSYESPNQREPIFDAKGIETVRRDTC 1609
Query: 1737 EAVAKIMEQSLRLFFEHQSLLEVKTYLQRQWKRILSGRVSLKDFIFAKEVRLGTYSARIS 1796
EAVAK MEQSLRLFFE +++ +VK+YL RQWKRILSGRVSL+DFIFAKEVRLGTYS R S
Sbjct: 1610 EAVAKTMEQSLRLFFEQKNISKVKSYLYRQWKRILSGRVSLQDFIFAKEVRLGTYSTRDS 1669
Query: 1797 S-LPPAAIVATKAMRVDRRAEPRYAERIPYVVIHGEPGARLVDMVVDPLEVLAIDSPFRI 1855
S LPPAAIVATK+M+ D R EPRYAER+PYVVIHGEPGARLVDMVVDPL +L +D+P+R+
Sbjct: 1670 SLLPPAAIVATKSMKADPRTEPRYAERVPYVVIHGEPGARLVDMVVDPLVLLDVDTPYRL 1729
Query: 1856 NDLYYINKQIIPALQRVFGLVGADLNQWFAEMPRPVREASVKHAF-TPNFQRTRIDYYYL 1914
NDLYYINKQIIPALQRVFGLVGADLNQWF EMPR R + + + N +TRIDY+YL
Sbjct: 1730 NDLYYINKQIIPALQRVFGLVGADLNQWFLEMPRLTRSSLGQRPLNSKNSHKTRIDYFYL 1789
Query: 1915 SKHCVLCGGLVQASARLCSQCSENEVAAATAVIGKTAKLEQEMQHLVSICQHCGGGDRLL 1974
SKHC+LCG +VQ SA+LC++C +N+ AAA ++ KT+KLE+EMQHL ++ LL
Sbjct: 1790 SKHCILCGEVVQESAQLCNRCLQNKSAAAATIVWKTSKLEREMQHLATVSTPKSNVYELL 1849
Query: 1975 ESGVKCTSISCLVFYERR 1992
++ V T+I L +R
Sbjct: 1850 QNLVTFTTIILLSLLSKR 1867
Score = 399 bits (1026), Expect = e-111
Identities = 206/366 (56%), Positives = 263/366 (71%), Gaps = 18/366 (4%)
Query: 1 MSDSESNSDVFSVRIVSVDHYMASPIPSIDISHSTFHRGKVNEVPVIRVYGSTPAGQKTC 60
M+DS+S S+VFS+RIVS+D+YMASPIP +I +S+F +VNEVPVIR+YGSTPAGQKTC
Sbjct: 1 MADSQSGSNVFSLRIVSIDYYMASPIPGYNICYSSFQGSEVNEVPVIRIYGSTPAGQKTC 60
Query: 61 LHIHGALPYLYVSCSDIPLQLDQEGDAYTYMVASSLEKALKLKGSADSSRQHVHGCSLVR 120
LHIH ALPYLY+ CS+IPL+ + D T ++ LEKALKLKG+A S RQH+H C +VR
Sbjct: 61 LHIHRALPYLYIPCSEIPLEHHKGVDGSTLALSLELEKALKLKGNAASKRQHIHDCEIVR 120
Query: 121 AKKFYGYCSFEEFFVKIYL----RYYPQDVSRAANLLLAGAVLNKSLQPHESHIPFILQF 176
AKKFYGY S EE FVKIYL Y+P DV+RAA+LLLAGAVL KSLQP+ESHIPFILQF
Sbjct: 121 AKKFYGYHSTEEAFVKIYLYPYSSYHPPDVARAASLLLAGAVLGKSLQPYESHIPFILQF 180
Query: 177 LVDYNLYGMGHLHLSKMRFRHPMPDSIHKK----SDINSQH------RKADPGAEACLES 226
LVDYNLYGMGH+H+SKM+FR P+P + D Q KA+ A A +
Sbjct: 181 LVDYNLYGMGHVHISKMKFRSPVPHHFRPRRFDLDDCPGQRIDEVAITKANSSAAASVSF 240
Query: 227 KIWMSSTISFDWMWSLPSES----GGSSNDNTHSPKRQSICELEGDISVDEILNQQFKMF 282
+W STI WMW+L ES S + + H +RQS+CELEGD + +ILNQQFKM+
Sbjct: 241 PVWSLSTIPGQWMWNLSEESDTPLSQSQHRHQHHYRRQSLCELEGDATSSDILNQQFKMY 300
Query: 283 SSLSQTSSNVNMVQSLVSIWEEQRKRSGIHEATMLSDSGKPLPEDVMKLLSSGLDFEKKL 342
+SLSQ S+ NMVQSLV+IWEE+ +R+G+H+A + D GKP DV++ +S + F L
Sbjct: 301 NSLSQAQSDTNMVQSLVAIWEEEYERTGVHDAPIPPDPGKPSAADVLQTMSDYVGFGNML 360
Query: 343 IQLCSE 348
++ ++
Sbjct: 361 KEMLNK 366
Score = 120 bits (302), Expect = 6e-27
Identities = 92/269 (34%), Positives = 137/269 (50%), Gaps = 14/269 (5%)
Query: 430 QLKRLKSQNLLKWLATSQAAEDINSDDELACETILSPLPPAATIDKMLEKANMAYESESQ 489
Q++ ++ L KW A+SQAAEDINSDDE+ ETILSPL P A+I+K+LE A+ Y S+SQ
Sbjct: 443 QIQDAEALGLFKWFASSQAAEDINSDDEILRETILSPLLPLASINKVLEMASTDYVSQSQ 502
Query: 490 QECQDILDSIDDMLALDLPKKKPSRSFDHNCPIEASSDNMIPQVDGSNDDEFSSPCASLA 549
+ECQDILDS +++ K+ S + + SSD +++ ++D SS +
Sbjct: 503 KECQDILDSQENLPDFGSSTKRALPSNPDSQNLRTSSDKQSLEIEVASDVPDSSTSNGAS 562
Query: 550 EISSAVEINSEYKRASENHLLHNTDTSTVNTDKRNRQWGSLPFSMTGKVNNDGEHATSQV 609
E S S+ SE N S N N WG LPF++T + D + +
