

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146329.6 + phase: 0
(486 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
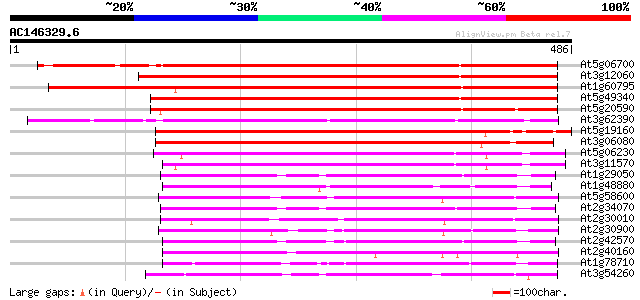
Score E
Sequences producing significant alignments: (bits) Value
At5g06700 unknown protein 496 e-140
At3g12060 hypothetical protein 490 e-139
At1g60795 unknown protein 431 e-121
At5g49340 putative protein 419 e-117
At5g20590 unknown protein 383 e-106
At3g62390 unknown protein 371 e-103
At5g19160 putative protein 320 1e-87
At3g06080 unknown protein 319 2e-87
At5g06230 unknown protein 284 7e-77
At3g11570 hypothetical protein 284 9e-77
At1g29050 unknown protein 272 3e-73
At1g48880 hypothetical protein 270 1e-72
At5g58600 unknown protein 270 1e-72
At2g34070 unknown protein 264 9e-71
At2g30010 unknown protein 262 4e-70
At2g30900 hypothetical protein 260 1e-69
At2g42570 unknown protein 255 4e-68
At2g40160 unknown protein 254 1e-67
At1g78710 hypothetical protein 251 6e-67
At3g54260 unknown protein 250 1e-66
>At5g06700 unknown protein
Length = 608
Score = 496 bits (1277), Expect = e-140
Identities = 233/450 (51%), Positives = 319/450 (70%), Gaps = 18/450 (4%)
Query: 25 NNSSSFSPPPSYTPQYSTNETTTHDKTSHSSNKDTELVVSSRSSSQISRKEGSGLAPKQS 84
NN+SS P S +TT + DTE V ++++S K + K +
Sbjct: 164 NNTSSH-------PLLSDKSSTTGSNNQSRTTADTETVNRNQTTSPAPSKAPVSVDLKTN 216
Query: 85 NQNDGQLFPQIASTIKPQQDKSFAPMPSPNARVDSKNVIQSDEHLRNNCDIYEGSWVLDD 144
+ ++ AS+ +Q K+ + S ++ + E L+N C+ ++G W+ DD
Sbjct: 217 SSSNSST----ASSTPKKQTKTVDLVSSVKQEIEKWS-----ESLKN-CEFFDGEWIKDD 266
Query: 145 SFPLYKSGSCPHIDEPFNCFLNGRRDNKYEKFRWQPKNCNMPRLNGKYMLKMLRGKRLVF 204
S+PLYK GSC IDE FNC NGR D ++K +W+PK C++PRLNG +L+MLRG+RLVF
Sbjct: 267 SYPLYKPGSCNLIDEQFNCITNGRPDKDFQKLKWKPKKCSLPRLNGAILLEMLRGRRLVF 326
Query: 205 VGDSLNRNMWDSLVCILSNSVKNKSRIFEASGRQEFQTEDSYSFIFTDYNCSIEFFRSPF 264
VGDSLNRNMW+SLVCIL SVK++++++EA GR F+ E YSF+F DYNC++EFF SPF
Sbjct: 327 VGDSLNRNMWESLVCILKGSVKDETKVYEARGRHHFRGEAEYSFVFQDYNCTVEFFVSPF 386
Query: 265 LVQEWEMPGKKGSKKETLRLDLVERSCDKYKNADVLIFNTGHWWTHEKTMEGKEYYQEGN 324
LVQEWE+ KKG+KKETLRLDLV +S ++YK ADV++FNTGHWWTHEKT +G++YYQEG+
Sbjct: 387 LVQEWEIVDKKGTKKETLRLDLVGKSSEQYKGADVIVFNTGHWWTHEKTSKGEDYYQEGS 446
Query: 325 HIHGQLNVEEAISKAFLTWSRWVDTNIDPKKTIVFFRGYSPSHFRGGEWDSGGRCESETE 384
+++ +L V EA KA TW RWV+ N++P K++VFFRGYS SHF GG+W+SGG C+SETE
Sbjct: 447 NVYHELAVLEAFRKALTTWGRWVEKNVNPAKSLVFFRGYSASHFSGGQWNSGGACDSETE 506
Query: 385 PMKTESTDLSENPPMMRTIESVIKGMKTPVVYLNISKMTDFRHDAHPSMYRNMNMSEETK 444
P+K + T L+ P M+ +E V++GMKTPV YLNI+++TD+R D HPS+YR ++SE+ K
Sbjct: 507 PIKND-TYLTPYPSKMKVLEKVLRGMKTPVTYLNITRLTDYRKDGHPSVYRKQSLSEKEK 565
Query: 445 KYMLKHQDCSHWCLPGVPDLWNEFVYAHLL 474
K L +QDCSHWCLPGVPD WNE +YA L+
Sbjct: 566 KSPLLYQDCSHWCLPGVPDSWNEILYAELI 595
>At3g12060 hypothetical protein
Length = 556
Score = 490 bits (1262), Expect = e-139
Identities = 218/363 (60%), Positives = 281/363 (77%), Gaps = 1/363 (0%)
Query: 112 SPNARVDSKNVIQSDEHLRNNCDIYEGSWVLDDSFPLYKSGSCPHIDEPFNCFLNGRRDN 171
SP + K+ ++ +C+ +EG WV DDS+PLYK GSC IDE FNC NGR D
Sbjct: 175 SPKRKQRRKSSLRKVIESLKSCEFFEGDWVKDDSYPLYKPGSCNLIDEQFNCISNGRPDV 234
Query: 172 KYEKFRWQPKNCNMPRLNGKYMLKMLRGKRLVFVGDSLNRNMWDSLVCILSNSVKNKSRI 231
++K +W+PK C++PRLNG +L+M+RG+RLVFVGDSLNRNMW+SLVCIL SVK++S++
Sbjct: 235 DFQKLKWKPKQCSLPRLNGGKLLEMIRGRRLVFVGDSLNRNMWESLVCILKGSVKDESQV 294
Query: 232 FEASGRQEFQTEDSYSFIFTDYNCSIEFFRSPFLVQEWEMPGKKGSKKETLRLDLVERSC 291
FEA GR +F+ E YSF+F DYNC++EFF SPFLVQEWE+ K G+KKETLRLDLV +S
Sbjct: 295 FEAHGRHQFRWEAEYSFVFKDYNCTVEFFASPFLVQEWEVTEKNGTKKETLRLDLVGKSS 354
Query: 292 DKYKNADVLIFNTGHWWTHEKTMEGKEYYQEGNHIHGQLNVEEAISKAFLTWSRWVDTNI 351
++YK AD+L+FNTGHWWTHEKT +G++YYQEG+ +H +L+V+EA KA TW RWVD N+
Sbjct: 355 EQYKGADILVFNTGHWWTHEKTSKGEDYYQEGSTVHPKLDVDEAFRKALTTWGRWVDKNV 414
Query: 352 DPKKTIVFFRGYSPSHFRGGEWDSGGRCESETEPMKTESTDLSENPPMMRTIESVIKGMK 411
