

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144928.8 + phase: 0 /pseudo
(359 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
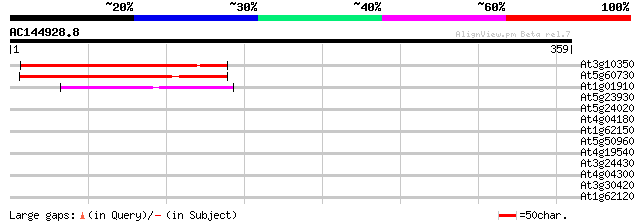
Score E
Sequences producing significant alignments: (bits) Value
At3g10350 putative ATPase 224 7e-59
At5g60730 ATPase - like protein 156 1e-38
At1g01910 arsA homolog (hASNA-I), putative 66 3e-11
At5g23930 putative protein 36 0.037
At5g24020 septum site-determining MinD (dbj|BAA90261.1) 33 0.18
At4g04180 vesicle transfer ATPase like protein 32 0.41
At1g62150 unknown protein 32 0.41
At5g50960 nucleotide-binding protein 32 0.70
At4g19540 ATP binding protein - like 31 1.2
At3g24430 mrp protein, putative 29 4.5
At4g04300 hypothetical protein 28 5.9
At3g30420 hypothetical protein 28 7.8
At1g62120 unknown protein 28 7.8
>At3g10350 putative ATPase
Length = 386
Score = 224 bits (570), Expect = 7e-59
Identities = 110/132 (83%), Positives = 123/132 (92%), Gaps = 1/132 (0%)
Query: 8 LSVKSVAAPTESISVFDDMVAGTERKYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVS 67
L VKSVA+PTE+IS FD+MV+GT+RKYYMLGGKGGVGKTSCAASLAV+FANNGHPTLVVS
Sbjct: 63 LQVKSVASPTETISEFDEMVSGTKRKYYMLGGKGGVGKTSCAASLAVRFANNGHPTLVVS 122
Query: 68 TDPAHSLSDSFAQDLTGGALVQVDGPDYPLFALEINPEKAREDFRDVAKQNGGSTGVKDF 127
TDPAHSLSDSFAQDLTGG LV V+GP+ PLFALEINPEKARE+FR ++ NGG TGVKDF
Sbjct: 123 TDPAHSLSDSFAQDLTGGMLVPVEGPEAPLFALEINPEKAREEFRSASQMNGG-TGVKDF 181
Query: 128 MDGMGLGMIVDQ 139
MDGMGLGM+V+Q
Sbjct: 182 MDGMGLGMLVEQ 193
>At5g60730 ATPase - like protein
Length = 393
Score = 156 bits (395), Expect = 1e-38
Identities = 81/133 (60%), Positives = 97/133 (72%), Gaps = 4/133 (3%)
Query: 7 LLSVKSVAAPTESISVFDDMVAGTERKYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVV 66
L V+S+A E S F++MV+ +RKYY+LGGKGGVGKTSCAASLAVKFA++GHPT+VV
Sbjct: 44 LFRVRSLATLAEGASHFNEMVSVNQRKYYLLGGKGGVGKTSCAASLAVKFASHGHPTIVV 103
Query: 67 STDPAHSLSDSFAQDLTGGALVQVDGPDYPLFALEINPEKAREDFRDVAKQNGGSTGVKD 126
STDPAHSLSDSF+QDL+GG L V G D PL ALEI P E +D K+ G VK+
Sbjct: 104 STDPAHSLSDSFSQDLSGGVLKPVQGVDSPLLALEITP----EIMKDEIKRQTGDKSVKN 159
Query: 127 FMDGMGLGMIVDQ 139
MD MGLGM +
Sbjct: 160 MMDSMGLGMFAGE 172
>At1g01910 arsA homolog (hASNA-I), putative
Length = 353
Score = 66.2 bits (160), Expect = 3e-11
Identities = 40/112 (35%), Positives = 63/112 (55%), Gaps = 4/112 (3%)
Query: 33 KYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSDSFAQDLT-GGALVQVD 91
K+ +GGKGGVGKT+C++ LA+ A+ L++STDPAH+LSD+F Q T LVQ
Sbjct: 20 KWVFVGGKGGVGKTTCSSILAICLASVRSSVLIISTDPAHNLSDAFQQRFTKSPTLVQGF 79
Query: 92 GPDYPLFALEINPEKAREDFRDVAKQNGGSTGVKDFMDGMGLGMIVDQTAKV 143
LFA+E++P +D +G + + + + G+ M + K+
Sbjct: 80 S---NLFAMEVDPTVETDDMAGTDGMDGLFSDLANAIPGIDEAMSFAEMLKL 128
>At5g23930 putative protein
Length = 457
Score = 35.8 bits (81), Expect = 0.037
Identities = 30/119 (25%), Positives = 47/119 (39%), Gaps = 10/119 (8%)
Query: 4 TVFLLSVKSVAAPTESISVFDDMVAGTERKYYMLGGKGGVGKTSCAASLAVKFANNGHPT 63
+ F S SV T+ +S+ D+ T Y++ G K + + S+ F G+P
Sbjct: 29 SAFTESFSSVVTTTKDLSLEDERKRKTFTVSYLIDSLGLTTKLAESISMKANFDEKGNPD 88
Query: 64 LVVSTDPAHSLSDSFAQDLTGGALVQVDGPDYPLFALEINPEKAREDFRDVAKQNGGST 122
V+ ++ DS + YP F +E NPEK K NG S+
Sbjct: 89 SVLKLLRSYGFKDSQISSIIS---------TYPRFLIE-NPEKTLRAKLHFLKLNGASS 137
>At5g24020 septum site-determining MinD (dbj|BAA90261.1)
Length = 326
Score = 33.5 bits (75), Expect = 0.18
Identities = 17/58 (29%), Positives = 29/58 (49%), Gaps = 3/58 (5%)
Query: 15 APTESISVFD---DMVAGTERKYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTD 69
+P S+ F+ ++ T R + GKGGVGKT+ A++ + A G + + D
Sbjct: 39 SPIRSVLQFNRKPELAGETPRIVVITSGKGGVGKTTTTANVGLSLARYGFSVVAIDAD 96
>At4g04180 vesicle transfer ATPase like protein
Length = 536
