

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144929.16 + phase: 0
(143 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
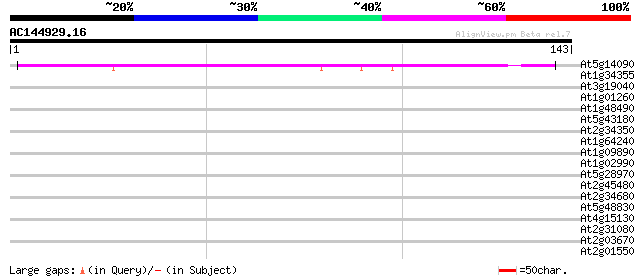
Score E
Sequences producing significant alignments: (bits) Value
At5g14090 unknown protein 63 5e-11
At1g34355 orkhead-associated domain-containing protein / FHA 28 1.7
At3g19040 hypothetical protein 28 2.2
At1g01260 transcription factor MYC7E, putative bHLH transcriptio... 28 2.2
At1g48490 IRE - like protein kinase 27 2.9
At5g43180 unknown protein 27 3.7
At2g34350 nodulin-like protein 27 4.9
At1g64240 hypothetical protein 27 4.9
At1g09890 hypothetical protein 27 4.9
At1g02990 overlap with bases 73023-78341 of BAC clone F10O3 (AC0... 27 4.9
At5g28970 putative protein 26 6.4
At2g45480 unknown protein 26 6.4
At2g34680 auxin-induced protein (AIR9) 26 6.4
At5g48830 unknown protein 26 8.3
At4g15130 CTP:phosphorylcholine cytidylyltransferase (AtCCT2) 26 8.3
At2g31080 putative non-LTR retroelement reverse transcriptase 26 8.3
At2g03670 putative AAA-type ATPase 26 8.3
At2g01550 putative non-LTR retroelement reverse transcriptase 26 8.3
>At5g14090 unknown protein
Length = 361
Score = 63.2 bits (152), Expect = 5e-11
Identities = 53/159 (33%), Positives = 80/159 (49%), Gaps = 25/159 (15%)
Query: 3 DAAVTETQLKLISYELEKFVEAD-EECFYESSGRISFRSNVTLSRKKTDGLETEDLGNKV 61
DA VTE LKLIS EL+KF+EA+ +E ++ SGR S + + + + +G++ E+ +
Sbjct: 126 DADVTENDLKLISNELDKFLEAEAKEGHHQPSGRNSDTNTIASTIEAIEGVDDEEDNQPM 185
Query: 62 ICPLQEYLLGSSFEIRE-KTEVRIERAS------VRETQVKQ--------------GRRS 100
PLQEY GS E+ E K + +RAS + E Q KQ +S
Sbjct: 186 KFPLQEYFFGSLIELPESKIAGKKDRASLGELFQITEVQDKQSENIYGKKKKQPNSAHKS 245
Query: 101 ALHIIKKMSNMVLSSSKSCNTYGNTADHATSTNEKLCKV 139
A H++KK+ + SS+ + D ST +K KV
Sbjct: 246 AKHLVKKVLKKIHPSSRGSVSGKPEVD---STKKKFQKV 281
>At1g34355 orkhead-associated domain-containing protein / FHA
Length = 1477
Score = 28.1 bits (61), Expect = 1.7
Identities = 22/94 (23%), Positives = 40/94 (42%), Gaps = 4/94 (4%)
Query: 37 SFRSNVTLSRKKTDGLETEDLGNKVICPLQEYLLGSSFEIREKTEVRIERASVRETQVKQ 96
S RS S + +G E D+G CP+ L + E T++ IE S ++ ++
Sbjct: 911 SGRSEKYYSLSEIEGEENTDIGRLSRCPIPSALAAKT---SEDTKL-IEELSSSDSGSQE 966
Query: 97 GRRSALHIIKKMSNMVLSSSKSCNTYGNTADHAT 130
+ H ++ + SS +CN + A+
Sbjct: 967 NQTPETHAVRDDVLCDMDSSSTCNIWSRRGKAAS 1000
>At3g19040 hypothetical protein