+PKK++VFFRGYSPSHF GG+W++GG C+ ETEP+K E T L+ M +E V++GMK
Sbjct: 415 NPKKSLVFFRGYSPSHFSGGQWNAGGACDDETEPIKNE-TYLTPYMLKMEILERVLRGMK 473
Query: 412 TPVVYLNISKMTDFRHDAHPSMYRNMNMSEETKKYMLKHQDCSHWCLPGVPDLWNEFVYA 471
TPV YLNI+++TD+R DAHPS+YR +S E K L +QDCSHWCLPGVPD WNE YA
Sbjct: 474 TPVTYLNITRLTDYRKDAHPSIYRKQKLSAEESKSPLLYQDCSHWCLPGVPDSWNEIFYA 533
Query: 472 HLL 474
LL
Sbjct: 534 ELL 536
>At1g60795 unknown protein
Length = 541
Score = 431 bits (1109), Expect = e-121
Identities = 203/445 (45%), Positives = 292/445 (65%), Gaps = 5/445 (1%)
Query: 34 PSYTPQYSTNETTTHDKTSHSSNKDTELVVSSRSSSQISRKEGSGLAPKQSNQNDGQLFP 93
PS+ + ++ TT + S + D+ + +EG+G N +
Sbjct: 90 PSFVVPDAGSKNTTLSEESKVPSFDSGQRSGETVKNSSLAEEGNGSVADDQNTLEANATT 149
Query: 94 QIASTIKPQQDKSFAPMPSPNARVDSKNVIQSDEH-LRNNCDIYEGSWVL--DDSFPLYK 150
+ ++ D + N ++ +V + +CDIY+GSWV D++ P Y
Sbjct: 150 SVGNSSSLVSDLGGRFVVPANTSKENGSVTEDRSRGSYEDCDIYDGSWVRADDETMPYYP 209
Query: 151 SGSCPHIDEPFNCFLNGRRDNKYEKFRWQPKNCNMPRLNGKYMLKMLRGKRLVFVGDSLN 210
GSCP+ID FNC NGR D+ Y K+RWQP C++PRLNG L+ LRGK+LVFVGDS+N
Sbjct: 210 PGSCPYIDRDFNCHANGRPDDAYVKWRWQPNGCDIPRLNGTDFLEKLRGKKLVFVGDSIN 269
Query: 211 RNMWDSLVCILSNSVKNKSRIFEASGRQEFQTEDSYSFIFTDYNCSIEFFRSPFLVQEWE 270
RNMW+SL+CIL +S+K+K R++E SGR+EF+ + Y+F F DYNC+++F SPF V+E
Sbjct: 270 RNMWESLICILRHSLKDKKRVYEISGRREFKKKGFYAFRFEDYNCTVDFVGSPFFVRESS 329
Query: 271 MPGKKGSKKETLRLDLVERSCDKYKNADVLIFNTGHWWTHEKTMEGKEYYQEGNHIHGQL 330
G G+ ETLRLD+++++ Y++AD+LIFNTGHWWTH+KT G+ YYQEGN ++ +L
Sbjct: 330 FKGVNGTTLETLRLDMMDKTTSMYRDADILIFNTGHWWTHDKTKLGENYYQEGNVVYPRL 389
Query: 331 NVEEAISKAFLTWSRWVDTNIDPKKTIVFFRGYSPSHFRGGEWDSGGRCESETEPMKTES 390
V EA +A +TW++WVD NID +T + FRGYS +HFRGG W+SGG+C ETEP+ S
Sbjct: 390 KVLEAYKRALITWAKWVDKNIDRSQTHIVFRGYSVTHFRGGPWNSGGQCHKETEPIFNTS 449
Query: 391 TDLSENPPMMRTIESVIKG-MKTPVVYLNISKMTDFRHDAHPSMYRNMNMSEETKKYMLK 449
L++ P M+ +E +++ MKTPV+Y+NIS++TDFR D HPS+YR + +E+ K+ +
Sbjct: 450 Y-LAKYPSKMKALEYILRDTMKTPVIYMNISRLTDFRKDGHPSIYRMVYRTEKEKREAVS 508
Query: 450 HQDCSHWCLPGVPDLWNEFVYAHLL 474
HQDCSHWCLPGVPD WN+ +Y LL
Sbjct: 509 HQDCSHWCLPGVPDTWNQLLYVSLL 533
>At5g49340 putative protein
Length = 457
Score = 419 bits (1076), Expect = e-117
Identities = 197/354 (55%), Positives = 257/354 (71%), Gaps = 3/354 (0%)
Query: 123 IQSDEHLRNNCDIYEGSWVLDDSFPLYKSGSCPHIDEPFNCFLNGRRDNKYEKFRWQPKN 182
I S+ +CDI++G+WV DDS P+Y G CP +++ FNCF NGR D+ + + RWQP
Sbjct: 90 IDSETKELASCDIFDGTWVFDDSEPVYLPGYCPFVEDKFNCFKNGRPDSGFLRHRWQPHG 149
Query: 183 CNMPRLNGKYMLKMLRGKRLVFVGDSLNRNMWDSLVCILSNSVKNKSRIFEASGRQ-EFQ 241
C++PR +GK MLKMLRGKR+VFVGDSLNRNMW+SLVC L +++++K+R+ + G+Q
Sbjct: 150 CSIPRFDGKKMLKMLRGKRVVFVGDSLNRNMWESLVCSLRSTLEDKNRVSKIIGKQSNLP 209
Query: 242 TEDSYSFIFTDYNCSIEFFRSPFLVQEWEMPGKKGSKKETLRLDLVERSCDK-YKNADVL 300
E Y F F D+ CSI+F +SPFLVQE E+ G ++ETLRLD+++RS K YKNAD++
Sbjct: 210 NEGFYGFRFNDFECSIDFIKSPFLVQESEVVDVYGKRRETLRLDMIQRSMTKIYKNADIV 269
Query: 301 IFNTGHWWTHEKTMEGKEYYQEGNHIHGQLNVEEAISKAFLTWSRWVDTNIDPKKTIVFF 360
IFNTGHWWTH+KT EGK YYQEGN ++ +L V+EA +KA TW+ WVD+NI+ KT VFF
Sbjct: 270 IFNTGHWWTHQKTYEGKGYYQEGNRVYERLEVKEAYTKAIHTWADWVDSNINSTKTRVFF 329
Query: 361 RGYSPSHFRGGEWDSGGRCESETEPMKTESTDLSENPPMMRTIESVIKGMKTPVVYLNIS 420
GYS SHFR G W+SGG+C+ ET P++ E T P MM+ +ESVI MKTPV Y+NI+
Sbjct: 330 VGYSSSHFRKGAWNSGGQCDGETRPIQNE-TYTGVYPWMMKVVESVISEMKTPVFYMNIT 388
Query: 421 KMTDFRHDAHPSMYRNMNMSEETKKYMLKHQDCSHWCLPGVPDLWNEFVYAHLL 474
KMT +R D HPS+YR T +QDCSHWCLPGVPD WN+ +YA LL
Sbjct: 389 KMTWYRTDGHPSVYRQPADPRGTSPAAGMYQDCSHWCLPGVPDSWNQLLYATLL 442
>At5g20590 unknown protein
Length = 485
Score = 383 bits (984), Expect = e-106
Identities = 170/357 (47%), Positives = 242/357 (67%), Gaps = 8/357 (2%)
Query: 123 IQSDEHL----RNNCDIYEGSWVL-DDSFPLYKSGSCPHIDEPFNCFLNGRRDNKYEKFR 177
++S EH CD+Y+GSWV DD +PLY+ GSCP++D+ F+C NGRRD+ Y +R