Sbjct: 563 ENSFRRYRKSDL-HTSEVMEYKNRSFSKSNKPS-NSVWGPLPFTLTKNLQKDFDSTNA-- 618
Query: 610 AHLFERETEDSSLSDYLTRNEIKNNKYIKRNVGEGASD---SKEVHSLVNCSLRDLMRRK 666
+ + +S Y NE+ +N + V E +D + + + L CSLRDLMR+K
Sbjct: 619 ----SDKLGLTKISSY-PMNEMTDNYIVP--VKEHQADVCNTIDRNVLAGCSLRDLMRKK 671
Query: 667 RSYRVEHDERESGTAKKLNLDRHGGTKTC 695
R E + ++K+ RHG C
Sbjct: 672 RLCHGESPVSQHMKSRKVRDSRHGEKNEC 700
>At5g63960 DNA polymerase III catalytic subunit
Length = 1081
Score = 283 bits (724), Expect = 7e-76
Identities = 211/725 (29%), Positives = 340/725 (46%), Gaps = 78/725 (10%)
Query: 1162 LLSIEVLAESRGDLLPDPQFDAVNIVALGFQKDCDATVEVLVLLHSKFVPCQRSLDGLSD 1221
+LS ++ R P+ + D V +A LV L + P R++ L
Sbjct: 306 VLSFDIECAGRKGHFPEAKHDPVIQIAN------------LVTLQGEDHPFVRNVMTLKS 353
Query: 1222 CK------VLNFTDEKHLLKEFTKIVSSSDPDILMGWEIQGSSLGFLAERASHLGFGLLN 1275
C V++F E+ +L + ++ DPDI++G+ I L +L ERA+ LG
Sbjct: 354 CAPIVGVDVMSFETEREVLLAWRDLIRDVDPDIIIGYNICKFDLPYLIERAATLGIEEFP 413
Query: 1276 DLSRTPSNSWINSQDIKTSEKSILEPDIPDTTSLDCCARESSIIEDEWGRTHASGVHVGG 1335
L R +K S + R+S+ + G + + G
Sbjct: 414 LLGR-----------VKNSRVRV---------------RDSTFSSRQQGIRESKETTIEG 447
Query: 1336 RIVLNLWRLIRGEVKLNLYSVEAVAEAVLRRKVPSLNHNVLTKWFSRGPGQARYQCIKYI 1395
R +L + I + KL+ YS+ +V+ L + ++H+++T G + R + Y
Sbjct: 448 RFQFDLIQAIHRDHKLSSYSLNSVSAHFLSEQKEDVHHSIITD-LQNGNAETRRRLAVYC 506
Query: 1396 VDRAKLNLEILNQLDMVNRTSELARVFGIDFFSVLSRGSQYRVESMFLRLAHTQNYVAIS 1455
+ A L +L++L + E+ARV G+ +L+RG +V S LR +N V
Sbjct: 507 LKDAYLPQRLLDKLMFIYNYVEMARVTGVPISFLLARGQSIKVLSQLLRKGKQKNLVL-- 564
Query: 1456 PGNQQVASQPAMECLPLVMEPESGFYSDPVVVLDFQSLYPSMIIAYNLCFCTCLGKVVPS 1515
P +Q S+ V+E +GFY P+ LDF SLYPS+++AYNLC+CT V P
Sbjct: 565 PNAKQSGSEQGTYEGATVLEARTGFYEKPIATLDFASLYPSIMMAYNLCYCTL---VTPE 621
Query: 1516 KTNTLGVSPFVPEQHILQDLKDQILLTPNGVMFVPSKIRRGVLPRLLEEILSTRIMVKQA 1575
L + P + + TP+G FV +++G+LP +LEE+L+ R K+A
Sbjct: 622 DVRKLNLPP------------EHVTKTPSGETFVKQTLQKGILPEILEELLTAR---KRA 666
Query: 1576 MKKLSPSEQVLQR-IFNARQLALKLISNVTYGYTAAGFSGRMPCAELADSIVQCGRCTLE 1634
L ++ L++ + + RQLALK+ +N YG+T A G++PC E++ S+ GR +E
Sbjct: 667 KADLKEAKDPLEKAVLDGRQLALKISANSVYGFTGATV-GQLPCLEISSSVTSYGRQMIE 725
Query: 1635 KAISFVNLH------EKWNAKVIYGDTDSMFVLLKGRTAKEAFQIGSEIASAITAMNPSP 1688
+ V ++NA+VIYGDTDS+ V + A +G E A I+ P
Sbjct: 726 QTKKLVEDKFTTLGGYQYNAEVIYGDTDSVMVQFGVSDVEAAMTLGREAAEHISGTFIKP 785
Query: 1689 VTLKMEKVYHPCFLITKKRYVGYSYENPNQIEPVFDAKGIETVRRDTCEAVAKIMEQSLR 1748
+ L+ EKVY P LI KKRY G + NP Q + + D KGIETVRRD C V ++ +SL
Sbjct: 786 IKLEFEKVYFPYLLINKKRYAGLLWTNPQQFDKM-DTKGIETVRRDNCLLVKNLVTESLN 844
Query: 1749 LFFEHQSLLEVKTYLQRQWKRILSGRVSLKDFIFAKEVRLGTYSARISSLPPAAIVATKA 1808
+ + +++ +L R+ L + K + + S +A +
Sbjct: 845 KILIDRDVPGAAENVKKTISDLLMNRIDLSLLVITKGLTKTGDDYEVKS--AHGELAERM 902
Query: 1809 MRVDRRAEPRYAERIPYVVIHGEPGARLVDMVVDPLEVLAIDSPFRINDLYYINKQIIPA 1868
+ D P +R+PYV+I GA+ + DP+ VL + P I+ YY+ QI
Sbjct: 903 RKRDAATAPNVGDRVPYVIIKAAKGAKAYERSEDPIYVLQNNIP--IDPNYYLENQISKP 960
Query: 1869 LQRVF 1873
L R+F
Sbjct: 961 LLRIF 965
Score = 56.2 bits (134), Expect = 2e-07
