

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146550.5 - phase: 0
(337 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
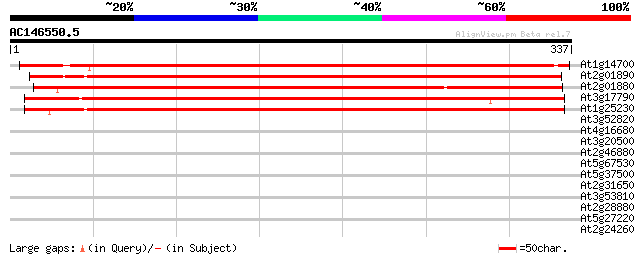
Score E
Sequences producing significant alignments: (bits) Value
At1g14700 purple acid phosphatase -like protein 411 e-115
At2g01890 purple acid phosphatase like protein 406 e-114
At2g01880 purple acid phosphatase like protein 405 e-113
At3g17790 acid phosphatase type 5 390 e-109
At1g25230 unknown protein 389 e-108
At3g52820 purple acid phosphatase (PAP22) 39 0.004
At4g16680 RNA helicase 32 0.64
At3g20500 purple acid phosphatase-like protein 32 0.64
At2g46880 unknown protein 30 1.4
At5g67530 cyclophilin (ROC23) 30 1.9
At5g37500 guard cell outward rectifying K+ channel 30 2.4
At2g31650 hypothetical protein 29 4.2
At3g53810 serine/threonine-specific kinase like protein 28 7.1
At2g28880 para-aminobenzoate synthase and glutamine amidotransfe... 28 7.1
At5g27220 putative protein 28 9.3
At2g24260 putative bHLH transcription factor (bHLH066) 28 9.3
>At1g14700 purple acid phosphatase -like protein
Length = 366
Score = 411 bits (1057), Expect = e-115
Identities = 197/333 (59%), Positives = 252/333 (75%), Gaps = 9/333 (2%)
Query: 7 QHMLSSVIIAVSVLLCLLSNHSSIAEKLPRFEHHLKPQQQ--SLNFLVVGDWGRKGNYNQ 64
+H +++ V L+ + S+HSS AE L+P + +++FLV+GDWGR+G+YNQ
Sbjct: 35 KHKPVNLVFYVYNLIIIFSSHSSTAE----LRRLLQPSKTDGTVSFLVIGDWGRRGSYNQ 90
Query: 65 SLVAHQMGIVGKNLNIEFVVSTGDNFYDDGLKGVDDPAFYESFVDIYTAPSLQQIWYNVL 124
S VA QMG +G+ L+I+FV+STGDNFYD+GL + DP F +SF +IYTAPSLQ+ WY+VL
Sbjct: 91 SQVALQMGEIGEKLDIDFVISTGDNFYDNGLTSLHDPLFQDSFTNIYTAPSLQKPWYSVL 150
Query: 125 GNHDYRGDVEAQLSPILRQKDNRWVCLRSFILDTDNVEFFFVDTTPFVEEYFTDPGKHTY 184
GNHDYRGDV AQLSP+LR DNRWVC+RSFI++ + V+ FFVDTTPFV++YF P KH Y
Sbjct: 151 GNHDYRGDVRAQLSPMLRALDNRWVCMRSFIVNAEIVDLFFVDTTPFVDKYFIQPNKHVY 210
Query: 185 DWKGVLPLESYRAELLKEVDSALVQSTAKWKIVVAHHPIKSAGPHGNTQELEEQLLPILK 244
DW GVLP ++Y LLKE+D AL +S AKWKIV+ HH IKSAG HGNT ELE+ LLPIL+
Sbjct: 211 DWSGVLPRQTYLNNLLKELDVALRESVAKWKIVIGHHTIKSAGHHGNTIELEKHLLPILQ 270
Query: 245 SNNVDAYINGHDHCLEHIIDKESGIHFFTSGGGSKAW-GGDIKPWDPKELKLYHDGQGFV 303
+N VD Y+NGHDHCLEHI +S I F TSGGGSKAW GGD+ +P+E++ Y+DGQGF+
Sbjct: 271 ANEVDLYVNGHDHCLEHISSVDSNIQFMTSGGGSKAWKGGDVNYVEPEEMRFYYDGQGFM 330
Query: 304 SMQISKTNASIVFYDVFGKVLHTWSMSKKLKEA 336
S+ +S+ +VFYDVFG VLH W K KEA