Sbjct: 127 VESVEHAVIEKMRGCDLYKGSWVKGDDEYPLYQPGSCPYVDDAFDCQRNGRRDSDYLNWR 186
Query: 178 WQPKNCNMPRLNGKYMLKMLRGKRLVFVGDSLNRNMWDSLVCILSNSVKNKSRIFEASGR 237
W+P C++PR N L LRGK L+ VGDS+NRN ++S++C+L + +KSR++E G
Sbjct: 187 WKPDGCDLPRFNATDFLVKLRGKSLMLVGDSMNRNQFESMLCVLREGLSDKSRMYEVHGH 246
Query: 238 QEFQTEDSYSFIFTDYNCSIEFFRSPFLVQEWEMPGKKGSKKETLRLDLVERSCDKYKNA 297
+ + F F DYNC++EF RS FLV+E +G+ TL +D +++S K+K A
Sbjct: 247 NITKGRGYFVFKFEDYNCTVEFVRSHFLVREGVRANAQGNTNPTLSIDRIDKSHAKWKRA 306
Query: 298 DVLIFNTGHWWTHEKTMEGKEYYQEGNHIHGQLNVEEAISKAFLTWSRWVDTNIDPKKTI 357
D+L+FNTGHWW H KT GK YY+EG++I+ + + EA ++ TW++W+D N++PKK +
Sbjct: 307 DILVFNTGHWWVHGKTARGKNYYKEGDYIYPKFDATEAYRRSLKTWAKWIDQNVNPKKQL 366
Query: 358 VFFRGYSPSHFRGGEWDSGGRCESETEPMKTESTDLSENPPMMRTIESVIKGMKTPVVYL 417
VF+RGYS +HFRGGEWDSGG C E EP+K S + P M+ ++ IK M+ PV+ L
Sbjct: 367 VFYRGYSSAHFRGGEWDSGGSCNGEVEPVKKGSI-IDSYPLKMKIVQEAIKEMQVPVILL 425
Query: 418 NISKMTDFRHDAHPSMYRNMNMSEETKKYMLKHQDCSHWCLPGVPDLWNEFVYAHLL 474
N++K+T+FR D HPS+Y N + KK + QDCSHWCLPGVPD+WN +YA LL
Sbjct: 426 NVTKLTNFRKDGHPSIYGKTN--TDGKKVSTRRQDCSHWCLPGVPDVWNHLIYASLL 480
>At3g62390 unknown protein
Length = 475
Score = 371 bits (953), Expect = e-103
Identities = 190/461 (41%), Positives = 267/461 (57%), Gaps = 17/461 (3%)
Query: 16 FSKVFSHIFNNSSSFSPPPSYTPQYSTNETTTHDKTSHSSNKDTELVVSSRSSSQISRKE 75
F+ S + ++SS P P Q + +S S L+ + +S +
Sbjct: 29 FTFFSSSLLKSNSSLYPTPEANFQIDLSPIAAISDSSVSPQASPILISTHFNSPE--NTS 86
Query: 76 GSGLAPKQSNQNDGQLFPQIASTIKPQQDKSFAPMPSPNARVDSKNVIQSDEHLRNNCDI 135
GS + G+ + I +PS N D K + +E CD+
Sbjct: 87 GSSKISVFEQKISGESLVKEVREIANLTSIKVIELPSNNGE-DKKTEKRIEE-----CDV 140
Query: 136 YEGSWVLDDSFPLYKSGSCPHIDEPFNCFLNGRRDNKYEKFRWQPKNCNMPRLNGKYMLK 195
+G WV D +PLY + SCP IDE F C NGR D Y +RW+P++C+ PR N ML+
Sbjct: 141 TKGKWVYDSDYPLYTNASCPFIDEGFGCQSNGRLDLNYMNWRWEPQDCHAPRFNATKMLE 200
Query: 196 MLRGKRLVFVGDSLNRNMWDSLVCILSNSVKNKSRIFEASGRQEFQTEDSYSFIFTDYNC 255
M+RGKRLVFVGDS+NRN W+S++C+L +VK+ R++E R+ + + +YSF F DY C
Sbjct: 201 MIRGKRLVFVGDSINRNQWESMLCLLFQAVKDPKRVYETHNRRITKEKGNYSFRFVDYKC 260
Query: 256 SIEFFRSPFLVQEWEMP-GKKGSKKETLRLDLVERSCDKYKNADVLIFNTGHWWTHEKTM 314
++EF+ + FLV+E GKK ++ETLR+D ++R+ ++K A++L+FNT HWW+H KT
Sbjct: 261 TVEFYVTHFLVREGRARIGKK--RRETLRIDAMDRTSSRWKGANILVFNTAHWWSHYKTK 318
Query: 315 EGKEYYQEGNHIHGQLNVEEAISKAFLTWSRWVDTNIDPKKTIVFFRGYSPSHFRGGEWD 374
G YYQEG+ IH +L+V A KA TWS WVD N+DPKKT VFFR +PSHF GGEW+
Sbjct: 319 SGVNYYQEGDLIHPKLDVSTAFKKALQTWSSWVDKNVDPKKTRVFFRSAAPSHFSGGEWN 378
Query: 375 SGGRCESETEPMKTESTDLSENPPMMRTIESVIKGMKTPVVYLNISKMTDFRHDAHPSMY 434
SGG C P+ T +E V+K M+TPV LN+S ++ +R DAHPS+Y
Sbjct: 379 SGGHCREANMPL--NQTFKPSYSSKKSIVEDVLKQMRTPVTLLNVSGLSQYRIDAHPSIY 436
Query: 435 RNMNMSEETKKYMLKHQDCSHWCLPGVPDLWNEFVYAHLLY 475
+ ++ QDCSHWCLPGVPD WN F+Y HLL+
Sbjct: 437 GTKPENRRSRAV----QDCSHWCLPGVPDTWNHFLYLHLLH 473
>At5g19160 putative protein
Length = 464
Score = 320 bits (819), Expect = 1e-87
Identities = 153/369 (41%), Positives = 227/369 (61%), Gaps = 15/369 (4%)
Query: 127 EHLRNNCDIYEGSWVLDDSFPLYKSGSCPHIDEPFNCFLNGRRDNKYEKFRWQPKNCNMP 186
E N CD++ G WV D+S+PLY+S C IDE F C GR D Y K+RWQP +C++P
Sbjct: 93 EESGNGCDLFNGKWVWDESYPLYQSKDCTFIDEGFRCTEFGRPDLFYTKWRWQPNHCDLP 152
Query: 187 RLNGKYMLKMLRGKRLVFVGDSLNRNMWDSLVCILSNSVKNKSRIFEASGRQEFQTEDSY 246
R + K ML+ LR KRLVFVGDS+ RN W+SL+C+L++++ NK+ ++E + R + +
Sbjct: 153 RFDAKLMLEKLRNKRLVFVGDSIGRNQWESLLCMLASAISNKNLVYEVNNRPITKHMGFF 212
Query: 247 SFIFTDYNCSIEFFRSPFLVQEWEMP-GKKGSKKETLRLDLVERSCDKYKNADVLIFNTG 305
F F DYNC++E++R+PFLV + P G K TL+L+ +E + DK+++AD+L+FNTG
Sbjct: 213 VFRFHDYNCTVEYYRAPFLVLQSRPPEGSPEKVKTTLKLETMEWTADKWRDADILVFNTG 272
Query: 306 HWWTHEKTMEGKEYYQEGNHIHGQLNVEEAISKAFLTWSRWVDTNIDPKKTIVFFRGYSP 365
HWW +EKT+ G Y+QEG + ++ +E A +A T +W+ +D KT VFFR ++P