Identities = 48/184 (26%), Positives = 84/184 (45%), Gaps = 21/184 (11%)
Query: 6 SNSDVFSVRIVSVDHYMASPIPSIDISHSTFHRGKVNEVPVIRVYGSTPAGQKTCLHIHG 65
SNS +F + +D +A SH G + P+IR++G T G C +HG
Sbjct: 90 SNSQIFQQ--LEIDSIIAE-------SHKELLPGSSGQAPIIRMFGVTREGNSVCCFVHG 140
Query: 66 ALPYLYVSCSDIPLQLDQEGDAYTYMVASSLEKALKLKGSADSSRQHVHGCSLVRAKKFY 125
PY Y++C P + + + + SLE ++ + V +V+ +
Sbjct: 141 FEPYFYIAC---PPGMGPDDISNFH---QSLEGRMRESNKNAKVPKFVKRIEMVQKRSIM 194
Query: 126 GYCSFE-EFFVKIYLRYYPQDVSRAANLLLAGAVLN----KSLQPHESHIPFILQFLVDY 180
Y + + F+KI + P V+ +L G ++ KS Q +ES+I F+L+F+VD
Sbjct: 195 YYQQQKSQTFLKITVA-LPTMVASCRGILDRGLQIDGLGMKSFQTYESNILFVLRFMVDC 253
Query: 181 NLYG 184
++ G
Sbjct: 254 DIVG 257
>At5g67100 DNA polymerase alpha 1
Length = 1492
Score = 128 bits (321), Expect = 4e-29
Identities = 166/704 (23%), Positives = 286/704 (40%), Gaps = 122/704 (17%)
Query: 1222 CKVLNFTD-EKHLLKEFTKIVSSSDPDILMGWEIQGSSLGFLAERASHLGFGLLNDLSRT 1280
C VL+ + E+ LL ++ D DIL+G I G L L +RA +
Sbjct: 640 CNVLSIENSERALLNRLFLELNKLDSDILVGHNISGFDLDVLLQRAQ---------ACKV 690
Query: 1281 PSNSWINSQDIKTSEKSILEPDIPDTTSLDCCARESSIIEDEWGRTHASGVHVGGRIVLN 1340
S+ W +K S P + ++ G T + GR++ +
Sbjct: 691 QSSMWSKIGRLKRS----FMPKLKGNSNYGS------------GATPGLMSCIAGRLLCD 734
Query: 1341 LWRLIRGEVKLNLYSVEAVAEAVLRRKVPSLNHNVLTKWFSRGPGQARYQCIKYIVDRAK 1400
R +K YS+ +++ L R + N + K F + + I+ A
Sbjct: 735 TDLCSRDLLKEVSYSLTDLSKTQLNRDRKEIAPNDIPKMFQSS--KTLVELIECGETDAW 792
Query: 1401 LNLEILNQLDMVNRTSELARVFGIDFFSVLSRGSQYRVESMFLRLAHTQNYVAISPGNQQ 1460
L++E++ L ++ T +L + G + L R+E L H++ ++ +Q+
Sbjct: 793 LSMELMFHLSVLPLTLQLTNISGNLWGKTLQGARAQRIEYYLLHTFHSKKFILPDKISQR 852
Query: 1461 V----ASQPAMECLP---------------------------------LVMEPESGFYSD 1483
+ +S+ M+ P LV+EP+ G Y
Sbjct: 853 MKEIKSSKRRMDYAPEDRNVDELDADLTLENDPSKGSKTKKGPAYAGGLVLEPKRGLYDK 912
Query: 1484 PVVVLDFQSLYPSMIIAYNLCFCTCLGKVVPSKTNTLGVSPFVPEQHILQDLKDQILLTP 1543
V++LDF SLYPS+I YN+CF T I +
Sbjct: 913 YVLLLDFNSLYPSIIQEYNICFTT-------------------------------IPRSE 941
Query: 1544 NGVMFVPSKIRRGVLPRLLEEILSTRIMVKQAMKKLSPSEQVLQRIFNARQLALKLISNV 1603
+GV +PS G+LP+L+E ++S R VK MKK + + RQ ALKL +N
Sbjct: 942 DGVPRLPSSQTPGILPKLMEHLVSIRKSVKLKMKK---ETGLKYWELDIRQQALKLTANS 998
Query: 1604 TYGYTAAGFS-GRMPCAELADSIVQCGRCTLEKAISFVNLHEKWNAKVIYGDTDSMFVLL 1662
YG GFS R LA+ I GR L++ + V H N +VIYGDTDS+ +
Sbjct: 999 MYG--CLGFSNSRFYAKPLAELITLQGRDILQRTVDLVQNH--LNLEVIYGDTDSIMIHS 1054
Query: 1663 KGRTAKEAFQIGSEIASAITAMNPSPVTLKM--EKVYHPCFLITKKRY--VGYSYENPNQ 1718
+E I S++ I +N LK+ + +Y L+ KK+Y V +++
Sbjct: 1055 GLDDIEEVKAIKSKV---IQEVNKKYRCLKIDCDGIYKRMLLLRKKKYAAVKLQFKDGKP 1111
Query: 1719 IEPVFDAKGIETVRRDTCEAVAKIMEQSLRLFFEHQSLLEVKTYLQRQWKRI----LSGR 1774
E + + KG++ VRRD +I + L S +V + + +I +G+
Sbjct: 1112 CEDI-ERKGVDMVRRDWSLLSKEIGDLCLSKILYGGSCEDVVEAIHNELMKIKEEMRNGQ 1170
Query: 1775 VSLKDFIFAKEVRLGTYSARISSLPPAAIVATKAMRVDRRAEPRYAERIPYVVIHGE--- 1831
V+L+ ++ K + + S P VA + + + + +PY++ + +
Sbjct: 1171 VALEKYVITKTLTKPPAAYPDSKSQPHVQVALRMRQRGYKEGFNAKDTVPYIICYEQGNA 1230
Query: 1832 ---PGARLVDMVVDPLEVLAIDSPFRINDLYYINKQIIPALQRV 1872