Length = 1700
Score = 27.7 bits (60), Expect = 2.2
Identities = 15/44 (34%), Positives = 23/44 (52%), Gaps = 6/44 (13%)
Query: 12 KLISYELEKFVEADEECFYESSGRISFRSNVTLSRKKTDGLETE 55
K IS ++ + +E DEE +S GRI KKTD ++ +
Sbjct: 137 KYISMDISELIEDDEEVLLKSHGRIDTHG------KKTDQIQLD 174
>At1g01260 transcription factor MYC7E, putative bHLH transcription
factor (AtbHLH013)
Length = 590
Score = 27.7 bits (60), Expect = 2.2
Identities = 20/75 (26%), Positives = 40/75 (52%), Gaps = 7/75 (9%)
Query: 20 KFVEADEECF-YESSGRISFRSNVTLSRKKTDGLETEDLGNKVICPLQEYLLGSSFEIRE 78
K +EA+ E Y S+ IS S++ + +T G ED+ ++ CPL+ + F E
Sbjct: 483 KVMEAERERLGYSSNPPISLDSDINV---QTSG---EDVTVRINCPLESHPASRIFHAFE 536
Query: 79 KTEVRIERASVRETQ 93
+++V + +++ +Q
Sbjct: 537 ESKVEVINSNLEVSQ 551
>At1g48490 IRE - like protein kinase
Length = 1235
Score = 27.3 bits (59), Expect = 2.9
Identities = 20/69 (28%), Positives = 33/69 (46%), Gaps = 1/69 (1%)
Query: 42 VTLSRKKTDGLETEDLGNKVICPLQE-YLLGSSFEIREKTEVRIERASVRETQVKQGRRS 100
V + R+K D L E G ++ +QE YL EK + E A + + V+ R S
Sbjct: 756 VVIDRRKFDALIVETFGTRIEKLIQEKYLQLCELMDDEKGTIIDEDAPLEDDVVRSLRTS 815
Query: 101 ALHIIKKMS 109
+H+ ++S
Sbjct: 816 PVHLRDRIS 824
>At5g43180 unknown protein
Length = 239
Score = 26.9 bits (58), Expect = 3.7
Identities = 17/52 (32%), Positives = 27/52 (51%), Gaps = 5/52 (9%)
Query: 23 EADEECFYESSGRISFRSNVTLSRKKTDGLETEDLGNKVIC---PLQEYLLG 71
E D +C Y+ +G SFR L ++ G +GN+V+C PL ++ G
Sbjct: 159 EGDCDCAYDITGTSSFREYTRLVLER--GFFMAMVGNRVMCVSIPLLLWMFG 208
>At2g34350 nodulin-like protein
Length = 2301
Score = 26.6 bits (57), Expect = 4.9
Identities = 16/54 (29%), Positives = 28/54 (51%), Gaps = 1/54 (1%)
Query: 8 ETQLKLISYELEKF-VEADEECFYESSGRISFRSNVTLSRKKTDGLETEDLGNK 60
+T+L L S EL K ++DEE ++ GR+ R RK+ +++ +K
Sbjct: 2134 KTRLALRSSELRKRKADSDEEAEFDVEGRLVIREGERSKRKELSDADSDAKSSK 2187
>At1g64240 hypothetical protein
Length = 938
Score = 26.6 bits (57), Expect = 4.9
Identities = 14/46 (30%), Positives = 23/46 (49%)
Query: 14 ISYELEKFVEADEECFYESSGRISFRSNVTLSRKKTDGLETEDLGN 59
IS E KF++ + C + + ++NV SR K+ G+ GN
Sbjct: 57 ISTECCKFLKEQQPCLCDVTKTSKIKTNVLSSRLKSCGIHNLKCGN 102
>At1g09890 hypothetical protein
Length = 477
Score = 26.6 bits (57), Expect = 4.9
Identities = 19/57 (33%), Positives = 31/57 (54%), Gaps = 3/57 (5%)
Query: 67 EYLLGSSFEIREKTEVRIERASVRETQVKQ-GRRSALHIIKKMSNMVLSSSKSCNTY 122