Sbjct: 331 SVHVSEAELRVVFYDVFGHVLHHW--KKTYKEA 361
>At2g01890 purple acid phosphatase like protein
Length = 335
Score = 406 bits (1044), Expect = e-114
Identities = 196/319 (61%), Positives = 241/319 (75%), Gaps = 2/319 (0%)
Query: 13 VIIAVSVLLCLLSNHSSIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQSLVAHQMG 72
+I ++ L+ +LS +S AE LPRF +P SL+FLVVGDWGR+G+YNQS VA QMG
Sbjct: 12 LIFSIFCLVIILSACNSTAE-LPRFVQPPEPDG-SLSFLVVGDWGRRGSYNQSQVALQMG 69
Query: 73 IVGKNLNIEFVVSTGDNFYDDGLKGVDDPAFYESFVDIYTAPSLQQIWYNVLGNHDYRGD 132
+GK+LNI+F++STGDNFYDDG+ D F +SF +IYTA SLQ+ WYNVLGNHDYRG+
Sbjct: 70 KIGKDLNIDFLISTGDNFYDDGIISPYDSQFQDSFTNIYTATSLQKPWYNVLGNHDYRGN 129
Query: 133 VEAQLSPILRQKDNRWVCLRSFILDTDNVEFFFVDTTPFVEEYFTDPGKHTYDWKGVLPL 192
V AQLSPILR D RW+CLRS++++ + V+ FFVDTTPFV+ YF +P H YDW+GVLP
Sbjct: 130 VYAQLSPILRDLDCRWICLRSYVVNAEIVDIFFVDTTPFVDRYFDEPKDHVYDWRGVLPR 189
Query: 193 ESYRAELLKEVDSALVQSTAKWKIVVAHHPIKSAGPHGNTQELEEQLLPILKSNNVDAYI 252
Y LL +VD AL +S AKWKIVV HH IKSAG HGNT ELE+QLLPIL++N VD YI
Sbjct: 190 NKYLNSLLTDVDVALQESMAKWKIVVGHHTIKSAGHHGNTIELEKQLLPILEANEVDLYI 249
Query: 253 NGHDHCLEHIIDKESGIHFFTSGGGSKAWGGDIKPWDPKELKLYHDGQGFVSMQISKTNA 312
NGHDHCLEHI SGI F TSGGGSKAW GD+ W+P+E++ Y+DGQGF+S+ S+
Sbjct: 250 NGHDHCLEHISSINSGIQFMTSGGGSKAWKGDVNDWNPQEMRFYYDGQGFMSVYTSEAEL 309
Query: 313 SIVFYDVFGKVLHTWSMSK 331
+VFYD G VLH WS K
Sbjct: 310 RVVFYDGLGHVLHRWSTLK 328
>At2g01880 purple acid phosphatase like protein
Length = 328
Score = 405 bits (1041), Expect = e-113
Identities = 194/321 (60%), Positives = 240/321 (74%), Gaps = 4/321 (1%)
Query: 15 IAVSVLLCLLSNH--SSIAEKLPRFEHHLKPQQQ-SLNFLVVGDWGRKGNYNQSLVAHQM 71
+ SV+L LS + KL R +H +K + SL+FLV+GDWGRKG +NQSLVAHQM
Sbjct: 5 VCFSVILMFLSIFFINGALSKLERLKHPVKKKSDGSLSFLVIGDWGRKGGFNQSLVAHQM 64
Query: 72 GIVGKNLNIEFVVSTGDNFYDDGLKGVDDPAFYESFVDIYTAPSLQQIWYNVLGNHDYRG 131
G+VG+ L+I+FV+S GDNFYDDGLKGV+DP+F SF IYT PSLQ+ WY+VLGNHDYRG
Sbjct: 65 GVVGEKLDIDFVISVGDNFYDDGLKGVNDPSFEASFSHIYTHPSLQKQWYSVLGNHDYRG 124
Query: 132 DVEAQLSPILRQKDNRWVCLRSFILDTDNVEFFFVDTTPFVEEYFTDPGKHTYDWKGVLP 191
+VEAQLS +L QKD RW C RSF+L + V+FFF DT PFVE+YFT+P HTYDW+ VLP
Sbjct: 125 NVEAQLSKVLTQKDWRWFCRRSFVLSSGMVDFFFADTNPFVEKYFTEPEDHTYDWRNVLP 184
Query: 192 LESYRAELLKEVDSALVQSTAKWKIVVAHHPIKSAGPHGNTQELEEQLLPILKSNNVDAY 251
Y + LL ++D + +S A WK VV HH IK+AG HG TQEL +QLLPIL+ N VD Y
Sbjct: 185 RNKYISNLLHDLDLEIKKSRATWKFVVGHHGIKTAGNHGVTQELVDQLLPILEENKVDLY 244