Sbjct: 273 HWWNYEKTIRGGCYFQEGEKVRMRMKIEHAYRRAMKTVMKWIQEEVDANKTQVFFRTFAP 332
Query: 366 SHFRGGEWDSGGRCESETEPMKTESTDLSENPPMMRTIESVIKGM--------KTPVVYL 417
HFRGG+W +GG C ET P S +E ++ ++ V+ + L
Sbjct: 333 VHFRGGDWRTGGTCHMETLPDFGASLVPAETWDHIKLLQDVLSSSLYYSNISETVKLKVL 392
Query: 418 NISKMTDFRHDAHPSMYRNMNMSEETKKYMLKHQDCSHWCLPGVPDLWNEFVYAHLLYIM 477
NI+ M R+D HPS+Y + ++ QDCSHWCLPGVPD WNE +YA L++
Sbjct: 393 NITAMAAQRNDGHPSLY-YLGLAGPAP---FHRQDCSHWCLPGVPDSWNELLYA--LFLK 446
Query: 478 KRNRGNPKT 486
+P++
Sbjct: 447 HEGYSSPRS 455
>At3g06080 unknown protein
Length = 469
Score = 319 bits (817), Expect = 2e-87
Identities = 155/351 (44%), Positives = 217/351 (61%), Gaps = 10/351 (2%)
Query: 127 EHLRNNCDIYEGSWVLDDSFPLYKSGSCPHIDEPFNCFLNGRRDNKYEKFRWQPKNCNMP 186
E + CD+++G WV D+S+PLY+S C +DE F C GR D Y ++RWQP++CN+P
Sbjct: 97 EESGSGCDVFDGDWVWDESYPLYQSKDCRFLDEGFRCSDFGRSDLFYTQWRWQPRHCNLP 156
Query: 187 RLNGKYMLKMLRGKRLVFVGDSLNRNMWDSLVCILSNSVKNKSRIFEASGRQEFQTEDSY 246
R + K ML+ LR KRLVFVGDS+ RN W+SL+C+LS++VKN+S I+E +G + +
Sbjct: 157 RFDAKLMLEKLRDKRLVFVGDSIGRNQWESLLCLLSSAVKNESLIYEINGSPITKHKGFL 216
Query: 247 SFIFTDYNCSIEFFRSPFLVQEWEMP-GKKGSKKETLRLDLVERSCDKYKNADVLIFNTG 305
F F +YNC++E++RSPFLV + P G G K +L+LD ++ + K+++ADVL+ NTG
Sbjct: 217 VFKFEEYNCTVEYYRSPFLVPQSRPPIGSPGKVKTSLKLDTMDWTSSKWRDADVLVLNTG 276
Query: 306 HWWTHEKTMEGKEYYQEGNHIHGQLNVEEAISKAFLTWSRWVDTNIDPKKTIVFFRGYSP 365
HWW KT Y+QEG + ++NV++A +A T +W+ T +D KT VFFR ++P
Sbjct: 277 HWWNEGKTTRTGCYFQEGEEVKLKMNVDDAYKRALNTVVKWIHTELDSNKTQVFFRTFAP 336
Query: 366 SHFRGGEWDSGGRCESETEPMKTESTDLSENPPMMRTIESVI-----KGMKTPVVYLNIS 420
HFRGG+W +GG C ET P S SE ++ + V+ + V LNI+
Sbjct: 337 VHFRGGDWKTGGTCHMETLPEIGTSLASSETWEQLKILRDVLSHNSNRSETVKVKLLNIT 396
Query: 421 KMTDFRHDAHPSMYRNMNMSEETKKYMLKHQDCSHWCLPGVPDLWNEFVYA 471
M R D HPS+Y L QDCSHWCLPGVPD WNE YA
Sbjct: 397 AMAAQRKDGHPSLY----YLGPHGPAPLHRQDCSHWCLPGVPDTWNELFYA 443
>At5g06230 unknown protein
Length = 413
Score = 284 bits (727), Expect = 7e-77
Identities = 145/367 (39%), Positives = 209/367 (56%), Gaps = 17/367 (4%)
Query: 125 SDEHLRNNCDIYEGSWVLDDSFP------LYKSGSCPHIDEPFNCFLNGRRDNKYEKFRW 178
S + + CD +G WV S L+ C +D F C +GR+D+ Y +RW
Sbjct: 54 SSQTVTKECDYSKGKWVRRASSSSSSVNGLFYGEECRFLDSGFRCHKHGRKDSGYLDWRW 113
Query: 179 QPKNCNMPRLNGKYMLKMLRGKRLVFVGDSLNRNMWDSLVCILSNSVKNKSRIFEASGRQ 238
QP C++PR N +L+ R R+VFVGDS+ RN W+SL+C+LS ++ NKS I+E +G
Sbjct: 114 QPHGCDLPRFNASDLLERSRNGRIVFVGDSIGRNQWESLMCMLSQAIPNKSEIYEVNGNP 173
Query: 239 EFQTEDSYSFIFTDYNCSIEFFRSPFLVQEWEMPGKKGSK-KETLRLDLVERSCDKYKNA 297
+ + S F N ++E+ RSPFLV P K + K T+R+D ++ +
Sbjct: 174 ITKHKGFLSMRFPRENLTVEYHRSPFLVVIGRPPDKSPKEIKTTVRVDEFNWQSKRWVGS 233
Query: 298 DVLIFNTGHWWTHEKTMEGKEYYQEGNHIHGQLNVEEAISKAFLTWSRWVDTNIDPKKTI 357
DVL+FN+GHWW +KT+ Y++EG ++ + V EA K+ TW WV +DP K+
Sbjct: 234 DVLVFNSGHWWNEDKTVLTGCYFEEGRKVNKTMGVMEAFGKSLKTWKSWVLEKLDPDKSY 293
Query: 358 VFFRGYSPSHFRGGEWDSGGRCESETEPMKTESTDLSENPPMMRTIESVIKGMK---TPV 414
VFFR YSP H+R G W++GG C++E EP +T+ L + I VI+ M+ + V
Sbjct: 294 VFFRSYSPVHYRNGTWNTGGLCDAEIEP-ETDKRKLEPDASHNEYIYKVIEEMRYRHSKV 352
Query: 415 VYLNISKMTDFRHDAHPSMYRNMNMSEETKKYMLKHQDCSHWCLPGVPDLWNEFVYAHLL 474
+LNI+ +T+FR D H S YR S + QDCSHWCLPGVPD WNE +YA LL
Sbjct: 353 KFLNITYLTEFRKDGHISRYREQGTSVDVP------QDCSHWCLPGVPDTWNEILYAQLL 406
Query: 475 YIMKRNR 481
+ R +
Sbjct: 407 SMNYRTK 413
>At3g11570 hypothetical protein
Length = 427
Score = 284 bits (726), Expect = 9e-77
Identities = 143/356 (40%), Positives = 203/356 (56%), Gaps = 14/356 (3%)
Query: 133 CDIYEGSWVL---DDSFPLYKSGSCPHIDEPFNCFLNGRRDNKYEKFRWQPKNCNMPRLN 189
CD G WV D Y C +D F C NGR+D+ + ++RWQP C++PR N
Sbjct: 79 CDYSYGRWVRRRRDVDETSYYGEECRFLDPGFRCLNNGRKDSGFRQWRWQPHGCDLPRFN 138
Query: 190 GKYMLKMLRGKRLVFVGDSLNRNMWDSLVCILSNSVKNKSRIFEASGRQEFQTEDSYSFI 249
L+ R R+VFVGDS+ RN W+SL+C+LS +V NKS I+E +G + + S