A + + P EV + S + ++ YY+ +QI P + R+
Sbjct: 1231 SSASSAGIAERARHPDEVKSEGSRWLVDIDYYLAQQIHPVVSRL 1274
>At1g49180 similar to MAP/ERK kinase kinase 3 gi|4505153
Length = 392
Score = 46.2 bits (108), Expect = 2e-04
Identities = 33/96 (34%), Positives = 48/96 (49%), Gaps = 6/96 (6%)
Query: 1399 AKLNLEILNQLDMVNRTSELARVFGIDFFSVLSRGSQYRVESMFLRLAHTQNYVAISPGN 1458
A L +L L + T E+ARV G+ ++SRG +V S LR A + +
Sbjct: 302 AYLPQRLLENLMYIYNTIEMARVTGVPTSYLISRGESIKVLSQLLRKAKQKTWFF----- 356
Query: 1459 QQVASQPAMECLPLVMEPESGFYSDPVVVLDFQSLY 1494
Q + S + L V+E +GFY +P+ LDF SLY
Sbjct: 357 QMLRSWNLNKELMKVLEARTGFY-EPIATLDFASLY 391
>At3g23900 unknown protein
Length = 989
Score = 38.1 bits (87), Expect = 0.053
Identities = 56/235 (23%), Positives = 87/235 (36%), Gaps = 46/235 (19%)
Query: 659 LRDLMRRKRSYRVEHDERESGTAKKLNLDRHGGTKTCLWPKLELETMQTDEVEKELQKNS 718
+++ R RS +E R S KL+ DR+ G++ ++ E +++
Sbjct: 754 MKERRGRSRSRSLETKNRSS-RKNKLDEDRNTGSRR----------RRSRSKSVEGKRSY 802
Query: 719 DHEVRSRDNLVCRKQSFPSDSDSFLNIS----------KDECLVQHERHYLEAGMVLKNS 768
+ E RSRD R+ S S S DE +H+RH + KNS
Sbjct: 803 NKETRSRDKKSKRRSGRRSRSPSSEGKQGRDIRSSPGYSDEKKSRHKRHSRSRSIEKKNS 862
Query: 769 ANQSSSSMHER------------------PVLLETSHLADSIDKSVACREKNTKDRTTYE 810
+ S HER PV E + S + +EK D YE
Sbjct: 863 SRDKRSKRHERLRSSSPGRDKRRGDRSLSPVSSEDHKIKKRHSGSKSVKEKPHSD---YE 919
Query: 811 K----HAASDAYTPNPSLDTHLRTDDDHKVRAPERCQETDSVASGSRQNSLVDDE 861
K A SD+ +L+ HL + D + E+ +E +S + DDE
Sbjct: 920 KVDDGDANSDSSQQERNLEGHLLSLDSMSSQDVEKSKENPPSSSSVERGDANDDE 974
>At4g31690 putative protein
Length = 461
Score = 36.6 bits (83), Expect = 0.15
Identities = 30/118 (25%), Positives = 53/118 (44%), Gaps = 13/118 (11%)
Query: 245 ESGGSSNDNTHSPKRQSICELEGDISVDEILNQQFKMFSSLSQTSSNVNMVQSLVSIWEE 304
E G S +N I E EG+ +V + ++ + SQ+ ++ N+ + VS+ +
Sbjct: 116 EEGEESGEN-------EISEKEGEENVQKESDKSSSDLNCFSQSVTHSNISRDAVSVPRD 168
Query: 305 QRKRSGI----HEATMLSDSGKPLPEDVMKLLSSGLDFEKKLIQLCSES--DTTLSCT 356
KRSG HE ++++ GK +V +S + C+E+ D SCT
Sbjct: 169 FVKRSGFSKGRHEIVLMNEEGKSWESEVKSYMSGAVYLVGGWTTFCTENKLDVGDSCT 226
>At5g37220 putative protein
Length = 214
Score = 35.0 bits (79), Expect = 0.45
Identities = 28/96 (29%), Positives = 47/96 (48%), Gaps = 5/96 (5%)
Query: 1490 FQSLYPSMIIAYNLC---FCTCLGKVVPSKTNTLGVSPFVPEQHILQDLKDQILLT-PNG 1545
F LY S ++ ++ FC LG+ + L P + E ++ +L ++L+T P+
Sbjct: 66 FSPLYISHLLQQSISHTRFCQDLGEKIAHNCAYLSGLPLIFEMFVVVELTQEVLITLPSA 125
Query: 1546 VMFVPSKIRRG-VLPRLLEEILSTRIMVKQAMKKLS 1580
+ VPSK VL RL+EE I +++ K S
Sbjct: 126 SVAVPSKAASSDVLQRLVEEQKVESIDLEEEKKTCS 161
>At1g50770 hypothetical protein
Length = 632
Score = 34.7 bits (78), Expect = 0.58
Identities = 50/228 (21%), Positives = 92/228 (39%), Gaps = 30/228 (13%)
Query: 641 VGEGASDSKEVHSLVNCSLRDLMRRK-RSYRVEHDERESGTAKKLNLDRHGGTKTCLWPK 699
+ + SK+ +S V RD + RVE D +SG +KL + G +T +
Sbjct: 233 IAQMVGPSKKKYSDVKNGGRDTSEPLGKKCRVEVDNNDSGPCQKLASTSYDGNETVPPSE 292
Query: 700 LELETMQTDEVEKELQKNSDHEVRSRDNLVCRKQSFPSDSDSFLNISKD-ECLVQHERHY 758
+E + TDE + KN +R + S++D + SK +CL+