Query: 252 INGHDHCLEHIIDKESGIHFFTSGGGSKAWGGDIKPWDPKELKLYHDGQGFVSMQISKTN 311
INGHDHCL+H I F TSGGGSKAW G ++PWDPKELKLY+DGQGF+S+ I+ +
Sbjct: 245 INGHDHCLQH-IGSHGKTQFLTSGGGSKAWRGHVQPWDPKELKLYYDGQGFMSLHITHSK 303
Query: 312 ASIVFYDVFGKVLHTWSMSKK 332
A ++YDV G VLH S+SK+
Sbjct: 304 AKFIYYDVSGNVLHRSSLSKR 324
>At3g17790 acid phosphatase type 5
Length = 338
Score = 390 bits (1003), Expect = e-109
Identities = 184/327 (56%), Positives = 249/327 (75%), Gaps = 4/327 (1%)
Query: 10 LSSVIIAVSVLLCLLSNHSSIAE-KLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQSLVA 68
L S ++S+LLC+ + ++ +L RF K S++F+V+GDWGR+G++NQSLVA
Sbjct: 8 LMSATASLSLLLCIFTTFVVVSNGELQRFIEPAK-SDGSVSFIVIGDWGRRGSFNQSLVA 66
Query: 69 HQMGIVGKNLNIEFVVSTGDNFYDDGLKGVDDPAFYESFVDIYTAPSLQQIWYNVLGNHD 128
+QMG +G+ ++++FVVSTGDNFYD+GL DP F +SF +IYTAPSLQ+ WY+VLGNHD
Sbjct: 67 YQMGKIGEKIDLDFVVSTGDNFYDNGLFSEHDPNFEQSFSNIYTAPSLQKQWYSVLGNHD 126
Query: 129 YRGDVEAQLSPILRQKDNRWVCLRSFILDTDNVEFFFVDTTPFVEEYFTDPGKHTYDWKG 188
YRGD EAQLS +LR+ D+RW+CLRSF++D + VE FFVDTTPFV+EY+T+ H+YDW+
Sbjct: 127 YRGDAEAQLSSVLREIDSRWICLRSFVVDAELVEMFFVDTTPFVKEYYTEADGHSYDWRA 186
Query: 189 VLPLESYRAELLKEVDSALVQSTAKWKIVVAHHPIKSAGPHGNTQELEEQLLPILKSNNV 248
V SY LL++++ +L S A+WKIVV HH ++S G HG+T+EL E+LLPILK N V
Sbjct: 187 VPSRNSYVKALLRDLEVSLKSSKARWKIVVGHHAMRSIGHHGDTKELNEELLPILKENGV 246
Query: 249 DAYINGHDHCLEHIIDKESGIHFFTSGGGSKAWGGDIKP--WDPKELKLYHDGQGFVSMQ 306
D Y+NGHDHCL+H+ D++S I F TSG GSKAW GDI P +PK LK Y+DGQGF+S +
Sbjct: 247 DLYMNGHDHCLQHMSDEDSPIQFLTSGAGSKAWRGDINPVTINPKLLKFYYDGQGFMSAR 306
Query: 307 ISKTNASIVFYDVFGKVLHTWSMSKKL 333
+ ++A IVFYDVFG++LH W SK+L
Sbjct: 307 FTHSDAEIVFYDVFGEILHKWVTSKQL 333
>At1g25230 unknown protein
Length = 339
Score = 389 bits (998), Expect = e-108
Identities = 187/326 (57%), Positives = 239/326 (72%), Gaps = 3/326 (0%)
Query: 10 LSSVIIAVSVLLC--LLSNHSSIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQSLV 67
+ S+ I +++L+C LLS + +L +H P S++FLV+GDWGR G YNQS V
Sbjct: 7 IGSLSIVMTLLICFLLLSLAPKLEAELATVQHAPNPDG-SISFLVIGDWGRHGLYNQSQV 65
Query: 68 AHQMGIVGKNLNIEFVVSTGDNFYDDGLKGVDDPAFYESFVDIYTAPSLQQIWYNVLGNH 127
A QMG +G+ ++I FVVSTGDN YD+G+K +DDPAF SF +IYT+PSLQ+ WY VLGNH
Sbjct: 66 ALQMGRIGEEMDINFVVSTGDNIYDNGMKSIDDPAFQLSFSNIYTSPSLQKPWYLVLGNH 125
Query: 128 DYRGDVEAQLSPILRQKDNRWVCLRSFILDTDNVEFFFVDTTPFVEEYFTDPGKHTYDWK 187