Sbjct: 139 ASDFLERSRNGRIVFVGDSIGRNQWESLLCMLSQAVSNKSEIYEVNGNPISKHKGFLSMR 198
Query: 250 FTDYNCSIEFFRSPFLVQEWEMP-GKKGSKKETLRLDLVERSCDKYKNADVLIFNTGHWW 308
F + N ++E+ R+PFLV P K T+R+D K+ +DVL+FNTGHWW
Sbjct: 199 FPEQNLTVEYHRTPFLVVVGRPPENSPVDVKMTVRVDEFNWQSKKWVGSDVLVFNTGHWW 258
Query: 309 THEKTMEGKEYYQEGNHIHGQLNVEEAISKAFLTWSRWVDTNIDPKKTIVFFRGYSPSHF 368
+KT Y+QEG ++ + V E K+ TW WV +D +++ VFFR +SP H+
Sbjct: 259 NEDKTFIAGCYFQEGGKLNKTMGVMEGFEKSLKTWKSWVLERLDSERSHVFFRSFSPVHY 318
Query: 369 RGGEWDSGGRCESETEPMKTESTDLSENPPMMRTIESVIKGMK---TPVVYLNISKMTDF 425
R G W+ GG C+++TEP +T+ + +P I I+ M+ + V +LNI+ +T+F
Sbjct: 319 RNGTWNLGGLCDADTEP-ETDMKKMEPDPIHNNYISQAIQEMRYEHSKVKFLNITYLTEF 377
Query: 426 RHDAHPSMYRNMNMSEETKKYMLKHQDCSHWCLPGVPDLWNEFVYAHLLYIMKRNR 481
R DAHPS YR E+ QDCSHWCLPGVPD WNE +YA LL + R +
Sbjct: 378 RKDAHPSRYREPGTPEDAP------QDCSHWCLPGVPDTWNEILYAQLLAMNYRTK 427
>At1g29050 unknown protein
Length = 380
Score = 272 bits (696), Expect = 3e-73
Identities = 131/344 (38%), Positives = 194/344 (56%), Gaps = 25/344 (7%)
Query: 131 NNCDIYEGSWVLDDSFPLYKSGSCPHIDEPFNCFLNGRRDNKYEKFRWQPKNCNMPRLNG 190
+ C++++G WV D S+P Y S CP ID F+C GR D ++ K+ WQP++C +PR +G
Sbjct: 59 SGCNLFQGRWVFDASYPFYDSSKCPFIDGEFDCLKFGRPDKQFLKYSWQPESCTIPRFDG 118
Query: 191 KYMLKMLRGKRLVFVGDSLNRNMWDSLVCILSNSVKNKSRIFEASGRQEFQTEDSYSFIF 250
L+ RGKR++FVGDSL+ NMW+SL C++ SV N F + + F
Sbjct: 119 GAFLRKYRGKRVMFVGDSLSLNMWESLACMIHASVPNAKTTF-------LKRTPLSTLTF 171
Query: 251 TDYNCSIEFFRSPFLVQEWEMPGKKGSKKETLRLDLVERSCDKYKNADVLIFNTGHWWTH 310
+Y ++ +R+P++V K L L +E D +KN DVL+FN+ HWWTH
Sbjct: 172 QEYGVTLYLYRTPYIVDI-----SKERVGRVLNLGAIEGGADAWKNMDVLVFNSWHWWTH 226
Query: 311 EKTMEGKEYYQEGNHIHGQLNVEEAISKAFLTWSRWVDTNIDPKKTIVFFRGYSPSHFRG 370
+ +G +Y ++G+ + +N +A K TW+RWVD N+D KT VFF+G SP+H+ G
Sbjct: 227 KGQSQGWDYIRDGSSLVRDMNRLDAFYKGLSTWARWVDQNVDTAKTRVFFQGISPTHYEG 286
Query: 371 GEWDSGGR-CESETEPMKTESTDLSENPPMMRTIESVIKGMKTPVVYLNISKMTDFRHDA 429
EW+ + C + +P+ S S PP + V+ MK PV L+I+ ++ R DA
Sbjct: 287 REWNEPRKTCSGQMQPLGGSSYP-SGQPPSSGVVSKVLSSMKKPVTLLDITTLSQLRKDA 345
Query: 430 HPSMYRNMNMSEETKKYMLKHQDCSHWCLPGVPDLWNEFVYAHL 473
HPS Y + DCSHWCLPG+PD WN+ +YA L
Sbjct: 346 HPSSYGGDGGT-----------DCSHWCLPGLPDTWNQLLYAAL 378
>At1g48880 hypothetical protein
Length = 485
Score = 270 bits (691), Expect = 1e-72
Identities = 134/342 (39%), Positives = 200/342 (58%), Gaps = 26/342 (7%)
Query: 133 CDIYEGSWVLDDSFPLYKSGSCPHIDEPFNCFLNGRRDNKYEKFRWQPKNCNMPRLNGKY 192
CDI++G+WV+DD++PLY + CP +++ FNC NGR ++Y K+RW+PK+C +PR +
Sbjct: 115 CDIFDGNWVVDDNYPLYNASECPFVEKGFNCLGNGRGHDEYLKWRWKPKHCTVPRFEVRD 174
Query: 193 MLKMLRGKRLVFVGDSLNRNMWDSLVCILSNSVKNKSRIFEASGRQEFQTEDSYSFIFTD 252
+LK LRGKR+VFVGDS++R W+SL+C+L +++K ++E +G + F+
Sbjct: 175 VLKRLRGKRIVFVGDSMSRTQWESLICMLMTGLEDKRSVYEVNGNNITKRIRFLGVRFSS 234
Query: 253 YNCSIEFFRSPFLVQ----EWEMPGKKGSKKETLRLDLVERSCDKYKNADVLIFNTGHWW 308
YN ++EF+RS FLVQ W P + K TL+LD+++ ++ +AD LIFNTG WW
Sbjct: 235 YNFTVEFYRSVFLVQPGRLRWHAPKR---VKSTLKLDVLDVINHEWSSADFLIFNTGQWW 291
Query: 309 THEKTMEGKEYYQEGNHIHGQLNVEEAISKAFLTWSRWVDTNIDPKKTIVFFRGYSPSHF 368
K E Y+Q GN + +++ A A TW+ W+++ +DP KT V FR + PSH
Sbjct: 292 VPGKLFETGCYFQVGNSLRLGMSIPAAYRVALETWASWIESTVDPNKTRVLFRTFEPSH- 350
Query: 369 RGGEWDSGGRCESETEPM-KTESTDLSENPPMMRTIESVIKGMKTPVVYLNISKMTDFRH 427
W C P TE D S M I+ V+K M PV L+++ M+ FR
Sbjct: 351 ----WSDHRSCNVTKYPAPDTEGRDKSIFSEM---IKEVVKNMTIPVSILDVTSMSAFRS 403
Query: 428 DAHPSMYRNMNMSEETKKYMLKHQDCSHWCLPGVPDLWNEFV 469
D H ++ + + DCSHWCLPGVPD+WNE +
Sbjct: 404 DGHVGLWSDNPLV----------PDCSHWCLPGVPDIWNEIL 435
>At5g58600 unknown protein
Length = 402
Score = 270 bits (690), Expect = 1e-72
Identities = 136/354 (38%), Positives = 208/354 (58%), Gaps = 22/354 (6%)
Query: 130 RNNCDIYEGSWVLDDSFPLYKSGSCPHIDEP-FNCFLNGRRDNKYEKFRWQPKNCNMPRL 188
R+ C ++ G+WV D+S+PLYK CP + EP F+C + GR D+ Y K+RWQP+NCN+P