Sbjct: 293 IEKKNEATDETGSKAGKNVVSPPETRKTCDGELDASKSNADMINDGSKQPKCLLH----- 347
Query: 759 LEAGMVLKNSANQSSSSMHERPVLLETSHLADSIDKSVACREKNTKDRTTYEKHAA---- 814
E G++ A S E+ E + D +D + N + ++++HA+
Sbjct: 348 -EDGIIAGEKA-----SSDEKLCGSEAAKEDDIVDDDIMI---NNNEPVSHQQHASGGTK 398
Query: 815 ----SDAY-----TPNPSLDTHLRTDDDHKVRAPERCQETDSVASGSR 853
S AY +P+ + H+ + + +RC+E +GS+
Sbjct: 399 GDETSRAYESVVLSPSDESNIHIAVAEGSQGH-EQRCEENGEEEAGSK 445
>At4g31680 putative protein
Length = 478
Score = 33.9 bits (76), Expect = 0.99
Identities = 26/113 (23%), Positives = 51/113 (45%), Gaps = 4/113 (3%)
Query: 241 SLPSESGGSSNDNTHSPKRQSICELEGDISVDEILNQQFKMFSSLSQTSSNVNMVQSLVS 300
S+ E G S + I E E + ++ + +Q + SQ+ ++ N+ LV+
Sbjct: 121 SISHEEGEESGEVGMIESSFKISEREVEENLQKEYDQSSADLNCFSQSVTHSNISNDLVT 180
Query: 301 IWEEQRKRSGI----HEATMLSDSGKPLPEDVMKLLSSGLDFEKKLIQLCSES 349
+ + KR+G+ HE ++++ GK +V +S + LCSE+
Sbjct: 181 LPRDFAKRNGLDKGMHEIVLMNEEGKSWESEVRSRMSGQVFIVGGWKSLCSEN 233
>At3g54870 kinesin-like protein
Length = 1070
Score = 33.9 bits (76), Expect = 0.99
Identities = 25/126 (19%), Positives = 55/126 (42%), Gaps = 11/126 (8%)
Query: 701 ELETMQTDEVEKELQKNSDHEVRSRDNLVCRKQSFPSDSDSF----------LNISKDEC 750
+L + E+EK L++ + + N V R + ++ L + KD+C
Sbjct: 495 KLRNSEKHELEKRLRECENSFAEAEKNAVTRSKFLEKENTRLELSMKELLKDLQLQKDQC 554
Query: 751 LVQHERHYLEAGMVLKNSANQSSSSMHERPVLLETSHLADSIDKSVACREKNTKDRTTYE 810
+ H++ ++ M LKN+ Q + L +TS + + + R ++ + R+T
Sbjct: 555 DLMHDKA-IQLEMKLKNTKQQQLENSAYEAKLADTSQVYEKKIAELVQRVEDEQARSTNA 613
Query: 811 KHAASD 816
+H ++
Sbjct: 614 EHQLTE 619
>At3g10320 unknown protein
Length = 494
Score = 33.9 bits (76), Expect = 0.99
Identities = 24/108 (22%), Positives = 50/108 (46%), Gaps = 13/108 (12%)
Query: 254 THSPKRQSICELEGDISVDEILNQQFKMF-----SSLSQTSSNVNMVQSLVSIWEEQRKR 308
THSP D++ D++L ++ K + +S+ +T + +V + ++ ++RK
Sbjct: 120 THSPSSSIFLYTSNDLTTDQVLQEKIKPYTRKWETSIMETIPELKLVTKDMKLFGDKRKC 179
Query: 309 SGIHE--ATMLSDSG------KPLPEDVMKLLSSGLDFEKKLIQLCSE 348
IHE A + S G + ++ L + F KK++ + +E
Sbjct: 180 EVIHEVPAVLFSTGGYTGNLYHEFNDGLIPLYITSKRFNKKVVFVIAE 227
>At5g14730 unknown protein
Length = 246
Score = 33.5 bits (75), Expect = 1.3
Identities = 34/153 (22%), Positives = 64/153 (41%), Gaps = 23/153 (15%)
Query: 532 QVDGSNDDEFSSPCASLAEIS--SAVEINSEYKRASENHLLHNTDTSTVNTDKRNRQWG- 588
+VDG ND EF L + S + + +KR+S++ D + +N +G
Sbjct: 43 KVDGENDKEFEFETLPLGDESLFHFSTMTTTFKRSSDDD-----DVADDRVTSKNLFYGG 97
Query: 589 ---------SLPFSMTGKVNNDGEHATSQVAHLF----ERETEDSSLSDYLTRNEIKNNK 635
S S++ ++D E+ + + F D SL + + +
Sbjct: 98 WSVDPPSLTSPSESLSSSDSDDSENISPSKFYCFWSPIRSPARDDSLKSKGSSRRCRIKE 157
Query: 636 YIKRNVGEGASDSKEVHSLVNCSLRDLMRRKRS 668
+++R+ +GA + V + CS +DL+RR S
Sbjct: 158 FLRRSHSDGAVST--VSATKRCSFKDLLRRSHS 188
>At3g23370 hypothetical protein
Length = 717
Score = 33.5 bits (75), Expect = 1.3
Identities = 64/336 (19%), Positives = 126/336 (37%), Gaps = 39/336 (11%)
Query: 628 RNEIKNNKYIKRNVGEGASDSKEVHSLVNCSLRDLMRRKRSYRVEHDERESGTAKKLNLD 687
R E +N + GE SD+ + HSL L D ++ +S E+DE A + NL