DYRGDVEAQLSPILR D+RW+C+RSFI+D + E FFVDTTPFV+ YF P TYDW
Sbjct: 126 DYRGDVEAQLSPILRSMDSRWICMRSFIVDAEIAELFFVDTTPFVDAYFLSPQDQTYDWS 185
Query: 188 GVLPLESYRAELLKEVDSALVQSTAKWKIVVAHHPIKSAGPHGNTQELEEQLLPILKSNN 247
GV P +SY +L E++ L +S+AKWKIVV HH IKSA HGNT+ELE LLPIL++N
Sbjct: 186 GVSPRKSYLQTILTELEMGLRESSAKWKIVVGHHAIKSASIHGNTKELESLLLPILEANK 245
Query: 248 VDAYINGHDHCLEHIIDKESGIHFFTSGGGSKAWGGDIKPWDPKELKLYHDGQGFVSMQI 307
VD Y+NGHDHCL+HI +S I F TSGGGSKAW G P+++K ++DGQGF+S++I
Sbjct: 246 VDLYMNGHDHCLQHISTSQSPIQFLTSGGGSKAWRGYYNWTTPEDMKFFYDGQGFMSVKI 305
Query: 308 SKTNASIVFYDVFGKVLHTWSMSKKL 333
+++ S+VFYDV G LH W SK L
Sbjct: 306 TRSELSVVFYDVSGNSLHKWDTSKML 331
>At3g52820 purple acid phosphatase (PAP22)
Length = 434
Score = 38.9 bits (89), Expect = 0.004
Identities = 56/263 (21%), Positives = 95/263 (35%), Gaps = 48/263 (18%)
Query: 35 PRFEHHLKPQQQSLNFLVVGDWGRKGNYNQSLVAHQMGIVGKNLNIEFVVSTGDNFYDDG 94
P F P + F +VGD G+ + + ++H + + + + GD Y D
Sbjct: 128 PEFSFKTPPSTFPVEFAIVGDLGQT-EWTAATLSHI-----NSQDYDVFLLPGDLSYAD- 180
Query: 95 LKGVDDPAFYESFVDIYTAPSLQQIWYNVLGNHDYRGDVEAQLSPIL-----RQKDNRWV 149
++SF + + ++ W GNH E + PI+ + + RW+
Sbjct: 181 ----THQPLWDSFGRLVEPLASKRPWMVTEGNH------EIEFFPIIEHTTFKSYNARWL 230
Query: 150 CLRSFILDTDNVEFFF----VDTTPFVEEYFTDPGKHTYDWKGVLPLESYRAELLKEVDS 205
+ T N+ + F V T D Y W +A+L K VD
Sbjct: 231 MPHTESFSTSNLYYSFDVAGVHTVMLGSYTDFDCESDQYQW--------LQADLAK-VD- 280
Query: 206 ALVQSTAKWKIVVAHHPIKSAGP--HGNTQELEEQLLPILKSNNVDAYINGHDHCLEHI- 262
+ T W +V+ H P + G + + E + +L + VD +GH H E
Sbjct: 281 ---RKTTPWVVVLLHAPWYNTNEAHEGEGESMREAMESLLFNARVDVVFSGHVHAYERFK 337
Query: 263 ------IDKESGIHFFTSGGGSK 279
D IH GG++
Sbjct: 338 RVYNNKADPCGPIHITIGDGGNR 360
>At4g16680 RNA helicase
Length = 883
Score = 31.6 bits (70), Expect = 0.64
Identities = 26/85 (30%), Positives = 35/85 (40%), Gaps = 16/85 (18%)
Query: 159 DNVEFFFVDTTPFVEEYF-TDPGKH-----------TYDWKGVLPLESYRAELLKEVDS- 205
D EF F D T FVEE + GKH + + LP+ YR ELLK ++
Sbjct: 179 DGYEFVFDDLTGFVEESSEAETGKHRGCYSKTAAEKAREGREFLPIHGYREELLKLIEEN 238
Query: 206 ---ALVQSTAKWKIVVAHHPIKSAG 227
+V T K ++ AG
Sbjct: 239 QVLVIVGETGSGKTTQIPQYLQEAG 263
>At3g20500 purple acid phosphatase-like protein
Length = 437
Score = 31.6 bits (70), Expect = 0.64
Identities = 56/261 (21%), Positives = 97/261 (36%), Gaps = 49/261 (18%)
Query: 38 EHHLK--PQQQSLNFLVVGDWGRKGNYNQSLVAHQMGIVGKNLNIEFVVSTGDNFYDDGL 95