Sbjct: 63 RSTCSLFLGTWVRDNSYPLYKPADCPGVVEPEFDCQMYGRPDSDYLKYRWQPQNCNLPTF 122
Query: 189 NGKYMLKMLRGKRLVFVGDSLNRNMWDSLVCILSNSVKNKSRIFEASGRQEFQTEDSYS- 247
NG L ++GK ++F GDSL +N W+SL+C++ +S S R E S
Sbjct: 123 NGAQFLLKMKGKTIMFAGDSLGKNQWESLICLIVSSAP--------STRTEMTRGLPLST 174
Query: 248 FIFTDYNCSIEFFRSPFLVQEWEMPGKKGSKKETLRLDLVERSCDKYKNADVLIFNTGHW 307
F F DY ++ F+++PFLV + GK+ L+LD + + + + +AD+LIFNTGHW
Sbjct: 175 FRFLDYGITMSFYKAPFLVDIDAVQGKR-----VLKLDEISGNANAWHDADLLIFNTGHW 229
Query: 308 WTHEKTMEGKEYYQEGNHIHGQLNVEEAISKAFLTWSRWVDTNIDPKKTIVFFRGYSPSH 367
W+H +M+G + Q GN + ++ A+ KA TW+ WV+T++D +T V F SP+H
Sbjct: 230 WSHTGSMQGWDLIQSGNSYYQDMDRFVAMEKALRTWAYWVETHVDRSRTQVLFLSISPTH 289
Query: 368 FRGGEW----DSGGR-CESETEPMKTESTDLSENPPMMRT-IESVIKGMKTPVVYLNISK 421
+W SG + C ETEP+ + +S +R+ I V+ GM P L+I+
Sbjct: 290 DNPSDWAASSSSGSKNCYGETEPITGTAYPVSSYTDQLRSVIVEVLHGMHNPAFLLDITL 349
Query: 422 MTDFRHDAHPSMYRNMNMSEETKKYMLKHQDCSHWCLPGVPDLWNEFVYAHLLY 475
++ R D HPS+Y + +S + + DCSHWCLPG+PD WN+ +Y L+Y
Sbjct: 350 LSSLRKDGHPSVYSGL-ISGSQRSRPDQSADCSHWCLPGLPDTWNQLLYTLLIY 402
>At2g34070 unknown protein
Length = 385
Score = 264 bits (674), Expect = 9e-71
Identities = 129/345 (37%), Positives = 199/345 (57%), Gaps = 26/345 (7%)
Query: 131 NNCDIYEGSWVLDDSFPLYKSGSCPHIDEPFNCFLNGRRDNKYEKFRWQPKNCNMPRLNG 190
+ C++++G WV D S+P Y S +CP ID F+C GR D ++ K+ WQP +C +PR +G
Sbjct: 63 SGCNLFQGRWVFDASYPFYDSSTCPFIDGEFDCLKFGRPDKQFLKYSWQPDSCTVPRFDG 122
Query: 191 KYMLKMLRGKRLVFVGDSLNRNMWDSLVCILSNSVKNKSRIFEASGRQEFQTEDSYSFIF 250
+ LK RGKR++FVGDSL+ NMW+SL C++ +SV N F + S F
Sbjct: 123 EAFLKKWRGKRVMFVGDSLSLNMWESLACMIHSSVPNTKTTF-------LKRTPLSSLTF 175
Query: 251 TDYNCSIEFFRSPFLVQEWEMPGKKGSKKETLRLDLVERSCDKYKNADVLIFNTGHWWTH 310
+Y+ ++ +R+P+LV K S L L +E D +KN D+L+FN+ HWWTH
Sbjct: 176 QEYDVTLFLYRTPYLVDI-----SKESVGRVLNLGAIEDGADAWKNMDLLVFNSWHWWTH 230
Query: 311 EKTM-EGKEYYQEGNHIHGQLNVEEAISKAFLTWSRWVDTNIDPKKTIVFFRGYSPSHFR 369
+G ++ ++G+ + ++ +A +K TW +WVD N++ +T VFF+G SP+H+
Sbjct: 231 TGVQSQGWDFIRDGSSLMRDMDRLDAFNKGLTTWGQWVDQNVNVSQTRVFFQGISPTHYM 290
Query: 370 GGEWDSGGR-CESETEPMKTESTDLSENPPMMRTIESVIKGMKTPVVYLNISKMTDFRHD 428
G EW+ + C + +P+ T ST + P + V+ M+TPV L+I+ ++ R D
Sbjct: 291 GREWNEPRKTCNGQMQPL-TGSTYPGGSLPAASIVSRVLSTMRTPVYLLDITTLSQLRKD 349
Query: 429 AHPSMYRNMNMSEETKKYMLKHQDCSHWCLPGVPDLWNEFVYAHL 473
AHPS Y + DCSHWCLPG+PD WN+ +YA L
Sbjct: 350 AHPSTYGGDGGT-----------DCSHWCLPGLPDTWNQLLYAAL 383
>At2g30010 unknown protein
Length = 398
Score = 262 bits (669), Expect = 4e-70
Identities = 136/358 (37%), Positives = 201/358 (55%), Gaps = 26/358 (7%)
Query: 131 NNCDIYEGSWVLDDSFPLYKSGSCPH--IDEPFNCFLNGRRDNKYEKFRWQPKNCNMPRL 188
++CD++ G WV D+++PLY+S C ID F+C GR D+ Y KFRW+P NCN+PR
Sbjct: 54 SSCDLFAGEWVRDETYPLYRSKECGRGIIDPGFDCQTYGRPDSDYLKFRWKPFNCNVPRF 113
Query: 189 NGKYMLKMLRGKRLVFVGDSLNRNMWDSLVCILSNSVKNKSRIFEASGRQEFQTEDSYS- 247
NG L+ +R K ++FVGDSL RN W+SL+C++S+S S ED S
Sbjct: 114 NGVKFLQEMRDKTIMFVGDSLGRNQWESLICMISSSA--------PSINTHIIHEDPLST 165
Query: 248 FIFTDYNCSIEFFRSPFLVQEWEMPGKKGSKKETLRLDLVERSCDKYKNADVLIFNTGHW 307
F DYN + F+R+P+LV ++ GK K + + +D + + ++ ADVL+FNTGHW
Sbjct: 166 FKILDYNVKVSFYRAPYLVDIDKINGKTTLKLDEISVD----ASNAWRTADVLLFNTGHW 221
Query: 308 WTHEKTMEGKEYYQEGNHIHGQLNVEEAISKAFLTWSRWVDTNIDPKKTIVFFRGYSPSH 367
W+H ++ G E + G +G ++ A+ K TWS WV I+ T VFF SP+H
Sbjct: 222 WSHTGSLRGWEQMETGGRYYGDMDRLVALRKGLGTWSSWVLRYINSPLTRVFFLSVSPTH 281
Query: 368 FRGGEWDS----------GGRCESETEPMKTESTDLSENPPMMRTIESVIKGMKTPVVYL 417
+ EW S G C +T P + S + I+ V+K MK+ V +
Sbjct: 282 YNPNEWTSRSKTSTITQGGKSCYGQTTPFSGTTYPTSSYVNQKKVIDDVVKEMKSHVSLM 341
Query: 418 NISKMTDFRHDAHPSMYRNMNMSEETKKYMLKHQDCSHWCLPGVPDLWNEFVYAHLLY 475
+I+ ++ R D HPS+Y +++ K+ + DCSHWCLPG+PD WN+ YA LLY
Sbjct: 342 DITMLSALRVDGHPSIYSG-DLNPSLKRNPDRSSDCSHWCLPGLPDTWNQLFYAALLY 398
>At2g30900 hypothetical protein