Sbjct: 215 RFETLSNSNSSMSTGESCSDAGDQHSLFGKMLSDPVQEIKSIPNENDEHRLSDAPQ-NLS 273
Query: 688 RHGGTKTCLWPKLELETMQTDEVEKELQKNSDHEVRSRDNLVC-----------RKQSFP 736
W ++ +++++ ++ S + S N V +FP
Sbjct: 274 SLTSVLLRDW----IDEQRSEDLAAIRERKSIFSLESSRNTVAPMAVGTMKKLMDSLNFP 329
Query: 737 SDSDSFLNISKDECLVQHERHYLEAGMVLKNSANQSSSSMHE---RPVLLETSHLA--DS 791
+D+ + + + E+ E ++NS + E ++ET L D+
Sbjct: 330 TDNGTDAEANSLNSNSREEKWISEGNSAMQNSDRFCAIEEEEPQGETFVMETQSLCSQDT 389
Query: 792 IDKSVACREKNTKD--RTTYEKHAASDA---YTPNPSLDTHLRTDDDHKVRAPERCQETD 846
+D + + K + +H+ + + P S+ H+ + A QE
Sbjct: 390 LDATTVNPKVTKKSFLALSAGEHSPNKVLLRFLPESSMKKHIVKAFSSQFGAVLHVQEIP 449
Query: 847 SVASGSRQNSLVDDEVSGKSKCIDKTSSGSIFFVQHDQMKSYEHAVGKSAASD--TRVLL 904
S+ +++L+ E + K K H + +Y V ++ D R+ +
Sbjct: 450 SIEGCIYKDALLTFETNTAVKKALKKG--------HVTVMNYNTVVEATSQEDMVERICI 501
Query: 905 TDKVDNQKLDKNLL---CKTVGSEPIVDDLKSNHMK 937
D + + + L+ +TV P+ D SN +K
Sbjct: 502 PDLIGDPDVPVALVKEPARTVKIHPLTHDFSSNQIK 537
>At2g07750 putative RNA helicase
Length = 845
Score = 33.5 bits (75), Expect = 1.3
Identities = 25/108 (23%), Positives = 46/108 (42%), Gaps = 7/108 (6%)
Query: 923 GSEPIVDDLKSNHMKLTKVV------IGNSSLVDKNLKSHLSLPTFPNN-LHLDEDDEMP 975
GS D + S + + ++V +G ++D + + S+ F N DE D++
Sbjct: 182 GSRSCSDSIDSTPIDVRRLVSATCDSMGKHRVLDSSRRGFSSMSRFKRNESSCDEGDDVD 241
Query: 976 GNALDVFLPISARNSQKQKKPWNKCVTIETPRSSGTKGVSTYYQNDGS 1023
LD P S + S ++K + + R+ G G + +ND S
Sbjct: 242 AKKLDTLSPFSPKFSGTKEKVKSSTSVVGVIRNKGLFGRRKFRKNDSS 289
>At5g40450 unknown protein
Length = 2910
Score = 33.1 bits (74), Expect = 1.7
Identities = 73/394 (18%), Positives = 148/394 (37%), Gaps = 63/394 (15%)
Query: 483 AYESESQQECQDILDSID-DMLALDLPKKKPSR-SFDHNCPIEASSDNM-IPQVDGSNDD 539
A E+E + DI + D D PK + S D P EA + + +P +D
Sbjct: 2474 ASEAEHEDPVDDIKSNDDRDFPTEQAPKDQSDEVSADETVPKEAIGEELKVPSSKVLDDI 2533
Query: 540 EFSS-------------PCASLAEI-------SSAVEINSEYKRASENHLLHNTDTSTVN 579
+ +S P +L+E+ + E + E+ +A E+ D N
Sbjct: 2534 QENSNTEAVTNFADRDLPVQNLSELIQSHQSPNQVEETSFEFNKAQEDKKEETVDALITN 2593
Query: 580 TDKRNRQWGSLPFSMTGKVNNDGEHATSQVAHLFERETEDSSLSDYLTRNEIKNNKY--- 636
+++ + K ++ EH + + + +E + + + +EIK + +
Sbjct: 2594 VQVQDQPKEDFEAAAIEKEISEQEHKLNDLTDV--QEDIGTYVKVQVPDDEIKGDGHDSV 2651
Query: 637 -IKRNVGEGASDSKEVHSLVNCSLRDLMRRKRSYRVEHDERES---GTAKKLNLDRHGGT 692
++ + +EV V + D ++ + S +++ E+ AK++ ++ G
Sbjct: 2652 AAQKEETSSIEEKREVEH-VKAEMEDAIKHEVSVEEKNNTSENIDHEAAKEI--EQEEGK 2708
Query: 693 KTCLWPKLELETMQTDEVEKELQKNSDHEVRSRDNLVCRKQSFPSDSDSFLNISKDECLV 752
+T + + E EKE+ + S + V+ D+ + + Q D +S ++SK +
Sbjct: 2709 QT------NIVKEEIREEEKEINQESFNNVKETDDAIDKTQPEIRDIESLSSVSKTQDKP 2762
Query: 753 QHERHYLEAGMVLKNSANQSSSSMHERPVLLETSHLADSIDKSVACREKNTKDRTTYEKH 812
+ E NQ + LE S + + + K +NTKD + K
Sbjct: 2763 EPEYEV----------PNQQKREITNEVPSLENSKIEEELQKKDE-ESENTKDLFSVVKE 2811
Query: 813 -----------AASDAYTPNPSLDTHLRTDDDHK 835
+ SD P + D+DH+
Sbjct: 2812 TEPTLKEPARKSLSDHIQKEPKTEEDENDDEDHE 2845
Score = 31.6 bits (70), Expect = 4.9
Identities = 92/424 (21%), Positives = 157/424 (36%), Gaps = 76/424 (17%)