Score = 32.3 bits (72), Expect = 0.41
Identities = 28/91 (30%), Positives = 39/91 (42%), Gaps = 11/91 (12%)
Query: 19 SISVFDDMVAGTERKY-------YMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTDPA 71
S V+DD+V GT K+ + G G GKTSCA +A G P L V +
Sbjct: 268 SPEVYDDIVRGTRSKFESNRPRAVLFEGPPGTGKTSCARVIA---NQAGIPLLYVPLEAV 324
Query: 72 HSLSDSFAQDLTGGALVQVDG-PDYPLFALE 101
S ++ L G Q + PD + L+
Sbjct: 325 MSKYYGESERLLGAVFSQANELPDGAIIFLD 355
>At1g62150 unknown protein
Length = 463
Score = 32.3 bits (72), Expect = 0.41
Identities = 21/76 (27%), Positives = 35/76 (45%), Gaps = 1/76 (1%)
Query: 2 IITVFLLSVKSVAAPTESISVFDDMVAGTERKYYMLGGKGGVGKTSCAASLAVKFANNGH 61
+ +VF S SVA + +S D Y++ G K + + S+ V F N G+
Sbjct: 31 VFSVFSNSFSSVATAAD-VSFRDSRKGNNFTVSYLVDSLGLASKLAESISMKVSFENKGN 89
Query: 62 PTLVVSTDPAHSLSDS 77
P V++ +H +DS
Sbjct: 90 PDTVLNLLRSHEFTDS 105
>At5g50960 nucleotide-binding protein
Length = 350
Score = 31.6 bits (70), Expect = 0.70
Identities = 19/51 (37%), Positives = 27/51 (52%), Gaps = 6/51 (11%)
Query: 25 DMVAGTER------KYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTD 69
D+VA ER K +L GKGGVGK++ +A L+ A H ++ D
Sbjct: 47 DLVAIAERMSTVKHKILVLSGKGGVGKSTFSAQLSFALAGMDHQVGLMDID 97
>At4g19540 ATP binding protein - like
Length = 313
Score = 30.8 bits (68), Expect = 1.2
Identities = 14/20 (70%), Positives = 16/20 (80%)
Query: 39 GKGGVGKTSCAASLAVKFAN 58
GKGGVGK+S A +LAV AN
Sbjct: 51 GKGGVGKSSTAVNLAVALAN 70
>At3g24430 mrp protein, putative
Length = 532
Score = 28.9 bits (63), Expect = 4.5
Identities = 22/67 (32%), Positives = 32/67 (46%), Gaps = 14/67 (20%)
Query: 40 KGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSDSFAQDLTGGALVQVDGPDYPLFA 99
KGGVGK++ A +LA A G + F D+ G +L + P+ +
Sbjct: 185 KGGVGKSTVAVNLAYTLAGMGARVGI------------FDADVYGPSLPTMVNPESRI-- 230
Query: 100 LEINPEK 106
LE+NPEK
Sbjct: 231 LEMNPEK 237
>At4g04300 hypothetical protein
Length = 286
Score = 28.5 bits (62), Expect = 5.9
Identities = 26/104 (25%), Positives = 46/104 (44%), Gaps = 23/104 (22%)
Query: 21 SVFDDMVAGTERKYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSDSFAQ 80
+VF+DM ++ + G GG+GKT +LA + G L +++ SL
Sbjct: 16 AVFNDMGG----VFFFVYGSGGIGKTFIWKTLAAVGRSKGQTCLNIASSGIASLL----- 66
Query: 81 DLTGGALVQVDGPDYPLFALEINPE-------KAREDFRDVAKQ 117
L GG + + F++ +NP+ K + D D+ K+
Sbjct: 67 -LEGGRIA------HYRFSIPLNPDEFSVCKIKPKSDLADLIKE 103
>At3g30420 hypothetical protein
Length = 837
Score = 28.1 bits (61), Expect = 7.8
Identities = 16/56 (28%), Positives = 27/56 (47%), Gaps = 3/56 (5%)
Query: 22 VFDDM---VAGTERKYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSL 74
++DD+ V E K + L G GG GKT ++ +NG + V++ +L
Sbjct: 356 IYDDVLKSVTNKEGKLFFLYGDGGTGKTFLYKTIISALRSNGKNVMPVASSAIAAL 411
>At1g62120 unknown protein
Length = 437
Score = 28.1 bits (61), Expect = 7.8
Identities = 14/43 (32%), Positives = 23/43 (52%)
Query: 35 YMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSDS 77
Y++ G K + + S+ V F N G+P V+S +H +DS
Sbjct: 61 YLVDSLGLATKVAESISMKVSFDNKGNPDSVLSLLRSHGFTDS 103
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.347 0.152 0.531
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,382,115
Number of Sequences: 26719
Number of extensions: 288214
Number of successful extensions: 1151
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 9
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 1138
Number of HSP's gapped (non-prelim): 13
length of query: 359
length of database: 11,318,596
effective HSP length: 100
effective length of query: 259
effective length of database: 8,646,696
effective search space: 2239494264
effective search space used: 2239494264
T: 11
A: 40
X1: 14 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC144928.8