Length = 367
Score = 260 bits (664), Expect = 1e-69
Identities = 141/352 (40%), Positives = 194/352 (55%), Gaps = 30/352 (8%)
Query: 130 RNNCDIYEGSWVLDDSFPLYKSGSCPHIDEPFNCFLNGRRDNKYEKFRWQPKNCNMPRLN 189
R C+IY+GSWV D S+PLY S +CP I+ FNC NGR D++Y K+RWQP CN+PR N
Sbjct: 39 RKFCNIYQGSWVYDKSYPLYDSKNCPFIERQFNCKSNGRPDSEYLKYRWQPSGCNLPRFN 98
Query: 190 G-KYMLKMLRGKRLVFVGDSLNRNMWDSLVCILSNSV--KNKSRIFEASGRQEFQTEDSY 246
G ++ ++++GK+L+FVGDSL+ N W SL C+L N+ N + SG F
Sbjct: 99 GLDFLGRIMKGKKLMFVGDSLSLNQWQSLTCLLHNAAPKANSTSTRSPSGLSVFS----- 153
Query: 247 SFIFTDYNCSIEFFRSPFLVQEWEMPGKKGSKKETLRLDLVERSCDKYKNADVLIFNTGH 306
F YN SI F R+ FLV P K+ ++LD + S +K ADVL+FN+ H
Sbjct: 154 ---FPAYNSSIMFSRNAFLVDIVGAPPKR-----VMKLDSIS-SGSLWKTADVLVFNSWH 204
Query: 307 WWTHEKTMEGKEYYQEGNHIHGQLNVEEAISKAFLTWSRWVDTNIDPKKTIVFFRGYSPS 366
WW H + + GN ++ A KA +TW++W+D NIDP KT VFF+G SP
Sbjct: 205 WWLHTDRKQPWDAIMSGNVTVKDMDRLVAYEKAMMTWAKWIDQNIDPSKTKVFFQGISPD 264
Query: 367 HFRGGEWD---SGGRCESETEPMKTESTDLSENPPMMRTIESVIKGMKTPVVYLNISKMT 423
H R EW G C ET+P+ S + M + VIK MK ++++ M+
Sbjct: 265 HGRAREWSKQGGKGSCIGETKPIMGSSYLAGPHAAEM-VVAKVIKTMKNQARLMDVTLMS 323
Query: 424 DFRHDAHPSMYRNMNMSEETKKYMLKHQDCSHWCLPGVPDLWNEFVYAHLLY 475
R D HPS+Y + DCSHWCL GVPD WN+ +Y+ L +
Sbjct: 324 QLRKDGHPSVYGFGGH---------RMADCSHWCLSGVPDSWNQLLYSELFH 366
>At2g42570 unknown protein
Length = 367
Score = 255 bits (651), Expect = 4e-68
Identities = 128/343 (37%), Positives = 192/343 (55%), Gaps = 27/343 (7%)
Query: 133 CDIYEGSWVLDDSFPLYKSGSCPHIDEPFNCFLNGRRDNKYEKFRWQPKNCNMPRLNGKY 192
C+ + G+WV D +PLY CP ID FNC GR DN Y K+RWQP +C++PR NG Y
Sbjct: 47 CNWFRGNWVYDVKYPLYDPYKCPFIDPQFNCKKYGRPDNAYLKYRWQPSSCSLPRFNGLY 106
Query: 193 MLKMLRGKRLVFVGDSLNRNMWDSLVCILSNSVKNKSRIFEASGRQEFQTEDSYSFIFTD 252
L+ +RGK+++FVGDSL+ NMW SL C++ + V N + + S F +
Sbjct: 107 FLRRMRGKKIMFVGDSLSTNMWQSLACLIHSWVPNTRYTL-------IRQKGLASLTFEE 159
Query: 253 YNCSIEFFRSPFLVQ-EWEMPGKKGSKKETLRLDLVERSCDKYKNADVLIFNTGHWWTHE 311
Y ++ +R+ FLV E G+ L+LD +++ + ++ DVLIFN+ HWWTH
Sbjct: 160 YGVTLLLYRTQFLVDLNVEKVGR------VLKLDSIKQG-NMWRGMDVLIFNSWHWWTHT 212
Query: 312 KTMEGKEYYQEGNHIHGQLNVEEAISKAFLTWSRWVDTNIDPKKTIVFFRGYSPSHFRGG 371
+ ++ +Y ++GN ++ +N A K TW+RWV+ +DP KT VFF G SP+H+ G
Sbjct: 213 EHIQPWDYMEDGNRLYKDMNRLVAFYKGMTTWARWVNAYVDPSKTKVFFNGVSPTHYEGK 272
Query: 372 EW-DSGGRCESETEPMKTESTDLSENPPMMRTIESVIKGMKTPVVYLNISKMTDFRHDAH 430
+W + C S+T+P P + V++ +K PV +L+I+ ++ R DAH
Sbjct: 273 DWGEPMNSCRSQTQPFYGRKYP-GGTPMAWVILNKVMRRLKKPVHWLDITGLSQLRKDAH 331
Query: 431 PSMYRNMNMSEETKKYMLKHQDCSHWCLPGVPDLWNEFVYAHL 473
PS + + DCSHWCLPG+PD WN Y+ L
Sbjct: 332 PSAFSGNHPG----------NDCSHWCLPGLPDTWNLLFYSTL 364
>At2g40160 unknown protein
Length = 427
Score = 254 bits (648), Expect = 1e-67
Identities = 122/356 (34%), Positives = 211/356 (59%), Gaps = 20/356 (5%)
Query: 133 CDIYEGSWVLDD-SFPLYKSGSCPHIDEPFNCFLNGRRDNKYEKFRWQPKNCNMPRLNGK 191
CD++ G WVLD+ + PLYK C + E C NGR D+KY+K+RWQP++C++PR + K
Sbjct: 77 CDVFTGKWVLDNVTHPLYKEDECEFLSEWVACTRNGRPDSKYQKWRWQPQDCSLPRFDSK 136
Query: 192 YMLKMLRGKRLVFVGDSLNRNMWDSLVCILSNSVKNKSRIFEASGRQEFQTEDSYSFIFT 251
+L+ LRGK+L+F+GDS++ N W S+VC++ + + + + + + + F
Sbjct: 137 LLLEKLRGKKLMFIGDSIHYNQWQSMVCMVQSVIPSGKKTLKHTAQMSI-------FNIE 189
Query: 252 DYNCSIEFFRSPFLVQ-EWEMPGKKGSKKETLRL-DLVERSCDKYKNADVLIFNTGHWWT 309
+YN +I F+ +PFLV+ + P K+ K + + + + + + + +K+AD LIFNT WWT
Sbjct: 190 EYNATISFYWAPFLVESNADPPDKRDGKTDPVIIPNSISKHGENWKDADYLIFNTYIWWT 249
Query: 310 HEKTME--GKEYYQEG-NHIHGQLNVEEAISKAFLTWSRWVDTNIDPKKTIVFFRGYSPS 366
T++ +E + +G + + ++ + + TW++W++ NI+P +T +FF SP+
Sbjct: 250 RHSTIKVLKQESFNKGDSKEYNEIGIYIVYKQVLSTWTKWLEQNINPSQTSIFFSSMSPT 309
Query: 367 HFRGGEW--DSGGRCESETEPM--KTESTDLSENPPMMRTIESVIKGMKTPVVYLNISKM 422
H R +W + G +CE ETEP+ ++ ++ N + + K K P+ +LNI+ M
Sbjct: 310 HIRSSDWGFNEGSKCEKETEPILNMSKPINVGTNRRLYEIALNATKSTKVPIHFLNITTM 369