Query: 446 SQAAEDINSDDELACETILSPLPPAATIDKMLEKANMAYESESQQECQDILDSIDDMLAL 505
S AED D+ + T SP ML + N E+++ + +D+ ++ L
Sbjct: 1752 SSEAEDDKPDEHVDSST--SP---------MLSEKN-DNETQTSKTSEDVCMQQEESGTL 1799
Query: 506 DLPKKKPSRSFDHNC------PIEASSDNMIPQVDGSNDDEFSSPCASLAEISSA----- 554
++PK + S+ IEA+SD +P D+ SS S + S
Sbjct: 1800 EVPKPEESKEDKSQEISETIEEIEATSDQTLPIETSHTDNTLSSELVSEQDDQSPKKVEE 1859
Query: 555 -----------VEINSEYKRASENHLLHNTDTSTVNTDKRNR---QWGS-LPFSMTGKVN 599
VE SE E NT +S + ++ + Q G LP + + +
Sbjct: 1860 IHEEEPKEAHDVEATSERNLPVETSDADNTLSSQLVSETKEEHKLQAGEILPTEIIPRES 1919
Query: 600 NDGEHATSQVAHLFERE------TEDSSLSDYLTRNEIKNNKYIKRNVGEGASDSKEVHS 653
+D + V+ L RE ED+ D N+I+ + I E ++K
Sbjct: 1920 SD----EALVSMLASREDDKVALQEDNCADDVRETNDIQEERSISVETEESVGETKPKEH 1975
Query: 654 LVNCSLRDLMRRKRSYRVEHDERESGT----AKKLNLDRHGGTKTCLW-----------P 698
+RD + + +E +S T AKK N + + +T P
Sbjct: 1976 --EDEIRDAHVETPTAPIILEENDSETLIAEAKKGNEEINETERTVALDHEEEFVNHEAP 2033
Query: 699 KLE----LETMQTDEVEKELQKNSDHEVRSRDNLVCRKQSFPSDSDSFLNISKDECLVQH 754
KLE ++ + E K + D + + + S SD D +E L +
Sbjct: 2034 KLEETKDEKSQEIPETAKATETTIDQTLPIGTSQADQTPSLVSDKDDQTPKQVEEILEEE 2093
Query: 755 --ERHYLEAGMVLKNSANQSSSSMHERPVLLETSHLADSIDKSVACREKNTKDRTTYEKH 812
E H ++A + ++ S E PV S LA D+ V +E + T E+H
Sbjct: 2094 TKETHKVQAEDIF-STETVPKESFIEAPV----SMLASGEDEPVTPQEGDYAANTQEERH 2148
Query: 813 AASD 816
+++
Sbjct: 2149 VSAE 2152
>At5g16500 protein kinase-like protein
Length = 636
Score = 33.1 bits (74), Expect = 1.7
Identities = 40/156 (25%), Positives = 65/156 (41%), Gaps = 17/156 (10%)
Query: 706 QTDEVEKELQKNSDHEVRSRDNLVCRKQSFPSDSDSFLNISKDECLVQHERHYLEAGMVL 765
+ +E EKE + + E S+ + + SD +S N KD+ + E+ LE
Sbjct: 407 EDEEEEKEQKAEKEEESTSKKRQEQEETATDSDDESDSNSEKDQ---EEEQSQLEKA--- 460
Query: 766 KNSANQSSSSMHERPVLLETSHLADSIDKSVACREKNTKDRTTYEKHAASDAYTPNPSLD 825
+ S++ SS S ER + ET+ A S+ S + +D EK ++ + N
Sbjct: 461 RESSSSSSDSGSERRSIDETNATAQSLKISYSNYSSEEEDN---EKLSSKSSCKSNEE-S 516
Query: 826 THLRTDDDHKVRAPERCQETDSVASGSRQNSLVDDE 861
T R D R + S + R NSL D+
Sbjct: 517 TFSRYDSG-------RDHDDSSRNTSMRINSLAHDD 545
>At3g53350 unknown protein
Length = 396
Score = 32.7 bits (73), Expect = 2.2
Identities = 31/120 (25%), Positives = 55/120 (45%), Gaps = 5/120 (4%)
Query: 614 ERETEDSSLSDYL--TRNEIKNNKYIKRNVGEGASDSKEVHSLVNCSLRDLMRRKRSYRV 671
E E+ S L + L + E+ ++ +KR E A D+K +N S + R
Sbjct: 75 ELESTISQLQEELKKAKEELNRSEALKREAQEEAEDAKHQLMDINASEDSRIEELRKLSQ 134
Query: 672 EHDERESGTAKKLNLDRHGGTKTCLWPKL-ELETMQTDEVEKELQ-KNSDHEVRSRDNLV 729
E D+ + + +HG T L + E++ +++ E E + + S +EVRS + LV
Sbjct: 135 ERDKTWQSELEAMQ-RQHGMDSTALSSAINEVQKLKSKLFESESELEQSKYEVRSLEKLV 193
>At3g22380 hypothetical protein
Length = 663
Score = 32.7 bits (73), Expect = 2.2
Identities = 50/223 (22%), Positives = 89/223 (39%), Gaps = 27/223 (12%)
Query: 445 TSQAAEDINSDDELACETILSPLPPAATIDKMLEKANMAYESESQQECQDILDSIDDMLA 504
TS ++ I+ +L T PLPP ++ K+ + +A + E + E ++L M+
Sbjct: 267 TSPSSSSISVRKKLPSGTKQKPLPPKSSSSKL--SSPVAVQDEIEIEIAEVLYG---MMR 321
Query: 505 LDLPKKKPSRSFD----HNCPIEASSDNMIPQVDGSNDDEFSSPCASLAEISSAVEINS- 559