E HLK P Q + F V GD G+ G + +S + H + GD Y D +
Sbjct: 129 EFHLKTPPAQFPITFAVAGDLGQTG-WTKSTLDHI-----DQCKYAVHLLPGDLSYADYM 182
Query: 96 KGVDDPAFYESFVDIYTAPSLQQIWYNVLGNHDYRGDVEAQLSPILRQK----DNRWVCL 151
+ +++F ++ + + W GNH E + P + + ++RW
Sbjct: 183 QHK-----WDTFGELVQPLASVRPWMVTQGNH------EKESIPFIVDEFVSFNSRWKMP 231
Query: 152 RSFILDTDNVEFFF--VDTTPFVEEYFTDPGKHT--YDWKGVLPLESYRAELLKEVDSAL 207
N+ + F + +TD +++ Y W LK S +
Sbjct: 232 YEESGSNSNLYYSFEVAGVHAIMLGSYTDYDRYSDQYSW-------------LKADLSKV 278
Query: 208 VQSTAKWKIVVAHHP-IKSAGPHGNT-QELEEQLLPILKSNNVDAYINGHDHCLEHIIDK 265
+ W IV+ H P S H + E+ ++ P+L ++ VD GH H E
Sbjct: 279 DRERTPWLIVLFHVPWYNSNNAHQHEGDEMMAEMEPLLYASGVDIVFTGHVHAYERTKRV 338
Query: 266 ESG-------IHFFTSGGGSK 279
+G +H GG++
Sbjct: 339 NNGKSDPCGPVHITIGDGGNR 359
>At2g46880 unknown protein
Length = 401
Score = 30.4 bits (67), Expect = 1.4
Identities = 19/55 (34%), Positives = 29/55 (52%), Gaps = 7/55 (12%)
Query: 81 EFVVSTGDNFYDDGLKGVDDPAFYESFVDIYTAPSLQQ--IWYNVLGNHDYRGDV 133
+ +V +GDN Y GL D A +D+ AP+++ W +LGNHD D+
Sbjct: 94 DLIVFSGDNVY--GLCETSDVA---KSMDMAFAPAIESGIPWVAILGNHDQESDM 143
>At5g67530 cyclophilin (ROC23)
Length = 595
Score = 30.0 bits (66), Expect = 1.9
Identities = 20/46 (43%), Positives = 21/46 (45%), Gaps = 4/46 (8%)
Query: 273 TSGGGSKAWGGDIKPWDPKELKLYHDGQGFVSMQIS--KTNASIVF 316
T GG WG K D KL H G+G VSM S TN S F
Sbjct: 402 TGKGGESIWGKPFK--DEPNSKLLHSGRGVVSMANSGPHTNGSQFF 445
>At5g37500 guard cell outward rectifying K+ channel
Length = 819
Score = 29.6 bits (65), Expect = 2.4
Identities = 42/183 (22%), Positives = 71/183 (37%), Gaps = 14/183 (7%)
Query: 143 QKDNRWVCLRSFILDTDNVEFFFVDTTPFVEEYFTD---PGKHTYDWKGVLPLESYRAEL 199
+KD R L + IL VE + T + Y P Y W G L L Y E
Sbjct: 189 EKDTRINYLFTRILKLLFVEVYCTHTAACIFYYLATTLPPENEGYTWIGSLKLGDYSYEN 248
Query: 200 LKEVDSALVQSTAKWKIVVAHHPIKSAGPHG-NTQELEEQLLPILKSNNVDAYINGHDHC 258
+E+D +TA + +V + H N +E+ ++ + + AY+ G+
Sbjct: 249 FREIDLWKRYTTALYFAIVTMATVGYGDIHAVNLREMIFVMIYVSFDMVLGAYLIGNITA 308
Query: 259 L-------EHIIDKESGIHFFTSGGGSKAWGGDIKPWDPKELKLYHDGQGFVSMQISKTN 311
L E DK + + F + K G D++ ++L +D ++ +
Sbjct: 309 LIVKGSNTERFRDKMNDLISFMN---RKKLGRDLRSQITGHVRLQYDSHYTDTVMLQDIP 365
Query: 312 ASI 314
ASI
Sbjct: 366 ASI 368
>At2g31650 hypothetical protein
Length = 514
Score = 28.9 bits (63), Expect = 4.2
Identities = 16/47 (34%), Positives = 24/47 (51%), Gaps = 1/47 (2%)