Query: 423 TDFRHDAHPSMYRNMN---MSEETKKYMLKHQDCSHWCLPGVPDLWNEFVYAHLLY 475
+++R D H S Y ++N M+ E K DC HWCLPG+PD WNE + +++Y
Sbjct: 370 SEYRKDGHTSFYGSINGKLMTPEQKLDPRTFADCYHWCLPGLPDSWNELLSLYIIY 425
>At1g78710 hypothetical protein
Length = 359
Score = 251 bits (641), Expect = 6e-67
Identities = 136/344 (39%), Positives = 196/344 (56%), Gaps = 26/344 (7%)
Query: 133 CDIYEGSWVLDDSF-PLYKSGSCPHIDEPFNCFLNGRRDNKYEKFRWQPKNCNMPRLNGK 191
C+IY+G W+ D+S PLY + +CP I +C GR D Y +RWQP C++PR NG+
Sbjct: 38 CNIYQGRWIYDNSSNPLYGTSTCPFIG--LDCQKFGRPDKNYLHYRWQPTGCDIPRFNGR 95
Query: 192 YMLKMLRGKRLVFVGDSLNRNMWDSLVCILSNSVKNKSRIFEASGRQEFQTEDSYSFIFT 251
L +GK+++FVGDSL+ NMW SL C+L +V N F+ + + +F
Sbjct: 96 DFLTRFKGKKILFVGDSLSNNMWVSLSCMLHAAVPNAKYTFQLN-------KGLSTFTIP 148
Query: 252 DYNCSIEFFRSPFLVQEWEMPGKKGSKKETLRLDLVERSCDKYKNADVLIFNTGHWWTHE 311
+Y S+ F ++ FLV ++ K ++ L+LD + R +++ +DV IFNT HWW+H
Sbjct: 149 EYGISVNFLKNGFLV---DLVSDK-TRGLILKLDSISRG-NQWLGSDVAIFNTFHWWSHT 203
Query: 312 KTMEGKEYYQEGNHIHGQLNVEEAISKAFLTWSRWVDTNIDPKKTIVFFRGYSPSHFRGG 371
+ +Y+Q G+ I ++N EA A TWS+W+D NIDP KT VF++G SP H GG
Sbjct: 204 GRAKTWDYFQTGDKIVKEMNRMEAFKIALTTWSKWIDHNIDPSKTRVFYQGVSPVHLNGG 263
Query: 372 EWDSGGR-CESETEPMKTESTDLSENPPMMRTIESVIKGMKTPVVYLNISKMTDFRHDAH 430
EW G+ C ET P++ S N ++SVI M PV L+++ MT+ R D H
Sbjct: 264 EWGKPGKTCLGETVPVQGPSYPGRPNEG-EAIVKSVIGRMAKPVELLDVTAMTEMRKDGH 322
Query: 431 PSMYRNMNMSEETKKYMLKHQDCSHWCLPGVPDLWNEFVYAHLL 474
PS+Y + DCSHWCLPGVPD WN+ +Y LL
Sbjct: 323 PSIYAGGGD---------RLNDCSHWCLPGVPDAWNQLLYTALL 357
>At3g54260 unknown protein
Length = 379
Score = 250 bits (639), Expect = 1e-66
Identities = 131/362 (36%), Positives = 202/362 (55%), Gaps = 26/362 (7%)
Query: 118 DSKNVIQSDEHLRNN-CDIYEGSWVLDDSFPLYKSGSCPHIDEPFNCFLNGRRDNKYEKF 176
D N +Q+ R CD G W D+++PLY S SCP++ +C NGR D+ Y+K+
Sbjct: 35 DPLNALQTRRERREERCDYSVGKWTFDETYPLYDS-SCPYLSSALSCQRNGRPDSYYQKW 93
Query: 177 RWQPKNCNMPRLNGKYMLKMLRGKRLVFVGDSLNRNMWDSLVCILSNSVKNKSRIFEASG 236
RW PK C++PR + L +RGKR++ VGDS+ RN W+SLVC++ + + + +G
Sbjct: 94 RWIPKACSLPRFDALKFLGKMRGKRIMLVGDSMMRNQWESLVCLVQSVLPTHRKKLTYNG 153
Query: 237 RQEFQTEDSYSFIFTDYNCSIEFFRSPFLVQEWEMPGKKG-SKKETLRLDLVERSCDKYK 295
+ SF D+ SIEF +P LV+ K+G +K L LD +E + ++
Sbjct: 154 -------PTMSFHSLDFETSIEFCWAPLLVEL-----KRGVDRKRVLHLDSIEDNARYWR 201
Query: 296 NADVLIFNTGHWWTHEKTMEGKEYYQEGNHIHGQLNVEEAISKAFLTWSRWVDTNIDPKK 355
DVL+F++ HWWTH + +YY +GN I ++ A + TW++WV+ N+DP K
Sbjct: 202 GVDVLVFDSAHWWTHSQRWSSWDYYMDGNKIFKAMDPMVAYERGLTTWAKWVEINLDPSK 261
Query: 356 TIVFFRGYSPSHFRGGEWDSGGRCESETEPMKTESTDLSEN-PPMMRTIESVIKGMKTPV 414
T V FR SP +SG C ++ P+ + S+ + P R + V++ MK V
Sbjct: 262 TKVIFRTVSPR-------ESGQMCYNQKHPLPSLSSSTKPHVPQQSRVLNKVLRTMKYRV 314
Query: 415 VYLNISKMTDFRHDAHPSMYRNMNMSEETKKYML--KHQDCSHWCLPGVPDLWNEFVYAH 472
+I+ M+ +R D HPS+++ M EE K + + DCSHWCLPGVPD+WNE + +
Sbjct: 315 YLYDITTMSAYRRDGHPSVFKRA-MHEEEKHHRIAGPSSDCSHWCLPGVPDIWNEMLSSI 373
Query: 473 LL 474
+L
Sbjct: 374 IL 375
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.317 0.132 0.412
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,283,963
Number of Sequences: 26719
Number of extensions: 570671
Number of successful extensions: 2011
Number of sequences better than 10.0: 78
Number of HSP's better than 10.0 without gapping: 56
Number of HSP's successfully gapped in prelim test: 22
Number of HSP's that attempted gapping in prelim test: 1764
Number of HSP's gapped (non-prelim): 89
length of query: 486
length of database: 11,318,596
effective HSP length: 103
effective length of query: 383
effective length of database: 8,566,539
effective search space: 3280984437
effective search space used: 3280984437
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 62 (28.5 bits)
Medicago: description of AC146329.6