+ K+ + D +E S P SN +LA SS+ +++
Sbjct: 322 MPSTSKQEAAGNDLTEAAKSTVEVKSRVSSPI---SNPQTLPQSSITLAANSSSSNVSAI 378
Query: 560 EYKRASENHLLHNTDTS----TVNTDKRNRQWGSLPFSMTGKVNNDGEHATSQVAHLFER 615
KR H+ + D S T+ ++ +PFS K + GE +S + +
Sbjct: 379 APKRKKPRHVKYEDDNSSRVTTIKSEAEAPSKSQVPFSNQLKSSGSGEGNSSVLDSIIPL 438
Query: 616 ETE-DSSLSDYLTRNEIKNNKYIK---------RNVGEGASDS 648
E ++SL N + ++ I R+ GEGA S
Sbjct: 439 TRESNASLDSEKKENNLSKDETILPKVESSSGFRSDGEGAKSS 481
>At1g77580 unknown protein
Length = 779
Score = 32.7 bits (73), Expect = 2.2
Identities = 73/315 (23%), Positives = 132/315 (41%), Gaps = 50/315 (15%)
Query: 440 LKWLATSQAAEDINSDDELA--CETILSPLPPAATIDKMLEKANMAYESESQQECQDILD 497
L +A S A E++ EL CE + TI++ L + A E+ S+Q+ LD
Sbjct: 182 LSQMAESVAKENVMLRHELLARCEEL-----EIRTIERDL--STQAAETASKQQ----LD 230
Query: 498 SIDDMLALDLPKKK------PSRSFDHNCPIEASSDNMIPQVDGSNDDEFSSPCA----S 547
SI + L+ +K S SF+ + ++ SD ++D S D ++S S
Sbjct: 231 SIKKVAKLEAECRKFRMLAKSSASFNDHRSTDSHSDGG-ERMDVSCSDSWASSTLIEKRS 289
Query: 548 LAEISSAVEINSEYKRASENHLLHNTDTSTVNTDKRNRQWGSLPFSMTGKVNNDGEHA-- 605
L SS++E++ L+ +T N S P S+T +V E++
Sbjct: 290 LQGTSSSIELDLMGDFLEMERLVALPETPDGNGK-------SGPESVTEEVVVPSENSLA 342
Query: 606 ------TSQVAHLFERETEDSSLSDYLTRNEIKNNKYIKRNVGEGASDSKEVHSLVNCSL 659
TS++ L E + E + NE+K N+ E A E ++
Sbjct: 343 SEIEVLTSRIKEL-EEKLEKLEAEKHELENEVKCNR-------EEAVVHIENSEVLTSRT 394
Query: 660 RDLMRRKRSYRVEHDERESGTAKKLNLDRHG-GTKTCLWPKLELETMQTDEVEKELQKNS 718
++L + E +E +S K N ++ + L ++E+ T +T E+E++L+K
Sbjct: 395 KELEEKLEKLEAEKEELKSEV--KCNREKAVVHVENSLAAEIEVLTSRTKELEEQLEKLE 452
Query: 719 DHEVRSRDNLVCRKQ 733
+V + C ++
Sbjct: 453 AEKVELESEVKCNRE 467
>At5g37380 unknown protein
Length = 431
Score = 32.3 bits (72), Expect = 2.9
Identities = 40/158 (25%), Positives = 64/158 (40%), Gaps = 27/158 (17%)
Query: 571 HNTDTSTVNTDKRNRQWGSLPFSMTGKVNNDGEHATSQVAHLFERETEDSSLSDYLTRNE 630
HNT + N N +W FS T + HA S V H +E+ +D + R E
Sbjct: 298 HNTTSGLFN----NSKW---TFSRT----SSAAHAASVVQHAYEKVKKDREQAKATARRE 346
Query: 631 IKNNKYIKRNVGEGASDSKEVHSLVNCSLRDL---MRRKRSYRVEHDERESGTAKKLNLD 687
KN K +++ + ++ + C D+ R+ +YRV + K +
Sbjct: 347 KKNAK--RKSTTDSSASGSSLKKRKVCLETDIGCSSGREVTYRVTGE-----NGKNMGKL 399
Query: 688 RHGGTKTCLWPKLELETMQTDEVEKELQKNSDHEVRSR 725
HG ++ KL L Q K ++N EV+SR
Sbjct: 400 EHGTKESA--DKLSLRRTQ----RKISKENVTREVKSR 431
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.133 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 45,510,465
Number of Sequences: 26719
Number of extensions: 2000099
Number of successful extensions: 5386
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 35
Number of HSP's that attempted gapping in prelim test: 5319
Number of HSP's gapped (non-prelim): 75
length of query: 2018
length of database: 11,318,596
effective HSP length: 114
effective length of query: 1904
effective length of database: 8,272,630
effective search space: 15751087520
effective search space used: 15751087520
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 68 (30.8 bits)
Medicago: description of AC146571.7