+ + GS+FE+ K E +IE + R+ Q G+ L+I K+ ++LS S TY
Sbjct: 71 DVIKGSNFEVIVKNEEQIELSFTRKWDPSQEGKAVPLNIDKRF--VMLSGSSGFYTY 125
>At1g02990 overlap with bases 73023-78341 of BAC clone F10O3
(AC006550). Genes in this region are annotated in the
record for clone F10O3, AC006550
Length = 1306
Score = 26.6 bits (57), Expect = 4.9
Identities = 26/91 (28%), Positives = 39/91 (42%), Gaps = 5/91 (5%)
Query: 45 SRKKTDGLE--TEDLGNKVICPLQEYLLGSSFEIREKTEVRIERASVRETQVKQGRRSAL 102
SR K++ LE TE L NK P Q +LG EI + R+ + T + R
Sbjct: 958 SRGKSNPLEVTTEQLKNKSASPAQVEVLGHDTEISNTKKQRLRNDNHSVTHDEGSRNQKQ 1017
Query: 103 HIIKKMSNMVLSSSK---SCNTYGNTADHAT 130
+ + ++ LS K + T N+ AT
Sbjct: 1018 NGSRHKDHVGLSPFKKESTSQTASNSIKEAT 1048
>At5g28970 putative protein
Length = 1172
Score = 26.2 bits (56), Expect = 6.4
Identities = 18/60 (30%), Positives = 32/60 (53%), Gaps = 8/60 (13%)
Query: 23 EADEECFYESSGRISFRSNVTLSRKKTDGLETEDLGNKVICPLQEYLLGSSFEIREKTEV 82
EAD+E G+I F S+ LS+KK LE +++ + ++ L S FE + + ++
Sbjct: 505 EADDE------GKIQFDSSANLSQKKQSPLEQDNIMEDTLETVEASL--SVFETQPEVKI 556
>At2g45480 unknown protein
Length = 431
Score = 26.2 bits (56), Expect = 6.4
Identities = 11/33 (33%), Positives = 20/33 (60%)
Query: 91 ETQVKQGRRSALHIIKKMSNMVLSSSKSCNTYG 123
E + +GR+ + +++ S + SS+K NTYG
Sbjct: 117 ERHMHRGRKRSRKLVESSSEVASSSTKYDNTYG 149
>At2g34680 auxin-induced protein (AIR9)
Length = 1661
Score = 26.2 bits (56), Expect = 6.4
Identities = 19/76 (25%), Positives = 32/76 (42%)
Query: 32 SSGRISFRSNVTLSRKKTDGLETEDLGNKVICPLQEYLLGSSFEIREKTEVRIERASVRE 91
SS R+S + VT+ R T G+ G + P Q S + ++ + ++SV
Sbjct: 67 SSLRVSGTTPVTIRRNSTGGVTENLAGTSKVLPKQVSTTASRTDPVRRSLPELRKSSVSS 126
Query: 92 TQVKQGRRSALHIIKK 107
K + +L KK
Sbjct: 127 LSAKTVSKPSLSESKK 142
>At5g48830 unknown protein
Length = 511
Score = 25.8 bits (55), Expect = 8.3
Identities = 19/66 (28%), Positives = 31/66 (46%), Gaps = 3/66 (4%)
Query: 62 ICPLQE---YLLGSSFEIREKTEVRIERASVRETQVKQGRRSALHIIKKMSNMVLSSSKS 118
+ P+Q+ Y G S + E EV ++S RET + +S ++K L + K
Sbjct: 110 VIPVQKTSTYSSGKSTIVEEIPEVGTSKSSGRETDFEGDLKSVWDVVKVKLLDSLDAIKR 169
Query: 119 CNTYGN 124
NT G+
Sbjct: 170 ENTLGS 175
>At4g15130 CTP:phosphorylcholine cytidylyltransferase (AtCCT2)
Length = 300
Score = 25.8 bits (55), Expect = 8.3
Identities = 17/81 (20%), Positives = 38/81 (45%), Gaps = 9/81 (11%)
Query: 47 KKTDGLETEDLGNKVICPLQEYLLGSSFEIREKTEVRIERASVRETQVKQGRRSALHI-I 105