Query: 98 VDDPAFYESFVDIYTAPSLQQIWYNVLGNHDYRG-DVEAQLSPILRQ 143
V+ P Y+S IY+ PS N +G+H V+AQ P++ Q
Sbjct: 18 VEAPIRYDSIESIYSIPSSALCCVNAVGSHSLMSKKVKAQKLPMIEQ 64
>At3g53810 serine/threonine-specific kinase like protein
Length = 677
Score = 28.1 bits (61), Expect = 7.1
Identities = 15/55 (27%), Positives = 28/55 (50%), Gaps = 7/55 (12%)
Query: 172 VEEYFTDPGKHTYDWKGVLPLESYRAELLKEVDSALVQSTAKWKIVVAHHPIKSA 226
+++Y + + T DWK R+ ++K V S L +W+ VV H +K++
Sbjct: 429 LDKYLYNNPETTLDWK-------QRSTIIKGVASGLFYLHEEWEQVVIHRDVKAS 476
>At2g28880 para-aminobenzoate synthase and glutamine
amidotransferase, a bifunctional enzyme like protein
Length = 919
Score = 28.1 bits (61), Expect = 7.1
Identities = 15/47 (31%), Positives = 27/47 (56%), Gaps = 10/47 (21%)
Query: 231 NTQELEEQLLPILKSN----------NVDAYINGHDHCLEHIIDKES 267
+T++LE+Q LP++ S+ + + YIN C+++I D ES
Sbjct: 617 STRKLEDQTLPVIDSSQSKTSFVPDKSREQYINDVQSCMKYIKDGES 663
>At5g27220 putative protein
Length = 1181
Score = 27.7 bits (60), Expect = 9.3
Identities = 19/64 (29%), Positives = 30/64 (46%), Gaps = 1/64 (1%)
Query: 42 KPQQQSLNFLVVGDW-GRKGNYNQSLVAHQMGIVGKNLNIEFVVSTGDNFYDDGLKGVDD 100
KP+QQ ++ R N +++ + +G V L+ E S N GL+G+ D
Sbjct: 813 KPEQQPPEAPIINSSDSRSTNVQETIASSHLGNVDVLLDPEGSTSFSPNEVFTGLQGMID 872
Query: 101 PAFY 104
PA Y
Sbjct: 873 PASY 876
>At2g24260 putative bHLH transcription factor (bHLH066)
Length = 350
Score = 27.7 bits (60), Expect = 9.3
Identities = 17/49 (34%), Positives = 26/49 (52%), Gaps = 4/49 (8%)
Query: 210 STAKWKIVVAH-HPIKSAGPHGNTQELEEQ---LLPILKSNNVDAYING 254
S+A W VV HP+ S G HG+ + Q ++P+ ++V A NG
Sbjct: 35 SSAPWPSVVDDAHPLPSDGFHGHDVDSRNQPIMMMPLNDGSSVHALYNG 83
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.137 0.427
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,375,375
Number of Sequences: 26719
Number of extensions: 392047
Number of successful extensions: 857
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 11
Number of HSP's that attempted gapping in prelim test: 843
Number of HSP's gapped (non-prelim): 19
length of query: 337
length of database: 11,318,596
effective HSP length: 100
effective length of query: 237
effective length of database: 8,646,696
effective search space: 2049266952
effective search space used: 2049266952
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 60 (27.7 bits)
Medicago: description of AC146550.5