K+T+G+ T D+ +++ +Y+L + + E+ Q++Q +R +++ +
Sbjct: 141 KRTEGISTSDIIMRIVKDYNQYVLRNPGPRYSREEL--------SCQLEQEKRLRVNMRL 192
Query: 106 KKMSNMVLSSSKSCNTYGNTA 126
KK+ V + T TA
Sbjct: 193 KKLQEKVKEQQEKIQTVAKTA 213
>At2g31080 putative non-LTR retroelement reverse transcriptase
Length = 1231
Score = 25.8 bits (55), Expect = 8.3
Identities = 23/88 (26%), Positives = 42/88 (47%), Gaps = 7/88 (7%)
Query: 4 AAVTETQLKLISYELEKFVEADEECFYESSGRISFRSNVTLSRKKTDGLE-----TEDLG 58
A + Q+++I LE+F EA + +I F NV+ ++ E T++LG
Sbjct: 552 AEASVAQIRIIRRVLERFCEASGQKVSLEKSKIFFSHNVSREMEQLISEESGIGCTKELG 611
Query: 59 NKVICP-LQEYLLGSSF-EIREKTEVRI 84
+ P LQ+ + +F E+ E+ R+
Sbjct: 612 KYLGMPILQKRMNKETFGEVLERVSARL 639
>At2g03670 putative AAA-type ATPase
Length = 603
Score = 25.8 bits (55), Expect = 8.3
Identities = 17/67 (25%), Positives = 28/67 (41%)
Query: 8 ETQLKLISYELEKFVEADEECFYESSGRISFRSNVTLSRKKTDGLETEDLGNKVICPLQE 67
+ L+ I+ E + F A+ E SG +S R N+ + +T K ++E
Sbjct: 479 DVDLRKIAEETDLFTGAELEGLCRESGTVSLRENIAATAVFNRHFQTAKSSLKPALTIEE 538
Query: 68 YLLGSSF 74
SSF
Sbjct: 539 VETYSSF 545
>At2g01550 putative non-LTR retroelement reverse transcriptase
Length = 1449
Score = 25.8 bits (55), Expect = 8.3
Identities = 28/103 (27%), Positives = 40/103 (38%), Gaps = 16/103 (15%)
Query: 18 LEKFVEADEECFYESSGRISFRSNVTLSRKKTDGLETEDLGNKVICPLQEYL-------- 69
+E F E +S F + + K GL E +GN V + YL
Sbjct: 682 VENFWRETEPIHMSTSSLFRFTKKLKALKPKLRGLAKEKMGNLVKRTREAYLSLCQAQQS 741
Query: 70 -----LGSSFEIREKTEVRIER-ASVRETQVKQGRRSALHIIK 106
+ EI + VR +R AS+ E +KQ S LH +K
Sbjct: 742 NSQNPSQRAMEIESEAYVRWDRIASIEEKYLKQ--VSKLHWLK 782
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.312 0.128 0.345
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,870,508
Number of Sequences: 26719
Number of extensions: 103090
Number of successful extensions: 243
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 12
Number of HSP's that attempted gapping in prelim test: 235
Number of HSP's gapped (non-prelim): 18
length of query: 143
length of database: 11,318,596
effective HSP length: 89
effective length of query: 54
effective length of database: 8,940,605
effective search space: 482792670
effective search space used: 482792670
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 55 (25.8 bits)
Medicago: description of AC144929.16


